Abstract
Background
Tamoxifen is frequently used for the treatment of hormone receptor positive breast cancer (BC). Mainly CYP2D6 is responsible for the transformation to therapeutically active metabolites, but CYP2C19, CYP2C9 and CYP2B6 also are involved. We investigated the impact of polymorphisms within the genes encoding these CYP enzymes on the relapse-free time (RFT) in patients with BC.
Methods
Ninety-nine patients with hormone receptor positive BC, who had undergone adjuvant tamoxifen therapy, were genotyped for seventeen common variants within the genes encoding CYP2D6, CYP2C9, CYP2C19 and CYP2B6 using TaqMan and PCR-RFLP technology. Kaplan–Meier and Cox regression analyses were performed to elucidate the impact of genetic variants on RFT. Furthermore, CYP2D6 metabolic activity was determined in a subset of 50 patients by assessing dextromethorphan/dextrorphan urinary excretion ratios. CYP2D6 activity was compared to the CYP2D6 allelic combinations to evaluate the predictive value of the CYP2D6 genotyping results on phenotype.
Results
Although a trend toward longer RFTs in carriers of CYP2D6 allele combinations encoding for extensive and ultrafast metabolizer phenotypes was observed, none of the investigated genetic variants had a statistically significant impact on RFT. The combined analysis of five major CYP2D6 variants was useful for the discrimination between poor and non-poor metabolizers.
Conclusions
Comprehensive CYP2D6 genotyping has a good predictive value for CYP2D6 activity. Common variants in CYP2C9, CYP2C19, CYP2D6, and CYP2B6 did not have a significant impact on the RFT in this cohort of patients with BC.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
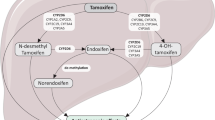
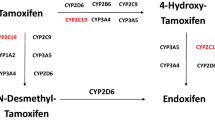
Breast cancer (BC) is one of the most frequent cancers in the Western world. Tamoxifen, a selective estrogen receptor modulator (SERM), is widely used as an efficient therapeutic option in the treatment and relapse prevention of estrogen receptor (ER)-positive BC [12]. Tamoxifen is transformed predominantly by the drug-metabolizing enzymes CYP3A4 and CYP2D6 into the therapeutically more efficient drug metabolites 4-hydroxytamoxifen (4-OH-tamoxifen) and endoxifen. By binding to the ER α (ERα) tamoxifen and its metabolites modulate, the estrogen-induced transcription of ERα target genes. The metabolites 4-OH-tamoxifen and endoxifen show up to 100 times higher affinity to the ERα than the parental compound [7]. As a result, the efficacy of tamoxifen strongly depends on its appropriate bioactivation by cytochrome P450 enzymes. CYP2D6 is highly polymorphic and shows a high interindividual variability in its activity. Currently, there are more than 100 genetic variants described for CYP2D6 [21]. Several genetic variants are known to cause an impaired enzyme activity (CYP2D6*10, *41) or to even induce a total loss of enzyme function (e.g., CYP2D6*4, *5, *6). Furthermore, gene duplications or multiplications of CYP2D6 that occur in 2–3 % of white individuals may lead to an exceptionally fast metabolism of CYP2D6 substrates. Genetic polymorphisms of CYP2D6 have been repeatedly associated with a changed therapeutic outcome in patients with BC treated with tamoxifen. Several studies could demonstrate that genetic variants causing an impaired CYP2D6 enzyme activity or even a loss of CYP2D6 function are associated with a shorter relapse-free time (RFT) upon tamoxifen therapy [10, 20]. Other studies, however, showed conflicting results. Especially in studies where the efficacy of tamoxifen in relation to the CYP2D6 metabolizer status was tested in an adjuvant setting, no clear relationship between CYP2D6 status and therapeutic outcome was observed [16, 18]. The latter studies are, however, controversial in the literature, due to the facts that DNA was isolated from tumor tissues and not from blood and that Hardy–Weinberg equilibria were partly not reached for several CYP2D6 genetic variants [14, 23]. Other enzymes involved in tamoxifen metabolism comprise CYP2C9, CYP2C19 and CYP2B6 [13]. These three enzymes are also involved in the formation of 4-OH-tamoxifen and endoxifen, but their contribution may depend on actual tamoxifen concentrations and on CYP2D6 activity.
In this study, we studied the impact of genetic variants in the genes encoding the drug-metabolizing enzymes CYP2C9, CYP2C19, CYP2B6 and CYP2D6 on the RFT in patients with BC treated with tamoxifen in an adjuvant setting. We chose to investigate genetic variants that have been demonstrated to have a significant impact on the metabolic efficacy of the enzymes in focus in order to evaluate to which extent these polymorphisms might be suitable to predict the therapeutic outcome with tamoxifen.
Patients and methods
Patients
Ninety-nine patients were included in this study. Patients were of Caucasian origin, diagnosed with early-stage ER-positive BC and treated with adjuvant tamoxifen. Patients were recruited at the Breast Center, Zurich, Switzerland between July 2009 and December 2012. Tumors were demonstrated to be ER positive in all patients and 91 % had a >20 % ER expression on the surface. The majority of cases also exhibited PR. Patients were treated with tamoxifen alone or sequentially with aromatase inhibitors (AIs) as the primary treatment choice. In case of a chemotherapeutic treatment (75.8 %), regimens comprised the combinations FEC (5-fluorouracil, epirubicin and cyclophosphamide), FEC-TXT (5-fluorouracil, epirubicin, cyclophosphamide and docetaxel) or EC-TAX (epirubicin, cyclophosphamide and paclitaxel). Data of primary BC diagnosis and disease recurrence were confirmed from the patients’ pathological and medical records. The study was approved by the local ethical committee of Zurich, and written informed consent was obtained. The demographic and disease-associated characteristics of all patients included in the study are shown in Table 1.
Genotyping
From all patients, a whole blood sample was collected in ethylenediaminetetraacetic acid vials. Genomic DNA was extracted using the QIAamp DNA Mini Kit (Qiagen, Hilden, Germany) according to the manufacturer’s recommendations. Extracted DNA was used to genotype all patients for 17 genetic variants within the genes encoding cytochrome P450 enzymes involved in the metabolism of tamoxifen. The genotyping comprised the allelic variants CYP2C9*2 and *3, CYP2C19*2, *3, *4, *6 and *17, CYP2B6*6, *7 and *16, CYP2D6*3, *4, *5, *6, *10 and *41 as well as analysis of CYP2D6 gene duplications/multiplications. The majority of variants were determined using TaqMan SNP assay technology (Agilent Technologies, Santa Clara, CA, USA). The variant CYP2D6*41 was analyzed using a conventional nested PCR approach followed by digestion of the PCR product with the restriction enzyme BsepMI. The forward primer used in the second PCR carried an artificial mutation in the area of the 3′ end, thereby creating a functional restriction site for BsepMI in interplay with the wild type form at the CYP2D6*41 locus. The determination of CYP2D6 duplications/multiplications was performed as described previously [24].
Genotypes of CYP2D6 were assigned to the expected corresponding metabolizer phenotypes according to the evaluation system by Gaedigk et al. [8]. The genotype/phenotype assignments are summarized in Table 2.
CYP2D6 phenotyping and CYP2D6 metabolizer type assignment
Phenotypes predicted on the basis of CYP2D6 genotypes were compared to the CYP2D6 phenotypes measured in a subset of patients with BC. Phenotypic data were obtained by high pressure liquid chromatography (HPLC) according to the methods described by Abdel-Rahman et al. [1] and Blake et al. [3]. HPLC-based phenotype was determined as follows: 6 h after the intake of 25 mg dextromethorphan (DM), a prototypical CYP2D6 substrate, the concentrations of methyl-dextrorphan (MD) and its O-demethylated metabolite dextrorphan (DX) were determined in the urine of 50 patients by reversed-phase HPLC with fluorescence detection. For this purpose, 1.25 ml of each urine sample was deglucuronidated by adding 500 μl phosphate buffer and 25 μl glucuronidase. The samples were incubated at 50 °C, cooled down, and subsequently centrifuged and included in the further analysis. The DM/DX ratios were calculated using the peak height concentrations of DM and DX, determined by HPLC. Patients were considered as CYP2D6 phenotypic poor metabolizer (PM), when exhibiting a DM/DX excretion ratio of >0.3, as described previously [4, 25]. Furthermore, patients were evaluated as CYP2D6 phenotypic intermediate metabolizers (IM), extensive metabolizer (EM) or UM, when showing DM/DX metabolic ratios of 0.03–0.3, 0.0003–(<0.03) and <0.0003, respectively.
Statistical analysis
The chi-square test and the Fisher’s exact test were used to compare categorical variables, such as the dichotomized response measure in relation to the abundance of different genetic variants (e.g., genotype frequencies) or any pathological molecular markers within the patient groups. Continuous data (e.g., age or duration of tamoxifen/AI intake) are given as median ± STD or mean plus time range and were compared using the Mann–Whitney U test. The Kruskal–Wallis test was performed to compare the observed DM/DX ratios in the predicted CYP2D6 phenotype groups.
The RFT was determined in patients who were taking tamoxifen for at least 12 months. The RFT was defined as the time interval in months between the time point of BC diagnosis and the occurrence of the first relapse of disease. A relapse was defined as the occurrence of an event such as either metastatic disease, local or distant recurrence of the tumor or a new invasive primary BC. The RFT was estimated using the Kaplan–Meier method and statistically analyzed using the log-rank test. A two-sided p value of <0.05 was considered statistically significant. The impact of any putative confounders on RFT, such as tumor stage, chemotherapy, menopausal or ER status was tested performing a Cox regression analysis. The program SPSS (version 21, SPSS Inc., Chicago, IL, USA) was used for statistical analysis.
Results
Ninety-nine patients were genotyped for 17 different polymorphisms in the genes encoding the drug-metabolizing enzymes CYP2C9, CYP2C19, CYP2D6 and CYP2B6 using leukocytic DNA. In case of CYP2D6, seven different genetic variants were determined, including gene duplications/multiplications and the most important variants (*3, *4, *5, *6, *10, *41) responsible for an impaired enzyme activity or the expression of null alleles. With the exception of the mutation CYP2C19*6, all variants were in Hardy–Weinberg-equilibrium. The variant CYP2C19*6 appeared only once in homozygous form in the cohort and was therefore calculated to be in Hardy–Weinberg-disequilibrium.
Fifteen patients developed a relapse of BC within the observational period. Table 1 demonstrates and compares the demographic and disease characteristics of the individuals showing a relapse with the patient group that stayed disease-free during the time of observation. In general, no significant differences were observed when comparing both groups with regard to age, disease stage, grade or menopausal status. A relapse was significantly less frequently observed in patients with >20 % ER expression (p < 0.001, chi-square test). The observational period and, consequently, the average time of tamoxifen intake was significantly longer in the relapse-free group (p < 0.001 and p = 0.003, respectively).
Eighty-six patients were successfully genotyped for CYP2D6 duplications/multiplications. In view of the obtained allele combinations for CYP2D6, an assignment of an activity score (AS) was possible in ninety-eight patients ([9], Table 2). As demonstrated in Table 2, no significant differences were detected in the overall frequencies of predicted CYP2D6 phenotypes in relation to the relapse status. Additionally, genotype-based predicted CYP2D6 phenotypes were correlated with the experimentally confirmed phenotype (urinary DM/DX metabolic ratio) in a subgroup of 50 patients, where both the genotypic and phenotypic information was available. As demonstrated in Fig. 1, the metabolic DM/DX ratios differed significantly between the predicted CYP2D6 phenotypes (p = 0.001, Kruskal–Wallis test). The prediction of phenotypes based on genotyping results was especially well reproducible for the discrimination between poor and non-PM (Table 3). A sensitive discrimination between intermediate, extensive and ultrafast metabolizers was, however, only possible to a limited extent (Fig. 1; Table 3).
CYP2D6 phenotypes were predicted on the basis of CYP2D6 activity scores, as calculated from CYP2D6 genotyping results. Predicted CYP2D6 phenotypes were correlated with measured CYP2D6 phenotypes, determined on the basis of urinary DM/DX ratios. Fifty patients were included, where both genotype and phenotype data were available. The plot represents median values with data ranges. The Kruskal–Wallis test detected significant differences in DM/DX ratios between the different AS score groups (p = 0.001)
All 99 patients were included into the analysis investigating, to which extent the genotypes of CYP2C9, CYP2C19, CYP2D6 and CYP2B6 influence the relapse time in patients with BC treated with tamoxifen. In several sub analyses, the ER status, the menopausal state and a concomitant chemotherapeutic treatment were considered additional putative risk factors and were also taken into consideration. In the case of ER status, it was assumed that an expression degree of >20 % would be an important prerequisite for an efficient therapy approach with antiestrogens, such as tamoxifen. The average duration of tamoxifen intake was 30 months (range of 12–77 months) and AIs 10 months (range of 0–64 months). The median follow-up time of patients was 46 months. To analyze the influence of genetic variants detected in CYP2C9, CYP2C19, CYP2B6 and CYP2D6 genes on the BC–RFT, the following approach was adopted. First, the frequencies of variant carriers (heterozygous plus homozygous individuals) were compared in the relapse and relapse-free groups before (Table 4) and after a logistic regression analysis stratifying for ER density, menopausal state, disease grade and stage and an additional chemotherapeutic treatment. Second, the RFTs were analyzed in relation to the investigated genotypes using the Kaplan–Maier method without and with stratification for ER and menopausal status (Cox regression analysis). No significant associations were observed between CYP2C9, CYP2C19 and CYP2B6 genotypes and the risk for a BC relapse before or after stratification for the different putative confounders. The ER state appeared to be an independent risk factor for the development of a disease relapse: 6 of 9 patients with an ER expression below or equal to 20 % had a relapse in comparison to 9 of 90 with an ER expression above 20 % (p < 0.001). As demonstrated in Fig. 2, a trend toward a lower relapse rate was observed for individuals carrying allelic combinations that induce an EM or UM genotype of CYP2D6 (p = 0.19). This trend was repeatedly observed, when stratifying for different risk confounders.
Kaplan–Meier analyses determining and comparing the RFT in months in individuals carrying CYP2D6 slow metabolizer (PMs plus IMs) and fast metabolizer genotypes (EMs plus UMs) (a), CYP2C19 wild type and CYP2C19*17 heterozygous (het) or homozygous (hom) (b), or CYP2B6 wild type and CYP2B6*6 heterozygous (het) or homozygous (hom) (c). Although not significant, a trend toward a more favorable disease outcome was observed for carriers of genotypes coding for CYP2D6 fast metabolizer phenotypes (p = 0.19). The comparisons in case of CYP2C19 and CYP2B6 polymorphisms were not significant (p ns)
Discussion
Several studies could demonstrate that interindividual differences in the activity of CYP2D6 may affect the therapeutic outcome in patients with BC receiving tamoxifen in an adjuvant setting [19, 20]. Besides CYP2D6, other drug-metabolizing enzymes play a role in the in vivo transformation of tamoxifen, but were, in this context, rarely investigated. Therefore, our aim was to study to which extent genetic polymorphisms in CYP2D6, CYP2C9, CYP2C19 and CYP2B6 either alone or in interplay with each other influence the relapse risk under treatment with tamoxifen in a cohort of Caucasian patients with BC.
Although not significant, we could show in concordance with previous studies [19, 20] that genotypes associated with normal or increased CYP2D6 activity lead to a favorable treatment outcome under tamoxifen, when taking the re-occurrence of BC as a primary endpoint. This trend toward lower relapse rates can be explained by the increased production of the potent metabolites 4-OH-tamoxifen and endoxifen. The variant CYP2C19*17 has been associated with a fast metabolizer phenotype of CYP2C19 due to an increased expression of CYP2C19 [2, 22]. CYP2C19*17 may, thus, lead to an augmented production of 4-OH-tamoxifen, which may improve the treatment outcome as reported by Schroth et al. [19]. In our study, however, we did not observe an effect of CYP2C19*17 on treatment outcome, neither when analyzing the impact of the *17 variant on the relapse rate alone nor in combination with fast metabolizer genotypes of CYP2D6.
Different genotypes of CYP2C9, CYP2C19 or CYP2B6 did not significantly alter the risk for a relapse of BC under treatment with tamoxifen, which underlines the stronger impact of CYP2D6 on tamoxifen pharmacokinetics. Of note, we used DNA from peripheral blood mononuclear cells (PBMCs) for our analysis, which more accurately reflects the DNA sequence of CYP genes in the liver, whereas DNA obtained from tumor samples may harbor additional cancer-induced mutations, as has been recently discussed [17].
Although several meta-analyses demonstrated that patients with CYP2D6 EM genotypes/phenotypes as compared to slow metabolizer geno- and phenotypes have a longer disease-free survival, the overall survival appeared not to be significantly influenced by the CYP2D6 metabolizer state in tamoxifen-treated BC patients. Furthermore, these studies provided inconclusive results with regard to potential treatment guidelines for tamoxifen based on CYP2D6 genotyping results. [11, 15, 26].
Instead of assigning four metabolizer phenotypes to genotypes, we tested the application of the CYP2D6 genotyping evaluation system of Gaedigk et al. [9], which weighs the impact of different CYP2D6 genotypes on CYP2D6 enzyme activity by assigning AS to the allelic combinations. By correlating predicted to observed CYP2D6 phenotypes, we could demonstrate that the AS system, if based on comprehensive CYP2D6 genotyping results, appears to be a useful approach to discriminate between slow and EM. Phenotypic poor and IM were especially well predicted. A finer discrimination especially between intermediate, extensive and ultrafast metabolizers is, however, not reliably possible based on this system. Since not all possible genetic variations in CYP2D6 were investigated in our study, misclassification may also arise from the assignment of the *1 allele to individuals who harbor a rare, activity-lowering CYP2D6 mutation. Additionally, CYP2D6 phenotyping by using urinary dextromethorphan-based metabolite ratios may not be the optimal phenotyping tool, because small activity differences cannot be reliably detected [5, 6], which in turn, may also lead to misclassifications.
Besides an estrogen receptor expression below 20 %, no other factor like demographics, menopausal state or chemotherapy regimen used were identified as a risk for a relapse.
Our study includes a limited number of individuals and was performed in a retrospective manner. Both aspects could potentially weaken the associations between genotypes and therapeutic outcome with tamoxifen especially in cases, where a polymorphism might have a weak impact on the BC recurrence risk. The data reported here should be validated in future by the performance of prospective studies assessing the impact of CYP2D6 variants on the therapeutic outcome of BC in large cohorts of tamoxifen-treated BC patients.
Abbreviations
- AS:
-
Activity score
- BC:
-
Breast cancer
- DFS:
-
Disease-free survival
- DM:
-
Dextromethorphan
- DX:
-
Dextrorphan
- ER:
-
Estrogen receptor
- EDTA:
-
Ethylenediaminetetraacetic acid
- EM:
-
Extensive metabolizer
- HPLC:
-
High pressure liquid chromatography
- IM:
-
Intermediate metabolizer
- PM:
-
Poor metabolizer
- PR:
-
Progesterone receptor
- PCR:
-
Polymerase chain reaction
- RFT:
-
Relapse-free time
- STD:
-
Standard deviation
- UM:
-
Ultrafast metabolizer
- 4-OH-tamoxifen:
-
4-Hydroxytamoxifen
References
Abdel-Rahman SM, Leeder JS, Wilson JT, Gaedigk A, Gotschall RR, Medve R, Liao S, Spielberg SP, Kearns GL (2002) Concordance between tramadol and dextromethorphan parent/metabolite ratios: the influence of CYP2D6 and non-CYP2D6 pathways on biotransformation. J Clin Pharmacol 42:24–29
Baldwin RM, Ohlsson S, Pedersen RS, Mwinyi J, Ingelman-Sundberg M, Eliasson E, Bertilsson L (2008) Increased omeprazole metabolism in carriers of the CYP2C19*17 allele; a pharmacokinetic study in healthy volunteers. Br J Clin Pharmacol 65:767–774
Blake MJ, Gaedigk A, Pearce RE, Bomgaars LR, Christensen ML, Stowe C, James LP, Wilson JT, Kearns GL, Leeder JS (2007) Ontogeny of dextromethorphan O- and N-demethylation in the first year of life. Clin Pharmacol Ther 81:510–516
Ducharme J, Abdullah S, Wainer IW (1996) Dextromethorphan as an in vivo probe for the simultaneous determination of CYP2D6 and CYP3A activity. J Chromatogr B Biomed Appl 678:113–128
Frank D, Jaehde U, Fuhr U (2007) Evaluation of probe drugs and pharmacokinetic metrics for CYP2D6 phenotyping. Eur J Clin Pharmacol 63:321–333
Fuhr U, Jetter A, Kirchheiner J (2007) Appropriate phenotyping procedures for drug metabolizing enzymes and transporters in humans and their simultaneous use in the “cocktail” approach. Clin Pharmacol Ther 81:270–283
Furr BJ, Jordan VC (1984) The pharmacology and clinical uses of tamoxifen. Pharmacol Ther 25:127–205
Gaedigk A, Ndjountche L, Divakaran K, Bradford LD, Zineh I, Oberlander TF, Brousseau DC, McCarver DG, Johnson JA, Alander SW, Riggs KW, Leeder JS (2007) Cytochrome P4502D6 (CYP2D6) gene locus heterogeneity: characterization of gene duplication events. Clin Pharmacol Ther 81:242–251
Gaedigk A, Simon SD, Pearce RE, Bradford LD, Kennedy MJ, Leeder JS (2008) The CYP2D6 activity score: translating genotype information into a qualitative measure of phenotype. Clin Pharmacol Ther 83:234–242
Goetz MP, Rae JM, Suman VJ, Safgren SL, Ames MM, Visscher DW, Reynolds C, Couch FJ, Lingle WL, Flockhart DA, Desta Z, Perez EA, Ingle JN (2005) Pharmacogenetics of tamoxifen biotransformation is associated with clinical outcomes of efficacy and hot flashes. J Clin Oncol 23:9312–9318
Lum DW, Perel P, Hingorani AD, Holmes MV (2013) CYP2D6 genotype and tamoxifen response for breast cancer: a systematic review and meta-analysis. PLoS ONE 8:e76648
Lyon E, Foster JG, Palomaki GE, Pratt VM, Reynolds K, Sabato MF, Scott SA, Vitazka P (2012) Laboratory testing of CYP2D6 alleles in relation to tamoxifen therapy. Genet Med 14:990–1000
Murdter TE, Schroth W, Bacchus-Gerybadze L, Winter S, Heinkele G, Simon W, Fasching PA, Fehm T, Eichelbaum M, Schwab M, Brauch H (2011) Activity levels of tamoxifen metabolites at the estrogen receptor and the impact of genetic polymorphisms of phase I and II enzymes on their concentration levels in plasma. Clin Pharmacol Ther 89:708–717
Nakamura Y, Ratain MJ, Cox NJ, McLeod HL, Kroetz DL, Flockhart DA (2012) Re: CYP2D6 genotype and tamoxifen response in postmenopausal women with endocrine-responsive breast cancer: the Breast International Group 1–98 trial. J Natl Cancer Inst 104:1264; author reply 1266–1268
Province MA, Goetz MP, Brauch H, Flockhart DA, Hebert JM, Whaley R, Suman VJ, Schroth W, Winter S, Zembutsu H, Mushiroda T, Newman WG, Michael Lee MT, Ambrosone CB, Beckmann MW, Choi JY, Dieudonne AS, Fasching PA, Ferraldeschi R, Gong L, Haschke-Becher E, Howell A, Jordan LB, Hamann U, Kiyotani K, Krippl P, Lambrechts D, Latif A, Langsenlehner U, Lorizio W, Neven P, Nguyen AT, Park BW, Purdie CA, Quinlan P, Renner W, Schmidt M, Schwab M, Shin JG, Stingl JC, Wegman P, Wingren S, Wu AH, Ziv E, Zirpoli G, Thompson AM, Jordan VC, Nakamura Y, Altman RB, Ames MM, Weinshilboum RM, Eichelbaum M, Ingle JN, Klein TE (2014) CYP2D6 genotype and adjuvant tamoxifen: meta-analysis of heterogeneous study populations. Clin Pharmacol Ther 95(2):216–227
Rae JM, Drury S, Hayes DF, Stearns V, Thibert JN, Haynes BP, Salter J, Sestak I, Cuzick J, Dowsett M (2012) CYP2D6 and UGT2B7 genotype and risk of recurrence in tamoxifen-treated breast cancer patients. J Natl Cancer Inst 104:452–460
Ratain MJ, Nakamura Y, Cox NJ (2013) CYP2D6 genotype and tamoxifen activity: understanding interstudy variability in methodological quality. Clin Pharmacol Ther 94:185–187
Regan MM, Leyland-Jones B, Bouzyk M, Pagani O, Tang W, Kammler R, Dell’orto P, Biasi MO, Thurlimann B, Lyng MB, Ditzel HJ, Neven P, Debled M, Maibach R, Price KN, Gelber RD, Coates AS, Goldhirsch A, Rae JM, Viale G (2012) CYP2D6 genotype and tamoxifen response in postmenopausal women with endocrine-responsive breast cancer: the breast international group 1–98 trial. J Natl Cancer Inst 104:441–451
Schroth W, Antoniadou L, Fritz P, Schwab M, Muerdter T, Zanger UM, Simon W, Eichelbaum M, Brauch H (2007) Breast cancer treatment outcome with adjuvant tamoxifen relative to patient CYP2D6 and CYP2C19 genotypes. J Clin Oncol 25:5187–5193
Schroth W, Goetz MP, Hamann U, Fasching PA, Schmidt M, Winter S, Fritz P, Simon W, Suman VJ, Ames MM, Safgren SL, Kuffel MJ, Ulmer HU, Bolander J, Strick R, Beckmann MW, Koelbl H, Weinshilboum RM, Ingle JN, Eichelbaum M, Schwab M, Brauch H (2009) Association between CYP2D6 polymorphisms and outcomes among women with early stage breast cancer treated with tamoxifen. JAMA 302:1429–1436
Sim SC, Ingelman-Sundberg M (2013) Update on allele nomenclature for human cytochromes P450 and the Human Cytochrome P450 Allele (CYP-allele) Nomenclature Database. Methods Mol Biol 987:251–259
Sim SC, Risinger C, Dahl ML, Aklillu E, Christensen M, Bertilsson L, Ingelman-Sundberg M (2006) A common novel CYP2C19 gene variant causes ultrarapid drug metabolism relevant for the drug response to proton pump inhibitors and antidepressants. Clin Pharmacol Ther 79:103–113
Stanton V, Jr. (2012) Re: CYP2D6 genotype and tamoxifen response in postmenopausal women with endocrine-responsive breast cancer: the Breast International Group 1–98 trial. J Natl Cancer Inst 104:1265–1266; author reply 1266–1268
Steijns LS, Van Der Weide J (1998) Ultrarapid drug metabolism: PCR-based detection of CYP2D6 gene duplication. Clin Chem 44:914–917
Trojan A, Vergopoulos A, Breitenstein U, Tausch C, Seifert B, Joechle W (2013) CYP2D6 phenotype indicative for optimized antiestrogen efficacy associates with outcome in early breast cancer patients. Cancer Chemother Pharmacol 71:301–306
Zeng Z, Liu Y, Liu Z, You J, Chen Z, Wang J, Peng Q, Xie L, Li R, Li S, Qin X (2013) CYP2D6 polymorphisms influence tamoxifen treatment outcomes in breast cancer patients: a meta-analysis. Cancer Chemother Pharmacol 72:287–303
Acknowledgments
This work was supported by the Swiss Tumor Institute, Zurich, Switzerland.
Conflict of interest
All authors declare that there is no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Mwinyi, J., Vokinger, K., Jetter, A. et al. Impact of variable CYP genotypes on breast cancer relapse in patients undergoing adjuvant tamoxifen therapy. Cancer Chemother Pharmacol 73, 1181–1188 (2014). https://doi.org/10.1007/s00280-014-2453-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00280-014-2453-5