Abstract
Porcine deltacoronavirus (PDCoV) is an emerging swine enteropathogenic coronavirus that caused diarrhea and/or vomiting in neonatal piglets worldwide. Coronaviruses nucleocapsid (N) protein is the most conserved structural protein for viral replication and possesses good antigenicity. In this study, three monoclonal antibodies (mAbs), 3B4, 4D3, and 4E3 identified as subclass IgG2aκ were prepared using the lymphocytic hybridoma technology against PDCoV N protein. Furthermore, the B-cell epitope recognized by mAb 4D3 was mapped by dozens of overlapping truncated recombinant proteins based on the western blotting. The polypeptide 28QFRGNGVPLNSAIKPVE44 (EP-4D3) in the N-terminal of PDCoV N protein was identified as the minimal linear epitope for binding mAb 4D3. And the EP-4D3 epitope’s amino acid sequence homology study revealed that PDCoV strains are substantially conserved, with the exception of the Alanine43 substitution Valine43 in the China lineage, the Early China lineage, and the Thailand, Vietnam, and Laos lineage. The epitope sequences shared high similarity (94.1%) with porcine coronavirus HKU15-155 (PorCoV HKU15), Asian leopard cats coronavirus (ALCCoV), sparrow coronavirus HKU17 (SpCoV HKU17), and sparrow deltacoronavirus. In contrast, the epitope sequences shared a very low homology (11.8 to 29.4%) with other porcine CoVs (PEDV, TGEV, PRCV, SADS-CoV, PHEV). Overall, the study will enrich the biological function of PDCoV N protein and provide foundational data for further development of diagnostic applications.
Key points
• Three monoclonal antibodies against PDCoV N protein were prepared.
• Discovery of a novel B-cell liner epitope (28QFRGNGVPLNSAIKPVE44) of PDCoV N protein.
• The epitope EP-4D3 was conserved among PDCoV strains.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coronaviruses (CoVs) are single positive-stranded, enveloped RNA viruses which belong to the order Nidovirales, family Coronaviridae, and subfamily Orthocoronavirinae comprising of four genera, Alpha-, Beta-, Gamma-, and Deltacoronavirus. CoVs are regarded as the largest proportion of bat-borne viruses that can infect mammals and birds, and cause respiratory, gastrointestinal, and neurological disease (Tian et al. 2022; Woo et al. 2012). With the ongoing Severe Acute respiratory Syndrome Coronavirus 2 (SARS-CoV-2) of the genus Beta-CoVs, the CoVs have been posing a serious threat to public health and economy on a global scale (Nguyen et al. 2022; Zhai et al. 2020). Porcine deltacoronavirus (PDCoV) as an emerging member in Delta-CoVs has been detected and reported in many countries’ swine population around the world since its first detected in 2009, in China Hong Kong (Le et al. 2018; Lee et al. 2016; Li et al. 2018a; Lorsirigool et al. 2017; Lorsirigool et al. 2016; Perez-Rivera et al. 2019; Wang et al. 2014; Woo et al. 2012). Its clinical symptoms are similar to those of the porcine epidemic diarrhea virus (PEDV), which causes diarrhea and/or vomiting in newborn piglets, but with milder virulence than PEDV(Wang et al. 2014). However, studies in recent years have reported PDCoV spillover occurrences into other species, with chickens, turkey poults, and calves being susceptible to PDCoV infection (Boley et al. 2020; Jung et al. 2017; Liang et al. 2019). Remarkably, the PDCoV mutated in the non-structural protein (Nsp) 15 and the spike (S) glycoprotein genes can infect children with mild illness (fever, cough, and abdominal pain) in Haiti (Lednicky et al. 2021). It is the first detected event that PDCoV of swine population successfully spilled into humans, suggesting that the PDCoV possesses the potential ability of cross-species transmission of CoVs (He et al. 2022b). Therefore, we urgently need to keep on conducting more researches to understand and prevent the PDCoV.
Among the four structural proteins of CoVs, the S protein and the nucleocapsid (N) protein are two main immunogens (Meyer et al. 2014). The CoVs S protein is a trimeric class I transmembrane protein which stimulates cell tropism and virus entry and induces neutralizing antibodies and protective immunity in the host system (He et al. 2006; Simmons et al. 2004). In order to monitor and evaluate the genetic evolution of PDCoVs well, PDCoV have been divided into four lineages based on the phylogenetic tree of the S protein because of its higher mutation rate, including the Thailand, Vietnam, and Laos lineage; the Early China lineage; the USA, Japan, and South Korea lineage; and the China lineage (He et al. 2020; Zhang et al. 2019). The N protein of the CoVs packages the viral genome RNA (gRNA) into a helical ribonucleocapsid (RNP) and plays an important role in transcription and replication in viral life cycle. (Chang et al. 2014; McBride et al. 2014). Since the PDCoV N protein exhibited the highest conservation level sharing 96.94 to 100.00% amino acid homologies, so it has been used as diagnostic marker and immunogen in the in the development of diagnostic applications. Wang et al. developed a N protein-based blocking enzyme-linked immunosorbent assays (ELISA) for the detection of PDCoV antibodies, the diagnostic sensitivity of the blocking ELISA were 98.7% (326/330), which is slightly lower than 100% (330/330) sensitivity by IFA identified 330 positive samples in 384 swine serum samples (Wang et al. 2022). Yu et al. developed an Epitope-ELISA diagnostic method based on SARS-CoV N protein that can successfully distinct from the SARS-CoV antibodies in serological detection of COVID-19 patients (Yu et al. 2022). In view of the abovementioned, it is attractive and effective that has been developing serological diagnostic method based on the CoVs N proteins.
Here, in order to have a better understanding of the biological function of PDCoV N protein, we purified recombinant His-tagged PDCoV N protein and prepared three monoclonal antibodies (mAbs) (3B4, 4D3, and 4E3) with the lymphocytic hybridoma technology. Due to the mAb 4D3 has the highest antibody affinity, accordingly, we further identified the precise B-cell linear epitope of mAb 4D3. Furthermore, homology analysis revealed that the sequence of novel liner B-cell epitope, EP-4D3 was highly conserved among PDCoV strains. The identification of novel epitope can assist us in comprehending the antigenic structure of viral protein and be used for further developing diagnostic methods for PDCoV.
Materials and methods
Virus strains and cells
PDCoV SD2018/10 (Genbank accession No.MN520194) and AH2019/H (Genbank accession No.MN520198) strains were preserved in our laboratory. The myeloma cell line SP 2/0 was cultured in Roswell Park Memorial Institute (RPMI) 1640 medium (HyClone, USA) with 15% fetal bovine serum (Gibco, USA) under a humidified 5% CO2 atmosphere at 37 °C. The LLC-PK1 cells were cultured in Dulbecco’s modified Eagle’s medium (DMEM) (Gibco, USA) with 10% fetal bovine serum (Gibco, USA) at 37 °C with 5% CO2. All culture media contained 1% penicillin–streptomycin antibiotics (NCM Biotech, China).
Expression and purification of PDCoV N protein
The PDCoV full-length N protein gene was subcloned into pET-32a ( +) vector (EcoR I and Xho I) with 6 × His-Tag, using the amplification primers listed in Table 1, and then transformed into E.coli BL21 (DE3). The recombinant His-PDCoV N protein was purified by Ni–NTA His-Bind Resin after induction at 16℃ for 12 h with 1 mM Isopropyl-beta-d-thiogalactopyranoside (IPTG), and evaluated by sodium dodecyl sulfate–polyacrylamide gel electrophoresis (SDS-PAGE) and western blotting.
Preparation and identification of mAbs against PDCoV N protein
Six- to-8week female BALB/c mice were immunized subcutaneously with 100 μg recombinant His-PDCoV N protein emulsified with freund complete adjuvant (Sigma-Aldrich, USA), followed by immunized with 100 μg recombinant His-N protein emulsified with incomplete adjuvant (Sigma-Aldrich, USA) every two weeks. One week after the third immunization, the antibody titer of mice was measured, once the antibody titer of immunized mice reached 10–6, 50 μg recombinant His-PDCoV N protein without adjuvant was injected intraperitoneally. Three days later, spleen B lymphocytes and SP 2/0 cells were collected and fused with polyethylene glycol 2000 (Sigma-Aldrich, USA). Hybridoma cells were selected in RPMI 1640 medium containing hypoxanthine-aminopterin-thymidine (HAT) (Sigma-Aldrich, USA). The positive cell clones were screened by indirect indirect ELISA, then subcloned by limited dilution method at least three rounds. To generate monoclonal antibodies, the obtained antibody-secreting cells were intraperitoneally injected into sensitized mice with incomplete adjuvant. The subtypes of monoclonal antibodies were identified using the monoclonal antibody isotyping determination kit (Biodragon Immunotechnologies, China).
Indirect ELISA detection of positive and monoclonal hybridoma cell lines
The purified recombinant His-PDCoV N protein was diluted to 2 μg/mL with bicarbonate buffer and coated overnight at 4℃ on ELISA plates. After five washes with phosphate buffer saline (PBS) supplemented with 0.05% Tween-20 (PBST), the plates were blocked for 1 h at 37℃ incubated with 1.0% bovine serum albumin (BSA). After another wash, 100 μL hybridoma supernatant was added and incubated at the condition of 37℃ for 1 h, while the SP 2/0 cells supernatants and mice positive serum were used as negative control and positive control, respectively. After washing with PBST five times, horseradish peroxidase (HRP)-conjugated goat anti-mouse IgG (1:10,000) (KPL, USA) was used as a secondary antibody incubating for 1 h at 37℃. Washing again, 100 μL 3,3′,5,5′ tetramethyl benzidine (TMB) color developing solution was added and reacted at 37℃ for 10 min before being terminated with 50 μL H2SO4 (2 M). The OD value was read at 450 nm with an enzyme reader (TECAN, Switzerland). The positive wells were screened out using P/N ≥ 2.1 as the positive criteria.
Western blotting
The recombinant N proteins with 6 × His-Tag or GST-Tag and native PDCoV N protein were transferred onto the nitrocellulose (NC) membrane after separation by SDS-PAGE. The NC membrane blocked with 5% skim milk for 1 h at 37℃ after washing five times with PBST, then incubated overnight at 4℃ with the monoclonal antibody (1:1000, 1:2500, 1:5000) as the primary antibody. After five washes with PBST, the HRP-conjugated goat anti-mouse (1:10,000) with 5% skim milk was used as the secondary antibody for 1 h at 37℃. The incubated substrate solution was exposed for development after being washed.
Indirect immunofluorescence assay (IFA)
PDCoV strains SD2018/10 and AH2019/H were respectively inoculated into LLC-PK1 cells at 24-well plates with MOI of 0.01. After 16 h post-inoculation, the cells were fixed with ice methanol at -20℃ for 10 min, washed three times with PBS, and then blocked at 37℃ for 1 h with 1.0% BSA. Following PBS washing, a monoclonal antibody (1: 1000) was used as the primary antibody and incubated at 37℃ for 2 h. After washing five times with PBS, fluorescein isothiocyanate (FITC)-labeled goat anti-mouse IgG (1: 400) (KPL, USA) was used as the secondary antibody and incubated at 37℃ for 1 h. After washing with PBS, 4′,6-diamidino-2-phenylindole (DAPI) (1:1000) was added to observe fluorescence under a fluorescence microscope.
Precise epitope of PDCoV N protein
In order to identify the precise epitope of the PDCoV N protein recognized by mAb 4D3, we truncated PDCoV N protein into a series of fragments or polypeptides for five rounds based on the prediction results of online B-cell epitope prediction website (http://www.iedb.org/). The first truncated three overlapped fragments were amplified and subcloned into pET-32a ( +) vector (EcoR I and Xho I) with 6 × His-Tag. Primers used for the truncated fragments were listed in Table 1. After the fourth round, the amino acids (aa) at the C-terminus and N-terminus of polypeptide aa 25–48 were deleted one by one until the smallest binding domain was recognized by the mAb 4D3 (Fig. 1). For certain shorter polypeptides, we employed complementary primer pairs for some shorter polypeptides that were annealed at 99℃ for 10 min and then cooled to room temperature for ligation. The truncated polypeptides from the second to the fifth round, were subcloned into pGEX-4 T-1 vector (EcoR I and Xho I) with GST-Tag. After accurately sequencing of recombinant plasmids, they were all induced expression in E.coli BL21 (DE3), after induction with 1 mM IPTG at 16℃ overnight, and tested the molecular weight of these polypeptides by SDS-PAGE. The reactivity of the truncated proteins for binding the mAb 4D3 was detected by western blotting.
Schematic diagram of the relative locations of truncated fragments of PDCoV N protein for epitope mapping. The blue fragment represented the complete PDCoV N protein. The orange fragments showed that the regions could react with mAb 4D3. The gray fragments were the corresponding regions which did not react with mAb 4D3
Sequence homology analysis
To analyze the sequence homology of the identified epitope among PDCoV strains, Delta-CoVs, and porcine CoVs, we used the DNAstar MegAlign (Madison, WI, USA) and MEGA7.0 software to compare the sequences of identified epitope and reference strains. All reference strain sequences in our study were obtained from Genbank (Tables S1, S2, and S3).
Spatial conformation
To understand the spatial distribution of the identified epitope, the three-dimensional (3D) model of PDCoV N protein was modeled with the SWISS-MODEL (https://swissmodel.expasy.org) online tools using the N-terminal RNA-binding domain of SARS-CoV-2 N protein as a template (Kang et al. 2020; Khan et al. 2020); then, the visualized analysis of the epitope was structured by the PyMOL software (https://pymol.org/2). The hydrophilicity, antigenicity, and surface possibility of the epitope were analyzed by the DNAstar Protean (Madison, WI, USA) software.
Results
Expression of recombinant proteins
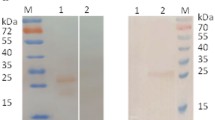
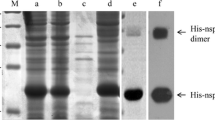
The PDCoV N gene was amplified based on the PDCoV SD2018/10 strain; then, the DNA fragment of complete PDCoV N protein was subcloned into prokaryotic expression vector the pET-32a ( +); the recombinant plasmid pET-32a-PDCoV N was then induced to express in E. coli BL21 (DE3) cells. As shown in SDS-PAGE (Fig. 2A), the recombinant His-PDCoV N successfully expressed in the supernatant (Lane 3); the molecular weight of purified His-PDCoV N protein is approximately 58 kDa (Lane 4). The western blotting (Fig. 2B) indicated that the recombinant His-PDCoV N protein was expressed as predicted and recognized specifically by anti-His mAb.
Analysis of recombinant His-PDCoV N protein by SDS-PAGE (A) and western blotting (B) with His-Tag mAb. Lane M: protein marker; Lane 1: sonication lysates of recombinant plasmid pET-32a-N transformed E. coli BL21 (DE3) without IPTG induction; Lane 2: precipitates of pET-32a-N transformed E. coli BL21 (DE3) with IPTG induction; Lane 3: cell supernatant of pET-32a-N transformed E. coli BL21 (DE3) with IPTG induction; Lane 4: purified recombinant His-N protein
Production and characterization of mAbs against the N protein
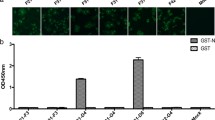
Purified recombinant His-PDCoV N protein was used as immunogen to immunize BALB/c mice for preparing the mAb against PDCoV N. After three immunizations, the mouse with the highest antibody titer assessed by specific indirect ELISA of PDCoV N was sacrificed for hybridoma production. The antibodies secreted by hybridoma cells in supernatant were analyzed by indirect ELISA and IFA, and then, the positive and growing well hybridoma cells were subcloned at least three times with the limiting limited dilution method to generate the monoclonal hybridoma cell lines. Isotype determination revealed that the heavy chain and light chain of three mAbs, 3B4, 4D3, and 4E3, were all belong to IgG2a and κ subtype. The antibody titer of our obtained three mAbs measured by the indirect ELISA were around 1 × 10–6, illustrating that the recombinant His-PDCoV N protein induced an effective immunological reaction in mice. As demonstrated by the results of IFA (Fig. 3A) and western blotting (Fig. 3B), the three mAbs could respectively and specifically recognize native N proteins from the LLC-PK1 cells respectively infected with PDCoV SD2018/10 and PDCoV AH2019/H, but not the mock sample, indicating that the prepared mAbs can be used for identification of viruses isolated and applied to other relevant molecular biology research.
Reactivity of three mAbs of PDCoV N by IFA (A) and western blotting (B). IFA was detected with mAb 3B4, 4D3, and 4E3 with 1:1000 dilution, respectively. Western blotting was performed using mAb 3B4, 4D3, and 4E3 at 1:1000, 1:2500, and 1:5000 dilutions, respectively. Lane M: protein marker; Lane 1: PDCoV SD2018/10 strain; Lane 2: PDCoV AH2019/10 strain; Lane 3: Mock
Epitope mapping of PDCoV N protein
Epitope mapping designed in Fig. 1, the complete N protein was truncated into three overlapped fragments, aa 1–142, aa 128–242, and aa 227–342, fused with His-Tag and expressed in E. coli BL21 (DE3). Western blotting results showed that only aa 1–142 could react with mAb 4D3 (Fig. 4A), indicating the epitope binding for the mAb 4D3 was located in the region of aa 1–142 of PDCoV N protein. During the second to fourth rounds of screening and identification, the polypeptide aa 25–48 fused with GST-Tag could be recognized by mAb 4D3 (Fig. 4B). To further determine the precise epitope, the N-terminal and C-terminal of polypeptide aa 25–48 were truncated one by one and fused with GST-Tag to induce expression in the fifth round. Ultimately, these results demonstrated that polypeptide 28QFRGNGVPLNSAIKPVE44 (EP-4D3) located in the N-terminal of PDCoV N protein was the minimal linear epitope for binding mAb 4D3 (Fig. 4C).
Epitope mapping of mAb 4D3 by western blotting. The expression of truncated fragments or polypeptides was detected by SDS-PAGE. A MAb 4D3 specifically reacted with truncated fragment aa 1–60 after the first two rounds of identification. B In the third and fourth round identification, the polypeptide aa 25–48 was recognized by mAb 4D3. C The polypeptide 28QFRGNGVPLNSAIKPVE.44 in the fifth round was the minimal epitope for binding mAb 4D3
Conservation analysis of epitope sequences
The conservation analysis of identified epitope EP-4D3 between diverse PDCoV strains (Table S1), together with other Delta-CoVs (Table S2) and porcine CoVs (Table S3) were respectively analyzed using the Megalign and MEGA7.0 software. Alignment results showed that the EP-4D3 epitope was a conserved epitope in all PDCoVs, despite the existence of two residues, valine43 and alanine43 that were simultaneously presented in the Thailand, Vietnam, and Laos lineage and China lineage. The amino acid at 43 site of the USA, Japan, and South Korea lineage was valine, but this position was alanine in the early China lineage (Fig. 5A). In comparison with other Delta-CoVs, EP-4D3 epitope sequence shared 35.3 to 94.1% sequence similarity with alignment sequences, and five residues (F29-G33V34P35-N37) in the identified epitope sequence were determined to be substantially conserved. The porcine coronavirus HKU15-155 (PorCoV HKU15), Asian leopard cats coronavirus (ALCCoV), sparrow coronavirus HKU17 (SpCoV HKU17), and sparrow deltacoronavirus all shared 94.1% sequence similarity with the identified epitope EP-4D3 sequence. Notably, the Alanine43 in the N epitope (28QFRGNGVPLNSAIKPAE44) of the bulbul coronavirus HKU11 (BuCoV HKU11), munia coronavirus HKU13 (MunCoV HKU13), SpCoV HKU17, sparrow deltacoronavirus, and night heron coronavirus HKU19 (NHCoV HKU19) were identical to those of the ALCCoV and PorCoV HKU15 (Fig. 5B). In addition, there was a low level of sequence homology of the EP-4D3 epitope among PDCoV and other porcine CoVs (PEDV, TGEV, PRCV, SADS-CoV, PHEV), ranging from 11.8 to 29.4%, with the valine34 and proline35 in the corresponding epitope sequences being relatively conserved in all alignment sequences (Fig. 5C).
Conservation analysis of identified epitope for mAb 4D3. A Amino acid alignment of diverse PDCoV strains, B comparison the B-cell epitope sequences with other Delta-CoVs, and C homology analysis with other porcine CoVs using the Megalign and MEGA7.0 software; the precise epitope 28QFRGNGVPLNSAIKPVE.44 were surrounded by black dotted box; the dark spot represented the same amino acid as the B-cell epitope of mAb 4D3
Spatial position of epitope binding
The spatial distribution of the identified epitope EP-4D3 (28QFRGNGVPLNSAIKPVE44) was located on a 3D model of the N-terminal of the PDCoV N protein constructed by the SWISS-MODEL and highlighted in red using PyMOL software (Fig. S1A and S1B). The structure of the identified epitope showed a typical B-cell epitope characterization with a high antigenic index and hydrophilicity (Fig. S1C).
Discussion
CoVs N protein is the most abundant and highly immunogenic viral protein in infected cells which can stimulate B lymphocytes to secrete specific and abundant antibodies against N protein in a short time (Seah et al. 2000), making CoV N protein as optimal candidate for diagnosis development. Therefore, the prepared specific monoclonal antibodies and identified novel B-cell linear epitope of PDCoV N are of great help to understand the antigenic structure of PDCoV N. Previous study revealed that PDCoV AH2019/H and SD2018/10 respectively belong to the USA, Japan, and South Korea lineage and the China lineage (He et al. 2020). In our study, all three generated mAbs (3B4, 4D3, and 4E3) can react to native N protein of PDCoV SD2018/10 and AH2019/H strains in IFA (Fig. 3A) and western blotting assay (Fig. 3B), which means that these three mAbs all have specific reactivity and can be widely used to identify PDCoV isolated strains in labs. The mAbs obtained in our study were identified as subclass IgG2aκ, and with an antibody titer of 1 × 10–6. So far, only few B-cell epitopes 59GTPIPPSYAFYY70, 251NFQAG-PDYER276, 309NKRETTLQQ317, and 326QDWEWDDA333 of N protein have been reported(Fu et al. 2020; Ren et al. 2022; Wei et al. 2021). Given that the highest affinity of antibody, we investigated the B-cell epitope of mAb 4D3. The polypeptide 28QFRGNGVPLNSAIKPVE44 (EP-4D3) in the N-terminal of PDCoV N protein was identified as the minimal linear epitope for binding mAb 4D3 (Fig. 1, Fig. 4, and Fig. S1).
Homology analysis revealed that EP-4D3 epitope sequences were highly conserved among PDCoV strains, with different amino-acid residues at the 43 position of EP-4D3 epitope sequence and sharing 94.1% sequence similarity between the first detected porcine deltacoronavirus PorCoV HKU15 (alanine43) and PDCoV SD2018/10 (valine43) which both belong to China lineage (Fig. 5A) (He et al. 2020). Previous studies discovered that the avain and mammalian CoVs of Deltacoronavirus shared similar comparable genome structures and characteristics, and that the avian CoVs serve as the gene source for Gammacoronavirus and Deltacoronavirus (Woo et al. 2009, 2012). It reported that real-time RT-PCR targeting the N protein gene in molecular investigation of PDCoV can cross-react with sparrow deltacoronavirus because of high sequence similarity between the two N proteins (Chen et al. 2018). One study found that PDCoV’s functional engagement of orthologous receptors offers credible evidence for PDCoV’s ancestor breaching the species barrier between avian and mammals (Li et al. 2018b). It is interesting that the N protein epitope sequences (28QFRGNGVPLNSAIKPAE44) of PorCoV HKU15, ALCCoV, SpCoV HKU17, and sparrow deltacoronavirus shared 100% sequence identity (Fig. 5B), once again suggesting that PDCoV may have originated from an avian-to-mammalian host-jump, which is consistent with the viewpoint that mammalian Delta-CoVs (PorCoV HKU15 and ALCCoV) may have evolved from recombination events within SpCoV HKU17, BuCoV HKU11, or other CoVs (Lau et al. 2018). In short, the amino acid homology analysis of EP-4D3 identified in this study gives us an insight into the evolution of CoVs.
Up to now, six porcine CoVs have been described, including PEDV (He et al. 2022a; Pensaert and de Bouck 1978), transmissible gastroenteritis virus (TGEV) (Laude et al. 1990), porcine respiratory coronavirus (PRCV) (Costantini et al. 2004), SADS-CoV (Zhou et al. 2018), porcine hemagglutinating encephalomyelitis virus (PHEV) (Roe and Alexander 1958), and PDCoV (Woo et al. 2012). PDCoV and SADS-CoV are regarded as emerging CoVs comparison to the other swine CoVs; there are few available diagnostic methods for the emerging swine CoVs at present. The EP-4D3 epitope sequence showed a relatively lower homology (11.8 to 29.4%) comparing with other porcine CoVs’ s corresponding sequences (Fig. 5C). It is noteworthy that the inter-genus and intra-genus cross-reactivity of porcine CoVs poses a significant obstacle to develop specifically diagnostic techniques (Gimenez-Lirola et al. 2017; Lin et al. 2015; Ma et al. 2016). Detected by the IFA and western blotting, the mAb 4D3 did not react to the native PEDV and TGEV strains in our lab (data not shown), which implying that this novel B-cell epitope could not be recognized by the PEDV and TGEV strains. Nevertheless, we also noticed that the novel B-cell epitope sequence contains a number of T-cell epitopes through IEDB online prediction, the two larger T-cell epitopes among which were 31GNGVPLNSAIKPVE44 and 32NGVPLNSAIKPVE44. Based on its high PDCoV-specificity and immunogenicity, the novel B-cell linear epitope recognized by mAb 4D3 can be exploited to develop epitope-based serological diagnostics method in the future.
In summary, we successfully prepared three mAbs (3B4, 4D3, and 4E3) against PDCoV N protein which showing good specificity and sensitivity, and identified a novel linear B-cell epitope 28QFRGNGVPLNSAIKPVE44 in the N-terminal of PDCoV N protein as the minimal linear epitope for binding mAb 4D3. Our findings added to the rapidly expanding field of PDCoV N protein antigenic structure and offered foundation data for further development of diagnostic applications.
Data availability
Authors can confirm that all relevant data are included in the article and/or its supplementary information files.
References
Boley PA, Alhamo MA, Lossie G, Yadav KK, Vasquez-Lee M, Saif LJ, Kenney SP (2020) Porcine deltacoronavirus infection and transmission in poultry, United States(1). Emerg Infect Dis 26(2):255–265. https://doi.org/10.3201/eid2602.190346
Chang CK, Hou MH, Chang CF, Hsiao CD, Huang TH (2014) The SARS coronavirus nucleocapsid protein–forms and functions. Antiviral Res 103:39–50. https://doi.org/10.1016/j.antiviral.2013.12.009
Chen Q, Wang L, Yang C, Zheng Y, Gauger PC, Anderson T, Harmon KM, Zhang J, Yoon KJ, Main RG, Li G (2018) The emergence of novel sparrow deltacoronaviruses in the United States more closely related to porcine deltacoronaviruses than sparrow deltacoronavirus HKU17. Emerg Microbes Infect 7(1):105. https://doi.org/10.1038/s41426-018-0108-z
Costantini V, Lewis P, Alsop J, Templeton C, Saif LJ (2004) Respiratory and fecal shedding of porcine respiratory coronavirus (PRCV) in sentinel weaned pigs and sequence of the partial S-gene of the PRCV isolates. Arch Virol 149(5):957–974. https://doi.org/10.1007/s00705-003-0245-z
Fu J, Chen R, Hu J, Qu H, Zhao Y, Cao S, Wen X, Wen Y, Wu R, Zhao Q, Ma X, Huang X (2020) Identification of a novel linear b-cell epitope on the nucleocapsid protein of porcine deltacoronavirus. Int J Mol Sci 21(2):648. https://doi.org/10.3390/ijms21020648
Gimenez-Lirola LG, Zhang J, Carrillo-Avila JA, Chen Q, Magtoto R, Poonsuk K, Baum DH, Pineyro P, Zimmerman J (2017) Reactivity of porcine epidemic diarrhea virus structural proteins to antibodies against porcine enteric coronaviruses: diagnostic implications. J Clin Microbiol 55(5):1426–1436. https://doi.org/10.1128/JCM.02507-16
He WT, Bollen N, Xu Y, Zhao J, Dellicour S, Yan Z, Gong W, Zhang C, Zhang L, Lu M, Lai A, Suchard MA, Ji X, Tu C, Lemey P, Baele G, Su S (2022a) Phylogeography reveals association between swine trade and the spread of porcine epidemic diarrhea virus in china and across the world. Mol Biol Evol 39(2):msab364. https://doi.org/10.1093/molbev/msab364
He WT, Hou X, Zhao J, Sun J, He H, Si W, Wang J, Jiang Z, Yan Z, Xing G, Lu M, Suchard MA, Ji X, Gong W, He B, Li J, Lemey P, Guo D, Tu C, Holmes EC, Shi M, Su S (2022b) Virome characterization of game animals in China reveals a spectrum of emerging pathogens. Cell 185(7):1117–1129 e8. https://doi.org/10.1016/j.cell.2022b.02.014
He WT, Ji X, He W, Dellicour S, Wang S, Li G, Zhang L, Gilbert M, Zhu H, Xing G, Veit M, Huang Z, Han GZ, Huang Y, Suchard MA, Baele G, Lemey P, Su S (2020) Genomic epidemiology, evolution, and transmission dynamics of porcine deltacoronavirus. Mol Biol Evol 37(9):2641–2654. https://doi.org/10.1093/molbev/msaa117
He Y, Li J, Heck S, Lustigman S, Jiang S (2006) Antigenic and immunogenic characterization of recombinant baculovirus-expressed severe acute respiratory syndrome coronavirus spike protein: implication for vaccine design. J Virol 80(12):5757–5767. https://doi.org/10.1128/JVI.00083-06
Jung K, Hu H, Saif LJ (2017) Calves are susceptible to infection with the newly emerged porcine deltacoronavirus, but not with the swine enteric alphacoronavirus, porcine epidemic diarrhea virus. Arch Virol 162(8):2357–2362. https://doi.org/10.1007/s00705-017-3351-z
Kang S, Yang M, Hong Z, Zhang L, Huang Z, Chen X, He S, Zhou Z, Zhou Z, Chen Q, Yan Y, Zhang C, Shan H, Chen S (2020) Crystal structure of SARS-CoV-2 nucleocapsid protein RNA binding domain reveals potential unique drug targeting sites. Acta Pharm Sin B 10(7):1228–1238. https://doi.org/10.1016/j.apsb.2020.04.009
Khan A, Tahir Khan M, Saleem S, Junaid M, Ali A, Shujait Ali S, Khan M, Wei DQ (2020) Structural insights into the mechanism of RNA recognition by the N-terminal RNA-binding domain of the SARS-CoV-2 nucleocapsid phosphoprotein. Comput Struct Biotechnol J 18:2174–2184. https://doi.org/10.1016/j.csbj.2020.08.006
Lau SKP, Wong EYM, Tsang CC, Ahmed SS, Au-Yeung RKH, Yuen KY, Wernery U, Woo PCY (2018) Discovery and sequence analysis of four deltacoronaviruses from birds in the Middle East reveal interspecies jumping with recombination as a potential mechanism for avian-to-avian and avian-to-mammalian transmission. J Virol 92(15):e00265-18. https://doi.org/10.1128/JVI.00265-18
Laude H, Rasschaert D, Delmas B, Godet M, Gelfi J, Charley B (1990) Molecular biology of transmissible gastroenteritis virus. Vet Microbiol 23(1–4):147–154. https://doi.org/10.1016/0378-1135(90)90144-k
Le VP, Song S, An BH, Park GN, Pham NT, Le DQ, Nguyen VT, Vu TTH, Kim KS, Choe S, An DJ (2018) A novel strain of porcine deltacoronavirus in Vietnam. Arch Virol 163(1):203–207. https://doi.org/10.1007/s00705-017-3594-8
Lednicky JA, Tagliamonte MS, White SK, Elbadry MA, Alam MM, Stephenson CJ, Bonny TS, Loeb JC, Telisma T, Chavannes S, Ostrov DA, Mavian C, De Rochars VMB, Salemi M, Morris JG (2021) Emergence of porcine delta-coronavirus pathogenic infections among children in Haiti through independent zoonoses and convergent evolution. medRxiv. https://doi.org/10.1101/2021.03.19.21253391
Lee JH, Chung HC, Nguyen VG, Moon HJ, Kim HK, Park SJ, Lee CH, Lee GE, Park BK (2016) Detection and phylogenetic analysis of porcine deltacoronavirus in Korean swine farms, 2015. Transbound Emerg Dis 63(3):248–252. https://doi.org/10.1111/tbed.12490
Li D, Feng H, Liu Y, Chen Y, Wei Q, Wang J, Liu D, Huang H, Su Y, Wang D, Cui Y, Zhang G (2018) Molecular evolution of porcine epidemic diarrhea virus and porcine deltacoronavirus strains in Central China. Res Vet Sci 120:63–69. https://doi.org/10.1016/j.rvsc.2018.06.001
Li W, Hulswit RJG, Kenney SP, Widjaja I, Jung K, Alhamo MA, van Dieren B, van Kuppeveld FJM, Saif LJ, Bosch BJ (2018) Broad receptor engagement of an emerging global coronavirus may potentiate its diverse cross-species transmissibility. Proc Natl Acad Sci U S A 115(22):E5135–E5143. https://doi.org/10.1073/pnas.1802879115
Liang Q, Zhang H, Li B, Ding Q, Wang Y, Gao W, Guo D, Wei Z, Hu H (2019) Susceptibility of chickens to porcine deltacoronavirus infection. Viruses 11(6):573. https://doi.org/10.3390/v11060573
Lin CM, Gao X, Oka T, Vlasova AN, Esseili MA, Wang Q, Saif LJ (2015) Antigenic relationships among porcine epidemic diarrhea virus and transmissible gastroenteritis virus strains. J Virol 89(6):3332–3342. https://doi.org/10.1128/JVI.03196-14
Lorsirigool A, Saeng-Chuto K, Madapong A, Temeeyasen G, Tripipat T, Kaewprommal P, Tantituvanont A, Piriyapongsa J, Nilubol D (2017) The genetic diversity and complete genome analysis of two novel porcine deltacoronavirus isolates in Thailand in 2015. Virus Genes 53(2):240–248. https://doi.org/10.1007/s11262-016-1413-z
Lorsirigool A, Saeng-Chuto K, Temeeyasen G, Madapong A, Tripipat T, Wegner M, Tuntituvanont A, Intrakamhaeng M, Nilubol D (2016) The first detection and full-length genome sequence of porcine deltacoronavirus isolated in Lao PDR. Arch Virol 161(10):2909–2911. https://doi.org/10.1007/s00705-016-2983-8
Ma Y, Zhang Y, Liang X, Oglesbee M, Krakowka S, Niehaus A, Wang G, Jia A, Song H, Li J (2016) Two-way antigenic cross-reactivity between porcine epidemic diarrhea virus and porcine deltacoronavirus. Vet Microbiol 186:90–96. https://doi.org/10.1016/j.vetmic.2016.02.004
McBride R, van Zyl M, Fielding BC (2014) The coronavirus nucleocapsid is a multifunctional protein. Viruses-Basel 6(8):2991–3018. https://doi.org/10.3390/v6082991
Meyer B, Drosten C, Muller MA (2014) Serological assays for emerging coronaviruses: challenges and pitfalls. Virus Res 194:175–183. https://doi.org/10.1016/j.virusres.2014.03.018
Nguyen NN, Houhamdi L, Hoang VT, Delerce J, Delorme L, Colson P, Brouqui P, Fournier PE, Raoult D, Gautret P (2022) SARS-CoV-2 reinfection and COVID-19 severity. Emerg Microbes Infect 11(1):894–901. https://doi.org/10.1080/22221751.2022.2052358
Pensaert MB, de Bouck P (1978) A new coronavirus-like particle associated with diarrhea in swine. Arch Virol 58(3):243–247. https://doi.org/10.1007/BF01317606
Perez-Rivera C, Ramirez-Mendoza H, Mendoza-Elvira S, Segura-Velazquez R, Sanchez-Betancourt JI (2019) First report and phylogenetic analysis of porcine deltacoronavirus in Mexico. Transbound Emerg Dis 66(4):1436–1441. https://doi.org/10.1111/tbed.13193
Ren H, Yan X, Liu L, Zhang Y, Li Q, Li X, Hu H (2022) Identification of two novel B-cell epitopes on the nucleocapsid protein of porcine deltacoronavirus. Virol Sin 37(2):303–306. https://doi.org/10.1016/j.virs.2022.01.025
Roe CK, Alexander TJ (1958) A Disease of Nursing Pigs Previously Unreported in Ontario. Can J Comp Med Vet Sci 22(9):305–307
Seah JN, Yu L, Kwang J (2000) Localization of linear B-cell epitopes on infectious bronchitis virus nucleocapsid protein. Vet Microbiol 75(1):11–16. https://doi.org/10.1016/s0378-1135(00)00202-9
Simmons G, Reeves JD, Rennekamp AJ, Amberg SM, Piefer AJ, Bates P (2004) Characterization of severe acute respiratory syndrome-associated coronavirus (SARS-CoV) spike glycoprotein-mediated viral entry. Proc Natl Acad Sci U S A 101(12):4240–4245. https://doi.org/10.1073/pnas.0306446101
Tian J, Sun J, Li D, Wang N, Wang L, Zhang C, Meng X, Ji X, Suchard MA, Zhang X, Lai A, Su S, Veit M (2022) Emerging viruses: Cross-species transmission of coronaviruses, filoviruses, henipaviruses, and rotaviruses from bats. Cell Rep 39(11):110969. https://doi.org/10.1016/j.celrep.2022.110969
Wang L, Byrum B, Zhang Y (2014) Detection and genetic characterization of deltacoronavirus in pigs, Ohio, USA. Emerg Infect Dis 20(7):1227–1230. https://doi.org/10.3201/eid2007.140296
Wang W, Zhang Y, Yang H (2022) Development of a nucleocapsid protein-based blocking ELISA for the detection of porcine deltacoronavirus antibodies. Viruses 14(8):1815. https://doi.org/10.3390/v14081815
Wei S, Shi D, Wu H, Sun H, Chen J, Feng L, Su M, Sun D (2021) Identification of a novel B cell epitope on the nucleocapsid protein of porcine deltacoronavirus. Virus Res 302:198497. https://doi.org/10.1016/j.virusres.2021.198497
Woo PC, Lau SK, Huang Y, Yuen KY (2009) Coronavirus diversity, phylogeny and interspecies jumping. Exp Biol Med (maywood) 234(10):1117–1127. https://doi.org/10.3181/0903-MR-94
Woo PC, Lau SK, Lam CS, Lau CC, Tsang AK, Lau JH, Bai R, Teng JL, Tsang CC, Wang M, Zheng BJ, Chan KH, Yuen KY (2012) Discovery of seven novel Mammalian and avian coronaviruses in the genus deltacoronavirus supports bat coronaviruses as the gene source of alphacoronavirus and betacoronavirus and avian coronaviruses as the gene source of gammacoronavirus and deltacoronavirus. J Virol 86(7):3995–4008. https://doi.org/10.1128/JVI.06540-11
Yu J, Qin Z, Liu X, He X, Yao J, Zhou X, Wen K, Yu N, Wu Q, Xiao W, Zhu L, Wan C, Zhang B, Zhao W (2022) High-specificity targets in SARS-CoV-2 N protein for serological detection and distinction from SARS-CoV. Comput Biol Med 143:105272. https://doi.org/10.1016/j.compbiomed.2022.105272
Zhai X, Sun J, Yan Z, Zhang J, Zhao J, Zhao Z, Gao Q, He WT, Veit M, Su S (2020) Comparison of severe acute respiratory syndrome coronavirus 2 spike protein binding to ACE2 receptors from human, pets, farm animals, and putative intermediate hosts. J Virol 94(15): e00831-20. https://doi.org/10.1128/JVI.00831-20
Zhang Y, Cheng Y, Xing G, Yu J, Liao A, Du L, Lei J, Lian X, Zhou J, Gu J (2019) Detection and spike gene characterization in porcine deltacoronavirus in China during 2016–2018. Infect Genet Evol 73:151–158. https://doi.org/10.1016/j.meegid.2019.04.023
Zhou P, Fan H, Lan T, Yang XL, Shi WF, Zhang W, Zhu Y, Zhang YW, Xie QM, Mani S, Zheng XS, Li B, Li JM, Guo H, Pei GQ, An XP, Chen JW, Zhou L, Mai KJ, Wu ZX, Li D, Anderson DE, Zhang LB, Li SY, Mi ZQ, He TT, Cong F, Guo PJ, Huang R, Luo Y, Liu XL, Chen J, Huang Y, Sun Q, Zhang XL, Wang YY, Xing SZ, Chen YS, Sun Y, Li J, Daszak P, Wang LF, Shi ZL, Tong YG, Ma JY (2018) Fatal swine acute diarrhoea syndrome caused by an HKU2-related coronavirus of bat origin. Nature 556(7700):255–258. https://doi.org/10.1038/s41586-018-0010-9
Funding
This work was supported by the National Key Research and Development Program of China (grant No. 2021YFD1801101), the Project of Sanya Yazhou Bay Science and Technology City, Grant No: SCKJ-JYRC-2022-08, and the BIC of Bioinformatics Center, Nanjing Agricultural University.
Author information
Authors and Affiliations
Contributions
S S conceived and designed the research. W H and XZ S wrote and formatted the manuscript. W H, XZ S, HF G, SK Z, YT Z, and ZW J performed the experiments. W H, ZW J, and XZ S analyzed all data. All authors have read and approved the manuscript.
Corresponding author
Ethics declarations
Ethics approval
The animal experiment and procedures were approved by the Institutional Animal Care and Use Committee of Nanjing Agricultural University (No. SYXK2017-0007).
Consent to participate
Not applicable.
Consent for publication
Not applicable.
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
He, W., Shi, X., Guan, H. et al. Identification of a novel linear B-cell epitope in porcine deltacoronavirus nucleocapsid protein. Appl Microbiol Biotechnol 107, 651–661 (2023). https://doi.org/10.1007/s00253-022-12348-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-022-12348-5