Abstract
2,3-Butanediol (2,3-BDO) is of considerable importance in the chemical, plastic, pharmaceutical, cosmetic, and food industries. The main bacterial species producing this compound are considered pathogenic, hindering large-scale productivity. The species Paenibacillus brasilensis is generally recognized as safe (GRAS) and is phylogenetically similar to P. polymyxa, a species widely used for 2,3-BDO production. Here, we demonstrate, for the first time, that P. brasilensis strains produce 2,3-BDO. Total 2,3-BDO concentrations for 15 P. brasilensis strains varied from 5.5 to 7.6 g/l after 8 h incubation at 32 °C in modified YEPD medium containing 20 g/l glucose. Strain PB24 produced 8.2 g/l of 2,3-BDO within a 12-h growth period, representing a yield of 0.43 g/g and a productivity of 0.68 g/l/h. An increase in 2,3-BDO production by strain PB24 was observed using higher concentrations of glucose, reaching 27 g/l of total 2,3-BDO in YEPD containing about 80 g/l glucose within a 72-h growth period. We sequenced the genome of P. brasilensis PB24 and uncovered at least six genes related to the 2,3-BDO pathway at four distinct loci. We also compared gene sequences related to the 2,3-BDO pathway in P. brasilensis PB24 with those of other spore-forming bacteria, and found strong similarity to P. polymyxa, P. terrae, and P. peoriae 2,3-BDO-related genes. Regulatory regions upstream of these genes indicated that they are probably co-regulated. Finally, we propose a production pathway from glucose to 2,3-BDO in P. brasilensis PB24. Although the gene encoding S-2,3-butanediol dehydrogenase (butA) was found in the genome of P. brasilensis PB24, only R,R-2,3- and meso-2,3-butanediol were detected by gas chromatography under the growth conditions tested here. Our findings can serve as a basis for further improvements to the metabolic capabilities of this little-studied Paenibacillus species in relation to production of the high-value chemical 2,3-butanediol.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Interest in the production of 2,3-butanediol (2,3-BDO) by a range of sugar (or citrate or carbon monoxide or H2 plus CO)-fermenting microbes is increasing because of the importance of this chemical to different industries. 2,3-BDO is commonly used as a liquid fuel additive, a softening and moistening agent, a solvent, a synthetic rubber precursor, an anti-freeze agent, in the manufacture of foods, and as a carrier for different drugs (Celinska and Grajek 2009; Garg and Jain 1995; Ji et al. 2009; Syu 2001). However, chemical synthetic methods for 2,3-BDO production face the problem of exhausted petroleum oil stocks, a primary feedstock in the production process (Ji et al. 2009).
Various bacterial species are considered efficient producers of 2,3-BDO such as Klebsiella pneumoniae, K. oxytoca, Enterobacter aerogenes, Serratia marcescens, and Paenibacillus polymyxa (Cho et al. 2015; Sabra et al. 2015). Except for P. polymyxa, the other species are all considered pathogenic (risk group 2) by the World Health Organization, WHO (US FDA/CFSAN 2015), making their use for 2,3-BDO production problematic. In contrast, the spore-forming bacteria that produce 2,3-BDO—such as those belonging to different Bacillus spp., Clostridium spp., and Paenibacillus spp.—do not cause disease in healthy humans (Biosafety level 1, BSL-1). Wild type B. licheniformis strains usually produce a mixture of S,S-2,3-BDO and meso-2,3-BDO isomers, with each isomer having its own unique applications (Qiu et al. 2016). Acetogenic members of the Clostridium genus (Clostridium autoethanogenum, C. ljungdahlii, and C. ragsdalei) use gases (CO alone or H2 plus CO) as the carbon and energy source for 2,3-BDO production (Köpke et al. 2011). P. polymyxa can utilize a broad spectrum of substrates for 2,3-BDO production, such as mannose, galactose, cellobiose, glycerol, a mixture of glucose, and xylose or a mixture of glucose and cellobiose (Jiang et al. 2015). In fact, P. polymyxa strains are being extensively studied to improve their industrial potential for 2,3-BDO production using renewable feedstocks (Okonkwo et al. 2017). The metabolic pathways of 2,3-BDO production by P. polymyxa strain ICGEB2008 were described by Adlakha et al. (2015).
Paenibacillus brasilensis was described by von der Weid et al. (2002) as nitrogen-fixing strains isolated from the rhizosphere of maize planted in Cerrado soil, Brazil. The strains were very homogeneous and shared a high level of relatedness with P. polymyxa and P. peoriae, with these latter comprising both nitrogen- and non-nitrogen-fixing strains. Fortes et al. (2008) and von der Weid et al. (2005) later demonstrated that some P. brasilensis strains produce antimicrobial substances active against bacteria and fungi, including phytopathogenic fungi that commonly cause diseases in maize. Like the other two Paenibacillus species mentioned above, P. brasilensis is typically worked with at a BSL-1, posing little to no threat of infection in healthy adults. However, as far as we know, no previous studies have assessed 2,3-BDO production in this species. Therefore, the purposes of this study were to (i) demonstrate production of pure or a mixture of 2,3-BDO isomers by P. brasilensis strains under different growth conditions; (ii) gain insight into the genes involved in 2,3-BDO production through genome sequencing and annotation of a representative strain of the species; and (iii) propose a production pathway from glucose to 2,3-BDO for P. brasilensis, confirming the presence of S,S-, R,R- and/or meso-2,3-BDO by gas chromatography. No genome sequences of P. brasilensis have previously been deposited at NCBI and our data can serve as a basis for further improvements to the 2,3-BDO production capabilities of this little-studied Paenibacillus species.
Materials and methods
Bacterial strains and growth conditions
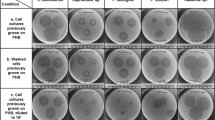
Fifteen Paenibacillus brasilensis strains previously described in von der Weid et al. (2002) (see Table 1) were stored aerobically either at room temperature on Trypticase Soy Broth (TSB) agar-containing slants supplemented with 1% CaCO3 (w/v) or at −80 °C in TSB with 20% glycerol. All strains were inoculated in TSB and incubated at 32 °C for 24 h. They were then further inoculated (about 3.0 × 106 UFC/ml) in YEPD medium (10 g/l glucose, 10 g/l yeast extract, 20 g/l peptone; pH 6.3; Adlakha and Yazdani 2015) and flasks were incubated at 32 °C with agitation (200 rpm) for 16 h. For 2,3-BDO analysis, we employed a modified YEPD medium (20 g/l glucose, 15 g/l yeast extract, 0.5 g/l K2HPO4, 2 g/l KH2PO4, 0.0225 g/l MnSO4, 0.3 g/l KCl; Adlakha and Yazdani 2015) inoculated with about 3.0 × 106 UFC/ml. Flasks were incubated at 32 °C with agitation (200 rpm). Metabolic end-products were analyzed by high-performance liquid chromatography (HPLC) after 8 h growth at 32 °C and agitation of 200 rpm in 250 ml-erlenmeyers with 100 ml of modified YEPD. A representative strain of P. brasilensis (PB24—deposited in the Culture Collection of the genus Bacillus and correlated genera—CCGB, Oswaldo Cruz Institute, IOC, FIOCRUZ, Rio de Janeiro, under the accession number LFB-FIOCRUZ #1431) was also grown in modified YEPD medium containing 20–80 g/l of glucose for up to 72 h in the same conditions described above. The number of viable bacterial cells (PB24) in the modified YEPD medium (20 g/l glucose) was determined by colony-forming units/ml (CFU/ml).
2,3-BDO production
Concentrations of 2,3-BDO, glucose, and other byproducts (mainly lactic acid) were determined as previously described in Petrov and Petrova (2009) using a high-performance liquid chromatography (HPLC) system (1260 INFINITY, Agilent Technologies, Santa Clara, CA) equipped with a refractive index (RI) detector and a Bio-Rad column for organic acids analysis (300 mm × 7.8 mm) under the following conditions: sample volume = 20 μl; mobile phase = 0.005 M H2SO4; flow rate = 0.6 ml/min; and column temperature = 45 °C. Statistical analysis for 2,3-BDO production was performed using Tukey’s pairwise test. Differences were considered significant if p < 0.01. Statistical test was performed using PAST software (Hammer et al. 2001).
The production of S,S- and/or R,R- and meso-2,3-BDO by P. brasilensis PB24 grown in the modified YEPD medium up to 72 h was verified by capillary gas chromatography with a chiral stationary phase (Restek Rt-bDEXsm, 30 m × 0.25 mm ID, 0.25 df). The analyses were made using a Shimadzu GC2010 chromatograph (oven temperature: isocratic, 100 °C, injector 200 °C, flame ionization detector, 230 °C). The three standards (S,S-, R,R-, and meso-2,3-BDO; Sigma-Aldrich) were used as control.
Carbohydrate utilization profile
The utilization of 49 carbohydrates was examined for P. brasilensis PB24 using the API50CH system (Biomérieux, France), according to the manufacturer’s instructions. Carbohydrate utilization was determined after incubation at 32 °C for 48 h (Seldin and Penido, 1986).
Whole genome sequencing (WGS), de novo genome assembly and sequence analyses
DNA from P. brasilensis PB24 was isolated according to the method described in Seldin et al. (1998). Cells from 6 × 60 ml cultures grown in TSB at 32 °C for 16 h were centrifuged (10,000×g, 10 min), resuspended in 2 ml of Tris–EDTA–NaCl buffer (Seldin and Dubnau 1985), and treated with 1 mg lysozyme (30 min, 37 °C), and 1% sodium dodecyl sulfate (10 min, 37 °C). Further purification steps were as described by Seldin and Dubnau (1985) and were completed using the ZR Fungal/Bacterial DNA MiniPrep™ system (Zymo Research, Irvine, CA). The DNA was quantified spectrophotometrically using a NanoDrop™ (Thermo Fisher Scientific, Waltham, MA) and a Qubit™ fluorimeter (Thermo Fisher Scientific). The P. brasilensis PB24 genome was sequenced by DNA Link Inc. (Seoul, Korea) using a PacBio RSII platform and two SMRT cells of P6-C4 chemistry with a 20-kb size-selected library. The reads were de novo-assembled with HGAP (version 2.3; Chin et al. 2013). Open reading-frame (ORF) prediction and amino acid translation were performed using the RAST server, version 2.0 (Aziz et al. 2008).
Search for 2,3-BDO-related genes
Functional annotation of the ORFs was performed using the SEED package (Overbeek et al. 2014) and the database FIGfam version 70 (Meyer et al. 2009). The text file containing MultiFASTA-formatted amino acid sequences was downloaded from RAST and submitted to KAAS (KEGG Automatic Annotation Server) to identify genes related to enzymes of the 2,3-BDO pathway (Moriya et al. 2007). The loci including genes related to the 2,3-BDO pathway have been deposited in GenBank (NCBI) under the accession numbers MF996568-MF996571.
2,3-BDO-coding genes in P. brasilensis PB24 and comparison to other Paenibacillus species
Gene sequences related to the 2,3-BDO pathway in P. brasilensis PB24 were aligned with those of other spore-forming bacterial species (Bacillus and Paenibacillus) using ClustalW (Larkin et al. 2007). Phylogenetic trees were constructed using the neighbor-joining method (Saitou and Nei 1987). The MEGA 7.0 software (Kumar et al. 2016) was used to calculate Jukes-Cantor distances, and bootstrap analyses were performed with 1000 replicates.
Synteny and genomic context of operons related to 2,3-BDO enzymes in P. brasilensis PB24 and other Paenibacillus species
In order to evaluate whether 2,3-BDO-related enzymes were evolutionarily acquired once and the synteny of all 2,3-BDO-related genes within the Paenibacillus genus, we used Microbial Genomic context Viewer (MGcV) (Overmars et al. 2013) to assess proteins related to the 2,3-BDO pathway in 12 Paenibacillus genomes: P. brasilensis PB24 (this study) and 11 genomes from the MGcV database (P. polymyxa CR1, P. polymyxa E681, P. polymyxa M1, P. polymyxa SC2, P. terrae HPL-003, Paenibacillus sp. Y412M10, Paenibacillus sp. JDR-2, P. mucilaginosus K02, P. mucilaginosus 3016, P. mucilaginosus KN414, and P. larvae subsp. larvae DSM 25430).
Analysis of regulatory regions upstream of 2,3-BDO-related genes
In order to evaluate whether the different operons related to enzymes belonging to the 2,3-BDO pathway could be co-regulated, regulatory regions (100 nucleotides upstream of the start codon) for each gene were analyzed with MAST (Motif Alignment & Search Tool; Bailey and Gribskov 1998).
Results
2,3-BDO production by Paenibacillus brasilensis
All of the 15 P. brasilensis strains we tested produced 2,3-BDO after 8 h cultivation at 32 °C in modified YEPD medium containing 20 g/l glucose using a bench shaker. We detected by HPLC the meso-form (meso-2,3-BDO) and R,R-2,3-BDO or S,S-2,3-BDO or their racemic mixture (R,R-/S,S-2,3-BDO represents any of the three products). The total amount of 2,3-BDO produced under these growth conditions varied from 5.5 to 7.6 g/l (Table 1).
We chose one representative strain of P. brasilensis (PB24) for further analyses. Production of 2,3-BDO by this strain was assessed over a 24-h growth period using the same conditions described above. PB24 produced 8.2 ± 0.06 g/l total 2,3-BDO within 12 h, and complete depletion of glucose was observed (Fig. 1). After 12 h growth, PB24 achieved a yield of 0.43 g/g 2,3-BDO and a productivity of 0.68 g/l/h 2,3-BDO. After 24 h, a reduced concentration of 2,3-BDO and a decrease of the number of viable cells (CFU/ml) were observed (Fig. 1). The same experiment was performed increasing the glucose concentration up to 80 g/l in modified YEPD medium and assessing 2,3-BDO production by HPLC over a 72-h growth period (Table 2). The highest 2,3-BDO production (27 g/l) was observed after 72 h in the medium containing about 80 g/l glucose. PB24 achieved a yield of 0.41 g/g 2,3-BDO and a productivity of 0.38 g/l/h 2,3-BDO in this latter condition.
The API 50CH profile (carbohydrate fermentation) was determined for strain PB24. Among the 49 different carbohydrates, PB24 produced acid from ribose, glucose, fructose, mannose, mannitol, methyl α-D-glucoside, amygdalin, arbutin, aesculin, salicin, cellobiose, maltose, lactose, melibiose, sucrose, threalose, melezitose, raffinose, starch, glycogen, gentibiose, and turanose. It was not able to utilize 27 of the other carbohydrates tested, including glycerol.
The Paenibacillus brasilensis PB24 genome
The genome of P. brasilensis PB24 was sequenced using the PacBio RSII platform and the 225,233 raw reads resulted in 180,815 quality-filtered trimmed reads yielding 1827 Mb. The DNA sequences were used to predict protein-coding genes related to 2,3-BDO production.
KAAS (KEGG) analysis identified at least six genes related to the 2,3-BDO pathway in four distinct loci (Fig. S1−S4). These genes can be classified as alsL (KEGG Orthology K01652, coding for acetolactate synthase) and alsS (K01653, coding for acetolactate synthase) in the same operon (locus MF996568, ORFs PB24_112 and PB24_111, respectively), with synteny conserved among all compared Paenibacillus species (Fig. S1). A second locus (MF996571) includes a second copy of alsL (K01652, PB24_5208) and neighboring alsD (K01575, PB24_5209 coding for acetolactate decarboxylase), with synteny conserved among P. brasilensis, P. terrae, and P. polymyxa (Fig. S2). A third locus (MF996569) contains the butB gene (K00004, PB24_3312 coding for R,R-2,3-butanediol dehydrogenase/meso-2,3-butanediol dehydrogenase/diacetyl reductase) and showed synteny between P. brasilensis and P. polymyxa CR1, whereas in the other strains their closest orthologs are annotated with other functions (Fig. S3). Finally, locus MF996570 includes the butA gene (K03366, PB24_4054 coding for S,S-2,3-butanediol dehydrogenase/meso-2,3-butanediol dehydrogenase/diacetyl reductase), with synteny conserved only between P. brasilensis and P. terrae HPL-003 among the 11 Paenibacillus genomes assessed from the MGcV database (Fig. S4).
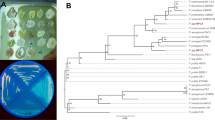
We constructed phylogenetic trees from the 2,3-BDO pathway-related gene sequences (alsS, alsD, butA and butB genes; Fig. S5−S8, respectively) generated from P. brasilensis PB24 and those of other spore-forming bacteria (Bacillus and Paenibacillus). Figure 2 shows partial phylogenetic trees (full trees shown in Fig. S5−S8), highlighting the relationships between strain PB24 and its closest relatives for these four genes. Considering the alsS gene, P. brasilensis PB24 clustered with P. polymyxa, and it clustered with P. peoriae, P. terrae, and P. polymyxa in the alsD tree. PB24 grouped with P. terrae in the butA phylogenetic tree and with P. peoriae and P. polymyxa in the butB tree.
Partial phylogenetic trees (full trees presented in Figs. S5–S8) highlighting the relationships of 2,3-BDO-related genes for closely-related members of the genus Paenibacillus and Bacillus. (a) alsS (acetolactate synthase); (b) alsD (acetolactate decarboxylase); (c) butA (diacetyl reductase); (d) butB (butanediol dehydrogenase). Phylogenetic trees were constructed using the neighbor-joining method, and bootstrap analyses were performed with 1000 replicates. The GenBank accession numbers of each species/strain used are shown in each tree
Our phylogenetic analyses revealed evident similarity between P. brasilensis and P. polymyxa 2,3-BDO-related genes. Genomic annotation of PB24 resulted in the identification of genes related to three butanediol isomers—R,R-2,3-butanediol (butB), S,S-2,3-butanediol (butA), and meso-2,3-butanediol (butA and butB).
We identified three conserved motifs (across all assessed Paenibacillus species and represented by different colors in Fig. 3) 100 bp upstream of the genes involved in 2,3-BDO production. Although these regulatory regions lie in distinct regions of the PB24 genome, they are probably co-regulated because of the presence of these conserved motifs. The presence of two motifs in regulatory regions upstream of als operons suggests that the enzymes codified by their genes could be part of different pathways of metabolism or be expressed under different conditions. However, the function of the observed motifs remains unknown. Again, there was high sequence similarity in these regions between PB24 and P. polymyxa and P. terrae.
Conserved motifs (red, blue and green) observed in regulatory regions (100 bases upstream of start codons) of genes related to the 2,3-BDO pathway in P. brasilensis PB24 and other Paenibacillus genomes from the MGcV database. Similar motifs in regulatory regions (motifs with the same color) suggest that 2,3-BDO-related genes are regulated in the same way and probably co-expressed despite considerable genomic distances between genes. The letters A, B, C, D, and E represent the 2,3-BDO-related gene sequences generated in this study
Proposed pathway from glucose to 2,3-BDO in P. brasilensis PB24
We propose a 2,3-BDO production pathway based on the 2,3-BDO-related genes found in the PB24 genome (Fig. 4), beginning with generation of S-2-acetolactate from two condensed pyruvate molecules by the enzyme acetolactate synthase, the two subunits of which are encoded by the alsL and alsS genes. S-2-acetolactate can be transformed into diacetyl by spontaneous decarboxylation in the presence of oxygen or into R-acetoin by the enzyme acetolactate decarboxylase encoded by the alsD gene. When diacetyl formation occurs in the presence of oxygen, the enzyme diacetyl reductase (encoded by butA) converts it into S-2-acetoin by a reduction reaction in the presence of NADPH. S-2-acetoin is converted to meso-2,3-BDO by the enzyme R-2,3-butanediol dehydrogenase (encoded by butB). In the presence of NADH, R-2,3-butanediol dehydrogenase converts diacetyl into R-2-acetoin.
Proposed 2,3-BDO production pathway for Paenibacillus brasilensis PB24 based on the 2,3-BDO-related genes encoded in its genome. The colored squares correspond to the conserved motifs shown in Fig. 4. The dashed lines correspond to the presence of butA encoding the S-2,3-butanediol dehydrogenase in PB24 genome. The absence of S,S-2,3-BDO production demonstrated by chiral gas chromatography suggests that S-2,3-butanediol dehydrogenase is not expressed in the growth conditions tested in this study
In the absence of oxygen, R,R-2,3-BDO is formed in the presence of NADH by the reduction of R-2-acetoin performed by R-2,3-butanediol dehydrogenase. This reaction is reversible so, in the presence of NAD+, R-2-acetoin can be formed from R,R-2,3-BDO. Moreover, R-2-acetoin can be converted into meso-2,3-BDO by S-2,3-butanediol dehydrogenase. Besides encoding diacetyl reductase, the butA gene is also responsible for the expression of S-2,3-butanediol dehydrogenase, which may direct the pathway for production of S,S-2,3-BDO and meso-2,3-BDO. To prove the simultaneous production of the three 2,3-BDO isomers during sugar fermentation in P. brasilensis, we used a chiral gas chromatography. Only the production of R,R-2,3-BDO and meso-2,3-BDO was observed (Fig. 5), suggesting that S-2,3-butanediol dehydrogenase is not expressed in the growth conditions tested in this study.
Chiral chromatogram of (a) the S,S-, R,R- and meso-2,3-BDO standards and (b) metabolic end-products produced by Paenibacillus brasilensis PB24 (see “Materials and methods” for conditions)
Discussion
A preliminary screening for 2,3-BDO production by P. brasilensis was performed in this study. No previous reports have shown 2,3-BDO production using P. brasilensis strains. Nevertheless, strains belonging to this species seems to be very industrially interesting because they are nitrogen-fixers, antimicrobial substance-producers and non-pathogenic for animals and humans (von der Weid et al. 2002, 2005). Although all strains produced 2,3-BDO in the conditions presented here, the screening was far from exhaustive, and new media and conditions should be tested in further studies as previously done for Bacillus atrophaeus NRS-213, B. mojavensis B-14698 and B. vallismortis B-14891 (Kallbach et al. 2017). B. vallismortis B-14891 has been shown to produce 60.4 g/l 2,3-BDO with an initial glucose concentration of 200 g/l within 55 h in a batch cultivation. Moreover, it can convert 14 different substrates obtained from residual biomass into 2,3-BDO (Kallbach et al. 2017). In this study, we only conducted bacterial cultivation in modified YEPD medium (Adlakha and Yazdani 2015) containing 20 g/l glucose for a short time period using a bench shaker to prove 2,3-BDO production by P. brasilensis strains, and also in YEPD containing up to 80 g/l glucose for 72 h using strain PB24. The highest amount of 2,3-BDO obtained (27 g/l—in medium containing about 80 g/l glucose) is still far from ideal. Therefore, controlling parameters such as initial glucose concentration, nitrogen source in culture medium, temperature and pH are still necessary to optimize the condition for the efficient production of 2,3-BDO. Moreover, the dissolved oxygen level has been determined as a key parameter in the effective production of 2,3-BDO by fermentation. Several parameters such as the oxygen transfer rate, oxygen transfer coefficient, and respiratory quotient have already been used to determine the optimal aerobic condition in several studies (Song et al., 2018). However, controlling those parameters in shake flask cultivation is a difficult task, and that oxygen may have been also a limiting factor in our experimental conditions.
Von der Weid et al. (2002) previously showed that P. brasilensis strains produced acid from ribose, glucose, fructose, mannose, mannitol, methyl α-D-glucoside, amygdalin, arbutin, aesculin, salicin, cellobiose, maltose, lactose, melibiose, sucrose, raffinose, starch, and glycogen. Besides all these carbohydrates, PB24 also produced acid from threalose, melezitose, gentibiose, and turanose. Different strains of Paenibacillus polymyxa have also been shown to produce R,R-2,3-BDO and a small amount of meso-2,3-BDO using as substrate mannose, galactose, cellobiose, glycerol, a mixture of glucose and xylose, and a mixture of glucose and cellobiose (Jiang et al. 2015), as well as lignocellulosic hydrolysate (Adlakha and Yazdani 2015). Therefore, P. brasilensis represents a promising species for improving large-scale production of 2,3-BDO with different substrates. One relevant point that should be considered is the reduction of 2,3-BDO concentration after 24 h growth of the representative strains of P. brasilensis (observed in Fig. 1). This could indicate that 2,3-BDO could be used as a carbon and energy source, as previously suggested by Ji et al. (2011). González et al. (2000) also demonstrated that Saccharomyces cerevisiae FY834α could grow on 2,3-BDO as the sole carbon and energy source. However, after 24 h of growth (with the total glucose depletion), the number of PB24 cells had already decreased as demonstrated by the number of viable cells (CFU/ml) (Fig. 1).
DNA sequences obtained from the genome of P. brasilensis PB24 were used to predict protein-coding genes related to 2,3-BDO production and to present regulatory regions in distinct regions of the genome studied. The identification of genes related to three butanediol isomers—R,R-2,3-butanediol (butB), S,S-2,3-butanediol (butA), and meso-2,3-butanediol (butA and butB)—suggests that PB24 can produce any of these isomers. Serratia sp. T241 also produces these three 2,3-butanediol stereoisomers, and its genome encodes three 2,3-butanediol dehydrogenases and one glycerol dehydrogenase involved in the formation of 2,3-BDO isomers (Zhang et al. 2016). In contrast, P. polymyxa almost exclusively produces the R,R-2,3-BDO isomer (over 98%) and only a small amount of meso-2,3-BDO (Yu et al. 2011). Likewise, only R,R-2,3- and meso-2,3-butanediol were detected by gas chromatography under the growth conditions tested here, although the gene encoding S-2,3-butanediol dehydrogenase (butA) was found in the genome of P. brasilensis PB24. Based on the 2,3-BDO-related genes found in the PB24 genome and the detection of only R,R-2,3- and meso-2,3-butanediol by gas chromatography, 2,3-BDO production pathway was proposed. With the depletion of dissolved oxygen in shake flask cultivation, the preferential production of R,R-2,3-BDO is expected. Modifications in regulatory regions could enhance 2,3-BDO production or facilitate selectivity for a specific isomer. Recently, Yang et al. (2017) presented a metabolic engineering approach, guided by systems and synthetic biology, for the improvement of microbial acetoin and 2,3-BDO production.
In conclusion, we demonstrate that P. brasilensis can be considered a novel producer of 2,3-BDO, based on the presence of genes involved in the 2,3-BDO pathway in its genome and production of 2,3-BDO in the conditions described here. However, various optimization strategies should be applied to establish the best medium components (including alternative substrates) and fermentation conditions for 2,3-BDO production. Moreover, since P. brasilensis poses little to no threat of infection in healthy adults and is typically worked with at a BSL-1, it is a promising alternative to other 2,3-BDO-producing microorganisms with industrial applications.
References
Adlakha N, Pfau T, Ebenhöh O, Yazdani SS (2015) Insight into metabolic pathways of the potential biofuel producer, Paenibacillus polymyxa ICGEB2008. Biotechnol Biofuels 8:159
Adlakha N, Yazdani SS (2015) Efficient production of (R,R)-2,3-butanediol from cellulosic hydrolysate using Paenibacillus polymyxa ICGEB2008. J Ind Microbiol Biotechnol 42:21–28
Aziz RK, Bartels D, Best AA, DeJongh M, Disz T, Edwards RA, Formsma K, Gerdes S, Glass EM, Kubal M, Meyer F, Olsen GJ, Olson R, Osterman AL, Overbeek RA, McNeil LK, Paarmann D, Paczian T, Parrello B, Pusch GD, Reich C, Stevens R, Vassieva O, Vonstein V, Wilke A, Zagnitko O (2008) The RAST server: rapid annotations using subsystems technology. BMC Genomics 9:75
Bailey TL, Gribskov M (1998) Combining evidence using p-values: application to sequence homology searches. Bioinformatics 14:48–54
Celinska E, Grajek W (2009) Biotechnological production of 2,3-butanediol–current state and prospects. Biotechnol Adv 27:715–725
Chin C-S, Alexander DH, Marks P, Klammer AA, Drake J, Heiner C, Clum A, Copeland A, Huddleston J, Eichler EE, Turner SW, Korlach J (2013) Nonhybrid, finished microbial genome assemblies from long-read SMRT sequencing data. Nat Methods 10:563–569
Cho S, Kim T, Woo HM, Lee J, Kim Y, Um Y (2015) Enhanced 2, 3-butanediol production by optimizing fermentation conditions and engineering Klebsiella oxytoca M1 through overexpression of acetoin reductase. PLoS One 10:e0138109
Fortes TO, Alviano DS, Tupinambá G, Padrón TS, Antoniolli AR, Alviano CS, Seldin L (2008) Production of an antimicrobial substance against Cryptococcus neoformans by Paenibacillus brasilensis Sa3 isolated from the rhizosphere of Kalanchoe brasiliensis. Microbiol Res 163:200–207
Garg SK, Jain A (1995) Fermentative production of 2,3-butanediol—a review. Bioresour Technol 51:103–109
González E, Fernández MR, Larroy C, Solà L, Pericàs MA, Parés X, Biosca JA (2000) Characterization of a (2R,3R)-2,3-butanediol dehydrogenase as the Saccharomyces cerevisiae YAL060W gene product. Disruption and induction of the gene J Biol Chem 275:35876–33585
Hammer Ø, Harper DAT, Ryan PD (2001) Paleontological statistics software package for education and data analysis. Palaeontol Electron 4:9–18
Ji XJ, Huang H, Du J, Zhu JG, Ren LJ, Hu N, Li S (2009) Enhanced 2,3-butanediol production by Klebsiella oxytoca using a two-stage agitation speed control strategy. Bioresour Technol 100:3410–3414
Ji XJ, Huang H, Ouyang PK (2011) Microbial 2,3-butanediol production: a state-of-the-art review. Biotechnol Adv 29:351–364
Jiang L-q, Fang Z, Zaho Z-l, He F, Li H-b (2015) 2,3-Butanediol and acetoin production from enzymatic hydrolysate of ionic liquid-pretreated cellulose by Paenibacillus polymyxa. Bioresources 10:1318–1329
Kallbach M, Horn S, Kuenz A, Prüße U (2017) Screening of novel bacteria for the 2,3-butanediol production. Appl Microbiol Biotechnol 101:1025–1033
Köpke M, Mihalcea C, Liew F, Tizard JH, Ali MS, Conolly JJ, Al-Sinawi B, Simpson SD (2011) 2,3-butanediol production by acetogenic bacteria, an alternative route to chemical synthesis, using industrial waste gas. Appl Environ Microbiol 77:5467–5475
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and Clustal X version 2.0. Bioinformatics 23:2947–2948
Meyer F, Overbeek R, Rodriguez A (2009) FIGfams: yet another set of protein families. Nucleic Acids Res 37:6643–6654
Moriya Y, Itoh M, Okuda S, Yoshizawa A, Kanehisa M (2007) KAAS: an automatic genome annotation and pathway reconstruction server. Nucleic Acids Res 35:182–185
Okonkwo CC, Ujor VC, Mishra PK, Ezeji TC (2017) Process development for enhanced 2,3-butanediol production by Paenibacillus polymyxa DSM 365. Fermentation 3:18
Overbeek R, Olson R, Pusch GD, Olsen GJ, Davis JJ, Disz T, Edwards RA, Gerdes S, Parrello B, Shukla M, Vonstein V, Wattam AR, Xia F, Stevens R (2014) The SEED and the rapid annotation of microbial genomes using subsystems technology (RAST). Nucleic Acids Res 42:D206–D214
Overmars L, Kerkhoven R, Siezen RJ, Francke C (2013) MGcV: the microbial genomic context viewer for comparative genome analysis. BMC Genomics 14:209
Petrov K, Petrova P (2009) High production of 2,3-butanediol from glycerol by Klebsiella pneumoniae G31. Appl Microbiol Biotechnol 84:659–665
Qiu Y, Zhang J, Li L, Wen Z, Nomura CT, Wu S, Chen S (2016) Engineering Bacillus licheniformis for the production of meso-2,3-butanediol. Biotechnol Biofuels 9:117
Sabra W, Groeger C, Zeng AP (2015) Microbial cell factories for diol production. Adv Biochem Eng Biotechnol 155:165–197
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Seldin L, Dubnau D (1985) Deoxyribonucleic acid homology among Bacillus polymyxa, Bacillus macerans, Bacillus azotofixans and other nitrogen-fixing Bacillus strains. Int J Syst Bacteriol 35:151–154
Seldin L, Penido EGC (1986) Identification of Bacillus azotofixans using API tests. Antonie Van Leeuwenhoek 52:403–409
Seldin L, Rosado AS, Cruz DW, Nobrega A, van Elsas JD, Paiva E (1998) Comparison of Paenibacillus azotofixans strains isolated from rhizoplane, rhizosphere and non-rhizosphere soil from maize planted in two different Brazilian soils. Appl Environ Microbiol 64:3860–3868
Song CW, Rathnasingh C, Park JM, Lee J, Hyohak Song H (2018) Isolation and evaluation of Bacillus strains for industrial production of 2,3-butanediol. J Microbiol Biotechnol 28:409–417
Syu MJ (2001) Biological production of 2,3-butanediol. Appl Microbiol Biotechnol 55:10–18
US FDA/CFSAN (2015) Inventory of GRAS notices: summary of all GRAS notices. https://www.fda.gov/food/ingredientspackaginglabeling/gras/; accessed in March 2015
von der Weid I, Artursson V, Seldin L, Jansson JK (2005) Antifungal and root surface colonization properties of GFP-tagged Paenibacillus brasilensis PB177. World J Microbiol Biotechnol 21:1591–1597
von der Weid I, Duarte GF, van Elsas JD, Seldin L (2002) Paenibacillus brasilensis sp. nov., a novel nitrogen-fixing species isolated from the maize rhizosphere in Brazil. Int J Syst Evol Microbiol 52:2147–2153
Yang T, Rao Z, Zhang X, Xu M, Xu Z, Yang ST (2017) Metabolic engineering strategies for acetoin and 2, 3-butanediol production: advances and prospects. Crit Rev Biotechnol 37:990–1005
Yu B, Sun J, Bommareddy RR, Song L, Zeng A-P (2011) Novel (2R,3R)-2,3-butanediol dehydrogenase from potential industrial strain Paenibacillus polymyxa ATCC 12321. Appl Environ Microbiol 77:4230–4233
Zhang L, Guo Z, Chen J, Xu Q, Lin H, Hu K, Guan X, Shen Y (2016) Mechanism of 2,3-butanediol stereoisomers formation in a newly isolated Serratia sp. T241. Sci Rep 6:19257
Acknowledgements
Thanks are due to Fábio Diniz for his technical support in the HPLC analyses. We are grateful to Prof. Apostolis Koutinas and Dr. Chrysanthi Pateraki from the Agricultural University of Athens, Attiki, Greece, for their helpful comments and guidance.
Funding
This study was supported by grants from the National Research Council of Brazil (CNPq), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES) and Fundação de Amparo à Pesquisa do Estado do Rio de Janeiro (FAPERJ). M.E.V. and D.M.G.F. were provided a grant by Petrobras (grant number: 2012/00320-2).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors have declared no competing interests.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Electronic supplementary material
ESM 1
(PDF 2181 kb)
Rights and permissions
About this article
Cite this article
Dias, B.d., Lima, M.E.d., Vollú, R.E. et al. 2,3-Butanediol production by the non-pathogenic bacterium Paenibacillus brasilensis. Appl Microbiol Biotechnol 102, 8773–8782 (2018). https://doi.org/10.1007/s00253-018-9312-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-9312-y