Abstract
Purpose
To compare a previous model-based image reconstruction (MBIR) with a newly developed deep learning (DL)-based image reconstruction for providing improved signal-to-noise ratio (SNR) in high through-plane resolution (1 mm) T2-weighted spin-echo (T2SE) prostate MRI.
Methods
Large-area contrast and high-contrast spatial resolution of the reconstruction methods were assessed quantitatively in experimental phantom studies. The methods were next evaluated radiologically in 17 subjects at 3.0 Tesla for whom prostate MRI was clinically indicated. For each subject, the axial T2SE raw data were directed to MBIR and to the DL reconstruction at three vendor-provided levels: (L)ow, (M)edium, and (H)igh. Thin-slice images from the four reconstructions were compared using evaluation criteria related to SNR, sharpness, contrast fidelity, and reviewer preference. Results were compared using the Wilcoxon signed-rank test using Bonferroni correction, and inter-reader comparisons were done using the Cohen and Krippendorf tests.
Results
Baseline contrast and resolution in phantom studies were equivalent for all four reconstruction pathways as desired. In vivo, all three DL levels (L, M, H) provided improved SNR versus MBIR. For virtually, all other evaluation criteria DL L and M were superior to MBIR. DL L and M were evaluated as superior to DL H in fidelity of contrast. For 44 of the 51 evaluations, the DL M reconstruction was preferred.
Conclusion
The deep learning reconstruction method provides significant SNR improvement in thin-slice (1 mm) T2SE images of the prostate while retaining image contrast. However, if taken to too high a level (DL High), both radiological sharpness and fidelity of contrast diminish.
Graphical abstract

Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
T2-weighted spin-echo (T2SE) imaging provides morphological information with excellent soft tissue contrast and spatial resolution and is one of the most commonly used pulse sequences in MRI, generally performed with a two-dimensional (2D) multislice acquisition [1] and a fast-spin-echo readout [2]. Although dependent on the anatomic region, the slice thickness is generally no smaller than 2–3 mm and is typically 3—8 × coarser than the in-plane resolution. Although alternatives have been studied for improved (≤ 2 mm) through-plane resolution with T2-weighted contrast, e.g., three-dimensional (3D) T2SE [3,4,5,6,7,8], or 3D T2-prepared SSFP [9, 10] with centric encoding [11], these in general have not replaced the 2D multislice T2SE technique.
T2SE slices thinner than the typical 3 mm can possibly be useful in prostate MRI. The improved lesion contrast and edge sharpness provided by reduced through-plane averaging could improve imaging of the conically shaped apex and better identify heterogeneity and encapsulation of typical nodules within the transition zone, improving the confidence in assessing extraprostatic extension [12], or in more accurately measuring volume such as used for PSA density [13]. T2SE images are usually used to outline prostate boundary and 3D lesion region of interest for a target biopsy using MRI and ultrasound fusion software. In a prospective study done at 1.5 T of 183 men with clinically low-risk prostate cancer, sensitivity in detecting small (defined as ≥ 0.5 cm3 but < 1 cm3) Gleason score 6 prostate cancerous lesions was significantly lower than for larger (≥ 1 cm3) lesions [14].
An alternative to the above 3D approaches for obtaining finer resolution in the slice select (Z) direction is to use 2D slices which have slice-to-slice overlap along Z. The reconstruction uses the overlapped slice data to account for the blur imposed by the slice profile [15]. This can be considered one approach within a class of “super-resolution” techniques which use multiple low resolution image sets to form a high-resolution set. Although studied in spin-echo multislice acquisition using data from multiple orientations [16,17,18,19], more recently investigators have developed specialized 2D data acquisitions from only one orientation to generate high through-plane resolution images along that orientation [20, 21].
As slice thickness is reduced in MRI, in general the signal-to-noise ratio (SNR) is also reduced. Resolution restoration in the slice direction using slice profile deblurring was done in the above-described overlapped slice acquisition cases [20, 21] by using deconvolution with Tikhonov regularization to control the degree of resolution recovery with SNR loss. To seek a more favorable resolution versus SNR operating point, an alternative, model-based image reconstruction (MBIR) was developed [22]. Although it provided improved SNR for thin-slice T2SE, we questioned if even further SNR improvement was possible. In this work, we studied whether a commercial deep learning (DL)-based algorithm could be applied in reconstructing the acquired overlapped 3-mm-thick slices for improved SNR prior to performing the original Tikhonov regularization reconstruction in generating the final 1-mm-thick slices.
Deep learning methods have previously been studied in the context of T2SE prostate MRI [23,24,25,26,27] for abutting, standard, 3-mm-thick slices. Rather than attempting to improve the current 3-mm acquisition, we sought to show the applicability of DL in generating high-quality images of thinner slices. Moreover, by reconstructing each data set using multiple levels of DL we illustrate the tradeoffs of SNR, sharpness and contrast fidelity in determining an operating point. We next describe the reconstruction algorithms and their evaluation in phantoms and in in vivo studies of T2SE prostate MRI.
Materials and methods
Reconstruction algorithms
Figure 1 shows a flow diagram of the various reconstruction methods evaluated. For each study, all reconstruction methods used as input the same data set, consisting of the raw k-space data from ≈80 3-mm thick slices with 1-mm slice-to-slice increment (2 mm slice-to-slice overlap). For the original method [21, 28], referred to in this work as “standard,” the raw data were reconstructed into images corresponding to overlapped 3-mm-thick slices and then filtered along the slice (Z) direction using the Tikhonov regularization (blue box at the top of Fig. 1). With the MBIR technique, the set of reconstructed images is passed to the MBIR algorithm described in [22], resulting in the image set identified as the combined blue and purple boxes. Briefly, the method is based on wavelet regularization along all three dimensions. Reconstruction time using auxiliary computation hardware is about three hours per T2SE image set [22]. The basis of the DL method studied here is described in [29]. It uses a convolutional network trained with supervised learning. For this work, the acquired raw data are passed to the vendor-provided DL image reconstruction software (AIR™ Recon DL GE Healthcare, Waukesha WI USA), designated by the bold arrow in Fig. 1, and the corresponding images are reconstructed using the three vendor-provided (L)ow, (M)edium, and (H)igh levels of DL. The resultant overlapped slices for each DL level are then directed to the same Tikhonov regularization method filtering process described in [21, 28] to unblur the 3-mm-wide slice profile in generating the final 1-mm-thick slices. Processing time using the vendor MRI computation hardware is about one minute per DL level. Reconstruction at all three levels results in the three data sets identified by the purple-only boxes in Fig. 1.
Description of data acquisition
The reconstruction methods were evaluated in phantom and human studies, all using the same T2SE acquisition technique. Data acquisition was done on a 3.0 Tesla whole-body research MRI scanner (Signa Premier™, GE Healthcare, Waukesha WI, USA). Data acquisition used the parameters for the super resolution T2SE prostate MRI study of Ref. [21], shown in Table 1. The sequence parameters TR, TE, and in-plane resolution approximately match the recommendations of Prostate Imaging-Reporting and Data Systems (PI-RADS) v2.1 [30]. In this work, the acquired slice thickness was 3 mm with a 1-mm slice-to-slice increment, or equivalently a 2-mm slice-to-slice overlap. The nominal acquisition orientation was axial, but the slice select direction was adjusted on a patient-specific basis to be aligned with the interface between the posterior aspect of the prostate capsule and the anterior rectal wall, as determined from sagittal localizer images. This tended to keep the prostate within the central region of the full set of axial slices. The number of slices was chosen to cover from the superior aspect of the seminal vesicles to inferior to the prostate apex, ranging from 78 to 102 slices for the patient group studied. Acquisition time ranged from 6:00 to 7:18 with mean 6:22. For reference, a conventional T2SE sequence with abutting 3 mm slices was also used which had identical parameters to those in Table 1, except that the typical number of slices was 30, NEX was doubled to 4, and the typical scan time was 4:40.
Phantom studies
The methods were compared for technical equivalence with respect to large-area contrast and high-contrast spatial resolution. Contrast was evaluated using the standardized NIST contrast phantom [31] which contains 19 mm diameter spherical vials with known T1 and T2 relaxation times. Spatial resolution was assessed using a phantom composed of polylactic acid (PLA) constructed using a 3D printer with vanes having spatial resolution ranging from 0.80 to 3.28 mm [22]. Replicates of the phantom were placed into a water bath at three orthogonal orientations, allowing simultaneous resolution measurement in all three directions in one scan. The contrast and resolution phantoms were each imaged using the pulse sequence of Table 1, and the acquired data sets were directed to the standard as well as the MBIR and DL L, M, and H reconstruction pathways of Fig. 1. Contrast was assessed using mean signal values within each vial expressed as a percentage of mean signal in the background water. The resolution phantom was also imaged using the conventional, abutting slice T2SE sequence. Resolution was assessed by direct observation of resolvable vanes in the phantom images.
Human studies
The algorithms were evaluated in 17 human studies, all men for whom prostate MRI was clinically indicated, all done under an IRB-approved protocol with all subjects giving written informed consent. The image data for this work was acquired for each subject in a separate research-specific examination without intravenous contrast and comprised primarily of the special purpose overlapped slice T2SE sequence of Table 1 and the similar, conventional T2SE acquisition using abutting 3-mm-thick slices. The research examination was done on a research MRI scanner within one day of the clinical MRI examination. Anti-spasmodic drugs were not used for the research-specific examination, but all subjects were instructed to fast for at least three hours before the MRI examination and to empty the rectum and void immediately prior to the examination. The 17 examinations were taken from 21 consecutive research examinations performed from March 2021 to September 2022; results from four studies were not evaluated due to patient motion. Demographics of the 17 participants are shown in Table 2 reflecting the approximate mix of our clinical practice.
Radiological evaluation
Results from the 17 human studies were evaluated independently on a thin client workstation (AW Server 3.2 Ext 3.0, GE Healthcare, Waukesha WI USA) by three fellowship-trained uroradiologists, each with over 10 years of experience in prostate MRI. No clinical information was provided, and the reviewers solely evaluated the reconstructed thin-slice T2SE images. For each study, the acquired raw data were converted into four series of 1-mm-thick T2SE slices using the four reconstruction pathways, the endpoints of which are identified by the purple boxes in Fig. 1 (MBIR, and DL L, M, H). Because it had previously been compared with MBIR and shown to be inferior in performance [22], the standard Tikhonov regularization reconstruction was not formally evaluated in this work. All four series evaluated for a study were presented to the evaluating radiologist simultaneously on a work station. All series for a study were normalized such that the ranges of reconstructed signal values were matched, allowing the same window and level to be used in comparing series. Any identification of the type of reconstruction pathway used was removed from each series, and the ordering of the four reconstruction pathways within each study was randomized.
Each series was evaluated using the following subjective evaluation criteria: (i) SNR within the prostate; (ii) SNR in structures outside the prostate; (iii) sharpness of the peripheral zone (PZ) to transition zone (TZ) interface; (iv) sharpness of the prostate wall / capsule; (v) sharpness of seminal vesicles and bladder and rectal walls; (vi) sharpness of any structures (nodules, cystic changes, areas of fibrosis) within the PZ; (vii) sharpness of any structures within the TZ; (viii) artifact; and (ix) overall image quality. These criteria were graded for each series using scoring (0: inadequate for diagnosis; 1: adequate for diagnosis; 2: more than adequate for diagnosis; 3: exceptional). An additional evaluation criterion was (x) fidelity of contrast graded according to (0: inadequate for diagnosis; 1: severe contrast loss or patchiness; 2: moderate contrast loss or patchiness; 3: negligible contrast loss or patchiness). Finally, each reviewer was asked to indicate his/her preferred series of the four generated for each study. For each measurement, the results for the three reviewers were averaged [32, 33], and then, scores for each series were compared pairwise, with significance evaluated using the two-sided Wilcoxon signed-rank test. Due to the four reconstruction pathways being compared pairwise, the Bonferroni correction was used [34], and significance was taken as p < 0.01 rather than 0.05. Reader-to-reader scores were compared using Cohen’s kappa [35] and multi-reader scores with Krippendorf’s alpha [36].
Results
With respect to contrast, five of the vials within the phantom have T2 values which span those relevant to prostate T2SE (46.4, 64.0, 96.8, 133, 190 ms) [37,38,39]. For each vial, the measurements across the five reconstruction pathways were indistinguishable, indicating that all pathways preserve large-area signals and thus the differences (contrasts) between them (see Supplemental Figure 1 which plots contrast versus vial number; standard deviations are well under 1% in all cases). For in-plane (phase and frequency) spatial resolution, there was no difference among results from any of the four pathways of Fig. 1 or the reference, abutting-slice T2SE. Results from MBIR and DL H (which is most prone to blurring) are shown in Fig. 2A, B and clearly demonstrate the 0.8-mm pattern in both the phase and frequency in-plane directions, consistent with the acquired resolution. In the slice direction, the MBIR (C), DL L (D), DL M (E), and DL H (F), all show modulation at 1 mm (e.g., black arrow, (D)). Note that 3-mm abutting slices when reformatted lose through-slice resolution just short of the 2.6-mm pattern as expected (G).
A Image of the resolution phantom demonstrating in-plane resolution in the phase (P) and frequency (F) directions for the MBIR acquisition method. Specific resolution values are indicated as known from the fiducial notches in the phantom. B Demonstration of in-plane resolution for DL H reconstruction pathway. Demonstration of resolution in the slice (S) direction for C MBIR, D DL L, E DL M, and F DL H reconstruction pathways. All demonstrate some signal modulation at 1.0 mm (e.g., black arrow (D)). G Resolution in the slice direction for abutting 3-mm-thick slices deteriorates somewhat coarser than the 2.6-mm fiducial pattern
Results for Subject #1 (see Supplemental Video 1) present the image series for the four reconstruction pathways evaluated radiologically as well as the series for the standard reconstruction (Tikhonov regularization) which was not evaluated as part of this work. Improvement in visual SNR is apparent, starting from standard reconstruction and then increasing progressively in MBIR and then DL L, M, and H.
Figures 3 and 4 show results of the radiological evaluation. Within each figure, the results for each reconstruction pathway are shown as a stacked plot. The color of a band within a stack indicates the score and the band height the number of times that score was assigned. The aggregate height in all cases is 17, equal to the number of studies. Red scores are unfavorable; green scores are favorable. Due to averaging three reviewer scores, average scores presented in the figures are quantized to 0.33. Any statistical significance between performance of two reconstruction pathways is indicated by a horizontal black line, with the arrowhead designating which pathway has the superior performance. For example, for SNR within the prostate (Fig. 3A), all three DL reconstructions were evaluated as superior to MBIR. Figure 3 presents the evaluations of SNR, artifact, and contrast fidelity. Figure 4 shows evaluation results for the five evaluation criteria related to sharpness.
Results of radiological evaluation for A SNR within the prostate, B SNR of structures outside the prostate, C fidelity of contrast, and D artifact for the four reconstruction pathways evaluated. For each pathway, there are 51 evaluations. Black horizontal arrows indicate statistical significance (p < 0.01) with the arrowhead indicating the superior pathway. For additional detail, see the text
Figure 5 shows the results of evaluation of overall diagnostic quality (A) and a summary of radiologist preference (B). As seen in (B), the DL M reconstruction pathway was preferred in 44 of the 51 evaluations (= 17 evaluations × three reviewers) and the MBIR reconstruction in none.
Krippendorf alpha values for the two aggregate SNR criteria, overall image quality, fidelity of contrast, and the five aggregate sharpness criteria were 0.23, 0.21, 0.35, and 0.02, the first three showing fair and the last slight agreement. Matrices comparing the three possible reader-to-reader comparison scoring for SNR, overall diagnostic quality, sharpness, fidelity of contrast, and associated κ-values are shown in Supplemental Figures 2-5.
Representative images from the radiological evaluation are presented in Figs. 6, 7 and 8. Accompanying the images from the single slice in Fig. 7 is a video showing full series for the DL M reconstructions (left) and the reference 3-mm abutting T2SE acquisition (right) (see Supplemental Video 2). The two series are spatially aligned, i.e., the DL M series generally increments three slices per one slice of the conventional T2SE series. Similarly, accompanying the still images of Fig. 8 is a video comparing the series from DL M and conventional abutting slice T2SE (see Supplemental Video 3).
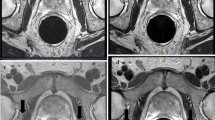
Images from Subject #4 reconstructed using A MBIR and B–D DL L-M-H. SNR of DL L-M-H was evaluated as superior to that of MBIR. Sharpness of the transition zone structures (e.g., dark nodule, as indicated with a yellow arrow in (A)), was evaluated as superior in DL M (C) versus others. Fidelity of contrast was evaluated as inferior in DL H (D) versus that of DL L (B) and DL M (C). The DL M series (C) was preferred by all three reviewers
Images from Subject #16 reconstructed using A MBIR and B–D DL L-M-H. SNR of DL L, M, and H was evaluated as superior to that of MBIR. Sharpness of peripheral zone structures (e.g., lobulated hypointense lesion, arrow, (A)) was rated higher in DL L, M, and H versus MBIR. Also shown in (E) is the best matching slice from the conventional, 3-mm abutting slice T2SE sequence. Side-by-side comparison of image series of 1 mm DL M (C) and 3-mm abutting slice T2SE (E) is available (see Supplemental Video 2)
Images from Subject #14 reconstructed using A MBIR, B–D DL L-M-H, and E conventional 3-mm abutting slice, all at the level of the apex. Hypointense lesion in peripheral zone (arrow, (A)) is seen in all images. SNR and Overall Diagnostic Quality of DL L, M, and H were evaluated as superior to those of MBIR. Ability of thin-slice reconstruction to visualize the lesion in multiple 1-mm slices though the suspect lesion in right apex posterior peripheral zone can be appreciated in comparison image series of DL M versus conventional 3-mm abutting slices (see Supplemental Video 3)
Discussion
We have shown how a deep learning (DL)-based MR image reconstruction algorithm can provide significant improvement in apparent signal-to-noise ratio (SNR) compared to a model-based image reconstruction (MBIR) which was previously evaluated and shown to be superior in apparent SNR to standard image reconstruction. These methods were evaluated in the context of thin-slice (approximating 1 mm thick) T2-weighted spin-echo (T2SE) images of the prostate, an application which is typically severely limited by SNR and thus one in which SNR improvement can be valuable. The three vendor-provided levels of DL were evaluated (L, M, H). All three levels provided visual SNR superior to that of MBIR.
In addition to SNR, the DL L and M reconstruction pathways provided superior performance to MBIR for fidelity of contrast, level of artifact, overall diagnostic quality, and four of the five criteria related to sharpness relevant to prostate MRI.
As is the case with a compressive sensing, wavelet-based, or deep learning algorithms in general, as the level of apparent noise reduction increases, the image sharpness and contrast can be disrupted. Thus, although the DL H pathway provided superior apparent SNR to MBIR and to DL L in the region outside the prostate (Fig. 3B), it was evaluated as inferior in sharpness to DL L and M for multiple sharpness criteria (e.g., structures in the transition and peripheral zones). Also, perhaps more importantly DL H was evaluated as having significantly inferior fidelity of contrast compared to all DL L and DL M (Fig. 3C).
The contrast and resolution measurements in phantoms showed indiscernible differences among various reconstruction pathways. This is to be expected and desirable in showing that no method had an intrinsic bias. For relatively large objects, in this case the 19-mm-diameter vials of the contrast phantom, one typically wants any reconstruction to faithfully preserve mean overall signal levels and thus their differences, i.e., contrast. Similarly, for the high-contrast resolution phantom, for all reconstruction pathways the intrinsic in-plane resolution of the acquisition (FOV/number of samples) was preserved, and the resolvable through-plane resolution approached 1.0 mm, markedly superior to that of 3-mm abutting slices. It was in situations of smaller objects or smaller signal differences such as encountered clinically that differences in performance of the reconstruction pathways were observed.
The presence of a horizontal arrow in Figs. 3, 4 and 5 indicates a statistically significant performance difference using p < 0.01. Further consideration of the inter-reader matrices can elucidate some of these. For SNR (Supplemental Figure 2A-C), the κ-values corresponding to the three matrices all indicate fair inter-reader agreement, further emphasizing the strong statistical significance of superior SNR in the DL reconstructions. The same holds for Overall Diagnostic Quality (Supplemental Figure 3A-C). Supplemental Figure 4A-C shows matrices for reviewer scores of the combined DL L and DL M as compared to MBIR for structures in the peripheral or transition zones. Although the κ-values and related Krippendorf alpha suggest only slight agreement, in no instance is there a negative score, and in all three matrices the vast majority of score pairs have at least one positive score. This is consistent with DL L and DL M having statistically significant sharpness overall versus MBIR (Fig. 4A, B). Sharpness improved in DL L and M not because of superior intrinsic resolution (shown to be equivalent for all pathways in Fig. 2) but was secondary to improved SNR. Finally, Supplemental Figure 5A-C includes matrices showing differential scores of Fidelity of Contrast of DL L and DL M compared to DL H. Positive scores indicate degraded fidelity with DL H. In all three cases, there is a clear preference for positive values, consistent with loss of contrast fidelity using DL H.
Although the DL L and M were both superior to MBIR for virtually all criteria, and superior to DL H for contrast fidelity, artifact, and multiple sharpness criteria, the DL M level of noise reduction was definitively selected (44 of 51 evaluations) as the preferred reconstruction pathway (Fig. 5B).
We believe these results are significant in that the marked increase in apparent SNR provided by the DL reconstruction may allow further improved thin-slice T2SE imaging. For example, SNR increase may allow reduction in the 20 cm FOV used in this work to 16 cm, which for the same in-plane sampling (320 × 280) would yield improved absolute in-plane resolution and with it the potentially improved depiction of nodule borders. Alternatively, the SNR improvement provided by DL reconstruction may allow increase in the acceleration factor above the R = 1.5 used in this work, permitting reduced acquisition time for the same resolution.
This work has limitations. The number of studies included (17) was limited. The work was performed at a single institution. The acquisition time used (6:00 or more) for a T2SE image set is longer than desired. However, these are all potentially correctable in further extensions of this work.
Conclusions
We have evaluated a deep learning (DL)-based image reconstruction for thin-slice (≈1 mm thick) super-resolution T2-weighted spin-echo MRI of the prostate and shown that the low and moderate levels of DL provide significantly improved performance in apparent SNR, sharpness, and retention of contrast fidelity versus a previously evaluated wavelet-based reconstruction method. Although the high level of DL also provides improved SNR, it can suffer from diminished radiological sharpness and contrast fidelity.
References
Crooks LE, Ortendahl DA, Kaufman L, et al. Clinical efficacy of nuclear magnetic resonance imaging. Radiology. 1983;146:123-8.
Hennig J, Nauerth A, Friedburg H. RARE imaging: a fast imaging method for clinical MR. Magn Reson Med. 1986;3:823-33.
Mugler JP, Bao S, Mulkern RV, et al. Optimized single slab three-dimensional spin-echo MR imaging of the brain. Radiology. 2000;216:891-9.
Busse RF, Hariharan H, Vu A, Brittain JH. Fast spin echo sequences with very long echo trains: design of variable refocusing flip angle schedules and generation of clinical T2 contrast. Magn Reson Med. 2006;55:1030-7.
Gold GE, Busse RF, Beehler C, et al. Isotropic MRI of the knee with 3D fast spin-echo extended echo-train acquistiion (XETA): initial experience. AJR. 2007;188:1287-93.
Rosenkrantz AB, Neil J, Kong X, et al. Prostate cancer: comparison of 3D T2-weighted with conventional 2D T2-weighted imaging for image quality and tumor detection. Am J Roentgenol. 2010;194:446-52.
Aiken AH, Mukherjee P, Green AJ, Glastonbury CM. MR imaging of optic neuropathy with extended echo-train acquisition fluid-attenuated inversion recovery. Am J Neuroradiol. 2011;32:301-5.
Mugler JP. Optimized three-dimensional fast-spin-echo MRI. J Magn Reson Img. 2014;39:745-67.
Srinivasan S, Wu HH, Sung K, Margolis DJA, Ennis DB. Fast 3D T2-weighted imaging usign variable flip angle transition into driven equilibrium (3D T2-TIDE) balanced SSFP for prostate imaging at 3T. Magn Reson Med. 2015;74:442-51.
Vidya Shankar R, Roccia E, Cruz G, et al. Accelerated 3D T2w-imaging of the prostate with 1-millimeter isotropic resolution in less than 3 minutes. Magn Reson Med. 2019;82:721-31.
Wilman AH, Riederer SJ. Improved centric phase encoding orders for three dimensional magnetization prepared MR angiography. Magn Reson Med. 1996;36:384-92.
Choi MH, Kim DH, Lee YJ, Rha SE, Lee JY. Image features of the PI-RADS for predicting extraprostatic extension of prostate cancer: systematic review and meta-analysis. Insights into Imaging. 2023;14:77.
Karademir I, Shen D, Peng Y, et al. Prostate volumes derived from MRI and volume-adjusted serum prostate-specific antigen: correlation with Gleason score of prostate cancer. 2013;201:1041-8.
Vargas HA, Akin O, Shukla-Dave A, et al. Performance characteristics of MR imaging in the evaluation of clincally low-risk prostate cancer: a prospective study. Radiology. 2012;265:478-87.
Greenspan H, Oz G, Kiryati N, Peled S. MRI inter-slice reconstruction using super-resolution. Magn Reson Img. 2002;20:437-46.
Shilling RZ, Robbie TQ, Bailloeul T, Mewes K, Mersereau RM, Brummer ME. A super-resolution framework for 3-D high-resolution and high-contrast imaging using 2-D multislice MRI. IEEE Trans Med Img. 2009;28:633-44.
Aganj I, Lenglet C, Yacoub E, Sapiro G, Harel N. A 3D wavelet fusion approach for the reconstruction of isotropic-resolution MR images from orthogonal anisotropic resolution scans. Magn Reson Med. 2012;67:1167-72.
Plenge E, Poot DHJ, Bernsen M, et al. Super-resolution methods in MRI: can they improve the trade-off between resolution, signal-to-noise ratio, and acquisition time? Magn Reson Med. 2012;68:1983-93.
Gholipour A, Afacan O, Aganj I, et al. Super-resolution reconstruction in frequency, image, and wavelet domains to reduce through-plane partial voluming in MRI. Med Phys. 2015;42:6919-32.
Okanovic M, Hillig B, Breuer F, Jakob P, Blaimer M. Time-of-flight MR-angiography with a helical trajectory and slice-super-resolution reconstruction. Magn Reson Med. 2018;80:1812-23.
Kargar S, Borisch EA, Froemming AT, et al. Use of kZ-space for high through-plane resolution in multi-slice MRI: application to prostate. Magn Reson Med. 2019;81:3691-704.
Borisch EA, Grimm RC, Froemming AT, Kawashima A, Trzasko JD, Riederer SJ. Model-based image reconstruction with wavelet sparsity regularization for through-plane resolution restoration in T2-weighted spin-echo prostate MRI. Magn Reson Med. 2023;89:454-68.
Wang X, Ma J, Bhosale P, et al. Novel deep learning-based noise reduction technique for prostate magnetic resonance imaging. Abdom Radiol. 2021;46:3378-86.
Gassenmaier S, Afat S, Nickel D, Mostapha M, Hermann J, Othman AE. Deep learning-accelerated T2-weighted imaging of the prostate: reduction of acquisition time and improvement of image quality. Eur J Radiol. 2021;137:109600.
Park JC, Park EJ, Park MY, Kim M, Kim JK. Fast T2-weighted imaging with deep learning-based reconstruction: evaluation of image quality and diagnostic performance in patients undergoing radical prostatectomy. J Magn Reson Img. 2022;55:1734-44.
Kelleher CB, Macdonald J, Jaffe TA, et al. A faster prostate MRI: comparing a novel denoised, single-average T2 sequence to the conventional multiaverage T2 sequence regarding lesion detection and PI-RADS score assessment. J Magn Reson Img. 2023;58:620-9.
Tong A, Bagga B, Petrocelli R, et al. Comparison of a deep learning-accelerated vs. conventional T2-weighted sequence in biparametric MRI of the prostate. J Magn Reson Img. 2023;58:1055–64.
Kargar S, Borisch EA, Froemming AT, et al. Modified acquisition strategy for reduced motion artifact in super resolution T2FSE multislice MRI: application to prostate. Magn Reson Med. 2020;84:2537-50.
Lebel RM. Performance characteristization of a novel deep learning-based MR image reconstruction pipeline. arXiv. 2020;arXiv:2008.06559.
Turkbey B, Rosenkrantz AB, Haider MA, et al. Prostate imaging reporting and data system version 2.1: 2019 update of prostate imaging reporting and data system version 2. Eur Urology. 2019;76:340-51.
Keenan KE, Stupic KF, Boss MA, et al. Multi-site, multi-vendor comparison of T1 measurement using ISMRM/NIST system phantom. 24th Annual Mtg, ISMRM. Singapore2016; p. 3290.
Senn S, Stevens L, Chaturvedi N. Repeated measures in clinical trials: simple strategies for analysis using summary measures. Statistics in Med. 2000;19:861-77.
Schober P, Vetter TR. Repeated measures designs and analysis of longitudinal data: if a first you do not succeed - try, try again. Anesth & Analg. 2018;127:569-74.
Rosner B. Fundamentals of Biostatistics. Sixth ed. Belmont CA: Thomson, 2006; p. 868.
Cohen J. A coefficeint of agreement for nominal scales. Educ Psych Meas. 1960;20:37-46.
Krippendorf K. Content analysis: an introduction to its methodology. Thousand Oaks CA, 2013: 221–250.
Lee CH. Quantitative T2-mapping using MRI for detection of prostate malignancy: a systematic review of the literature. Acta Radiologica. 2019;60:1181-9.
Mai J, Abubrig A, Lehmann T, et al. T2 mapping in prostate cancer. Invest Radiol. 2019;54:146-52.
Klingebiel M, Schimmoller L, Weiland E, et al. Value of T2 mapping MRI for prostate cancer detection and classification. J Magn Reson Img. 2022;54:413-22.
Acknowledgements
We would like to acknowledge Kathy J. Brown and Corey C. Woxland, R.T., for assistance with the human studies and Nicholas B. Larson, Ph.D., for consultation on statistical evaluation.
Funding
National Institutes of Health RR018898; National Institutes of Health R01 EB031790; Mayo Discovery-Translation Program; Mayo Imaging Biomarker Program; General Electric Healthcare.
Author information
Authors and Affiliations
Contributions
All authors contributed to the study conception and design. Material preparation, data collection, and analysis were performed by S. J. Riederer and E. A. Borisch. The first draft of the manuscript was written by S. J. Riederer, and all authors commented on previous versions of the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
SJR and EAB are inventors of a US patent related to the technology discussed. ATF, AK, and NT declare no relevant financial or non-financial interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Supplementary file6 (MP4 39946 kb)
Supplementary file7 (MP4 42759 kb)
Supplementary file8 (MP4 30339 kb)
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Riederer, S.J., Borisch, E.A., Froemming, A.T. et al. Comparison of model-based versus deep learning-based image reconstruction for thin-slice T2-weighted spin-echo prostate MRI. Abdom Radiol 49, 2921–2931 (2024). https://doi.org/10.1007/s00261-024-04256-1
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00261-024-04256-1