Abstract
Although there is effective chemotherapy for many patients with leukemia, 20% of children and up to 65% of adults relapse. Novel therapies are needed to treat these patients. Leukemia cells are very sensitive to the proteasome inhibitor bortezomib (VELCADE®, PS-341), which enhances the in vitro cytotoxic effects of dexamethasone and doxorubicin in multiple myeloma. To determine if bortezomib enhances the cytotoxicity of agents used in leukemia, we employed an in vitro tetrazolium-based colorimetric assay (MTT) to evaluate the cytotoxic effects of bortezomib alone and in combination with dexamethasone, vincristine, doxorubicin, cytarabine, asparaginase, geldanamycin, trichostatin A, and the bcl-2 inhibitor HA14.1. We demonstrated that primary leukemia lymphoblasts and leukemia cell lines are sensitive to bortezomib, with an average IC50 of 12 nM. Qualitative and quantitative bortezomib-drug interactions were evaluated using the universal response surface approach (URSA). Bortezomib was synergistic with dexamethasone in dexamethasone-sensitive leukemia cells, and additive with vincristine, asparaginase, cytarabine, and doxorubicin. The anti-leukemic activity of bortezomib was also additive with geldanamycin and HA14.1, and additive or synergistic with trichostatin A. These results were compared to analysis using the median-dose effect method, which generated complex drug interactions due to differences in dose-response curve sigmoidicities. These data suggest bortezomib could potentiate the cytotoxic effects of combination chemotherapy in patients with leukemia.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Although there have been substantial advances in pediatric leukemia therapy during the past 30 years, approximately 20% of children and up to 65% of adults relapse following initial therapy [1]. Survival for many patients with recurrent leukemia is guarded despite aggressive chemotherapy and stem cell transplant. Relapsed leukemia is often resistant to conventional chemotherapeutic agents and novel therapeutic approaches are needed to improve outcome for these patients.
Bortezomib is an inhibitor of the 26S proteasome, a multi-subunit protease that controls the degradation of short-lived regulatory proteins involved in essential cell processes such as cell cycle regulation, cell differentiation, transcriptional regulation, and apoptosis [2, 3]. Proteasome substrates are targeted for degradation by conjugation with ubiquitin, a highly regulated process controlled by the ubiquitin-conjugating complex [4]. Proteasome inhibition results in apoptosis in many malignant cell types. Although not fully elucidated, the cytotoxic effects of bortezomib may be related to the inhibition of NF-κB [5, 6].
Although bortezomib can induce apoptosis in vitro [7, 8] and in vivo [9–11] in a variety of malignant cell types, non-malignant cells appear resistant to the cytotoxic effects of bortezomib [12–14]. Bortezomib may also potentiate the cytotoxic effects of chemotherapy in a variety of tumor types. Bortezomib enhances the cytotoxicity of dexamethasone, melphalan, and doxorubicin in multiple myeloma [15] and other hematologic malignancies in vitro [16–18]. Bortezomib also enhances the cytotoxic effects of histone-deacetylase (HDAC) inhibitors such as phenyl butyrate and trichostatin A [18–20], as well as bcl-2 inhibitors such as HA14.1 [21]. NF-κB activates the expression of several anti-apoptotic proteins, including members of the bcl-2 family (bcl-XL) and inhibitors of apoptosis (XIAP) [22]; and NF-κB activation can attenuate the pro-apoptotic response to chemotherapy and ionizing radiation [23]. Furthermore, inhibition of NF-κB can sensitize chemoresistant multiple myeloma and Hodgkin’s disease cells to apoptosis [6, 23, 24]. The in vitro effects of bortezomib-drug combinations in leukemia, however, have not been well characterized.
Bortezomib has shown clinical efficacy in the treatment of multiple myeloma and non-Hodgkin’s lymphoma (NHL), as evidenced by a 35% overall response rate in a Phase 3 study of heavily pretreated multiple myeloma patients (SUMMIT trial) [25, 26] and a 55% overall response rate in a phase 2 trial of bortezomib in indolent NHL [27, 28]. Bortezomib may also clinically enhance the cytotoxicity of other chemotherapy agents, such as dexamethasone and doxorubicin [25]. However, the efficacy of bortezomib in combination with cytotoxic chemotherapy agents in leukemia is unknown.
Bortezomib has been shown to induce apoptosis in leukemia cell lines [29, 30] and in nude mice with leukemic xenografts [31]. Prior reports have shown that pre-B and T-cell acute lymphoblastic leukemia (ALL) cell lines are very sensitive to bortezomib, with an average IC50 of 10 nM [29]. In this study we investigated the effects of bortezomib in combination with a variety of agents in leukemic cell lines including: dexamethasone, cytarabine, doxorubicin, vincristine, asparaginase, the heat shock protein (HSP) inhibitor geldanamycin, the histone-deacetylase (HDAC) inhibitor trichostatin A, and the bcl-2 inhibitor HA14.1.
Materials and methods
Chemicals
RPMI-1640 cell culture medium, phosphate-buffered saline (PBS), dextrose, sodium pyruvate, sodium chloride, sodium bicarbonate, HEPES, and gentamicin were purchased from Gibco (Grand Island, NY, USA); fetal calf serum and high-glucose RPMI-1640 cell culture medium were purchased from American Type Culture Collection (ATCC, Rockville, MD, USA); penicillin/streptomycin was purchased from InVitrogen (Carlsbad, CA, USA); Hanks-buffered saline solution (HBSS) was purchased from HyClone (Logan, UT, USA), and Lymphoprep for mononuclear cell isolation was purchased from Greiner Bio-One (Monroe, NC, USA). Asparaginase (Asp), dexamethasone (Dexa), cytarabine (AraC), doxorubicin (Doxo), geldanamycin (Gld), trichostatin A (TsA), vincristine (VCR), methyl-tetrazolium (MTT) and dimethyl sulfoxide (DMSO) were purchased from Sigma-Aldrich (St Louis, MO, USA). HA14.1 was purchased from EMD BioSciences (San Diego, CA, USA).
Bortezomib was provided by Millennium Pharmaceuticals (Cambridge, MA, USA). Stock solutions of bortezomib (0.5 mg/ml) were prepared in ethanol/ascorbic acid. Stock solution of dexamethasone, geldanamycin, trichostatin A, and HA14.were prepared in DMSO. Drug dilutions were freshly prepared from concentrated stock solutions. Drugs were serially diluted over a 106-fold range of concentrations to determine single agent IC50s.
Cell lines
The human T-cell ALL cell lines Jurkat, Molt 4, Molt 3, CEM, and HSB2; the human pre-B ALL cell lines JM1, RS4, and Reh; the human B-cell lymphoblastoid cell line SB; the CML line K562; and the AML cell lines KG1 (erythroleukemia), GDM1 (myelomonocytic leukemia) and THP1 (monocytic leukemia) were purchased from ATCC (Rockville, MD, USA). The multiple myeloma cell line MM1.S was kindly provided by Dr Steven T. Rosen (Northwestern University, Chicago, IL, USA). Reh, SB, JM1, HSB-2 and MM1.S cell lines were cultured in RPMI-1640 supplemented with 2 mM glutamine and 10% fetal calf serum. Molt-4, Molt-3, CEM, RS4, and GDM-1 cells were grown in RPMI-1640 supplemented with 1.5 g/l sodium bicarbonate, 4.5 g/l glucose, 10 mM HEPES, 1.0 mM sodium pyruvate, and 10% fetal calf serum (high-glucose RPMI). THP-1 cells were grown in high-glucose RPMI supplemented with 0.05 mM 2-mercaptoethanol. K562 and KG-1 cells were grown in Iscove’s modified Dulbecco’s medium (IMDM) supplemented with 4 mM l-glutamine, 1.5 g/l sodium bicarbonate, and either 10% (K562) or 20% (KG-1) fetal calf serum. For the MTT cytotoxicity assay (see below) cell lines were maintained at a density of 0.5–1×105 cells/ml and incubated at 37°C in a humidified atmosphere containing 5% CO2.
Culture of primary leukemic cells
Primary leukemia cells from pediatric patients were obtained from peripheral blood, leukapheresis, or bone marrow specimens under an IRB-approved protocol. All samples were obtained from pediatric patients newly diagnosed with leukemia prior to the administration of chemotherapy. Data from four representative patients are shown. The first patient was a 10-year-old male diagnosed with pre-B ALL. Patient 2 was a 17-year-old male diagnosed with AML, M1 subtype. Patient 3 was a 2-year-old male with pre-B ALL. Patient 4 was a 5-month-old male with infant ALL. Lymphoblasts were isolated using Lymphoprep and cultured in RPMI-1640 supplemented with 20% fetal calf serum, penicillin/streptomycin and gentamicin. During the MTT assay, cell viability was confirmed by Trypan blue exclusion at 48 and 72 h and noted to be >90% in the absence of drug. Cells were maintained at a density of 1×106 cells/ml and incubated at 37°C in a humidified atmosphere containing 5% CO2.
Cytotoxicity assay
The growth inhibition effect of bortezomib was assessed using the [3-(4,5-dimethyl-thiazol-2yl)-2,5-diphenyl-tetrazolium bromide] (MTT) colorimetric dye reduction method [32, 33]. One hundred thirty-five μl of exponentially growing lymphoblasts were plated at a density of 0.5×104 to 1×104 cells/well in 80 wells of a 96-well microtiter plate. After 16 h, drug at specified concentrations was added to each well. After 72 h of continuous drug exposure (unless otherwise indicated), 15 μl of MTT (final concentration 0.5 mg/ml) was added to each well and the plates were incubated for 4 h at 37°C. Medium was replaced with 150 μl of DMSO to solubilize the formazan, and the optical density (OD) was measured at 550 nm using a microplate spectrophotometer (Anthos Analytical, Durham, NC, USA). Cell survival was calculated by subtracting the background OD of media alone, then dividing the OD of test wells by the OD of the control (untreated) wells. For single drug assays, replicates of six wells were used for each drug concentration and two replicate plates were performed for each experiment. Primary cell drug concentrations were done in triplicate.
In combination experiments, the two drugs were added simultaneously unless otherwise indicated. In each 96-well plate, cells were exposed to medium alone (controls), single drug alone, or different concentrations of the combination of two drugs, for a total of 36 concentration pairs per plate. Combination effects were analyzed using non-constant drug combination ratios at single drug IC10, IC25, IC75, IC90 and IC100 drug concentrations. All combination drug concentrations were tested in four replicate plates. Each bortezomib-drug combination was tested in at least two separate experiments and in at least two different leukemia cell lines. In sequential combination experiments, cells were pretreated with the one drug (drug A) for 24 h before administration of the second drug (drug B) (total drug A treatment time, 72 h; total drug B treatment time, 48 h).
Analysis of drug effects
Synergy was assessed using the Universal Response Surface Analysis (URSA) approach of Greco et al. [34], which involves a parametric model relating the concentrations of the two drugs to measured cell survival. Initial estimates of the concentration of each drug required to produce 50% inhibition of control cell growth (IC50) and the slope parameter (m) signifying the sigmoidicity of the dose–effect curve were obtained by fitting Eq. 1 to the data from single-drug growth inhibition assays:
where D is the drug concentration, Econ is the cell survival in the absence of drug, and E is the measured effect (cell survival). Estimates of the parameters IC50 and m were obtained using the ADAPT II software [35] (maximum likelihood estimation option, 10% CV error model). Subsequently, the two-drug interaction model in Eq. 2 was implemented in the ADAPT II software and fitted to the data from the combination experiments (maximum likelihood estimation, 10% CV error model):
where Da is the concentration of drug A, Db is the concentration of drug B, Econ is the effect (cell survival) in the absence of drug, and E is the measured effect (cell survival). In Eq. 2, IC50a and IC50b are the median effective drug concentrations of each drug, ma and mb represent the slope parameters of each drug, and α represents the drug interaction parameter (these five parameters were estimated from the cell survival data for each combination experiment). When α is positive, Loewe synergy is indicated; when α is negative, Loewe antagonism is indicated. The interaction is considered additive if the 95% confidence interval (CI) around α encompassed zero.
Median dose effect analysis
In vitro combination effects were also analyzed using the median-dose effect analysis method [36] using a commercially available software program (Calcusyn version 1.2, Biosoft, Ferguson, MO, USA). Each drug interaction is defined using the median effect equation:
where fa is the fraction of cells affected by drug, fu the fraction of cells unaffected by drug, D is the drug concentration, Dm is the IC50, and m is the slope parameter signifying the sigmoidicity of the dose–effect curve. The median effect analysis method defines a combination index (CI) for each two-drug interaction using a fractional product equation;
where Dalone is the dose of each drug alone required for a given effect (fa), Dcomb is the dose of each drug in the combination required for a given effect (fa), and α = 0 if the effects of the two drugs are mutually exclusive (i.e. Dalone1, Dalone2, and the mixture of the two drugs yield parallel lines in the median effect plot [log (fa/fu) vs log (D)]) and α=1 if the effects of the two drugs are mutually non-exclusive (i.e. Dalone1 and Dalone2 yield parallel lines in the median-effect plot but the slope of the two-drug combination is not parallel to the single-drug lines). CI values above 1.1 are considered antagonistic, values between 0.9 and 1.1 are considered additive, and CI values less than 0.9 are considered synergistic. When exclusivity of effects could not be established (i.e. Dalone1 and Dalone2 had non-parallel slopes in the median effect plot) the data were analyzed using the more stringent mutually non-exclusive assumptions. Each drug combination was tested with drug A plus drug B at five constant-ratio combinations (1:1, 1:2, 2:1, 1:4 and 4:1) at the predicted IC50s. Four to six serial dilutions were made at each constant-ratio combination and the combination index analyzed for at least two different fixed ratios as noted. The bortezomib + dexamethasone combination was tested at ratios of 1:1 (equipotent) and 1:2 (mutually non-exclusive equation); bortezomib + doxorubicin at potency ratios of 1:1, 1:2, and 1:4 (mutually exclusive equation); bortezomib + cytarabine at ratios of 1:1, 2:1, and 1:4, (RS4 cell line), 1:1, 1:2, and 2:1 (Reh cell line), or 1:1, 1:2, and 1:4 (Jurkat cell line) (mutually non-exclusive equation). Correlation coefficients for the median effect plots were >0.9.
Results
Bortezomib growth inhibition in cell lines and primary cells
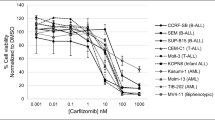
Five T-cell ALL (Fig. 1a) and four pre-B or B-cell ALL cell lines (Fig. 1b) were very sensitive to bortezomib, with an average IC50 of 7.5 nM (range 2–26 nM). AML cell lines were also sensitive to bortezomib with an average IC50 of 19 nM (range 12–23 nM) (Fig. 1c), as were K562 erythroleukemia and MM1.S multiple myeloma cell lines with an with an IC50 of 20 nM and 9 nM, respectively (Fig. 1c). IC50 values were not significantly different following exposure to bortezomib for 48 h or 72 h (data not shown).
Bortezomib in vitro growth inhibition assays. MTT cytotoxicity assays were performed with bortezomib at the concentrations noted. Cell survival is plotted as a function of bortezomib drug concentration. Leukemic cells tested include (a) the T-cell ALL cell lines Jurkat, Molt-4, HSB-2, CEM and Molt3; (b) the pre-B ALL cell lines JM-1, Reh, and RS4, and the B-cell lymphoblastoid cell line SB; (c) the AML cell lines KG-1, GDM1, and THP1; the multiple myeloma cell line MM1.S; and the CML cell line K562; and (d) representative primary leukemic lymphoblasts obtained from pediatric patients with AML and ALL (see Methods for clinical information)
Primary leukemia cells were also sensitive to the cytotoxic effects of bortezomib, with an average IC50 of 23 nM (range 10–30 nM) (Fig. 1d). As with the leukemia cell lines, there was no difference in bortezomib sensitivity with 48 h versus 72 h incubation times (data not shown). Primary ALL and AML cells were sensitive to bortezomib with IC50 values of 10 nM and 30 nM, respectively. Primary infant ALL cells are also sensitive to bortezomib, with an IC50 of 26 nM.
IC50 comparisons of bortezomib and other chemotherapy agents
Figure 2 shows the IC50 for bortezomib and five chemotherapy agents commonly used in childhood leukemia treatment, including cytarabine, dexamethasone, doxorubicin, asparaginase, and vincristine; as well as four other agents, including the pyrimidine antagonist gemcitabine, the heat shock protein (HSP) inhibitor geldanamycin, the histone deacetylase (HDAC) inhibitor trichostatin A, and the bcl-2 inhibitor HA14.1.
Distribution of the IC50 values of several chemotherapy agents as a function of drug and leukemic cell line. Cell lines tested include the T cell ALL cell lines Jurkat, Molt-4, CEM, Molt-3, and HSB-2; the pre-B ALL cell lines Reh, RS4 and JM1, the B-lymphoblastoid cell line SB2, and the AML cell line THP-1. Each data point is an average of at least 2 determinations from separate experiments. Dexa dexamethasone, Doxo doxorubicin, Asp E. coli asparaginase, Trich A histone deacetylase inhibitor trichostatin A, HA14.1 =bcl-2 inhibitor HA14.1
ALL cell lines exhibited variable sensitivity to dexamethasone. Although several cell lines were very resistant to dexamethasone (IC50 > 10 μM), other cell lines were either intermediately sensitive (HSB2; IC50 = 145 nM) or very sensitive (RS4; IC50 = 1.5 nM) to dexamethasone. Similarly, there was marked variability between cell lines in the IC50s for asparaginase (0.4–350 milliunits) and trichostatin A (8 nM–580 nM). Geldanamycin and gemcitabine were relatively potent in ALL cell lines, with average IC50s of 10 nM and 5 nM, respectively. The bcl-2 inhibitor HA14.1, in contrast, was less potent, with an average IC50 of 17 μM. Bortezomib, with an average IC50 of 12 nM (range 2–30 nM), compares favorably in potency to these agents.
Combination effects of bortezomib and other chemotherapy agents
Table 1 and Fig. 3 summarize the results of the combination drug experiments analyzed by the URSA method. Most interactions between bortezomib and other chemotherapy agents were additive; two interactions, however, were synergistic. Bortezomib was synergistic with dexamethasone in the dexamethasone-sensitive pre-B ALL cell line (RS4), with an alpha of 3.2±1.3 (95% confidence interval 1.9–4.5). Synergism was limited to dexamethasone-sensitive cells; bortezomib was additive with dexamethasone in dexamethasone-resistant cell lines (Jurkat, Molt 4) and cell lines with intermediate dexamethasone sensitivity (HSB2). Sequential addition of bortezomib (before or after dexamethasone) did not affect the additive bortezomib-dexamethasone drug interactions (data not shown). However, sequential addition of bortezomib before or after dexamethasone in the dexamethasone-sensitive RS4 cell line changed the synergistic bortezomib-dexamethasone interaction into an additive interaction (data not shown). Thus, bortezomib enhanced dexamethasone-sensitivity in leukemic cells (if given simultaneously), but did not make dexamethasone-resistant cells sensitive to dexamethasone.
Estimates of the synergism-antagonism parameter (α±95% confidence interval) obtained by fitting Eq. 2 to the full data set from each experiment with weighted non-linear regression. Each data point is from a separate 96-well plate growth inhibition assay done in quadruplicate; each experiment was repeated at least twice. The shaded area brackets those interactions demonstrating Loewe additivity. Data points above the grey box demonstrate Loewe synergism. There were no cases of Loewe antagonism
Bortezomib was also synergistic with the HDAC inhibitor TsA in the AML cell line THP1, with an alpha of 5.0+3.2 (95% confidence interval 1.8–8.2). Bortezomib synergism with this HDAC inhibitor was not universal since the bortezomib-TsA combination was additive in the pre-B ALL cell line JM1 and in two other AML cell lines (KG1 and GDM1, data not shown). There were no antagonistic interactions between bortezomib and the chemotherapy agents tested.
We also analyzed bortezomib combinations using the median effect method of Chou and Talalay [36]. Using this modeling method, many of the two-drug interactions involving bortezomib were mixed, with different interactions (additive, synergistic, or antagonistic) noted at different drug concentrations (Fig. 4). For example, the combination index (CI) for the bortezomib-doxorubicin combination (Fig. 4a) was less than 1.0, denoting synergistic interactions for most, but not all, concentrations. Bortezomib/cytarabine combinations (Fig. 4b) also appeared synergistic at high concentrations, but were additive or antagonistic at lower drug concentrations. Bortezomib-dexamethasone interactions in RS4 cells (Fig. 4c), which were synergistic using the URSA method of analysis, were synergistic only at high concentrations using median effect analysis and were additive or antagonistic at lower concentrations.
Median effect analysis: Combination index (CI) as a function of the fraction affected (Fa) for equipotent fixed-ratio combinations of the following agents: (a) bortezomib and doxorubicin (molar ratio 1:0.4) in the pre-B ALL cell line JM1, (b) bortezomib and cytarabine (molar ratio 1:1.5) in the pre-B ALL cell line RS4, and (c) bortezomib and dexamethasone (molar ratio 1:1) in the dexamethasone-sensitive cell line RS4. Potency ratios were determined from single-drug IC50 s; the CI was also calculated on at least one other fixed ratio combination for each drug combination (see methods for details) with similar results (data not shown). CI values above 1.1 are considered antagonistic, values between 0.9 and 1.1 are considered additive, and values less than 0.9 are considered synergistic. Drug concentration increases on the X-axis (i.e. 0.5 represents the IC50 of each drug, 0.9 represents the IC90 of each drug)
Discussion
We have demonstrated that both primary leukemia cells and leukemia cell lines are very sensitive to the proteasome inhibitor bortezomib in vitro, with an average IC50 of 12 nM. Primary leukemic cells isolated from patients with ALL, AML and infant ALL were also very sensitive to bortezomib. Leukemia cells, however, rapidly develop resistance to cytotoxic chemotherapies when used as single agents in vivo and most treatment regimens combine several chemotherapy agents to prevent the development of drug resistance. Thus, if bortezomib is to play a major role in leukemia treatment it is likely to be in the context of combination regimens.
Bortezomib has been a clinically effective drug in multiple myeloma and non-Hodgkin’s lymphoma [26, 27, 37]. It is also undergoing clinical trials in a variety of solid tumors [38, 39]. Bortezomib can enhance the effectiveness of both standard chemotherapeutic agents and biologic molecules in a variety of malignant cell types. Although no drug-interaction modeling was performed, bortezomib appeared to increase the cytotoxic effects of doxorubicin, melphalan, dexamethasone, bcl-2 inhibitors, and HDAC inhibitors in multiple myeloma in vitro [15, 19, 21]. Bortezomib also increased Philadelphia-positive chronic myelogenous leukemia (CML) cell sensitivity to HDAC inhibitors [40, 41] and flavopiridol [18]; increased lymphoma cell sensitivity to dexamethasone [42] and gemcytabine [43]; and increased myelomonocytic leukemic (AML) cell sensitivity to flavopiridol [17]. There have been several reports of bortezomib potentiation of dexamethasone in multiple myeloma both in vitro [13, 44] and in vivo [37]. Synergistic interactions have also been reported between dexamethasone and bortezomib in primary effusion lymphomas [42]. In contrast to previous observations in multiple myeloma, however, our data did not show that bortezomib could make dexamethasone-resistant leukemia cells sensitive to dexamethasone. Although bortezomib was unable to overcome dexamethasone resistance in leukemia cell lines, there have been case reports of the effectiveness of the bortezomib–dexamethasone combination in the treatment of relapsed ALL [45].
In our study, there was also synergy between bortezomib and the HDAC inhibitor trichostatin A in the AML cell line THP1. In vitro synergistic interactions have been previously reported with the combination of bortezomib and HDAC inhibitors in multiple myeloma [19, 41], CML [41], and gastrointestinal adenomas [20]. These data suggest that the combination of bortezomib and HDAC inhibitors may warrant further clinical study.
Drug combination effects with bortezomib in hematologic malignancies have been examined using a variety of analysis methods, including the fractional product method [40, 46], the isobologram method [47], the median dose effect method of Chou and Talalay [18, 21, 41, 42, 48, 49] and the universal response surface approach (URSA) method refined by Greco et al. [34] which we used in this study. There are significant differences between these drug combination analysis methods. Differences in conclusions between the median effect and URSA analysis methods are common. Greco et al. reanalyzed 136 data sets using their parametric model fitting approach and noted that only 38 of the 136 data sets (28%) showed close agreement in the final conclusions between the Chou and Greco analysis [50].
The URSA is an empiric modeling method which uses weighted, parametric, non-linear regressions to simultaneously fit all the experimental data to a fully combined-action model, after which the interaction parameters are estimated [50]. The median effect model, in contrast, is a mechanistic modeling method that employs equations derived from mass action enzyme kinetics [36]. The merits of these two analysis methods have undergone extensive critiques [48, 50–53] and it is not our goal to reanalyze the effectiveness of different drug combination analysis methods. However, there are several important differences between these modeling methods that can explain the different conclusions drawn by the URSA and the median effect modeling in this study.
Both Greco [50] and Berenbaum [51] have noted that the median effect method of Chou and Talalay relies on the assumption that mutually exclusive interactions (i.e. drug interactions occurring at the same binding site) have dose–response curves with equal sigmoidicities (m). However, in complex cell systems, it is often difficult to tell if dose–response curve sigmoidicity accurately reflects binding site avidity [51]. Unequal sigmoidicities could be due to different drug binding characteristics (as proposed by Chou) or could be due to other factors inherent in a complex cell system, such as unequal drug metabolism, altered binding site access, or drug efflux mechanisms [51].
When single-agent dose–response curves have different sigmoidicities (i.e. m1 ≠ m2) mutual exclusivity using the median effect method cannot be established and an alternative mutually non-exclusive analysis is used [48]. However, investigators have pointed out that the equation used to determining mutually non-exclusive interactions overestimates the combination index and incorrectly categorizes interactions that are synergistic as either additive or even antagonistic at low-fractional effects (fa) [50, 51]. We believe this accounts for the “antagonistic” interactions seen in bortezomib/cytarabine and bortezomib/dexamethasone combinations at low fa’s (Table 2 and Fig. 4).
Greco also notes that drugs with unequal dose–response curve sigmoidicities will have a curved median effect plot in which large differences in slope parameters have a profound effects on the curvature of the median effect plot (which should be linear) [50]. Because of this curvature in the median effect plot, the median effect analysis will also overestimate synergy at high-fractional effects. As is evident from Fig. 5, the sigmoidicity of the bortezomib dose–response curve differs from the other agents studied in most cell lines. Because of the limitations of the median effect method in analyzing drugs with different dose–response curve sigmoidicities, we believe the URSA method provides a more robust analysis of bortezomib-drug interactions.
Distribution of the dose-response curve slope parameters (−m) for bortezomib, dexa dexamethasone, asp asparaginase, AraC cytarabine, doxo doxorubicin and the histone-deacetylase inhibitor trichostatin A. Slope parameters for bortezomib ranged from −2 to −5.8; each m value was determined from six data points done in quadruplicate. Large negative slope values indicate a steep dose–response curve
Different analysis methods have been employed in prior studies examining bortezomib/drug combinations. Although several papers noting bortezomib potentiation do not use modeling methods, [13, 15, 43, 54] several others have noted synergistic interactions between bortezomib and other chemotherapy agents (flavopiridol, the bcl-2 inhibitor HA14-1, the HDAC inhibitor suberoylanilide hydroxamic acid (SAHA), and gemcitabine) using the median effect method of Chou and Talalay [18, 21, 41, 42, 42, 49]. As noted above, however, the Chou method can overestimate synergy at high-fractional effects (fa) and the bortezomib drug combination interactions noted in prior reports may in fact be additive if analyzed using the more rigorous URSA method.
Regardless of the analysis method used, bortezomib appears to potentiate the effectiveness of combination chemotherapy in leukemia. The results of our study suggest that bortezomib is at least additive with several chemotherapy agents commonly used in leukemia treatment and may be synergistic with dexamethasone under certain conditions. Bortezomib has non-overlapping toxicities with myelosuppressive agents used to treat adults with hematologic malignancies [11, 55] and a phase 1 trial of bortezomib in pediatric patients with solid tumors showed minimal bortezomib toxicity in children [37, 56] These clinical results, combined with the in vitro combination analysis presented here, suggests that the combination of bortezomib and cytotoxic chemotherapies warrants further clinical investigation in leukemia.
References
SEER Cancer Statistic Review, 1973–1999 Bethesda, MD: National Cancer Institute, 2000;467–482
Hochstrasser M (1995) Ubiquitin, proteasomes, and the regulation of intracellular protein degradation. Curr Opin Cell Biol 7:215–223
Spataro V, Norbury C, Harris AL (1998) The ubiquitin-proteasome pathway in cancer. Br J Cancer 77:448–455
Ciechanover A (1994) The ubiquitin-proteasome proteolytic pathway. Cell 79:13–21
Karin M, Cao Y, Greten FR et al (2002) NF-kappaB in cancer: from innocent bystander to major culprit. Nat Rev Cancer 2:301–310
Hideshima T, Chauhan D, Richardson P et al (2002) NF-kappa B as a therapeutic target in multiple myeloma. J Biol Chem 277:16639–16647
Shinohara K, Tomioka M, Nakano H et al (1996) Apoptosis induction resulting from proteasome inhibition. Biochem J 317(Pt 2):385–388
Drexler HC (1997) Activation of the cell death program by inhibition of proteasome function. Proc Natl Acad.Sci USA 94:855–860
Adams J, Palombella VJ, Sausville EA et al (1999) Proteasome inhibitors: a novel class of potent and effective antitumor agents. Cancer Res 59:2615–2622
Papandreou CN, Daliani DD, Nix D et al (2004) Phase I trial of the proteasome inhibitor bortezomib in patients with advanced solid tumors with observations in androgen-independent prostate cancer. J Clin Oncol 22:2108–2121
Orlowski RZ, Stinchcombe TE, Mitchell BS et al (2002) Phase I trial of the proteasome inhibitor PS-341 in patients with refractory hematologic malignancies. J Clin Oncol 20:4420–4427
Masdehors P, Merle-Beral H, Magdelenat H et al (2000) Ubiquitin-proteasome system and increased sensitivity of B-CLL lymphocytes to apoptotic death activation. Leuk Lymphoma 38:499–504
Hideshima T, Richardson P, Chauhan D et al (2001) The proteasome inhibitor PS-341 inhibits growth, induces apoptosis, and overcomes drug resistance in human multiple myeloma cells. Cancer Res 61:3071–3076
Soligo D, Servida F, Delia D et al (2001) The apoptogenic response of human myeloid leukaemia cell lines and of normal and malignant haematopoietic progenitor cells to the proteasome inhibitor PSI. Br J Haematol 113:126–135
Mitsiades N, Mitsiades CS, Richardson PG et al (2002) The proteasome inhibitor PS-341 potentiates sensitivity of multiple myeloma cells to conventional chemotherapeutic agents: therapeutic applications. Blood 101:2377–2380
Pajonk F, Pajonk K, McBride WH (2000) Apoptosis and radiosensitization of Hodgkin cells by proteasome inhibition. Int J Radiat Oncol Biol Phys 47:1025–1032
Dai Y, Rahmani M, Grant S (2003) Proteasome inhibitors potentiate leukemic cell apoptosis induced by the cyclin-dependent kinase inhibitor flavopiridol through a SAPK/JNK- and NF-kappaB-dependent process. Oncogene 22:7108–7122
Dai Y, Rahmani M, Pei XY et al (2004) Bortezomib and flavopiridol interact synergistically to induce apoptosis in chronic myeloid leukemia cells resistant to imatinib mesylate through both Bcr/Abl-dependent and -independent mechanisms. Blood 104:509–518
Mitsiades CS, Mitsiades NS, McMullan CJ et al (2004) Transcriptional signature of histone deacetylase inhibition in multiple myeloma: biological and clinical implications. Proc Natl Acad Sci USA 101:540–545
Adachi M, Zhang Y, Zhao X et al (2004) Synergistic effect of histone deacetylase inhibitors FK228 and m-carboxycinnamic acid bis-hydroxamide with proteasome inhibitors PSI and PS-341 against gastrointestinal adenocarcinoma cells. Clin Cancer Res 10:3853–3862
Pei XY, Dai Y, Grant S (2003) The proteasome inhibitor bortezomib promotes mitochondrial injury and apoptosis induced by the small molecule Bcl-2 inhibitor HA14-1 in multiple myeloma cells. Leukemia 17:2036–2045
Wang CY, Mayo MW, Korneluk RG et al (1998) NF-kappaB antiapoptosis: induction of TRAF1 and TRAF2 and c-IAP1 and c-IAP2 to suppress caspase-8 activation. Science 281:1680–1683
Wang CY, Cusack JC Jr, Liu R et al (1999) Control of inducible chemoresistance: enhanced anti-tumor therapy through increased apoptosis by inhibition of NF-kappaB. Nature Medicine 5:412–417
Izban KF, Ergin M, Huang Q et al (2001) Characterization of NF-kappaB expression in Hodgkin’s disease: inhibition of constitutively expressed NF-kappaB results in spontaneous caspase-independent apoptosis in Hodgkin and Reed-Sternberg cells. Mod.Pathol 14:297–310
Richardson PG, Hideshima T, Mitsiades C et al (2004) Proteasome inhibition in hematologic malignancies. Ann Med 36:304–314
Richardson PG, Sonneveld P, Schuster MW et al (2005) Bortezomib or high-dose dexamethasone for relapsed multiple myeloma. N Engl J Med 352:2487–2498
O’Connor OA, Wright J, Moskowitz C et al (2005) Phase II clinical experience with the novel proteasome inhibitor bortezomib in patients with indolent non-Hodgkin’s lymphoma and mantle cell lymphoma. J Clin Oncol 23:676–684
Goy A, Younes A, McLaughlin P et al (2005) Phase II study of proteasome inhibitor bortezomib in relapsed or refractory B-cell non-Hodgkin’s lymphoma. J Clin Oncol 23:667–675
An WG, Hwang SG, Trepel JB et al (2000) Protease inhibitor-induced apoptosis: accumulation of wt p53, p21WAF1/CIP1, and induction of apoptosis are independent markers of proteasome inhibition. Leukemia 14:1276–1283
Yu C, Rahmani M, Dent P et al (2004) The hierarchical relationship between MAPK signaling and ROS generation in human leukemia cells undergoing apoptosis in response to the proteasome inhibitor Bortezomib. Exp Cell Res 295:555–566
Tan C, Waldmann TA (2002) Proteasome inhibitor PS-341, a potential therapeutic agent for adult T- cell leukemia. Cancer Res 62:1083–1086
Mosmann T (1983) Rapid colorimetric assay for cellular growth and survival: application to proliferation and cytotoxicity assays. J Immunol Methods 65:55–63
Carmichael J, DeGraff WG, Gazdar AF et al (1987) Evaluation of a tetrazolium-based semiautomated colorimetric assay: assessment of chemosensitivity testing. Cancer Res 47:936–942
Greco WR, Park HS, Rustum YM (1990) Application of a new approach for the quantitation of drug synergism to the combination of cis-diamminedichloroplatinum and 1-beta-D-arabinofuranosylcytosine. Cancer Res 50:5318–5327
DiArgenio DZ, Schumitsky A (1997)ADAPT II Users guide: pharmacokinetic/pharmacodynamic systems analysis software. (4). Los Angeles, CA, Biomedical systems resource
Chou T, Talalay C (1984) Quantitative analysis of dose effect relationship: the combined effects of multiple drugs or enzyme inhibitors. Adv Enzyme Regul 22:27–55
Richardson PG, Barlogie B, Berenson J et al (2003) A phase 2 study of bortezomib in relapsed, refractory myeloma. N Engl J Med 348:2609–2617
Appleman LJ, Ryan DP, Clark JW, Eder JP, Fishman M, Cusack JC Jr, Fidias P, Supko JG, Guerciolini R, Esseltine D, Kashala O (2003) Phase I dose escalation study of bortezomib and gemcitabine safety and tolerability in patients with advanced solid tumors. Proc ASCO 22:A839
Cusack JC (2003) Rationale for the treatment of solid tumors with the proteasome inhibitor bortezomib. Cancer Treat Rev 29(Suppl 1):21–31
Gatto S, Scappini B, Pham L et al (2003) The proteasome inhibitor PS-341 inhibits growth and induces apoptosis in B positive cell lines sensitive and resistant to imatinib mesylate. Haematologica 88:853–863
Yu C, Rahmani M, Conrad D et al (2003) The proteasome inhibitor bortezomib interacts synergistically with histone deacetylase inhibitors to induce apoptosis in Bcr/Abl+ cells sensitive and resistant to STI571. Blood 102:3765–3774
An J, Sun Y, Fisher M et al (2004) Antitumor effects of bortezomib (PS-341) on primary effusion lymphomas. Leukemia 18:1699–1704
Zheng B, Georgakis GV, Li Y et al (2004) Induction of cell cycle arrest and apoptosis by the proteasome inhibitor PS-341 in Hodgkin disease cell lines is independent of inhibitor of nuclear factor-kappaB mutations or activation of the CD30, CD40, and RANK receptors. Clin Cancer Res 10:3207–3215
Chauhan D, Li G, Auclair D et al (2004) 2-Methoxyestardiol and bortezomib/proteasome-inhibitor overcome dexamethasone-resistance in multiple myeloma cells by modulating Heat Shock Protein-27. Apoptosis 9:149–155
Brown RE, Bostrom B, Zhang PL (2004) Morphoproteomics and bortezomib/dexamethasone-induced response in relapsed acute lymphoblastic leukemia. Ann.Clin Lab Sci 34:203–205
Webb JL (1963) effect of more than one inhibitor. Enzymes and metabolic inhibitors ed Vol 1; New York: Academic Press, 66-79-487-512
Faessel HM, Slocum HK, Jackson RC et al (1998) Super in vitro synergy between inhibitors of dihydrofolate reductase and inhibitors of other folate-requiring enzymes: the critical role of polyglutamylation. Cancer Res 58:3036–3050
Chou JH (1991) Quantitation of synergism and antagonism of two or more drugs by computerized analysis. Synergism and antagonism in chemotherapy pp 223–241
Pei XY, Dai Y, Grant S (2004) Synergistic induction of oxidative injury and apoptosis in human multiple myeloma cells by the proteasome inhibitor bortezmib and histone deacetylase inhibitors. Clin Cancer Res 10:3839–3852
Greco WR, Bravo G, Parsons JC (1995) The search for synergy: a critical review from a response surface perspective. Pharmacol Rev 47:331–385
Berenbaum MC (1989) What is synergy? Pharmacol Rev 41:93–141
Zoli W, Ricotti L, Tesei A et al (2001) In vitro preclinical models for a rational design of chemotherapy combinations in human tumors. Crit Rev Oncol Hematol 37:69–82
Chang TT, Chou TC (2000) Rational approach to the clinical protocol design for drug combinations: a review. Acta Paediatr Taiwan 41:294–302
Bonvini P, Dalla RH, Vignes N et al (2004) Ubiquitination and proteasomal degradation of nucleophosmin-anaplastic lymphoma kinase induced by 17-allylamino-demethoxygeldanamycin: role of the co-chaperone carboxyl heat shock protein 70-interacting protein. Cancer Res 64:3256–3264
Richardson PG, Hideshima T, Mitsiades C et al (2004) Proteasome inhibition in hematologic malignancies. Ann Med 36:304–314
Blaney SM, Bernstein M, Neville K et al (2004) Phase I study of the proteasome inhibitor bortezomib in pediatric patients with refractory solid tumors: a Children’s Oncology Group study (ADVL0015). J Clin Oncol 22:4752–4757
Chou TC, Talalay P (1984) Quantitative analysis of dose-effect relationships: the combined effects of multiple drugs or enzyme inhibitors. Adv Enzyme Regul 22:27–55
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Horton, T.M., Gannavarapu, A., Blaney, S.M. et al. Bortezomib interactions with chemotherapy agents in acute leukemia in vitro. Cancer Chemother Pharmacol 58, 13–23 (2006). https://doi.org/10.1007/s00280-005-0135-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00280-005-0135-z