Abstract
The immunodominant antigen A, IsaA, of Staphylococcus aureus was found to include a putative soluble lytic transglycosylase domain in its C-terminal region. Since the presence of this distinctive domain suggested that the protein might participate in peptidoglycan turnover, as indicated in Gram-negative bacteria, its cellular location was investigated. The protein was found not only in the culture supernatant but also in the cell wall fraction. To estimate its physiological role for the bacterium, its cell surface distribution was studied by immunoelectron microscopy. Protein A-gold particles binding to the immune complex were mainly located on the septal region of the bacterial cell surface. These data suggested that IsaA might be involved in bacterial cell separation through a preferential interaction with peptidoglycan chain.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
We had previously found a 29 kDa protein in the culture supernatant of Staphylococcus aureus strain Cowan I and cloned its corresponding gene (accession no. AB042839). During our study, this protein was reported as the immunodominant antigen A, IsaA, by Lorenz et al. [12], though there was one amino acid difference at residue 172. They isolated this protein from methicillin-resistant Staphylococcus aureus, and showed that its antibody was highly detectable in serum from patients with staphylococcal infection [12]. However, the function of the protein has not been established. Recently, it was shown that IsaA might be regulated by the essential two-component system YycG and YycF [3]. This system is present in low G+C Gram-positive bacteria and has been reported to be essential for cell growth in Bacillus subtilis, Staphylococcus aureus, and Streptococcus pneumoniae [5, 11, 13, 18]. That is, the inactivation of the yycF/yycG genes is lethal in some bacteria [3, 5, 6, 13]. Moreover, yycF/yycG genes are expressed during exponential growth and shut off as the cells enter stationary phase [5]. In S. aureus, it has been shown that YycF bound specifically to the promoter region of the IsaA gene [3]. These data suggest an important role for IsaA in staphylococcal growth or survival. Therefore, we attempted to establish the protein’s function for the bacterial cells. For this purpose, a search of the conserved domain database for the deduced amino acid sequence of IsaA was performed, and its cellular distribution was also studied.
Materials and Methods
Production of IsaA recombinant protein
Chromosomal DNA isolated from S. aureus strain Cowan I was used for amplification by PCR, using primers 5′-CGCTGCTGAAGTAAACGTTGATC-3′ and 5′-ATGAAGGAATTAGAATCCCCAAGC-3′. The PCR product was cloned into pGEM-T Easy. The isolated plasmid DNA was digested with SpeI and EcoRI, and this fragment was then ligated into pProEX HTb (Life Technologies) that was digested with the same enzymes. The resulting construct was transformed into Escherichia coli DH5α for the production of recombinant protein with an N-terminal histidine tag. Induction of the recombinant plasmid was performed with 0.6 mM isopropyl-β-thiogalactopyranoside (IPTG) for 4 h at 37°C. After the treatment, the bacterial cells were collected by centrifugation and used for the isolation of inclusion bodies by the BugBuster Protein Extraction Reagent (Novagen). The isolated inclusion bodies were treated with the Protein Refolding Kit (Novagen) and the recombinant protein was purified by affinity chromatography using a Ni-NTA resin (Qiagen).
Preparation of IsaA antiserum
The purified recombinant protein (240 μg) mixed with complete Freund’s adjuvant was subcutaneously injected into a rabbit. This procedure was repeated four times at 3-week intervals with incomplete Freund’s adjuvant. The rabbit was bled 2 weeks after the last injection. Purified IgG from the serum and a preimmune serum was obtained using the MabTrap Kit (Amersham Bioscience).
Isolation of the bacterial cell fractions
A S. aureus spontaneous protein-A-deficient mutant, strain 2PF-18 [17], was used for the isolation of the bacterial cell fractions. The bacterial cells were precultured in Trypto-soya broth (Eikenkagaku), and then inoculated into 250 mL of the same fresh medium for additional incubation with continuous shaking for 4 h at 37°C to the mid-log phase. The culture was centrifuged and cell fractions were generally prepared according to Sugai et al. [16]. Briefly, the bacterial cells were washed twice with 0.9% NaCl and suspended in 6 mL of digestion buffer, 30% raffinose in 0.05 M Tris-HCl (pH 7.5) with 0.145 M NaCl, containing 1 mg of lysostaphin (Wako Chemical), and 75 units of Benzonase (Novagen), 1 mM phenylmethylsulfonyl fluoride (PMSF). The cell mixture was incubated for 1 h at 37°C with gentle shaking. After centrifugation at 8000g for 10 min, the supernatant, the cell wall fraction and the protoplasts were collected. The protoplasts were resuspended in 6 mL of 0.05 M Tris-HCl (pH 7.5), containing 75 units of Benzonase, with Vortex mixing and then placed on ice for 20 min. The mixture was spun at 2000g for 10 min, and the collected supernatant was subjected to centrifugation at 40,000g for 15 min to obtain the cell membrane pellet and cytoplasmic fraction. The culture supernatant and the cell wall fraction were concentrated utilizing an Ultrafree-15 Filter Unit with a 5 kDa molecular weight cutoff.
Detection of IsaA in the bacterial cell fractions
The concentrated supernatant (30 μg of protein), the cell wall fraction (8.3 μg), the cell membrane fraction (10 μg), and the cytoplasmic fraction (10 μg) were subjected to SDS-PAGE, and then blotted onto a PVDF membrane. Nonspecific binding sites were saturated with 3% bovine serum albumin (BSA) in 10 mM Tris-HCl (pH 7.5) with 0.15 M NaCl. The membranes were incubated with IsaA antiserum (diluted 1:3000) or the preimmune rabbit serum as a control for 2 h at room temperature, and washed with 20 mM Tris-HCl (pH 7.5) containing 0.5 M NaCl, 0.05% Tween 20, and 0.2% Triton X-100. The washed membranes were incubated with an alkaline phosphatase conjugated anti-rabbit IgG, F(ab′)2 as a secondary antibody (Sigma) for 1 h at room temperature. After washing, the membranes were developed with NBT/BCIP (Promega) as a substrate for the conjugated enzyme. The entire experiment was replicated once more.
Immunoelectron microscopy
The IsaA cell distribution was carried out according to Diaz et al. [1]. Briefly, S. aureus strain 2PF-18 cells in the exponential phase were collected and washed twice with phosphate-buffered saline (PBS). The washed cells were incubated for 1 h at 4°C with anti-IsaA IgG or preimmune serum IgG as a control, 1 mg/mL, in PBS containing 0.5% BSA. After washing twice with PBS, the collected cells were incubated with 1:25 diluted protein A–colloidal gold conjugate (10 nm particles, Sigma) for 1 h at 4°C. The cells were extensively washed with PBS and fixed with 0.5% glutaraldehyde. Drops of the cell suspension were deposited onto carbon-coated copper grids and examined under a transmission electron microscope. The same experiment using IsaA antiserum and preimmune serum was also performed.
Protein assay
A protein assay was performed with a BCA (bicinchoninic acid) protein assay reagent (Pierce), using BSA as a standard. Briefly, 0.1 mL of each standard or unknown protein sample was mixed with 2 mL of Working Reagent for the protein assay. For blanks, the diluent was used. After incubation at 60°C for 30 min, the absorbance at 562 nm of the mixture was measured. All assays were performed in duplicate.
Results and Discussion
Sequence similarity of IsaA
IsaA was encoded by a 699 bp open reading frame (Fig. 1). The deduced amino acid sequence indicated that IsaA possessed no Gram-positive bacterial cell wall sorting signals [15] and was probably a secretory protein consisting of 233 amino acids, including an N-terminal signal peptide of 29 amino acids (Fig. 1). A search of the conserved domain database at the National Center for Biotechnology Information showed that IsaA possessed a sequence similar to the soluble lytic transglycosylase (SLT) domain in its C-terminal region (Fig. 1). This domain is present in a number of prokaryotic, eukaryotic, and phage proteins [2, 10, 14]. Among the bacterial proteins, the IsaA SLT motif showed a high similarity to E. coli soluble lytic transglycosylase Slt70. Comparison of the corresponding region with SLT consensus sequences from the database and SLT70 is shown in Fig. 2. This domain consists of three motifs that form the lytic transglycosylase catalytic region [14]. A spacer region between motifs II and III was lacking in IsaA and homology in motif III was low, but all these motifs were found in the staphylococcal protein. Moreover, a glutamate, which had been shown to be involved in SLT70 catalytic activity, and an adjacent conserved serine residue in motif I were also maintained in IsaA [19]. Bacterial lytic transglycosylases degrade murein via cleavage of the bond between N-acetylmuramic acid and N-acetylglucosamine [4, 7, 9, 20, 21]. This type of enzyme is generally found in Gram-negative bacteria [14] and is considered an autolytic enzyme involved in the controlled turnover of the bacterial cell wall [8]. For such a role, the enzymes need to function on the bacterial cell wall. In fact Slt70 was shown to be specifically located on the cell wall with a strong interaction with murein structure [22]. Therefore, the location of IsaA was investigated in isolated staphylococcal cellular fractions.
Alignment of the presumptive IsaA SLT domain. The deduced amino acid sequence of IsaA was compared with the SLT consensus sequence from the conserved domain database and the SLT domain of the E. coli soluble lytic transglycosylase, Slt70. A black or gray background indicates amino acids that are identical for all or two out of the three sequences, respectively. The glutamic acid residue marked with a star in motif I is the catalytic site in Slt70. The three motifs in the alignment are described in reference [14].
Detection of IsaA in the bacterial cell fractions
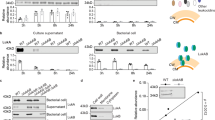
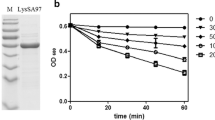
To obtain a specific antiserum against IsaA, a recombinant protein was prepared. The open reading frame for the gene, except for the signal peptide region, was ligated into an expression vector. The recombinant protein carrying an N-terminal polyhistidine tag was obtained from isolated inclusion bodies by affinity chromatography (Fig. 3). The antiserum prepared to the recombinant protein was used for the detection of IsaA in the cell fractions from S. aureus strain 2PF-18 by Western blotting. After development of the blotted membranes, a distinct band with approximately 26 kDa molecular weight was found in the cell wall fraction and the culture supernatant, but barely detectable in the cell membrane and not detectable in the cytoplasmic fractions (Fig. 4). The antiserum did not react to lysostaphin used for the cell wall preparation. Preimmune rabbit serum showed no detectable bands with any of these fractions. Therefore, it was concluded that IsaA existed in the cell wall fraction and in the culture supernatant. This result also indicated that IsaA was not tethered to the cell membrane, because it was easily removed from the cells by lysostaphin treatment. IsaA probably works on the staphylococcal cell wall with a preferential interaction with murein or its accessory components. There was a slight difference in the molecular size of IsaA between the two strains used in this study, but a minor variation in molecular mass of this protein has also been reported in some strains [12].
Detection of IsaA in the bacterial cell fractions. The culture supernatant (lanes 4 and 9), cell wall fraction (lanes 2, 5 and 10), cell membrane fraction (lanes 6 and 11), and cytoplasmic fraction (lanes 7 and 12) were applied to a 12.5% polyacrylamide gel. After blotting, the membranes were stained with Coomassie brilliant blue (lanes 1 to 3), or reacted with IsaA antiserum (lanes 4 to 8) or pre-immune serum (lanes 9 to 12). Lysostaphin used for preparation of the cell wall fraction was also tested for its reactivity to the antiserum (lanes 3 and 8). Molecular weights (kDa) of the standard proteins (lane 1) are shown on the left.
Distribution of IsaA on the bacterial cell surface
The cell surface distribution of IsaA was studied by using immunoelectron microscopy. The intact bacterial cells were incubated with IgG from the antiserum or the preimmune rabbit serum and then with colloidal-gold-labeled protein A. Binding particles were mainly found in the septal region of dividing cells (Fig. 5A). Control experiments using preimmune rabbit serum showed no distinct binding of the gold particles to the bacterial cell (Fig. 5B).
These results suggest that IsaA may be involved in cell separation, although its role is not defined yet. It is currently unclear whether IsaA really possesses cell wall hydrolyzing activity. Further studies, including a characterization of IsaA, are required to establish the participation of IsaA in the staphylococcal murein degradation process.
Literature Cited
E Díaz E García C Ascaso E Méndez R López JF García (1989) ArticleTitleSubcellular localization of the major pneumococcal autolysin: A peculiar mechanism of secretion in Escherichia coli J Biol Chem 264 1238–1244
BW Dijkstra A-MWT Thunnissen (1994) ArticleTitle“Holy” protein II: The soluble lytic transglycosylase Curr Opin Struct Biol 4 810–813
S Dubrac T Msadek (2004) ArticleTitleIdentification of genes controlled by the essential YycG/YycF two-component system of Staphylococcus aureus J Bacteriol 186 1175–1181
H Engel B Kazemier W Keck (1991) ArticleTitleMurein-metabolizing enzymes from Escherichia coli: Sequence analysis and controlled overexpression of the slt gene, which encodes the soluble lytic transglycosylase J Bacteriol 173 6773–6782
C Fabret JA Hoch (1998) ArticleTitleA two-component signal transduction system essential for growth of Bacillus subtilis: Implications for anti-infective therapy J Bacteriol 180 6375–6383
K Fukuchi Y Kasahara K Asai K Kobayashi S Moriya N Ogasawara (2000) ArticleTitleThe essential two-component regulatory system encoded by yycF and yycG modulates expression of the ftsAZ operon in Bacillus subtilis Microbiology 146 1573–1583
JV Höltje D Mirelman N Sharon U Schwarz (1975) ArticleTitleNovel type of murein transglycosylase in Escherichia coli J Bacteriol 124 1067–1076
JV Höltje (1998) ArticleTitleGrowth of the stress-bearing and shape-maintaining murein sacculus of Escherichia coli Microbiol Mol Biol Rev 62 181–203
W Keck FB Wientjes U Schwarz (1985) ArticleTitleComparison of two hydrolytic murein transglycosylases of Escherichia coli Eur J Biochem 148 493–497
EV Koonin E Rudd (1994) ArticleTitleA conserved domain in putative bacterial and bacteriophage transglycosylases Trends Biochem Sci 19 106–107
R Lange C Wagner A Saizieu Particlede N Flint J Molnos M Stieger P Caspers M Kamber W Keck KE Amrein (1999) ArticleTitleDomain organization and molecular characterization of 13 two-component systems identified by genome sequencing of Streptococcus pneumoniae Gene 237 223–234
U Lorenz K Ohlsen H Karch M Hecker A Thiede J Hacker (2000) ArticleTitleHuman antibody response during sepsis against targets expressed by methicillin resistant Staphylococcus aureus FEMS Immunol Med Microbiol 29 145–153
PK Martin T Li D Sun DP Biek MB Schmid (1999) ArticleTitleRole in cell permeability of an essential two-component system in Staphylococcus aureus J Bacteriol 181 3666–3673
AR Mushegian KJ Fullner EV Koonin EW Nester (1996) ArticleTitleA family of lysozyme-like virulence factors in bacterial pathogens of plants and animals Proc Natl Acad Sci USA 93 7321–7326
O Schneewind P Model VA Fischetti (1992) ArticleTitleSorting of protein A to the staphylococcal cell wall Cell 70 267–281
M Sugai T Akiyama H Komatsuzawa Y Miyake H Suginaka (1990) ArticleTitleCharacterization of sodium dodecyl sulfate-stable Staphylococcus aureus bacteriolytic enzymes by polyacrylamide gel electrophoresis J Bacteriol 172 6494–6498
M Sugai H Komatsuzawa T Akiyama Y-M Hong T Oshida Y Miyake T Yamaguchi H Suginaka (1995) ArticleTitleIdentification of end-ß-N-acetylglucosaminidase and N-acetylmuramyl-L-alanine amidase as cluster-dispersing enzyme in Staphylococcus aureus J Bacteriol 177 1491–1496
JP Throup KK Koretke AP Bryant KA Ingraham AF Chalker Y Ge et al. (2000) ArticleTitleA genomic analysis of two-component signal transduction in Streptococcus pneumoniae Mol Microbiol 35 566–576
A-MWH Thunnissen AJ Dijkstra KH Kalk HJ Rozeboom H Engel W Keck BW Dijkstra (1994) ArticleTitleDoughnut-shaped structure of a bacterial muramidase revealed by X-ray crystallography Nature 367 750–753
A-MWT Thunnissen HJ Rozeboom KH Kalk BW Dijkstra (1995) ArticleTitleStructure of the 70-kDa soluble lytic transglycosylase complexed with bulgecin A. Implications for the enzymatic mechanism Biochemistry 34 12729–12737
EJ Asselt ParticleVan A-MWT Thunnissen BW Dijkstra (1999) ArticleTitleHigh resolution crystal structures of Escherichia coli lytic transglycosylase Slt70 and its complex with a peptidoglycan fragment J Mol Biol 291 877–898
B Walderich JV Höltje (1991) ArticleTitleSubcellular distribution of the soluble lytic transglycosylase in Escherichia coli J Bacteriol 173 5668–5676
Acknowledgments
We are grateful to Dr. S. Yamada, Kawasaki Medical College, for S. aureus 2PF-18; Dr. S. Sakurai, Kohno Clinical Medicine Research Institute, for the N-terminal analysis; and Ms. C. Sasaki, Institute for Ultrastructural Morphology, Postgraduate, St. Marianna University School of Medicine, for technical assistance. We also thank Dr. I. Kondo, Jikei University School of Medicine, and Dr. T. Wadström, Lund University, for critical suggestions.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Sakata, N., Terakubo, S. & Mukai, T. Subcellular Location of the Soluble Lytic Transglycosylase Homologue in Staphylococcus aureus. Curr Microbiol 50, 47–51 (2005). https://doi.org/10.1007/s00284-004-4381-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00284-004-4381-9