Abstract
A biphenyl-utilizing bacterium isolated from polychlorinated biphenyls (PCBs)-contaminated soils grew on tryptic soy at temperatures between 4 and 40°C. The Gram-negative rod bacterium formed yellow colonies on nutrient agar and it denitrified nitrate to nitrogen. Analysis of cellular fatty acids showed that it was most closely related to Hydrogenophaga taeniospiralis. At 5°C, biphenyl-grown cells cometabolically degraded di- and trichlorinated isomers of PCBs in 10 ppm of Aroclor 1248. At 30°C, PCBs that were removed included a congener with four chlorine substituents. At 5°C, cells transformed 2,4′-dichlorobiphenyl (2,4′-DCB) and accumulated ortho-chlorinated meta-cleavage product as a stable metabolite. Analysis of extracts of culture supernatant by gas chromatography–mass spectrometry indicated that products of transformation of 2,4′-DCB included 2- and 4-chlorobenzoic acid (2- and 4-CBA), suggesting that (chloro)biphenyl-degrading upper-pathway enzymes of the bacterium are active at low temperature. The bacterium Hydrogenophaga sp. IA3-A is a PCB-degrading psychrotolerant strain.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Polychlorinated biphenyls (PCBs) are a group of 209 isomers with a biphenyl nucleus substituted with 1 to 10 chlorine atoms. Although banned in several countries in the 1970s and 1980s, PCBs are ubiquitous [11] and are among the most persistent organic pollutants. Several Arctic and sub-Arctic sites are contaminated with PCBs. Because of the remote location of these sites, treatment using conventional cleanup technologies is very expensive, and bioremediation is a cost-saving alternative [18].
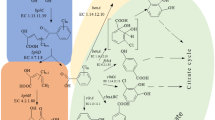
Microbial degradation of PCBs has been reported previously [1, 3, 4, 9, 16, 18]. PCBs transforming bacterial strains utilize the same sets of enzymes employed in the catabolism of biphenyl [1]. Bacteria able to grow on biphenyl usually have the ability to cometabolize various PCB congeners [13]. Biphenyl is often required as carbon source and as an inducer of the necessary enzymes. As illustrated in Fig. 1, low chlorine-containing PCBs are degraded through a four step meta-cleavage pathway to generate a five carbon compound and chlorobenzoic acid. In most cases, the chlorobenzoic acids accumulate as dead-end metabolites [1, 7].
Upper pathway for bacterial aerobic degradation of (chloro)biphenyl. Compounds: I, (Chloro)biphenyl; II, 2,3-dihydroxy-4-phenylhexa-2,4-diene (dihydrodiol compound); III, 2,3-dihydroxybiphenyl; IV, 2-hydroxy-6-oxo-6-phenylhexa-2,4-dienoic acid (meta-cleavage compound); V, 2-hydroxypenta-2,4-dienoate; VI, (chloro)benzoic acid. Enzyme activities: BphA, biphenyl dioxygenase; BphB, dihydrodiol dehydrogenase; BphC, 2,3-dihydroxybiphenyl dioxygenase; BphD, 2-hydroxy-6-oxo-6-phenylhexa-2,4-dienoic acid hydrolase [8, 12]. Numbering scheme for compound I: 2, and 2′, ortho-; 6 and 6′, ortho-; 4 and 4′, para-; 3 and 3′, meta-; 5 and 5′, meta-position.
Bioremediation of PCB-contaminated sites in cold climates will require the use of cold-adapted microorganisms. Several studies have reported the degradation of PCBs by mesophilic bacterial strains [1, 3, 4, 9, 10, 13, 20]. Information on PCB-degrading cold-adapted bacteria is scarce. The few cold-adapted bacteria that degrade PCBs belong to the genus Pseudomonas [16, 18]. Additionally, production of metabolite at low temperature by PCB-degrading cold-adapted bacteria is yet to be reported. In this study, biodegradation of PCBs in Aroclor 1248 at low temperature by a psychrotolerant bacterium is investigated. Adaptation of (chloro)biphenyl-degrading upper-pathway enzymes of the bacterium to low temperature was confirmed by analysis of metabolite production from 2,4′-dichlorobipheynyl (2,4′-DCB) at 5°C. To our knowledge, psychrotolerant strains of members of the genus Hydrogenophaga are yet to be characterized for their potential to attack PCBs or other organic contaminants.
Materials and Methods
Organism and growth conditions
The organism was isolated from PCB-contaminated soil in western Newfoundland (48.32°N, 58.33°W), Canada [14]. Soil (5 g) was transferred into 50 mL of minimal salt medium (MSM) [9] containing 0.5% biphenyl (wt/vol) (Sigma). The culture was incubated at 30°C at 200 rpm. After 7 days, the culture was streaked on MSM solidified with 1.5% Bacto-agar (MSA, Difco) in Petri plates. Biphenyl was placed in the lid of the plates and incubated at 30°C. Colonies that formed on MSA were repeatedly transferred on trypticase soy agar (TSA, Difco). An isolate that could be maintained on biphenyl-containing MSA after several transfers on TSA was chosen for further investigation. Growth of the isolate on tryptic soy broth (TSB, Difco) at 4, 10, 15, 20, 25, 30, 35, 40, and 45°C was determined turbidometrically. The isolate was characterized morphologically and biochemically. Its identity was confirmed using the API 20E test kits (bioMerieux Inc., St. Laurent, QE, Canada) and by analyses of cellular fatty acid profile using the MIDI/Hewlett-Packard Microbial Identification System (Analytical Services, VT, USA).
Cometabolism of PCBs
Cells were grown in MSM containing 0.5% biphenyl (wt/vol) at 30°C at 200 rpm to the exponential phase and harvested by centrifugation (15,000 g for 15 min at 4°C). Cells were washed with 50 mM sodium phosphate buffer (pH 7.5), and then resuspended in the same buffer. Two milliliters of cell suspension (final OD600 of 2.0) was dispensed into a 25-mL glass tube. Cells in control cultures were inactivated with two drops of 70% perchloric acid. Thereafter, acetone solution of Aroclor 1248 (Crescent Chemical Co., NY, USA) was added into experimental and control cultures to a final concentration of 10 ppm, followed by the addition of an internal standard, 2,2′,4,4′,6,6′-hexachlorobiphenyl (Ultra Scientific, RI, USA) to a final concentration of 4 ppm. Cultures were incubated at 5 or 30°C on a shaker at 200 rpm for 48 h. Metabolites production at low temperature was monitored by adding 500 μM of 2,4′-DCB (Crescent Chemical Co., NY, USA) into 4 mL of cell suspensions (final OD600 of 1.0) followed by incubation at 5°C on a shaker at 200 rpm for 72 h. 2,4′-DCB transformation was assessed in comparison with acid-inactivated cells. The accumulation and identity of the meta-cleavage metabolite produced from 2,4′-DCB were determined by UV spectrometry [20]. After the incubation period, reactions were stopped with two drops of 70% perchloric acid.
Extraction of PCBs and detection of metabolites
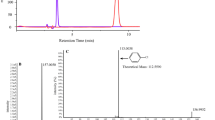
PCBs were extracted as described [3]. Polar metabolites produced from 2,4′-DCB were extracted from culture supernatants (pH 3) with two volumes of ethyl acetate, evaporated and derivatized with pyridine and N-O-bis-(trimethylsilyl)-trifluoroacetamide containing 1% trimethylchloro- silane (TMS, Sigma) according to Maltseva et al. [15]. Hexane extracts and TMS-derivatized samples (1 μL) were analyzed by gas chromatography using a Varian CP-3800 connected to a Varian Saturn 2000 ion trap mass spectrometer (Varian Inc., CA, USA). The capillary column was ZB-5 (5% phenyl polysiloxane) (30 m × 0.25 mm × 0.25 μm; Phenomenex, CA, USA). The carrier gas was helium at a flow rate of 1 mL min−1. PCBs in hexane extracts and metabolites in derivatized samples were analyzed using the temperature program of Bedard et al. [3] and Maltseva et al. [15], respectively. The number of chlorine substituent of peaks corresponding to congeners in Aroclor 1248 was determined by monitoring ions at the m/z of PCB congeners. Congeners were assigned by comparing the retention time and/or number of chlorine substituent of peaks to published data [2, 4, 6]. 2- and 4-Chlorobenzoic acid (2- and 4-CBA, Ultra Scientific) were identified by GC-MS in comparison with the authentic standards.
Oxygen uptake by whole cells
Uptake of oxygen in the presence of biphenyl (250 μM), 2,4′-DCB (250 μM), benzoic acid (2 mM), 2-CBA (2 mM), and 4-CBA (2 mM) by cells grown on biphenyl (0.5%, wt/vol) or benzoic acid (4 mM) was measured using a Clark-type oxygen electrode (Yellow Springs Instrument Co., OH, USA) connected to a 2-mL reaction chamber held at 30°C. The mixture contained 50 mM sodium phosphate buffer (pH 7.5), washed cells (final OD600 of 1.0), and substrate. Oxygen uptake values were corrected for endogenous respiration.
Enzyme assays
Cell-free extracts were prepared from 2 g of washed cells suspended in 4 mL of 50 mM 2-(N-morpholino)ethanesulfonic acid buffer (pH 6.0) containing EDTA (1 mM), dithiothreitol (1 mM), and glycerol (15%, vol/vol). The suspension was sonicated with constant cooling for 6 min, followed by centrifugation at 45,000 g for 1 h at 4°C. The supernatant was used for enzyme assay. The activity of biphenyl dioxygenase was followed spectrophotometrically at 30°C by a decrease in absorbance of NADPH at 340 nm due to the presence of extracts of cells grown on either biphenyl, TSB, succinate, or succinate plus 2,4′-DCB. The reactions were followed in duplicate. Protein concentration was determined by the method of Bradford [5], with bovine serum albumin as a standard.
Results and Discussion
Identity of the psychrotolerant bacterial isolate
Seven bacteria capable of growing on biphenyl were isolated from PCB-contaminated soils [14]. However, only one of the two cold-adapted isolates obtained could be maintained on biphenyl after growth on TSA. Thus, this cold-adapted bacterium was chosen for further investigation. The bacterium grew on TSB at low temperature and its optimum temperature was above 20°C, indicating that it is a psychrotolerant strain [19]. The Gram-negative rod-shaped bacterium was motile. It occurred singly or in pairs, and was non-spore-forming. It formed circular, smooth, opaque, convex, and pale-yellow colonies with entire margin when cultured on nutrient agar. It was oxidase positive and exhibited a weak positive catalase reaction. It reduced nitrate and denitrified nitrate to nitrogen. It did not produce indole or hydrogen sulfide. It hydrolyzed gelatin, while DNA, starch, urea, lipid, and casein were not hydrolyzed. It utilized sucrose, mannitol, D-xylose, pyruvate, succinate, acetate, and citrate, but it did not utilize lactose, D-fructose, maltose, p-hydroxybenzoate, or formate. It did not produce acid from oxidative-fermentative media containing glucose, mannitol, or maltose. It was negative for lysine, arginine, and ornithine decarboxylase. It grew on TSB at temperatures between 4°C and 40°C, and growth was not observed at 45°C. Its optimum temperature was 30°C.
Fatty acids, containing 10–18 carbon atoms, were detected in the cell membrane of the bacterium. The predominant fatty acids include closely eluting cis-7-hexadecenoic acid and 2-hydroxy-iso-pentadecanoic acid (47.75%), hexadecanoic acid (27.4%), and cis-7-octadecenoic acid (11.78%). Other fatty acids detected were 3-hydroxydecanoic acid (2.03%), dodecanoic acid (3.07%), tetradecanoic acid (1.86%), pentadecanoic acid (3.43%), cis-6-pentadecenoic acid (0.67%), heptadecanoic acid (1.22%), and cycloheptadecanoic acid (0.8%). Results of cellular fatty acid analysis suggested that it was most closely related to Hydrogenophaga taeniospiralis, with a similarity index (SI) of 0.669 [14]. The SI after comparing the fatty acids in the cell membrane of the isolate to those of Acidovorax konjaci (SI, 0.507), Hydrogenophaga pseudoflava (SI, 0.451), or Acidovorax avenae (SI, 0.399) was lower. Its ability to denitrify nitrate to nitrogen together with other physiological characteristics support the results obtained from fatty acid analysis. However, unlike a previously described strain of H. taeniospiralis [22], 3-hydroxydecanoic acid was detected in the cell membrane of the bacterium. This variation could be a result of exposure of the bacterium to PCBs present in the soil it was originally isolated from. A change in fatty acid composition was reported in a bacterial strain after exposure to toxic aromatic compounds [21]. The bacterium was designated as Hydrogenophaga sp. IA3-A. Attempts to identify the bacterium with the API 20E test kits did not give a conclusive result.
Cometabolism of PCBs
After 48 h, only 5 and 14 congeners of the 32 PCB congeners that were resolved by GC-MS were removed by biphenyl-grown resting cells incubated with Arolcor 1248 at 5°C and 30°C, respectively (Table 1). PCBs that were removed at 30°C include 2,4-, 2,5-, 2,3′-DCB, 2,2′,5-, 2,2′,4-, 2,2′,3-, 2,4′,6-, 2,3′,4-, 2,4′,5-, 2,4,4′-, 2′,3,4-trichlorobiphenyl (TCB), 2,2′,3,4-, 2,3,4′,6- and 2,3′,4′,6-TCB. At 5°C, only 2,3′-DCB, 2,2′,4-, 2,2′,3-, 2,4′,6-, and 2,4,4′-TCB were removed. At low temperature, there was insignificant or no removal of congeners with a ring substituted at 2-ortho- and 5-meta- position, while removal of congeners with this pattern of chlorination was moderate at 30°C. A similar observation was made for congeners with 2-ortho, 3-meta, plus 4-para-substitution at 5°C and 30°C. Among the TeCBs, the congener that was significantly removed at 30°C is more likely to be 2,3,4′,6-tetrachlorobiphenyl (TeCB), since this congener has a free 2,3- position (Fig. 1) on the para-substituted ring that was amenable to dioxygenation. These data suggest that cells of strain IA3-A attacked PCBs using a 2,3-dioxygenase. Most PCB-degrading bacteria are known to utilize a 2,3-dioxygenase [3]. However, removal of 2,2′,3,4- and 2,3′,4′,6-tetrachlorobiphenyl (TeCB) was also possible, assuming the bacterium could oxidize a ring with 2-ortho substitution. A comparison of the pattern of chlorination of congeners that were removed at 5°C and 30°C indicated that except for those with 2-ortho- plus 5-meta- or 2-ortho, 3-meta, plus 4-para substitution that were slightly or moderately removed at 30°C, the pattern was similar at both temperatures. The resistance of some congeners with these patterns of chlorination to degradation at low temperature was shown in previous studies on psychrotolerant bacteria that degraded Aroclor 1221 and 1242 [16, 18]. Because most of the congeners removed in the present study are lightly chlorinated, the bacterium will be more useful in the aerobic phase of the anaerobic–aerobic bioremediation process, where highly chlorinated congeners are anaerobically dechlorinated to produce lightly chlorinated congeners that can be degraded aerobically. Removal of ortho- plus para-substituted 2,2′,4- and 2,4,4′-TCB, typical byproducts of anaerobic dechlorination of PCBs, supports this conclusion.
Production of metabolites at low tempera- ture
Supernatants and ethyl acetate extracts of supernatants of culture incubated with 2,4′-DCB at 5°C for 72 h were analyzed by UV spectrometry and GC-MS to confirm whether cells could transform the congener at low temperature to metabolites of chlorobiphenyl degradation pathway. Three major metabolites were identified. A persistent yellow-colored metabolite with λmax at 397 nm was identified as ortho-substituted meta-cleavage product [20]. 2- and 4-CBAs were identified by GC-MS by comparing the mass spectrum and retention times of the metabolites to those of the authentic standards. In all cases, the mass spectrum and the retention times of the extracted metabolites were in good agreement with those of the standards (data not shown). None of these metabolites were present in the supernatants or extracts of supernatants of control cultures. These data agree with the results of previous studies on mesophilic strains that accumulated meta-cleavage product as a result of oxidation of the para-substituted ring of 2,4′-DCB [15]. The presence of 2- and 4-CBA in culture supernatants indicated that cells of Hydrogenophaga sp. IA3-A oxidized both rings of 2,4′-DCB. Several studies have reported CBAs as dead-end metabolites during transformation of PCBs [1, 7, 15]. However, the identity of the metabolites produced at low temperature by psychrotolerant PCB-degrading bacterial strains was not reported in previous studies [16, 18].
Oxygen uptake by whole cells
The rate of oxidation of biphenyl (147.44 ± 23.31 nmol/min/mg protein) by cells grown on biphenyl was three times greater than the rate of oxidation of 2,4′-DCB (48.12 ± 11.13 nmol/min/mg protein) by these cells, whereas the rate of oxidation of benzoic acid (11.29 ± 2.61 nmol/min/mg protein), present at a much higher concentration, was 13 times less than that of biphenyl. Biphenyl-grown cells failed to oxidize 2- or 4-CBA. Cells grown on benzoic acid could oxidize benzoic acid (13.05 ± 0.99 nmol/min/mg protein) or biphenyl (41.29 ± 0.97 nmol/min/mg protein), but these cells oxidized 2-CBA (1.52 ± 0.08 nmol/min/mg protein), 4-CBA (1.36 ± 1.44 nmol/min/mg protein), and 2,4′-DCB (1.52 ± 1.42 nmol/min/mg protein) at a similar but very low rate. Generally, results of oxygen uptake experiments support the conclusion that 2- and 4-CBAs are dead-end metabolites during the transformation of 2,4′-DCB by cells of Hydrogenophaga sp. IA3-A. Insignificant uptake of oxygen by benzoic acid-grown cells in the presence of 2- or 4-CBA indicated that benzoic acid–degrading enzymes in cells of strain IA3-A are different from those required to degrade 2- or 4-CBA. It is also possible that these compounds were present at a concentration that was toxic to cells.
Enzymatic studies
Extracts of cells grown on biphenyl, succinate, succinate plus 2,4′-DCB, or TSB contained the activity of biphenyl dioxygenase (Table 2). Although 2,4′-DCB was not a growth substrate for cells, activity of the enzyme was higher in extracts of cells grown on succinate with low amount of 2,4′-DCB than in extracts of cells grown on succinate only, whereas biphenyl dioxygenase was induced to a much higher level by growth on biphenyl. There was a relatively moderate level of constitutive enzyme production after growth on TSB, whereas constitutive production of the enzyme was much lower by growth on succinate only. Inductions of biphenyl dioxygenase gene of psychrotolerant bacterium, Pseudomonas strain Cam-1, by substrates other than biphenyl have been reported [17]. 2- and 4-Chlorobiphenyl induced the gene to a level greater than the basal level [17]. This is consistent with the results obtained in the present study that the activity of biphenyl dioxygenase was higher in the extracts of cells grown on succinate and 2,4′-DCB than in the extracts of cells grown on succinate only; however, induction of biphenyl dioxygenase in cells of psychrotolerant bacteria by a dichlorinated PCB congener has not been reported before.
The results of the present study suggest that biodegradation of PCBs is possible if Hydrogenophaga sp. IA3-A is used as part of a consortia with cold-adapted CBA-degrading bacteria. To our knowledge, cold-adapted bacteria that are capable of degrading CBAs are yet to be described. Therefore, further investigations of cold-adapted bacteria active against PCBs or CBAs at low temperature is required for successful bioremediation of PCB-contaminated sites in cold climates. The significance of meta-cleavage product with respect to degradation of ortho- plus para-substituted PCBs (e.g., 2,4′-DCB) at low temperature needs to be clarified, because this metabolite accumulated in culture supernatants.
Literature Cited
Ahmed M, Focht DD (1973) Degradation of polychlorinated biphenyls by two species of Achromobacter. Can J Microbiol 19:47–52
Albro PW, Corbett JT, Schroeder JL (1981) Quantitative characterization of polychlorinated biphenyl mixtures (Aroclors 1248, 1254 and 1260) by gas chromatography using capillary columns. J Chromatogr 136:147–153
Bedard DL, Untermann R, Bopp LH, Brennan MJ, Haberl ML, Johnson C (1986) Rapid assay for screening and characterizing microorganisms for the ability to degrade polychlorinated biphenyls. Appl Environ Microbiol 51:761–768
Bedard DL, Wagner RE, Brennan MJ, Haberl ML, Brown JF Jr (1987) Extensive degradation of Aroclors and environmentally transformed polychlorinated biphenyls by Alcaligenes eutrophus H850. Appl Environ Microbiol 53:1094–1102
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilising the principle of protein-dye binding. Anal Biochem 72:248–254
Frame GM, Cochrane JW, Bowadt SS (1996) Complete PCB congener distributions for 17 Aroclor mixtures determined by 3 HRGC systems optimized for comprehensive, quantitative, congener-specific analysis. J High Resol Chromatogr 19:657–668
Furukawa K (1994) Molecular genetics and evolutionary relationship of PCB-degrading bacteria. Biodegradation 5:289–300
Furukawa K, Arimura N (1987) Purification and properties of 2,3-dihydroxybiphenyl dioxygenase from polychlorinated biphenyl-degrading Pseudomonas pseudoalcaligenes and Pseudomonas aeruginosa carrying the cloned bphC gene. J Bacteriol 169:924–927
Furukawa K, Matsumura F, Tonomura K (1978) Alcaligenes and Acinetobacter strains capable of degrading polychlorinated biphenyls. Agric Biol Chem 4:545–548
Furukawa K (1982) Microbial degradation of polychlorinated biphenyls. In: Chakrabarty AM (ed) Biodegradation and detoxification of environmental pollutants. Boca Raton, Fla.: CRC Press Inc, pp 33–57
Kalmaz EV, Kalmaz GD (1979) Transport, distribution, and toxic effects of polychlorinated biphenyls in ecosystems. Rev Ecol Model 6:223–251
Kikuchi Y, Nagata Y, Hinata M, Kimbara K, Fukuda M, Yano K, Takagi M (1994) Identification of the bphA4 Gene encoding ferredoxin reductase involved in biphenyl and polychlorinated biphenyl degradation in Pseudomonas sp. Strain KK102. J Bacteriol 176:1689–1694
Kohler H-PE, Kohler-Staub D, Focht DD (1988) Cometabolism of polychlorinated biphenyls: Enhanced transformation of Aroclor 1254 by growing bacteria cells. Appl Environ Microbiol 54:1940–1945
Lambo AJ, Patel TR (2005) Isolation and characterization of a biphenyl-utilizing psychrotrophic bacterium, Hydrogenophaga faeniospiralis IA3-A, that cometabolize dichlorobiphenyls and polychlorinated biphenyl congenes in Arodor 1221. J Basic Microbiol (in press)
Maltseva VO, Tsoi TV, Quensen JF III, Fukuda M, Tiedje JM (1999) Degradation of anaerobic reductive dechlorination products of Aroclor 1242 by four aerobic bacteria. Biodegradation 10:363–371
Master ER, Mohn WW (1998) Psychrotolerant bacteria isolated from Arctic soil that degrade polychlorinated biphenyls at low temperatures. Appl Environ Microbiol 64:4823–4829
Master ER, Mohn WW (2001) Induction of bphA, encoding biphenyl dioxygenase, in two polychlorinated biphenyl-degrading bacteria, psychrotolerant Pseudomonas strain Cam-1 and mesophilic Burkholderia strain LB400. Appl Environ Microbiol 67:2669–2676
Mohn WM, Westerberg K, Cullen WR, Reimer KJ (1997) Aerobic biodegradation of biphenyl and polychlorinated biphenyls by Arctic soil microorganisms. Appl Environ Microbiol 63:3378–3384
Morita RY (1975) Psychrophilic bacteria. Bacteriol Rev 39:144–167
Seeger M, Timmis KN, Hofer B (1995) Conversion of chlorobiphenyls into phenylhexadienoates and benzoates by the enzymes of the upper pathway for polychlorinated degradation encoded by the bph locus of Pseudomonas sp. Strain LB400. Appl Environ Microbiol 61:2654–2658
Tsitko IV, Zaitsev GM, Lobanok AG, Salkinoja-Salonen MS (1999) Effect of aromatic compounds on fatty acid composition of Rhodococcus opacus. Appl Environ Microbiol 65:853–855
Willems A, Busse J, Goor M, Pot B, Falsen E, Jantzen E, Hoste B, Gillis M, Kersters K, Auling G, De Ley J (1989) Hydrogenophaga, a new genus of hydrogen–oxidizing bacteria that includes Hydrogenophaga flava comb. nov. (formerly Pseudomonas flava), Hydrogenophaga palleronii (formerly Pseudomonas palleronii), Hydrogenophaga pseudoflava (formerly Pseudomonas pseudoflava and Pseudomonas carboxydoflava) and Hydrogenophaga taeniospiralis (formerly Pseudomonas taeniospiralis). Int J Syst Bacteriol 39:319–333
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Lambo, A.J., Patel, T.R. Cometabolic Degradation of Polychlorinated Biphenyls at Low Temperature by Psychrotolerant Bacterium Hydrogenophaga sp. IA3-A. Curr Microbiol 53, 48–52 (2006). https://doi.org/10.1007/s00284-005-0194-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-005-0194-8