Abstract
Fucosyllactoses have gained much attention owing to their multiple functions, including prebiotic, immune, gut, and cognition benefits. In this study, human milk oligosaccharide (HMO) 2′-fucosyllactose (α-L-Fuc-(1,2)-D-Galβ-1,4-Glu, 2′FL) and its isomer 3′-fucosyllactose (α-L-Fuc-(1,3)-D-Galβ-1,4-Glu, 3′FL) with potential prebiotic effect were synthesized efficiently by a novel recombinant α-L-fucosidase. An α-L-fucosidase gene (PbFuc) from Pedobacter sp. CAU209 was successfully cloned and expressed in Escherichia coli (E. coli). The deduced amino acid sequence shared the highest identity of 36.8% with the amino sequences of other reported α-L-fucosidases. The purified α-L-fucosidase (PbFuc) had a molecular mass of 50 kDa. The enzyme exhibited specific activity (26.3 U/mg) towards 4-nitrophenyl-α-L-fucopyranoside (pNP-FUC), 3′FL (8.9 U/mg), and 2′FL (3.4 U/mg). It showed the highest activity at pH 5.0 and 35 °C, respectively. PbFuc catalyzed the synthesis of 3′FL and 2′FL through a transglycosylation reaction using pNP-FUC as donor and lactose as acceptor, and total conversion ratio was up to 85% at the optimized reaction conditions. The synthesized mixture of 2′FL and 3′FL promoted the growth of Lactobacillus delbrueckii subsp. bulgaricus NRRL B-548, L. casei subsp. casei NRRL B-1922, L. casei subsp. casei AS 1.2435, and Bifidobacterium longum NRRL B-41409. However, the growths of E. coli ATCC 11775, S. enterica AS 1.1552, L. monocytogenes CICC 21635, and S. aureus AS 1.1861 were not stimulated by the mixture of 2′FL and 3′FL. Overall, our findings suggest that PbFuc possesses a great potential for the specific synthesis of fucosylated compounds.
Key Points | |
• A novel α-L-fucosidase (PbFuc) from Pedobacter sp. was cloned and expressed. | |
• PbFuc showed the highest hydrolysis activity at pH 5.0 and 35 °C, respectively. | |
• It was used for synthesis of 3′-fucosyllactose (3′FL) and 2′-fucosyllactose (2′FL). | |
• The mixture of 3′FL and 2′FL promoted the growth of some Lactobacillus sp. and Bifidobacteria sp. |
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fucosyllactoses (FLs), an essential group of human milk oligosaccharides (HMOs), have gained much attention owing to their various beneficial health effects, such as gut maturation (Vandenplas et al. 2018; Yu et al. 2013), pathogens resistance (Yu et al. 2016), immunity improvement (Zehra et al. 2018; Xiao et al. 2018), and nervous system development (Matthies et al. 1996; Vázquez et al. 2015). They usually have a core lactose, and the fucose residues are attached to the terminal galactose residues through α(1→2) glycosidic linkage, or to the glucose units at the reducing end through α(1→3) glycosidic linkage (Saumonneau et al. 2016). 2′FL synthesized by chemical and biotechnological methods has been enlisted in generally regarded as safe (GRAS) (FDA, 2019) and obtained approval from European Union (NFregulation, European Food Safety Authority) as an ingredient in infant formula, dietary supplements, and medical foods (Bych et al. 2019). Thus, much amount of fucosyllactose is required to meet the market.
Enzymatic production of functional oligosaccharides has acquired great interest owing to the specific synthesis of glycoside linkages and mild reaction conditions (Hayes and Pietruszka 2017). Glycosyltransferases (EC 2.4) and glycosidases (EC 3.2.1) can be used for glycoside synthesis. Currently, glycosidases have been much studied since they often possess high availability and activity when compared with glycosyltransferases (Bojarová and Kren 2011; Zeuner et al. 2014). α-L-Fucosidases (EC 3.2.1.51) are exo-glycoside hydrolases that catalyze the release of L-fucosyl residues from oligosaccharides and glycoconjugates. They are widely distributed in animals, plants, and microorganism (Guzmán-Rodríguez et al. 2019). So far, most of reported α-L-fucosidases are from bacteria, such as Bacteroides sp., Paenibacillus sp., Tannerella sp., Xanthomonas sp., Thermotoga sp., and Bifidobacterium sp. (Lammerts van Bueren et al. 2010; Benešová et al. 2013; Zeuner et al. 2018; Sela et al. 2012; Rodríguez-Díaz et al. 2011). There are few reports on α-L-fucosidases from Pedobacter sp. According to the amino acid sequence similarities, α-L-fucosidases are divided into three glycoside hydrolase (GH) families, viz. GH families 29, 95, and 151 (Henrissat 1991; Lombard et al. 2014; Guzmán-Rodríguez et al. 2018a). GH family 29 α-L-fucosidases are typical retaining enzymes which catalyze the reactions through a two-step double-displacement mechanism. GH family 95 α-L-fucosidases are inverting enzymes catalyzing the reactions by a single-displacement mechanism. The reaction mechanisms of GH family 151 α-L-fucosidases are still unknown. Due to these mechanistic differences, only GH family 29 α-L-fucosidases are potential candidates that show transglycosylation activity. However, those α-L-fucosidases are usually less efficient to synthesize fucosyllactose, such as GH family 29 α-L-fucosidases from Fusarium graminearum (Zeuner et al. 2018), Alcaligenes sp. (Murata et al. 1999), Thermotoga maritima (Guzmán-Rodríguez et al. 2018), and soil-derived metagenome library (Lezyk et al. 2016). Therefore, the discovery of novel α-L-fucosidases with high transfucosylation activities is very important for fucosyllactose synthesis.
3′-fucosyllactose (3′FL) is an isomer of 2′FL and 3FL in human milk oligosaccharides, which has not been found in nature (Takamura et al. 1981; Murata et al. 1999). However, L-Fuc-(1,3)-D-Gal linkage naturally exists as part of human milk oligosaccharides (non-asaccharide) (Yamashita et al. 1976). 3′FL has been successfully synthesized through chemical modification of lactose which is unsuitable for large-scale production because of extreme complexity, harmful chemical reagents, and low product yield (Takamura et al. 1981). Enzymatic synthesis of 3′FL has been carried out by α-L-fucosidase from Alcaligenes sp. with a yield of 34%, using p-nitrophenyl α-L-fucopyranoside and lactose as the substrates (Murata et al. 1999). So far, few studies have been done on the functionalities of 3′FL. Miyauchi et al. have reported that antiserum against 3′-O-a-L-fucosyllactose-p-isothiocyanato-phenethylamine-BSA could preferentially react with adenocarcinoma and embryonal carcinoma cells (Miyauchi et al. 1982). 2′FL and 3FL have been reported to play a critical role in the establishment of a healthy gut microbiome by selectively stimulating the growth of specific Bifidobacterium sp. and Bacteroides sp. (Ashida et al. 2009; Asakuma et al. 2011; Yun et al. 2019) while also suppressing the growth of pathogens such as Clostridium perfringens, Salmonella enterica serovar Typhimurium, Enterobacter cloacae, and Escherichia coli (Yu et al. 2013; Yun et al. 2019). However, the prebiotic effect of 3′FL is unknown.
In this report, a novel α-L-fucosidase gene (PbFuc) from Pedobacter sp. CAU209 was successfully cloned and expressed in E. coli. PbFuc was biochemically characterized and used to synthesize 2′FL and 3′FL. In addition, the prebiotic effect of the mixture of 2′FL and 3′FL towards selected probiotics was investigated.
Materials and methods
Materials and chemicals
E. coli DH5α and BL21 (DE3) from Biomed (Beijing, China) were used for plasmid propagation and as hosts for gene cloning and expression, respectively. pET-28 a (+) was obtained from Novagen (Madison, WI, USA). Restriction endonuclease and T4 DNA ligase were purchased from New England Biolabs (Ipswich, MA, USA). Taq DNA polymerase was obtained from Biolabs (Tokyo, Japan).
4-Nitrophenyl-α-L-fucopyranoside (pNP-FUC) was purchased from Megazyme (Chicago, USA). 2′FL and 3FL were obtained from Elicityl (Crolles, France). β-D-Lactose and fucoidan were purchased from Sigma-Aldrich (Steinheim, Germany). Fructooligosaccharide (FOS, Raftilose P95), degree of polymerization (DP) was 2–7, was obtained from Orafti Group (Tienen, Belgium). Ni2+ Sepharose was obtained from GE Healthcare (Uppsala, Sweden). Bio-Gel P2 was obtained from Bio-Rad Laboratories (Hercules, CA, USA). Selected probiotics used in this study included L. rhamnosus AS 1.2466, L. delbrueckii subsp. bulgaricus NRRL B-548, L. acidophilus NRRL B-4495, L. casei subsp. casei NRRL B-1922, L. brevis NRRL B-4527 (ATCC 14869), L. Rod NRRL B-4391, L. casei subsp. casei AS 1.2435, L. delbrueckii subsp. Lactis AS 1.2132 (JCM 1248), L. reuteri CICC-6132, B. adolescentis ATCC 15703, B. infantis NRRL B-41661, B. breve NRRL B-41408, B. longum NRRL B-41409, and B. bifidum NRRL B-41410. Pathogens including E. ATCC 11775, Samonella enterica AS 1.1552, Listeria monocytogenes CICC 21635, and Staphylococcus aureus AS 1.1861 were used in this study. These probiotics were kept at − 80 °C in MRS broth supplemented with 15% (v/v) glycerol, while pathogens were kept at − 80 °C in Luria-Bertani (LB) medium supplemented with 15% (v/v) glycerol. All other chemicals used were of analytical grade obtained from commercial sources.
Cloning of an α-L-fucosidase gene from Pedobacter sp. CAU209 and sequence analysis
Two primers, PbFuc-F (5′-CCGGAATTCCAGGATTACACACCTACAGCCGC-3′) and PbFuc-R (5′-ATAAGAATGCGGCCGCCTATCCAATCTCCAAAACAATCACCTG-3′), were designed to amplify an α-L-fucosidase gene (PbFuc) from the Pedobacter sp. CAU209 without the signal sequence. The restriction enzyme sites for EcoRI and NotI were underlined and located at the 5′-end and 3′-end, respectively. The PCR mixture (50 μL) contained 10 μL of 5× TransStart FastPfu buffer, 4 μL dNTPs mix (2.5 mM), 1.0 μL Aga16A-up/dn (10 pmol/μL), 1 μL genemic DNA (300 ng), 1 μL 5× TransStart FastPfu DNA polymerase (2.5 U/μL), and 33 μL H2O. PCR conditions were as follows: 5 min at 95 °C, followed by 30 cycles of 20 s at 95 °C, 20 s at 58 °C, and 30 s at 72 °C, with a final extension at 72 °C for 10 min. The PCR product was purified by AxyPrep DNA Gel Extraction Kit (AxyGen, America), and then cloned into pET-28a (+) vector in EcoRI and NotI sites. Subsequently, the recombinant plasmid was transformed into E. coli BL21 (DE3) competent cells.
The potential signal peptide was predicted at the online tool SignalP 4.1 server (Petersen et al. 2011). Nucleotide and predicted amino acid sequences were analyzed with the ExPASy Proteomics tools (Artimo et al. 2012). The multiple sequence alignment analysis of PbFuc and other GH family 29 α-L-fucosidases was carried out by Clustal Omega (Sievers et al. 2014).
Expression and purification of PbFuc
A single E. coli transformant was cultured in LB medium containing 50 μg/mL kanamycin at 37 °C, until the optical density (OD) of the broth at 600 nm reached A600 = 0.6–0.8, then isopropyl-β-D-thiogalactopyranoside (IPTG, 1 mM) was added to induce protein expression, followed by incubation at 20 °C for 16 h. E. coli cells were harvested by centrifugation (10,000 rpm, 5 min, 4 °C) and were re-suspended in buffer A (20 mM Tris-HCl buffer pH 8.0 containing 0.5 M NaCl and 20 mM imidazole) and then lysed by sonication. The supernatant as crude enzyme was collected by centrifugation (11,000 rpm, 20 min, 4 °C), and then loaded on a nickel-iminodiacetic acid column (Ni-IDA: 1 × 5 cm, GE Life Sciences) at 0.5 mL/min. Unbound proteins were washed off with buffer A at 1 mL/min, then the bound proteins were eluted with buffer B (20 mM Tris-HCl buffer pH 8.0, 0.5 M NaCl, 100 mM imidazole). The proteins were collected and dialyzed against 20 mM Tris-HCl buffer pH 7.0 for 12 h. The protein purity was confirmed by SDS-PAGE and the concentrations of protein were quantified by the Lowry method with Bovine serum albumin (BSA) standard (Lowry et al. 1951).
Enzyme assay
Hydrolysis activity of PbFuc was determined using 100 μL of 10 mM pNP-FUC, 100 μL of 50 mM citrate buffer pH 5.0, and 10 μL appropriate diluted enzyme (Paper et al. 2013). The reaction was carried out at 35 °C for 20 min, and stopped by adding 200 μL of 2 M sodium carbonate, then the absorbance was measured at 405 nm. One unit of enzyme activity was defined as the amount of enzyme required to release 1 μmol pNP per minute at the above conditions.
Characterization of the purified PbFuc for hydrolytic activity
The optimal pH of PbFuc was measured by assaying enzyme activity at pH values ranging from pH 3.0 to 12.0 in 50 mM different buffers: citrate buffer (■) pH 3.0–6.0; phosphate buffer (○) pH 6.0–8.0; Tris-HCl buffer (▼) pH 6.0–9.0; 2-cyclohexylaminoethanesulfonic acid (CHES) buffer pH (△) 8.0–10.0; 3-cyclohexylaminopropane sulfonic acid (CAPS) buffer (*) pH 10.0–11.0; and Na2HPO4-NaOH buffer (●) pH 11.0–12.0. The pH stability of PbFuc was evaluated by incubating the enzyme in different buffers at 25 °C for 30 min and assaying residual activity under the standard conditions. The optimal temperature of PbFuc was determined by measuring enzyme activity at different temperatures from 25 to 50 °C. Thermostability of PbFuc was studied by assaying residual activity after incubation of the enzyme at 35 to 50 °C for 30 min. The thermal denaturing half-life of the enzyme was evaluated at 35, 40, and 45 °C for 4 h, samples were withdrawn at different intervals, and residual activities of treated enzymes were determined by the standard assay.
The impact of metal ions and other chemical reagents on the hydrolysis activity of PbFuc was assayed after incubation of the enzyme in the presence of Ag+, Ca2+, Fe3+, Cr3+, Co2+, Mg2+, Mn2+, Zn2+, Cu2+, Ni2+, Na+, K+, Ba2+, EDTA, β-mercaptoethanol, and SDS. The enzyme was incubated in 50 mM citrate buffer pH 5.0 with 1 mM metal ions and reagents at 25 °C for 30 min. The residual activities were then measured according to the standard method; the enzyme without any treatment was used as control.
Substrate specificity of PbFuc was determined by measuring the enzyme activity using 100 μL of 10 mM different substrates (pNP-FUC, 2′FL, 3FL, 3′FL, xyloglucan, and fucoidan), 100 μL of 50 mM citrate buffer pH 5.0, and 10 μL enzyme. The reactions were carried out at 35 °C for 20 min, and stopped by boiling for 10 min. The amount of released fucose was analyzed by HPLC (Agilent HPLC-1260 Infinity) equipped with BP-800 Pb2+ column at a flow rate of 0.8 mL/min at 80 °C, and was monitored with a refractive index detector (RID) at 35 °C. One unit of enzyme activity was defined as the amount of enzyme required to release 1 μmol fucose per minute at the above conditions. Xyloglucan (XyG) from apple was prepared from NaOH extracts of apple cell walls (Renard et al. 1992).
The kinetic parameters of PbFuc towards pNP-FUC were determined by measuring the enzyme activities with different concentrations of the substrate in 50 mM citrate buffer (pH 5.0) at 35 °C for 5 min. The Vmax and Km values were calculated using the Graphpad Prism program 7.04.
Transglycosylation activity of PbFuc
The transglycosylation activity of PbFuc was explored using 10 mM pNP-FUC as glycosyl donor, 0.5 U/mL purified PbFuc, and 50 mM different saccharides as acceptors (glucose, galactose, fructose, arabinose, fucose, ribose, xylose, rhamnose, glucuronic acid, galacturonic acid, maltose, or lactose). The reactions were carried out in 50 mM citrate buffer (pH 5.0) at 35 °C for 3 h and stopped by boiling for 10 min. The results were analyzed by thin-layer chromatography (TLC), which was performed on Silica gel 60 (Merck, Germany) plates. The samples were applied to the plates and developed twice with 2:1:1 n-butanol-acetic-water aqueous. Samples on the TLC plate were visualized by spraying with 5% H2SO4 in methanol and heating at 120 °C.
Isolation, identification, and synthesis optimization of transglycosylation products
To further identify the synthesized saccharides by PbFuc from pNP-FUC and lactose, products were purified by a Bio-gel P2 column (1.2 × 110 cm, MQ water; 0.3 mL/min, Pharmacia, Germany); oligosaccharide fractions were collected after analysis by TLC, then concentrated and freeze-dried. The purity of oligosaccharide was analyzed by HPLC; HPLC analysis was performed using an Agilent-1260 Infinity system equipped with a refractive index detector (RID) and waters XBridge amide column (3.5 μm, 4.6 × 250 mm). The column was maintained at 45 °C and eluted with acetonitrile-water mobile phases (72/28, v/v) at a flow rate of 0.5 mL/min. To determine the molecular weight of purified product, mass spectra (MS) analysis was performed through Thermo Scientific™ Q Exactive™ equipped with positive-ion mode infusion/offline electrospray ionization (ESI). The structure of purified oligosaccharide was analyzed by NMR. Briefly, the sample was dissolved in D2O and then transferred into a NMR tube. One-dimensional spectra were recorded on a 500 MHz Varian VNMR SYSTEM™ at 298 K at 500 MHz for 1H and 125 MHz for 13C, respectively. Chemical shifts were expressed in parts per million (ppm) referenced to DSS (4,4-dimethyl-4-silapentane-1-sulfonic acid).
To optimize the synthesis of 3′FL and 2′FL, the reaction conditions including temperature, pH, enzyme dosage, reaction time, and lactose concentration were investigated. The effect of temperature on the synthesis of 3′FL and 2′FL was determined in 50 mM citrate buffer (pH 5.0) for 3 h at 20–50 °C. The effect of pH on the synthesis of 3′FL and 2′FL was evaluated at 35 °C using different buffers (pH 4.0–10.0). The effect of enzyme dosage on the synthesis of 3′FL and 2′FL was studied from 0.05 to 5 U/mL. The influence of reaction time on the synthesis of 3′FL and 2′FL was studied at 0.5 to 24 h, respectively. Finally, the effect of different lactose concentration (0.05–1.0 M) was evaluated for the synthesis of 3′FL and 2′FL.
The reaction was terminated by boiling for 10 min and samples were filtered by 0.22 μm filter, then the products were analyzed by a HPLC system (Agilent-1260 Infinity) equipped with a refractive index detector (RID) and waters XBridge amide column (3.5 μm, 4.6 × 250 mm). The column was maintained at 45 °C and eluted with acetonitrile-water mobile phases (72/28, v/v) at a flow rate of 0.5 mL/min. The conversion ratio (yield) represented the ratio of the amount of the mixture of 3′FL and 2′FL (mM) to the amount of donor (mM, pNP-FUC).
Growth test of probiotics and pathogens using the mixture of 3′FL and 2′FL as the sole carbon source
To obtain suitable amount of 3′FL and 2′FL, the synthesis reaction (200 mL) was carried out at the optimized conditions. The mixture of 3′FL and 2′FL was collected, concentrated, and freeze-dried.
To investigate the prebiotic effect of the mixture of 3′FL and 2′FL, nine Lactobacillus sp. and five Bifidobacterium sp. strains were incubated in Man, Rogosa, and Sharpe (MRS) medium without glucose supplemented with 1% (w/v) the mixture of 3′FL and 2′FL, 2′FL, fructooligosaccharide (FOS, DP 2–7, Raftilose P95), or pure water (no sugar) as the sole carbon source. Fermentation broth was prepared in 96 Microwell Plate with a total volume of 250 μL in each well containing 100 μL bacteria in MRS medium without glucose, 100 μL of 2% carbohydrate solution (w/v) or pure water, and 50 μL of mineral oil (Hoeflinger et al. 2015). The anaerobic fermentation was performed at 37 °C for 72 h. To study the influence of the mixture of 3′FL and 2′FL on the growth of pathogens, E. coli ATCC 11775, S. enterica AS 1.1552, L. monocytogenes CICC 21635, and S. aureus AS 1.1861 were incubated in the M9 broth medium at 37 °C for 48 h supplemented with 1% (w/v) of the mixture of 3′FL and 2′FL, 2′FL, FOS, or pure water (Yun et al. 2019). To measure the growth of bacteria, optical density (OD) of each culture media was recorded at 595 nm (OD595) with an automated microplate reader (Multiskan FC, Thermo Electron Corporation) at different times during incubation.
Accession number
The gene sequence (PbFuc) is available at the GenBank database under accession number MN902190.
Results
Cloning and sequence analysis of PbFuc
An α-L-fucosidase gene (PbFuc) was amplified from the genomic DNA of Pedobacter sp. CAU209. It is 1326 bp in length and encodes a protein of 441 amino acids. The molecular mass and isoelectric point of deduced protein were predicted to be 50.46 kDa and 8.58, respectively. The amino acid sequence of PbFuc was compared with different GH family 29 α-L-fucosidases (Fig. 1). PbFuc shared relatively high identity (36.8%) with the α-L-fucosidase isoenzyme 1 (α-L-f1wt, PDB entry 6GN6) from P. thiaminolyticus (Benešová et al. 2013), followed by the α-L-fucosidases from Homo sapiens (31.6%, GenBank accession number P04066, Ohira et al. 2003), H. sapiens (31.3%, GenBank accession number Q9BTY2, Clark et al. 2003), Macaca fascicularis (31.1%, GenBank accession number Q60HF8, Fisher and Aronson 1989), and B. thetaiotaomicron (BtFuc2970, 29.6%, PDB entry 2WVT, Lammerts van Bueren et al. 2010). Two conserved glutamic acid (E260) and aspartic acid (D204) residues were considered the catalytic residues in PbFuc (Shaikh et al. 2013). The results indicated that PbFuc should be a novel α-L-fucosidase.
Multiple amino acid sequence alignment of PbFuc with six GH family 29 α-L-fucosidases. Numbers on the left are the residue numbers of first amino acid for each line. Identical and similar amino acids are shaded black and gray, respectively. Two catalytic residues—glutamic acid residue (E260) and aspartic acid residue (D204)—are marked by black stars. Listed sequences are the α-L-fucosidases from Pedobacter sp. CAU209 (PbFuc), Homo sapiens (UniProtKB/Swiss-Prot: Q9BTY2), H. sapiens (UniProtKB/Swiss-Prot: P04066), Macaca fascicularis (UniProtKB/Swiss-Prot: Q60HF8), Rattus norvegicus (UniProtKB/Swiss-Prot: P17164), P. thiaminolyticus (α-L-f1wt, PDB code: 6GN6), and B. thetaiotaomicron VPI-5482 (BtFuc2970, PDB code: 2WVT)
Expression and purification of PbFuc
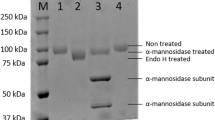
The α-L-fucosidase gene (PbFuc) was successfully expressed in E. coli BL21 (DE3) as a soluble intracellular protein. The enzyme was purified by His-tag Ni-IDA affinity chromatography with a yield of 47.1% (Table 1). SDS-PAGE analysis of purified protein showed that the molecular mass of PbFuc was approximately 50 kDa (Fig. 2). The specific activity of purified PbFuc was 26.3 U/mg using pNP-FUC as substrate (Table 1).
Characterization of the purified PbFuc for hydrolytic activity
PbFuc was optimally active at pH 5.0 in 50 mM citrate buffer (Fig. 3a), and it exhibited broad pH stability from pH 4.0 to 11.0 (Fig. 3b). The optimal temperature of PbFuc was 35 °C (Fig. 3c) and the enzyme was stable up to 40 °C (Fig. 3d). The half-lives of PbFuc at 35 °C, 40 °C, and 45 °C were determined to be 555 min, 51 min, and 2.8 min, respectively (Fig. 3e). The substrate specificity of PbFuc towards different glycosides with fucose residues was examined (Table 2). The enzyme showed high specific activity towards pNP-FUC (26.3 U/mg). It also exhibited hydrolysis activity towards 3′FL (8.3 U/mg) and 2′FL (3.0 U/mg), while PbFuc did not show detectable activity towards 3FL, XyG, and fucoidan.
Optimal pH (a), pH stability (b), optimal temperature (c), thermostability (d), and thermal denaturing half-lives (e) of the purified α-L-fucosidase from Pedobacter sp. CAU209. Symbols for optimal pH and pH stability: citrate buffer (■) pH 4.0–6.0; phosphate buffer (○) pH 6.0–8.0; Tris-HCl buffer (▼) pH 6.0–9.0; CHES buffer (△) pH 8.0–10.0; CAPS buffer (*) pH 10.0–11.0, Na2HPO4-NaOH buffer (●) pH 11.0–12.0. Symbols for thermal inactivation: 35 °C (●), 40 °C (▲), 45 °C (□). All experiments were performed with three parallels
The effect of various metal ions and chemical reagents on the activity of PbFuc was studied (Table 3). Ag+ completely inhibited the enzyme activity, whereas Ca2+, Fe3+, Cr3+, Co2+, EDTA, β-mercaptoethanol, and SDS inhibited enzyme activity by 15.9%, 13.6%, 10.9%, 10.7%, 14.1%, 14.9%, and 25.9%, respectively. Mg2+, Mn2+, Zn2+ Cu2+, Ni2+, Na+, K+, and Ba2+ did not influence the enzyme activity of PbFuc. The kinetic parameters Km and Vmax of PbFuc towards pNP-FUC were determined to be 0.77 mM and 27.8 μmol/min/mg, respectively (data not shown).
Transglycosylation activity of PbFuc
The transglycosylation activity of PbFuc was evaluated using pNP-FUC as donor and ten sugars as acceptors. PbFuc demonstrated its ability to catalyze the transglycosylation of some acceptors including glucose, galactose, arabinose, ribose, xylose, lactose, and maltose to yield new oligosaccharides detected on TLC (Fig. S1).
Fucosyllactose synthesis, purification, and identification
Among various fucosyl-oligosaccharides synthesized by PbFuc, the oligosaccharides produced using lactose as acceptor were analyzed by HPLC (Fig. 4a), then purified and analyzed by MS and NMR. The positive-ion electrospray ionization (ESI) mass spectrum of purified oligosaccharides showed a peak of [M+Na]+ ion at m/z value of 511.1611 (Fig. 4b), which is consistent with the molecular weight of fucosyllactose (488.44). 1H NMR (Fig. 4c) and 13C NMR (Fig. S2) spectrometric data for the product suggested that the oligosaccharides were composed of 3′-fucosyllactose (α-L-Fuc-(1,3)-D-Galβ-1,4-Glu, 83%) and 2′-fucosyllactose (α-L-Fuc-(1,2)-D-Galβ-1,4-Glu,17%).
HPLC chromatogram of the transglycosylation reaction products with PbFuc (a). MS spectra of the purified oligosaccharides synthesized by PbFuc (b). Mass spectra was recorded on an instrument (Thermo Scientific™ Q Exactive™) equipped with an ESI source in positive-ion mode. One-dimensional 1H NMR spectra of oligosaccharides synthesized by PbFuc (c). One-dimensional 1H and NMR spectra were performed on a 500 MHz Varian VNMR SYSTEM™ at 298 K. Chemical shifts are expressed in ppm referenced to DSS (4,4-dimethyl-4-silapentane-1-sulfonic acid)
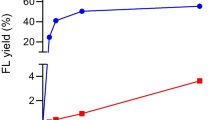
The optimal temperature, pH, enzyme dosage, reaction time, and lactose concentration on the 3′FL and 2′FL synthesis were investigated (Fig. 5). The results indicated that the temperature did not show obvious effect on the synthesis of 2′FL and 3′FL (Fig. 5a), whereas pH, enzyme dosage, reaction time, and lactose concentration remarkably affected the synthesis of 2′FL and 3′FL (Fig. 5b–e). The optimized conditions for fucosyllactose synthesis by PbFuc were 25 to 40 °C (Fig. 5a), pH 8.5 (Fig. 5b), 0.5 U/mL (Fig. 5c), 3 h (Fig. 5d), and 700 mM (Fig. 5e), respectively. At the optimized conditions, 85% total conversion ratio of 3′FL (70.5%) and 2′FL (14.5%) from pNP-FUC was achieved, respectively.
Effects of temperature (a), pH (b), enzyme dosage (c), reaction time (d), and lactose concentrations (e) on the synthesis of 3′FL and 2′FL by PbFuc. The effect of temperature on the synthesis of 3′FL and 2′FL was determined at 20–50 °C in 50 mM citrate buffer pH 5.0. The effect of pH on the synthesis of 3′FL and 2′FL was determined at 35 °C using different buffers with pH ranging from 4.0 to 10.0. The effect of enzyme dosage (0.05–5 U/mL) on the synthesis of 3′FL and 2′FL was determined in 50 mM Tris-HCl buffer pH 8.5 at 35 °C. The effect of reaction time on the synthesis of 3′FL and 2′FL was determined in 50 mM Tris-HCl buffer (pH 8.5) with PbFuc (0.5 U/mL); samples were withdrawn at different times within 24 h. The effect of lactose concentrations (0.05–1 M) on the synthesis of 3′FL and 2′FL was determined in 50 mM Tris-HCl buffer (pH 8.5) at 35 °C with 0.5 U/mL PbFuc
Growth test of probiotics and pathogens using the mixture of 3′FL and 2′FL as carbon source
Among the tested probiotics, three Lactobacillus and one Bifidobacterium strains exhibited higher biomass concentration (OD595) in the mixture of 3′FL and 2′FL medium than those of FOS and non-sugar control. The optical densities at 595 nm of L. casei subsp. casei AS 1.2435 (Fig. 6a), L. casei subsp. casei NRRL B-1922 (Fig. 6b), L. delbrueckii subsp. bulgaricus NRRL B-548 (Fig. 6c), and B. longum NRRL B-41409 (Fig. 6d) were 0.362, 0.375, 0.467, and 0.403 at 48 h in the mixture of 3′FL and 2′FL medium, respectively, but were 0.290, 0.320, 0.182, and 0.333 at 48 h in the FOS medium, respectively. Other probiotic strains, including L. rhamnosus AS 1.2466, L. acidophilus NRRL B-4495, L. brevis NRRL B-4527 (ATCC 14869), L. rod NRRL B-4391, L. delbrueckii subsp. Lactis AS 1.2132 (JCM 1248), L. reuteri CICC-6132, B. infantis NRRL B-41661, B. adolescentis ATCC 15703, and B. breve NRRL B-41408, grew poorly in the mixture of 3′FL and 2′FL medium, whereas B. bifidum NRRL B-41410 did not grow in the mixture of 3′FL and 2′FL medium (data not shown). While supplementation of 2′FL in the MRS medium promoted the growth of L. casei subsp. casei AS 1.2435 (Fig. S3a), and weakly promoted the growths of B. longum NRRL B-41409 (Fig. S3d), L. rhamnosus AS 1.2466, L. reuteri CICC-6132, and B. adolescentis ATCC 15703 (data not shown), other probiotic strains almost did not grow in the 2′FL medium (data not shown).
Cell growth of L. casei subsp. casei AS 1.2435 (a), L. casei subsp. casei NRRL B-1922 (b), L. delbrueckii subsp. bulgaricus NRRL B-548 (c), and B longum NRRL B-41409 (d) with the mixture of 3′FL and 2′FL (●), or FOS (□), or no sugar (○) as the sole carbon source. Cell density (OD595) was monitored. All experiments were performed with three parallels
For pathogenic strains, none of four pathogens could utilize the mixture of 3′FL and 2′FL (Fig. 7) or 2′FL (Fig. S4). The optical densities at 595 nm of E. coli ATCC 11775 (Fig. 7a), S. enterica AS 1.1552, (Fig. 7b), L. monocytogenes CICC 21635 (Fig. 7c), and S. aureus AS 1.1861 (Fig. 7d) were 0.091, 0.074, 0.086, and 0.095 after incubation in the mixture of 3′FL and 2′FL medium for 24 h, while were 0.179, 0.186, 0.199, and 0.109 when incubated in the FOS medium for 24 h, and were 0.091, 0.085, 0.085, and 0.089 when incubated with non-sugar MRS broth for 24 h. Also, the optical densities at 595 nm of four pathogens in 2′FL medium were close to non-sugar groups (Fig. S4).
Cell growth of E. coli ATCC 11775 (a), S. enterica AS 1.1552 (b), L. monocytogenes CICC 21635 (c), and S. aureus AS 1.1861 (d) with the mixture of 3′FL and 2′FL (●), or FOS (□), or no sugar (○) as the sole carbon source. Cell density (OD595) was monitored. All experiments were performed with three parallels
Discussion
Enzymatic production of fucosyllactoses has acquired much interest owing to the specific synthesis of glycoside linkages and mild reaction conditions (Hayes and Pietruszka 2017). However, the reported α-L-fucosidases are usually less efficient due to the hydrolysis activity towards synthesized products (Guzmán-Rodríguez et al. 2019). Thus, the discovery of novel α-L-fucosidases with high transfucosylation activities is very important for fucosyllactose synthesis. In this study, we described a novel α-L-fucosidase (PbFuc) from Pedobacter sp. CAU209 and the synthesis of 2′FL and 3′FL by an efficient enzymatic method. Also, the prebiotic effect of the mixture of 2′FL and 3′FL was investigated.
A putative α-L-fucosidase candidate gene (PbFuc) was cloned from Pedobacter sp. CAU209; the amino acid sequence of PbFuc shared relatively high similarities with several known GH family 29 α-L-fucosidases, especially with the highest identity of 36.8% to that of the α-L-fucosidase (α-L-f1wt) from P. thiaminolyticus (Fig. 1), suggesting that it should be a new member of GH family 29 α-L-fucosidases. It has been reported that GH 29 family α-L-fucosidases have two catalytic residues, including a highly conserved nucleophile Asp residue and a structurally conserved general acid/base Glu residue (Sulzenbacher et al. 2004; Lammerts van Bueren et al. 2010; Sakurama et al. 2012). In PbFuc, aspartic acid (D204) and glutamic acid (E260) were predicted to be the putative nuclophilic and acid/base residues, respectively (Fig. 1). Based on the phylogenetic relationship clustering and substrate specificity, the GH family 29 has been further divided into two subgroups (GH 29A and GH 29B). GH family 29A α-L-fucosidases usually have relatively diverse substrate specificities, while GH 29B α-L-fucosidases display specific activity towards α-1,3/4 linkages (Cao et al. 2014; Lezyk et al. 2016; Zeuner et al. 2018). PbFuc exhibited hydrolysis activity towards pNP-FUC and α-1,2/3 linkages, indicating that it should belong to GH29A, which is similar to several α-L-fucosidases from Alcaligenes sp., X. campestris, T. maritima, and T. forsythia (Zeuner et al. 2018).
Generally, most of GH 29A α-L-fucosidases have relatively low molecular masses. PbFuc had a molecular mass of 50 kDa determined by SDS-PAGE (Fig. 2), which is similar to some bacterial α-L-fucosidases, such as TfFuc1 from T. forsythia (51 kDa), TmαFuc from T. maritima (52 kDa), and iso1 from P. thiaminolyticus (51.2 kDa) (Zeuner et al. 2018; Benešová et al. 2013). The optimal pH of PbFuc was 5.0 (Fig. 3a), which is comparable with α-L-fucosidases from F. oxysporum strain 0685 (Paper et al. 2013), and X. campestris (Dupoiron et al. 2015). PbFuc was stable within a wide pH range 4.0–11.0 (Fig. 3b), which is more stable than most of the reported α-L-fucosidases from B. thetaiotaomicron (pH 4–9, Sakurama et al. 2012), F. oxysporum strain 0685 (pH 4–6, Paper et al. 2013), and Thermus sp. strain Y5 (pH 4–7.5, Eneyskaya et al. 2001). Typically, bacterial α-L-fucosidases have optimal temperature of 30-45 °C. PbFuc showed optimal activity at 35 °C (Fig. 3c), which is similar to those of α-L-fucosidases from B. thetaiotaomicron (37 °C, Sakurama, et al. 2012), and X. campestris pv. campestris (37 °C, Dupoiron et al. 2015). The Km value of PbFuc for pNP-FUC was 0.72 mM, which is lower than that of α-L-fucosidase from B. thetaiotaomicron (BT 2970, 1.5 mM, Akurama et al. 2012), whereas is higher than those of the α-L-fucosidases (iso1 and iso2) from P. thiaminolyticus (0.44 and 0.52 mM, Benešová et al. 2013, 2015), Mfuc5 from soil-derived metagenome library (0.13 mM, Lezyk et al. 2016), and TmαFuc from T. maritima (0.034 mM, Zeuner et al. 2018).
Many GH family 29 α-L-fucosidases with transglycosylation activities can be used to produce fucosylated oligosaccharides (Liu et al. 2016). Recently, several natural fucosyl substrates have been explored as donors in enzymatic synthesis of fucosyllactose, such as fucosylated XyG, fucoidan, and fucosylated chondroitin sulfate (Guzmán-Rodríguez et al. 2019). However, only few GH family 29 α-L-fucosidases possessed the ability to transfer fucose from XyG to lactose, and the product yields were low, such as α-L-fucosidase FgFCO1 from F. graminearum (Zeuner et al. 2018) and Mfuc5 from a soil metagenome (Lezyk et al. 2016). In our study, PbFuc displayed no hydrolytic activity towards XyG or fucoidan (Table 2). Therefore, the transglycosylation activity of PbFuc was only studied using pNP-FUC. PbFuc was capable of catalyzing transglycosylation reaction using pNP-FUC as donor and different sugars as acceptors (Fig. S1). The transglycosylation property is similar to several α-L-fucosidases from P. thiaminolyticus (Benešová et al. 2013, 2015), Alcaligenes sp. (Murata et al. 1999), and Thermus sp. strain Y5 (Eneyskaya et al. 2001). Especially, PbFuc was efficient in synthesizing fucosyllactoses when pNP-FUC and lactose were used, 85% of pNP-FUC was converted into 3′FL and 2′FL under the optimized conditions (Fig. 5). The transglycosylation efficiency for 2′FL synthesis (14.5%) by PbFuc is close to that of α-L-fucosidase FgFCO1 from F. graminearum (14%, Zeuner et al. 2018), higher than that of α-L-fucosidase from T. maritima (6.4%), T. forsythia (0.7%), and soil-derived metagenome library (Mfuc2, 0.4%, Lezyk et al. 2016). And also its transglycosylation efficiency for 3′FL synthesis (70.5%) is much higher than that of α-L-fucosidases from Alcaligenes sp. (34%, Murata et al. 1999). In addition, the transglycosylation efficiency (85%) for total fucosyllactose synthesis by PbFuc is higher than that of most reported wild-type α-L-fucosidases, such as α-L-fucosidases from T. maritima (32.5%, Guzmán-Rodríguez et al. 2018 b), L. rhamnosus GG (25% Escamilla-Lozano et al. 2019), soil-derived metagenome library (3.9%, Lezyk et al. 2016), and is comparable with that of α1,2-L-fucosynthase (AfcA) mutants, using β-L-fucosyl fluoride and lactose to produce 2′FL with a yield of 88% (Sugiyama et al. 2016). So far, fucosyllactoses including 2′FL, 3FL, and 2′,3-difucosyllactose (DFL) have been successfully produced by cell factory approaches (Faijes et al. 2019). However, 3′FL has never been produced by cell factory approach since the lack of suitable glycosyltransferase for the specific synthesis of Fuc-α1,3-Gal linkage in vivo. Thus, compared with the cell factory methods and other reported α-L-fucosidases, PbFuc possessed a unique advantage in synthesizing 3′FL due to its efficient transglycosylation activity.
Fucosyl-oligosaccharides such as 2′FL, 3FL, and DFL have promoted the growth of some Bifidobacterium spp. and Lactobacillus spp. (Yu et al. 2013). Also many in vitro studies suggested that 2′FL promote growth of certain Bifidobacteria sp., such as B. longum subsp. Infantis (LoCascio et al. 2007). However, the prebiotic effect of 3′FL still remains unknown. Supplementation of the mixture of 3′FL and 2′FL in the media significantly promoted the growth of B. longum NRRL B-41409, L. delbrueckii subsp. bulgaricus NRRL B-548, L. casei subsp. casei NRRL B-1922, and L. casei subsp. casei AS 1.2435, whereas supplementation of 2′FL in the MRS medium only promoted the growth of L. casei subsp. casei AS 1.2435 (Fig. S3a), and weakly promoted the growths of B. longum NRRL B-41409 (Fig. S3d). The results indicated that the mixture of 3′FL and 2′FL synthesized in this study possessed a broader proliferative effect on the selected probiotics (Fig. 6) when compared with commercial 2′FL (Fig. S3). Recently, several GH families 29 and 95 α-L-fucosidases from B. longum sp. and L. casei sp. have been demonstrated to be related to the fucosylated HMOs metabolism (LoCascio et al. 2007; Sela et al. 2012; Rodríguez-Díaz et al. 2011). Furthermore, growth curves of four pathogenic bacteria in the mixture of 3′FLand 2′FL medium (Fig. 7) and 2′FL medium (Fig. S4) are close to non-sugar control, suggesting that these pathogens could not utilize 3′FL or 2′FL. This finding is in agreement with previous conclusion that 2′FL, 3FL, and other fucosylated HMOs are not considered ideal energy resource for broad-spectrum pathogenic bacteria (Yu et al. 2013; Hoeflinger et al. 2015). Thus, the mixture of 3′FL and 2′FL synthesized in our study can be used as ideal prebiotic for certain beneficial bacteria and gut maturation.
In conclusion, a novel α-L-fucosidase gene (PbFuc) from the Pedobacter sp. CAU209 was cloned and heterologously expressed in E. coli. PbFuc efficiently synthesized 3′FL and 2′FL using pNP-FUC and lactose owning to its transfucosylation activity. A high total conversion ratio of 85% was obtained under the optimized reaction conditions. Metabolic characteristics of the mixture of 3′FLand 2′FL in bacterial growth assay illustrated its potential to improve gut microbial environment. Thus, this work furnishes a novel and efficient enzymatic strategy to produce valuable functional fucosylated oligosaccharide.
References
Artimo P, Jonnalagedda M, Arnold K, Baratin D, Csardi G, de Castro E, Duvaud S, Flegel V, Fortier A, Gasteiger E, Grosdidier A, Hernandez C, Ioannidis V, Kuznetsov D, Liechti R, Moretti S, Mostaguir K, Redaschi N, Rossier G, Xenarios I, Stockinger H (2012) ExPASy: SIB bioinformatics resource portal. Nucleic Acids Res 40:W597–W603. https://doi.org/10.1093/nar/gks400
Asakuma S, Hatakeyama E, Urashima T, Yoshida E, Katayama T, Yamamoto K, Kumagai H, Ashida H, Hirose J, Kitaoka M (2011) Physiology of consumption of human milk oligosaccharides by infant gut-associated bifidobacteria. J Biol Chem 286:34583–34592. https://doi.org/10.1074/jbc.M111.248138
Ashida H, Miyake A, Kiyohara M, Wada J, Yoshida E, Kumagai H, Katayama T, Yamamoto K (2009) Two distinct α-L-fucosidases from Bifidobacterium bifidum are essential for the utilization of fucosylated milk oligosaccharides and glycoconjugates. Glycobiology 19:1010–1017. https://doi.org/10.1093/glycob/cwp082
Benešová E, Lipovová P, Dvořáková H, Králová B (2013) α-L-Fucosidase from Paenibacillus thiaminolyticus: its hydrolytic and transglycosylation abilities. Glycobiology 23:1052–1065. https://doi.org/10.1093/glycob/cwt041
Benešová E, Lipovová P, Krejzová J, Kovaľová T, Buchtová P, Spiwok V, Králová B (2015) Alpha-L-fucosidase isoenzyme iso2 from Paenibacillus thiaminolyticus. BMC Biotechnol 15:36. https://doi.org/10.1186/s12896-015-0160-x
Bojarová P, Kren V (2011) Glycosidases in carbohydrate synthesis: when organic chemistry falls short. Chimia (Aarau) 65:65–70
Bych K, Miks MH, Markus TJ, Hederos J, Vigsnæs LK, Becker P (2019) Production of HMOs using microbial hosts—from cell engineering to large scale production. Curr Opin Biotechnol 56C:130–137. https://doi.org/10.1016/j.copbio.2018.11.003
Cao HN, Walton JD, Brumm P, Phillips GN (2014) Structure and substrate specificity of a eukaryotic fucosidase from Fusarium graminearum. J Biol Chem 289:25624–25638. https://doi.org/10.1074/jbc.M114.583286
Clark HF, Gurney AL, Abaya E, Baker K, Baldwin D, Brush J, Chen J, Chow B, Chui C, Crowley C, Currell B, Deuel B, Dowd P, Eaton D, Foster J, Grimaldi C, Gu Q, Hass PE, Heldens S, Huang A, Kim HS, Klimowski L, Jin Y, Johnson S, Lee J, Lewis L, Liao D, Mark M, Robbie E, Sanchez C, Schoenfeld J, Seshagiri S, Simmons L, Singh J, Smith V, Stinson J, Vagts A, Vandlen R, Watanabe C, Wieand D, Woods K, Xie MH, Yansura D, Yi S, Yu G, Yuan J, Zhang M, Zhang Z, Goddard A, Wood WI, Godowski P, Gray A (2003) The secreted protein discovery initiative (SPDI), a large-scale effort to identify novel human secreted and transmembrane proteins: a bioinformatics assessment. Genome Res 13(10):2265–70. https://doi.org/10.1101/gr.1293003
Dupoiron S, Zischek C, Ligat L, Carbonne J, Boulanger A, Dugé de Bernonville T, Lautier M, Rival P, Arlat M, Jamet E, Lauber E, Albenne C (2015) The N-Glycan cluster from Xanthomonas campestris pv. Campestris. J Biol Chem 290(10):6022-36. https://doi.org/10.1074/jbc.M114.624593
Eneyskaya EV, Kulminskaya AA, Kalkkinen N, Nifantiev NE, Arbatskii NP, Saenko A, Chepurnaya OV, Arutyunyan AV, Shabalin KA, Neustroev KN (2001) An α-L-fucosidase from Thermus sp. with unusually broad specificity. Glycoconj J 18:827–834
Escamilla-Lozano Y, Guzmán-Rodríguez F, Alatorre-Santamaría S, García-Garibay M, Gómez-Ruiz L, Rodríguez-Serrano G, Cruz-Guerrero A (2019) Synthesis of fucosyl-oligosaccharides using α-l-fucosidase from Lactobacillus rhamnosus GG. Molecules 24(13):2402. https://doi.org/10.3390/molecules24132402
Fisher KJ, Aronson NN Jr (1989) Isolation and sequence analysis of a cDNA encoding rat liver alpha-L-fucosidase. Biochem J 264(3):695-701. https://doi.org/10.1042/bj2640695
Guzmán-Rodríguez F, Alatorre-Santamaría S, Gómez-Ruiz L, Rodríguez-Serrano G, García-Garibay M, Cruz-Guerrero A (2018) Synthesis of a fucosylated trisaccharide via transglycosylation by α-L-fucosidase from Thermotoga maritima. Appl Biochem Biotechnol 186:681–691. https://doi.org/10.1007/s12010-018-2771-x
Guzmán-Rodríguez F, Alatorre-Santamaría S, Gómez-Ruiz L, Rodríguez-Serrano G, García-Garibay M, Cruz-Guerrero A (2019) Employment of fucosidases for the synthesis of fucosylated oligosaccharides with biological potential. Biotechnol Appl Biochem 66:172–191. https://doi.org/10.1002/bab.1714
Hayes MR, Pietruszka J (2017) Synthesis of glycosides by glycosynthases. Molecules 22:1434. https://doi.org/10.3390/molecules22091434
Henrissat B (1991) A classification of glycosyl hydrolases based on amino acid sequence similarities. Biochem J 280:309–316. https://doi.org/10.1042/bj2800309
Hoeflinger JL, Davis SR, Chow J, Miller MJ (2015) In vitro impact of human milk oligosaccharides on Enterobacteriaceae growth. J Agric Food Chem 63:3295–3302. https://doi.org/10.1021/jf505721p
Lammerts van Bueren A, Ardèvol A, Fayers-Kerr J, Luo B, Zhang Y, Sollogoub M, Blériot Y, Rovira C, Davies GJ (2010) Matthieu sollogoub analysis of the reaction coordinate of α-L-Fucosidases: a combined structural and quantum mechanical approach. J Am Chem Soc 132:1804–1806. https://doi.org/10.1021/ja908908q
Lezyk M, Jers C, Kjaerulff L, Gotfredsen CH, Mikkelsen MD, Mikkelsen JD (2016) Novel α-L-fucosidases from a soil metagenome for production of fucosylated human milk oligosaccharides. PLoS One 11:1–18. https://doi.org/10.1371/journal.pone.0147438
Liu S, Kulinich A, Cai Z, Ma HY, Du YM, Lv YM, Liu L, Voglmeir J (2016) The fucosidase-pool of Emticicia oligotrophica: biochemical characterization and transfucosylation potential. Glycobiology 26:871–879. https://doi.org/10.1093/glycob/cww030
LoCascio RG, Ninonuevo MR, Freeman SL, Sela DA, Grimm R, Lebrilla CB, Mills DA, German JB (2007) Glycoprofiling of bifidobacterial consumption of human milk oligosaccharides demonstrates strain specific, preferential consumption of small chain glycans secreted in early human lactation. J Agric Food Chem 55:8914–8919. https://doi.org/10.1021/jf0710480
Lombard V, Golaconda RH, Drula E, Coutinho PM, Henrissat B (2014) The carbohydrate-active enzymes database (CAZy) in 2013. Nucleic Acids Res 42:D490–D495. https://doi.org/10.1093/nar/gkt1178
Lowry OH, Rosebrough NJ, Farr AL, Randall RJ (1951) Protein measurement with the folin phenol reagent. J Biol Chem 193:265–275
Matthies H, Staak S, Krug M (1996) Fucose and fucosyllactose enhance in-vitro hippocampal long-term potentiation. Brain Res 725:276–280. https://doi.org/10.1016/0006-8993(96)00406-4
Miyauchi T, Yonezawa S, Takamura T, Chiba T, Tejima S, Ozawa M, Sato E, Muramatsu T (1982) A new fucosyl antigen expressed on colon adenocarcinoma and embryonal carcinoma cells. Nature 299:168–169. https://doi.org/10.1038/299168a0
Murata T, Morimoto S, Zeng X, Watanabe S, Usui T (1999) Enzymatic synthesis of alpha-L-fucosyl-N-acetyllac-tosamines and 3′-O-alpha-L-fucosyllactose utilizing alpha-L-fucosidases. Carbohydr Res 320:192–199. https://doi.org/10.1016/s0008-6215(99)00156-1
Ohira M, Morohashi A, Nakamura Y, Isogai E, Furuya K, Hamano S, Machida T, Aoyama M, Fukumura M, Miyazaki K, Suzuki Y, Sugano S, Hirato J, Nakagawara A (2003) Neuroblastoma oligo-capping cDNA project: toward the understanding of the genesis and biology of neuroblastoma. Cancer Lett. 197(1-2):63-8. https://doi.org/10.1016/s0304-3835(03)00085-5
Paper JM, Scott-Craig JS, Cavalier D, Faik A, Wiemels RE, Borrusch MS, Bongers M, Walton JD (2013) α-Fucosidases with different substrate specificities from two species of Fusarium. Appl. Microbiol. Biotechnol 97:5371–5380. https://doi.org/10.1007/s00253-012-4423-3
Petersen TN, Brunak S, Von Heijne G, Nielsen H (2011) SignalP 4.0: discriminating signal peptides from transmembrane regions. Nat Methods 8:785–786. https://doi.org/10.1038/nmeth.1701
Renard CM, Lomax JA, Boon JJ (1992) Apple-fruit xyloglucans: a comparative study of enzyme digests of whole cell walls and of alkali-extracted xyloglucans. Carbohydr Res 232:303–320. https://doi.org/10.1016/0008-6215(92)80062-6
Rodríguez-Díaz J, Monedero V, Yebra MJ (2011) Utilization of natural fucosylated oligosaccharides by three novel α-L-fucosidases from a probiotic Lactobacillus casei strain. Appl Environ Microbiol 77:703–705. https://doi.org/10.1128/AEM.01906-10
Sakurama H, Tsutsumi E, Ashida H, Katayama T, Yamamoto K, Kumagai H (2012) Differences in the substrate specificities and active-site structures of two α-L-fucosidases (glycoside hydrolase family 29) from Bacteroides thetaiotaomicron. Biosci Biotechnol Biochem 76:1022–1024. https://doi.org/10.1271/bbb.111004
Saumonneau A, Champion E, Peltier-Pain P, Molnar-Gabor D, Hendrickx J, Tran V, Hederos M, Dekany G, Tellier C (2016) Design of an α-L-transfucosidase for the synthesis of fucosylated HMOs. Glycobiology 26:261–269. https://doi.org/10.1093/glycob/cwv099
Sela DA, Garrido D, Lerno L, Wu S, Tan K, Eom HJ, Joachimiak A, Lebrilla CB, Mills DA (2012) Bifidobacterium longum subsp. infantis ATCC 15697 α-fucosidases are active on fucosylated human milk oligosaccharides. Appl Environ Microbiol 78:795–803. https://doi.org/10.1128/AEM.06762-11
Shaikh FA, Lammerts van Bueren A, Davies GJ, Withers SG (2013) Identifying the catalytic acid/base in GH29 α-L-fucosidase subfamilies. Biochemistry 52:5857–5864. https://doi.org/10.1021/bi400183q
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus Li KW, Lopez R, McWilliam H, Remmert M, Söding J, Thompson JD, Higgins DG (2014) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539–539. https://doi.org/10.1038/msb.2011.75
Sugiyama Y, Gotoh A, Katoh T, Honda Y, Yoshida E, Kurihara S, Ashida H, Kumagai H, Yamamoto K, Kitaoka M, Katayama T (2016) Introduction of H-antigens into oligosaccharides and sugar chains of glycoproteins using highly efficient 1,2-α-L-fucosynthase. Glycobiology 26:1235–1247. https://doi.org/10.1093/glycob/cww085
Sulzenbacher G, Bignon C, Nishimura T, Tarling CA, Withers SG, Henrissat B, Bourne Y (2004) Crystal structure of Thermotoga maritima α-L-fucosidase insights into the catalytic mechanism and the molecular basis for fucosidases. J Biol Chem 279:13119–13128. https://doi.org/10.1074/jbc.M313783200
Takamura T, Chiba T, Tejima S (1981) Chemical modification of lactose. XV. Syntheses of O-α-and O-β-L-Fucopyranosyl-(1→3)-O-β-D-galactopyranosyl-(1→4)-D-glucopyranoses (3′-O-α-and 3′-O-β-L-Fucopyranosyllactoses). Carbohydr Res 84:53–60. https://doi.org/10.1002/chin.198139330
Vandenplas Y, Berger B, Carnielli VP, Ksiazyk J, Lagström H, Luna MS, Migacheva N, Mosselmans JM, Picaud JC, Possner M, Singhal A, Wabitsch M (2018) Human milk oligosaccharides: 2’-fucosyllactose (2’FL) and lacto-N-Neotetraose (LNnT) in infant formula. Nutrients 10:1–12. https://doi.org/10.3390/nu10091161
Vázquez E, Barranco A, Ramírez M, Gruart A, Delgado-García JM, Martínez-Lara E, Blanco S, Martín MJ, Castanys E, Buck R, Prieto P, Rueda R (2015) Effects of a human milk oligosaccharide, 2′-fucosyllactose, on hippocampal long-term potentiation and learning capabilities in rodents. J Nutr Biochem 26:455–465. https://doi.org/10.1016/j.jnutbio.2014.11.016
Xiao L, LeusinkMuis T, Kettelarij N, vanArk I, Blijenberg B, Hesen NA, Stahl B, Overbeek SA, Garssen J, Folkerts G, Van't Land B (2018) Human milk oligosaccharide 2′-fucosyllactose improves innate and adaptive immunity in an influenza-specific murine vaccination model. Front Immunol 9:452. https://doi.org/10.3389/fimmu.2018.00452
Yamashita K, Tachibana Y, Kobata A (1976) Oligosaccharides of human milk. Isolation and characterization of three new disialylfucosyl hexasaccharides. Arch Biochem Biophys 174:582–591. https://doi.org/10.1016/0003-9861(76)90387-8
Yu ZT, Chen C, Kling DE, Liu B, McCoy JM, Merighi M, Heidtman M, Newburg DS (2013) The principal fucosylated oligosaccharides of human milk exhibit prebiotic properties on cultured infant microbiota. Glycobiology 23:169–177. https://doi.org/10.1093/glycob/cws138
Yu ZT, Nanthakumar NN, Newburg DS (2016) The human milk oligosaccharide 2′-fucosyllactose quenches Campylobacter jejuni-induced inflammation in human epithelial cells HEp-2 and HT-29 and in mouse intestinal mucosa. J Nutr 146:1980–1990. https://doi.org/10.3945/jn.116.230706
Yun EJ, Liu JJ, Lee JW, Kwak S, Yu S, Kim KH, Jin YS (2019) Biosynthetic routes for producing various fucosyl-oligosaccharides. ACS Synth Biol 8:415–424. https://doi.org/10.1021/acssynbio.8b00436
Zehra S, Khambati I, Megan V, Mian MF, Buck R, Paul F (2018) Human milk oligosaccharides attenuate antigen-antibody complex induced chemokine release from human intestinal epithelial cell lines. J Food Sci 83:499–508. https://doi.org/10.1111/1750-3841.14039
Zeuner B, Jers C, Mikkelsen JD, Meyer AS (2014) Methods for improving enzymatic trans-glycosylation for synthesis of human milk oligosaccharide biomimetics. J Agric Food Chem 62:9615–9631. https://doi.org/10.1021/jf502619p
Zeuner B, Muschiol J, Holck J, Lezyk M, Gedde MR, Jers C, Mikkelsen JD, Meyer AS (2018) Substrate specificity and transfucosylation activity of GH 29 α-L-fucosidases for enzymatic production of human milk oligosaccharides. New Biotechnol 41:34–45. https://doi.org/10.1016/j.nbt.2017.12.002
Funding
This work was supported by the National Natural Science Foundation of China, grant numbers 31630096 and 31822037.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declared that they have no conflict of interest.
Human and animal rights
This article does not contain any studies with human participants or animals.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 788 kb)
Rights and permissions
About this article
Cite this article
Shi, R., Ma, J., Yan, Q. et al. Biochemical characterization of a novel α-L-fucosidase from Pedobacter sp. and its application in synthesis of 3′-fucosyllactose and 2′-fucosyllactose. Appl Microbiol Biotechnol 104, 5813–5826 (2020). https://doi.org/10.1007/s00253-020-10630-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-020-10630-y