Abstract
Two archaeal trehalase-like genes, Saci1250 and Saci1816, belonging to glycoside hydrolase family 15 (GH15) from the acidophilic Crenarchaeon Sulfolobus acidocaldarius were expressed in Escherichia coli. The gene products showed trehalose-hydrolyzing activities, and the names SaTreH1 and SaTreH2 were assigned to Saci1816 and Saci1250 gene products, respectively. These newly identified enzymes functioned within a narrow range of acidic pH values at elevated temperatures, which is similar to the behavior of Euryarchaeota Thermoplasma trehalases. SaTreH1 displayed high KM and kcat values, whereas SaTreH2 had lower KM and kcat values despite a high degree of identity in their primary structures. A mutation analysis indicated that two glutamic acid residues in SaTreH1, E374 and E574, may be involved in trehalase catalysis because SaTreH1 E374Q and E574Q showed greatly reduced trehalose-hydrolyzing activities. Additional mutations substituting G573 and H575 residues with serine and glutamic acid residues, respectively, to mimic the TVN1315 sequence resulted in a decrease in trehalase activity and thermal stability. Taken together, the results indicated that Crenarchaea trehalases adopt active site structures that are similar to Euryarchaeota enzymes but have distinct molecular features. The identification of these trehalases could extend our understanding of the relationships between the structure and function of GH15 trehalases as well as other family enzymes and will provide insights into archaeal trehalose metabolism.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Trehalose is a stable disaccharide that consists of two glucose units linked primarily by an α,α-(1 → 1)-linkage, and it has been found in a wide variety of organisms, including archaea, bacteria, yeast, fungi, insects, crustaceans, invertebrates, and plants. In these organisms, trehalose is a source of carbon energy and plays a protective role against various stress conditions. Additionally, trehalose has gained attention for its ability to reduce protein aggregate accumulation by stimulating cellular autophagy (Sarkar et al. 2007). Recently, DeBosch et al. (2016) demonstrated that trehalose inhibits glucose transport and subsequently activates kinase pathways to induce autophagy. Trehalose is expected to be useful in the field of pharmaceuticals as well as in the food and cosmetic industries (Argüelles 2000; Higashiyama 2002; Elbein et al. 2003).
Trehalase (trehalose-glucohydrolase, EC 3.2.1.28) hydrolyzes trehalose to produce two glucose molecules, and it is found in bacteria and eukarya. Trehalases have been classified into glycoside hydrolase 37 (GH37) and GH65 families in the Carbohydrate-Active enZymes database (CAZy, http://www.cazy.org/). Recently, we identified two archaeal trehalose-hydrolyzing enzymes (TreHs), TVN1315 and Ta0286, from thermoacidophilic Euryarchaeota Thermoplasma volcanium and T. acidophilum, respectively (Sakaguchi et al. 2015). These enzymes are assigned to the GH15 family, which also contains glucoamylases (GAs), glucodextranases (GDases), dextran dextrinase, and isomaltase. We showed that GH15 family trehalases functioned within a narrow pH range with higher KM values than those of other GH37 and GH65 family trehalases.
GH37, GH65, and GH15 family enzymes primarily have catalytic domains with (α/α)6-barreled structures, although the sequence homologies among them are low. GH37 and GH65 yeast trehalases have two catalytic residues, aspartic acid and glutamic acid, but in GH15 trehalases, two glutamic acid residues are likely involved in the catalysis (Maicas et al. 2016; Sakaguchi et al. 2015). In addition, GH15 trehalases lack two trehalase signature consensus motifs (i.e., motif 1 and motif 2) as well as other trehalase motifs found in GH37 trehalases (Horlacher et al. 1996; Barraza and Sánchez 2013). GH15 trehalases seem to have different active site structures from those of GH37 and GH65 trehalases. Comparative studies on GH15 trehalases and GAs may be more meaningful for the elucidation of their structure-function relationships, because comparison of the amino acid sequences showed that these archaeal trehalases are more similar to GAs than to GH37 and GH65 trehalases, and the general structure of the substrate-binding site in the GH15 archaeal trehalases likely resembles those in GAs (Sakaguchi et al. 2015). However, the amino acid residues that discriminate between trehalose and other sugar substrates in GH15 archaeal trehalases are poorly understood.
To explore the possibility of similar trehalose-hydrolyzing enzymes in another archaeon and expand our understanding on archaeal trehalases and GH15 family enzymes, we searched for sequences similar to those of archaeal trehalases from another phylum. A BLAST search suggested two candidate genes, Saci1250 and Saci1816, in acidophilic Crenarchaeon Sulfolobus acidocaldarius. In this study, we attempted to express the Saci1250 and Saci1816 genes in Escherichia coli and characterize the recombinant enzymes. Newly cloned genes were expressed in their active forms. The purified enzymes could hydrolyze trehalose under a narrow acidic pH range and presented distinct activities and affinities toward trehalose.
Materials and methods
Chemical reagents and genetic engineering
Chemical reagents and oligosaccharide substrates were obtained from Wako Pure Chemicals (Osaka, Japan) and Sigma-Aldrich (St. Louis, MO, USA) unless otherwise specified.
Sulfolobus acidocaldarius (NBRC No. 15157) genomic DNA was purchased from the Biological Resource Center (NBRC), National Institute of Technology and Evaluation (Tokyo, Japan). The BL21(DE3) E. coli strain was used for recombinant protein expression (Novagen, Madison, WI, USA), and the DH5α (TOYOBO, Osaka, Japan) and MV1184 (TAKARA BIO, Shiga, Japan) strains were utilized for genetic engineering. Luria-Bertani (LB) medium was used to culture the E. coli used in the genetic engineering and for preculturing prior to protein expression. During protein expression in E. coli, 2× YT liquid medium was used. Ampicillin (50–100 μg/ml) or kanamycin (100 μg/ml) was added to the media as needed.
Cloning trehalase-like genes from S. acidocaldarius and constructing expression plasmids
To amplify the S. acidocaldarius Saci1250 and Saci1816 genes using S. acidocaldarius genomic DNA as a template, PCR reactions were performed as previously described (Sakaguchi et al. 2015). The primer sequences for Saci1250 gene were as follows: the forward primer was 5′-GGAATTCCATATGAATTACGCCTTACTTTCAAACG-3′ and the reverse primer was 5′-CCGCTCGAGTCTTACTTCATCAAACTCCAGTATACTTAATACC-3′ (NdeI site and XhoI site are boldfaced). The primer sequences for Saci1816 gene were as follows: the forward primer was 5′-GGAATTCCATATGTTAGGTATGAAACCTCTAGGATTTATC-3′ and the reverse primer was 5′-CCGCTCGAGTGTACTTAGACCTTTTACTTCCTTCAGAG-3′ (NdeI site and XhoI site are boldfaced). The primers were purchased from Eurofins Genomics (Tokyo, Japan). After trimming the amplified PCR products with NdeI and XhoI, the products were inserted into the pET21a vector (Novagen) to yield pET-Saci1250 and pET-Saci1816 expression vectors. These gene products had C-terminal tails with a hexahistidine tag (His-tag) derived from the vector. The nucleotide sequences were verified by DNA sequencing (Eurofins Genomics). The Saci1250 and Saci1816 nucleotide sequences are available in the DDBJ/EMBL/GenBank database under the accession numbers LC330867 and LC330868, respectively.
Expression, purification, enzyme activity, and protein assays
Each gene product was expressed in E. coli BL21(DE3) and purified. Pre-cultures of transformants grown overnight at 37 °C in LB medium were inoculated into 2× YT medium containing 50 μg/ml ampicillin. The cells were grown at 37 °C for approximately 18 h, then harvested by centrifugation and suspended in TS buffer [20 mM Tris-HCl, 0.5 M NaCl (pH 7.5)] containing Complete Mini (EDTA-free) proteinase inhibitors (Roche Diagnostics, Mannheim, Germany). Following sonication, the crude extracts and the supernatants obtained by centrifugation for 20 min at 20,000×g were assayed for enzymatic activity, and the expression of gene products was judged by SDS-polyacrylamide gel electrophoresis (SDS-PAGE) after staining with Coomassie brilliant blue R-250 (CBB R-250) (Laemmli 1970). The supernatants were treated at 65 °C for 30 min to remove heat-labile E. coli proteins. After centrifugation, the samples were subjected to affinity chromatography using a Ni-Sepharose™ 6 Fast flow column (GE Healthcare, Buckinghamshire, UK) with TS buffer. The bound proteins were eluted with a stepwise increase in imidazole concentration (0, 50, and 500 mM) in TS buffer. The enzymes were eluted in 500 mM imidazole in TS buffer, and each enzyme was confirmed to migrate as a single band by SDS-PAGE analysis. The fractions containing the purified enzymes were dialyzed against TS buffer, pooled at 4 °C, and used for subsequent experiments.
Enzymatic activities were measured at 50 °C in 50 mM acetate buffer (pH 4.0) for SaTreH1 (Saci1816 gene product) and 50 mM Gly-HCl buffer (pH 3.7) for SaTreH2 (Saci1250 gene product), with 90 mM trehalose as a substrate using a previously described method (Sakaguchi et al. 2015). The amount of glucose liberated from the substrates was measured using the F-kit D-glucose (Roche Diagnostics Gmbh, Mannheim, Germany) based on a glucose standard curve as a reference. One unit of enzyme was defined as the amount of the enzyme that hydrolyzed 1 μmol of trehalose in 1 min. Protein concentrations were measured with the micro-assay method (Bio-Rad Laboratories, Hercules, CA, USA) using bovine serum albumin as a standard protein.
Substrate specificity and pH and temperature profile
The substrate specificity of recombinant enzymes was assessed at 50 °C in 50 mM acetate buffer (pH 4.0) using 9 mM trehalose, nigerose (Hayashibara Biochemical Laboratories, Okayama, Japan), maltose, isomaltose, turanose, malturose, or sucrose for up to 3 h. The effect of pH on the enzymatic activities toward 90 mM trehalose at 50 °C was measured at a range of pH values (from pH 3.0 to 5.0) using 50 mM Gly-HCl buffer (pH 3.0–4.0) and acetate buffer (pH 3.3–5.0). To study the pH stability of the enzymes, the enzyme solutions were added to various buffers (pH 3–11) at 4 °C for 60 min. After the treatment, treated enzyme solutions were dialyzed against TS buffer and the activities of SaTreH1 and SaTreH2 toward 90 mM trehalose were measured at 50 °C and pH 4.0 or pH 3.7, respectively. The effect of temperature on the enzymatic activity toward 90 mM trehalose at pH 4.0 or pH 3.7 was examined at 30, 40, 50, 60, 70, or 80 °C. To determine the heat stability of the enzymes, the enzyme solutions were incubated at various temperatures (30–90 °C) for 60 min and then cooled rapidly, and then the remaining activity toward 90 mM trehalose was measured at 50 °C. Differential scanning fluorimetry (DSF) was performed, and the melting temperature (Tm) values were estimated as previously reported (Niesen et al. 2007; Sakaguchi et al. 2015). The experiments were performed in triplicate.
Kinetic analysis of the trehalose hydrolysis
The initial rates of trehalose hydrolysis at various trehalose concentrations were measured in 50 mM acetate (pH 4.0, SaTreH1) and 50 mM Gly-HCl (pH 3.7, SaTreH2). The kinetic parameters Vmax and KM for trehalose were estimated as previously reported (Sakaguchi et al. 2015). The apparent kcat values were estimated using molecular masses of 64.6 kDa (SaTreH1, Saci1816 gene product) and 67.6 kDa (SaTreH2, Saci1250 gene product), which were deduced from the amino acid sequences of the enzymes. The apparent Ki values were estimated according to previous reports (Mertens et al. 2010; Sakaguchi et al. 2014). The experiments were performed in triplicate.
Construction of mutant expression plasmids and the expression, purification, and analysis of the mutant proteins
Site-directed mutagenesis of the Saci1816 gene was performed using the ODA-PCR method (Mutan®-Super Express Km; TAKARA BIO). The oligonucleotide primers for mutagenesis were used as follows (mismatched bases are boldfaced): SaTreH1 E374Q, 5′-GCTGGAATGTGGCAGGACAGAGGAG-3′; SaTreH1 G523S, 5′-GGACGTTGTACCTAATCAGCGAACATATTGATGTAGAG-3′; SaTreH1 E524Q, 5′-ACGTTGTACCTAATCGGTCAGCATATTGATGTAGAGAAC-3′; SaTreH1 H525E, 5′-TTGTACCTAATCGGTGAAGAAATTGATGTAGAGAACATG-3′. The nucleotide sequences of the gene, including the mutation site and other regions, were confirmed by DNA sequencing. For these mutants, the expression, purification, activity measurements, and DSF analyses were performed as described above.
Results
Comparison of Saci1250 and Saci1816 gene products with other GH15 family enzymes
The deduced amino acid sequence encoded by Saci1816 gene exhibited 44% identity and 61% similarity with that of the Saci1250 gene. The amino acid sequence of TVN1315, a trehalase from T. volcanium, exhibited 34 and 35% identity with that of Saci1250 and Saci1816 gene products, respectively. The phylogenetic tree analysis with relevant GH15 family enzyme members is shown in Fig. 1. The analysis yielded two major branches: eukaryotic GAs and bacterial and archaeal enzymes, including GAs, GDases, and trehalases. As expected, Saci1250 and Saci1816 gene products were relatively similar to each other among the characterized GH15 family members and categorized close to the members of prokaryotic enzymes. Therefore, the structures of the two Sulfolobus gene products were also assumed to have N-terminal β-sandwiched and C-terminal (α/α)6-barreled structures similar to other prokaryotic GH15 family members (Aleshin et al. 2003). The primary structures of the putative catalytic domains of the Sulfolobus gene products showed 50% identity with each other.
Phylogenetic relationship of related enzyme sequences. The phylogenetic tree was constructed using MEGA7 (Kumar et al. 2016) and the neighbor-joining method. Bootstrap values based on 1000 replicate are shown at nodes, and the accession number, enzyme name, and organism name are shown. SaTreH2 (Saci1250 gene product) and SaTreH1 (Saci1816 gene product) are indicated by a closed circle
Soluble protein expressions in E. coli and substrate specificity assays for the Sulfolobus gene products
Saci1250 and Saci1816 gene products were expressed as soluble proteins in E. coli BL21(DE3), and the supernatants of cell homogenates specifically hydrolyzed trehalose. When protein expression was induced by isopropyl β-d-thiogalactopyranoside (IPTG; 0.01 or 0.1 mM) in the present expression system, most of the expressed proteins were recovered as insoluble proteins, and only a small amount of Saci1250 and Saci1816 gene products were detected in the supernatant. Saci1816 gene product in the supernatant showed a detectable trehalase activity, whereas the trehalase activity of Saci1250 gene product could not be confirmed at this stage using the Saci1250 gene product-containing extract (data not shown). Protein expression without IPTG induction could substantially improve the quantities of soluble proteins recovered in the supernatants for both Saci1250 and Saci1816 gene products. The expressed gene products were purified via heat treatment and subsequent affinity chromatography to generate single bands in SDS-PAGE analysis (Fig. 2). The N-terminal sequence analysis of the purified Saci1250 and Saci1816 gene products showed MNYALLSN and MLGMKPLGFIGNGLT, respectively, which coincided with the expected N-terminal sequence of the respective genes (Fig. S1). Slight differences between the expected molecular masses of the C-terminal (His-tag)-added Saci1250 gene product (67.6 kDa) and Saci1816 gene product (64.6 kDa) and the apparent molecular masses estimated from the mobilities on SDS-PAGE were observed, which is consistent with Thermoplasma TreHs (Sakaguchi et al. 2015). The discrepancy may be resulted from the hydrophobicity and/or isoelectric point of the protein (Shirai et al. 2008), because the presence of the correct N-terminal and C-terminal His-tag regions was confirmed by a N-terminal sequence analysis and successful affinity chromatography purification, respectively. Based on gel filtration analysis, both gene products may be dimeric proteins (data not shown).
SDS-PAGE analysis of the purified proteins. Lane M, molecular weight marker (Precision Plus Protein™ Standards, Bio-Rad Laboratories); Saci1816, Saci1816 gene product (SaTreH1); and Saci1250, Saci1250 gene product (SaTreH2). The numbers in the margin indicate the molecular masses (kDa) of the proteins in the molecular weight marker
Both purified gene products hydrolyzed trehalose to yield two glucose units, although they had different specific activities, with Saci1816 gene product exhibiting higher trehalose-hydrolyzing activity than Saci1250 gene product. In addition, both gene products were highly specific for trehalose as substrate, and showed little or no activity toward other disaccharides (i.e., nigerose, maltose, isomaltose, turanose, malturose, and sucrose) even after a 3-h reaction at 50 °C. Based on these observations, we identified Saci1816 and Saci1250 gene products from S. acidocaldarius as an archaeal trehalose-degrading enzyme member, S. acidocaldarius TreH, namely SaTreH1 and SaTreH2, respectively (Fig. S1).
pH and temperature effects on SaTreH1 and SaTreH2 and steady-state kinetic analysis
Figure 3a, b illustrates the effects of pH on SaTreH1 and SaTreH2 activity. The trehalose-hydrolyzing activities of SaTreH1 and SaTreH2 were maximal at pH 3.7 (SaTreH2) and pH 4.0 (SaTreH1), and more than half of their maximal activities were observed over a range between pH 3.5 and pH 4.5 and between pH 3.3 and pH 4.0, respectively. The optimal temperature for the enzymatic activity with 90 mM trehalose in 50 mM acetate buffer at pH 4.0 (SaTreH1) or 50 mM Gly-HCl buffer at pH 3.7 (SaTreH2) was approximately 60 °C for SaTreH1 and approximately 70 °C for SaTreH2 (Fig. 3c, d). It is noted that SaTreH1 and SaTreH2 seem to begin partial, though reversible, inactivation at temperatures above 50 and 60 °C, respectively, because each Arrhenius plot was apparently linear below 50 °C but began to deviate from linearity above 50 or 60 °C (Fig. S2A and B). For this reason and also to compare their properties with related Thermoplasma TreHs under the same condition, we carried out the pH- and temperature-related experiments at 50 °C.
Effects of pH and temperature on the activities of SaTreH1 (a, c) and SaTreH2 (b, d). a, b Activity was measured at 50 °C in appropriate buffers at different pH values as follows: 50 mM Gly-HCl buffer (pH 3.0–4.0, circle) and 50 mM acetate buffer (pH 3.3–4.5, square). c, d Activities of SaTreH1 (closed square) or SaTreH2 (open circle) were measured at various temperatures (30–70 °C) in 50 mM acetate buffer (pH 4.0) or 50 mM Gly-HCl buffer (pH 3.7), respectively. The average values for SaTreH1 (closed symbol) and SaTreH2 (open symbol) with error bars are presented. Experiments were performed in triplicate
SaTreH1 and SaTreH2 were stable over a wide pH range. Figure 4a, b shows that SaTreH1 and SaTreH2 retained more than 80% activity after a 60-min incubation at pH 3.5–10 and pH 3.5–11, respectively. We then examined the heat stability of the enzymes and determined the enzymatic activities at 50 °C after 60 min of heat treatment (30 to 90 °C). The results are shown in Fig. 4c, d. SaTreH1 was nearly stable for 60 min at temperatures ranging from 30 to 70 °C, although the activity decreased at and above 80 °C after 60 min. The active site of SaTreH2 showed higher stability than that of SaTreH1 over a range of temperatures (30–90 °C). The DSF analysis could not estimate the apparent protein melting temperature (Tm) values because the inflection points of the transition curves could be observed only above 95 °C, suggesting that the Tm values of SaTreH1 and SaTreH2 were higher than 95 °C. A difference between the optimal temperatures and Tm values was also observed with Thermoplasma TreHs (Sakaguchi et al. 2015).
Stability of trehalase activity, SaTreH1 (a, c) and SaTreH2 (b, d), at different pH values (a, b) and temperatures (c, d). a, b Trehalases were incubated at 4 °C for 60 min at various pH values as follows: 50 mM Gly-HCl buffer (pH 3–4, circle), 50 mM acetate buffer (pH 4–6, square), 50 mM Tris-HCl buffer (pH 7–9, triangle), and 50 mM carbonate-NaOH buffer (pH 10–11, rhombus). After the treatment, treated enzyme solutions were dialyzed against TS buffer and the activities of SaTreH1 (closed symbol) and SaTreH2 (open symbol) toward 90 mM trehalose were measured at 50 °C and pH 4.0 or pH 3.7, respectively. c, d The trehalases were incubated at various temperatures (30 to 90 °C) for 60 min, after which the activity of SaTreH1 (closed square) and SaTreH2 (open circle) were measured at 50 °C. Experiments were performed in triplicate, and the relative values with error bars are shown
We determined the steady-state kinetic parameters of SaTreH1 and SaTreH2 trehalases at 60 or 70 °C, respectively, and at 50 °C. In 50 mM acetate buffer (pH 4.0), the SaTreH1 reaction obeyed Michaelis-Menten kinetics (Fig. S3A), and the kcat and KM values at 60 °C were 102.5 ± 6.10 s−1 and 54.5 ± 6.03 mM and those at 50 °C were 77.0 ± 1.24 s−1 and 41.8 ± 0.67 mM respectively. The KM values and the Michaelis-Menten-type behavior were similar to those of Thermoplasma TreHs (40–49 mM) and Mycobacterium trehalase (20 mM) (Sakaguchi et al. 2015; Carroll et al. 2007). However, SaTreH2 behaved differently from previously described trehalases at 70 and 50 °C in 50 mM Gly-HCl buffer (pH 3.7) and presented kcat values of 3.43 ± 0.03 and 1.60 ± 0.02 s−1 and KM values of 3.47 ± 0.18 and 2.39 ± 0.06 mM, respectively. The KM values were similar to that of E. coli GH37 trehalases and GH65 acidic trehalase, although the kcat values were significantly lower (Table 1). SaTreH2 activity was inhibited with elevated concentrations of trehalose at 50 °C, with apparent Ki value of 260 mM, but the inhibition was attenuated at 70 °C (Fig. S3B and C). Similar inhibition at elevated substrate concentrations was reported on fungal GA, AmyC (Mertens et al. 2010).
Involvement of Glu374, Glu524, Gly523, and His525 in SaTreH1 trehalase activity
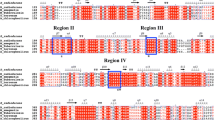
Based on our knowledge of GH15 family enzymes and archaeal trehalases, which possess five conserved regions in the primary structures and have two catalytic residues conserved in regions 3 and 5, we found that two amino acid residues of SaTreH1, Glu374 and Glu524, may correspond to conserved catalytic residues (Figs. 5 and S1). To examine the possible involvement of the two amino acid residues in trehalase activity, we constructed two mutants, E374Q and E524Q, in which Glu374 or Glu524 of SaTreH1 was substituted with a glutamine residue one at a time. The SaTreH1 E374Q and E524Q mutants were expressed in E. coli and purified similarly to wild-type SaTreH1 (Fig. 6). The Tm values of both mutants were higher than 95 °C. The SaTreH1 E374Q and E524Q mutants exhibited greatly reduced hydrolytic activities of 0.001 and 0.04 U/mg, respectively, with 90 mM trehalose at 50 °C in 50 mM acetate buffer (pH 4.0) compared with 48.4 U/mg of SaTreH1. These results indicate that these glutamic acid residues may be involved in SaTreH1 activity and suggest that E382 and E543 of SaTreH2, which present corresponding locations, are similarly involved in trehalase activity (Figs. 5 and S1).
Sequence alignments of the conserved regions (1–5) of archaeal trehalases with selected deduced sequences of relevant genes. The aligned sequences are as follows: SaTreH2; putative gene products of three SaTreH2-related genes (accession numbers: BAB65993, AKA78347, and ADX86426, see Fig. S4); SaTreH1; putative gene products of five SaTreH1-related genes (accession numbers: ADX86414, AKA72943, BAB66158, ABP95480, and ABP95157, see Fig. S4); Ta0286; and TVN1315. Amino acid residues that are identical in seven or more of the aligned enzymes are shaded. The “number sign” symbol denotes the putative catalytic residues
SDS-PAGE analysis of mutant SaTreH1 trehalases. a Lane M, molecular weight marker (Precision Plus Protein™ Standards, Bio-Rad Laboratories); lane 1, E374Q; lane 2, E524Q; lane 3, G523S; and lane 4, H525E. The numbers in the margin indicate the molecular masses (kDa) of the proteins in the molecular weight marker
Notably, SaTreH1 has a -Gly-Glu-His- (GEH) sequence at position 523–525, which is unique among GH15 family enzymes. Thermoplasma TreHs and Clostridium sp. GA have SEE and SEQ sequences, respectively, at corresponding positions. To investigate the role of the glycine and/or histidine residues in the active site of SaTreH1 and the possible significance of differences in these positions between Sulfolobus and Thermoplasma TreHs, we constructed two mutants in which Gly523 or His525 was substituted with serine or glutamic acid residues, respectively, thereby mimicking the TVN1315 sequence. The mutants were expressed, purified (Fig. 6), and examined for their trehalose-hydrolyzing activities. The activity of SaTreH1 G523S (50.0 U/mg) was similar to that of wild-type SaTreH1, although the activity of H525E mutant was reduced to 7.4 U/mg with 90 mM trehalose at 50 °C in 50 mM acetate buffer (pH 4.0). Kinetic parameters of these mutants toward trehalose at 50 °C in 50 mM acetate buffer (pH 4.0) were as follows: the kcat and KM values of G523S were 98.3 ± 2.65 s−1 and 71.6 ± 3.36 mM, and those of H525E were 8.70 ± 0.15 s−1 and 6.43 ± 0.53 mM, respectively. Both mutants showed lower Tm values at approximately 89 °C than did the wild-type SaTreH1, E374Q, or E524Q mutants in the DSF analysis. These results indicate that SaTreH1 His525 is involved in the catalytic reaction and substrate recognition and that both amino acid residues contribute to SaTreH1 stability, although interactions between these residues and other amino acid residues remain unidentified. The configuration of the general backbone as well as the substrate-binding sites and catalytic residues are likely consistent in SaTreH1 and Thermoplasma TreHs, although additional amino acid residues that are not conserved between the two may be required to generate the proper active site structure of Sulfolobus TreHs and confer additional thermal stability to the Sulfolobus enzyme.
Discussion
Recently, we reported two trehalose-hydrolyzing enzymes from Euryarchaeota Thermoplasma species expressed in E. coli as recombinant proteins and proposed a third route of trehalose degradation in archaea, which directly hydrolyzes trehalose to glucose molecules and may regulate trehalose concentration. In this study, we found two archaeal trehalase-like genes in thermoacidophilic Crenarchaeon S. acidocaldarius and demonstrated that they could hydrolyze trehalose under a narrow acidic pH range and elevated temperatures, and they showed distinct specific activities.
SaTreH1 and SaTreH2 (Sulfolobus TreHs) are similar to each other (Figs. 1, 5, and S1). A phylogenic analysis of archaeal GH15 trehalase-like sequences in the CAZy database, which includes a number of functionally unidentified sequences, suggests that the sequences can be divided into Crenarchaeota and Euryarchaeota enzyme groups, and the Crenarchaeota enzyme group is further divided into two subgroups: SaTreH1 and SaTreH2 (Fig. S4). Although the former and the latter subgroup members may have high KM and kcat and low KM and kcat values, respectively, similar to SaTreH1 and SaTreH2, their identification as trehalases has yet to be established.
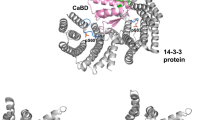
In yeast Saccharomyces cerevisiae, two trehalases, Nth1 and Nth2 (77% identity at the protein level), are also known as neutral trehalases belonging to GH37 family. Evidence has been accumulated indicating that Nth1 is activated through cAMP-dependent kinase (PKA) phosphorylation, Ca2+ binding, and the 14-3-3 protein binding (Uno et al. 1983; Ortiz et al. 1983; Franco et al. 2003; Panni et al. 2008; Veisova et al. 2012; Kopecka et al. 2014; Alblova et al. 2017). Nth2 also showed trehalose hydrolyzing activity at neutral pH range (Jules et al. 2008). However, the activation process of Nth2 has not been reported despite of its high identity with Nth1 at protein level.
SaTreH2 shares 44% identity with SaTreH1 and possesses putative catalytic residues at positions corresponding to those of SaTreH2, but it exhibits low trehalose-hydrolyzing activity. In addition, SaTreH2 has a significantly lower KM value than SaTreH1. The reason for this difference remains unclear at present. A number of amino acid residues are conserved among GH15 archaeal TreHs (Figs. 5 and S1) in five conserved regions. These amino acid residues are likely involved in forming active site and recognizing the substrate. However, some of these conserved residues are not conserved in SaTreH2. These SaTreH2-unique residues as well as yet unidentified amino acid residue(s) in SaTreH2 may contribute to enhanced trehalose affinity but negatively affect the catalytic reaction. Minor changes not involving catalytic residues may lead to profound changes in catalytic properties. Thus, the chitinolytic activity of human acidic mammalian chitinases (AMCase) is significantly lower than that of its mouse counterpart, although Okawa et al. (2016) reported that human AMCase activity was increased to a level comparable to its mouse homolog through amino acid-substitutions in a conserved region of the mammalian AMCase, not including the catalytic residues. It should be noted in this context that SaTreH1 mutant H525E showed lower kcat and KM values than those of SaTreH1.
Considering the activation process for yeast neutral trehalase described above, phosphorylation as well as interaction with regulatory protein(s) in Sulfolobus sp. may activate SaTreH2, because potential protein kinase/phosphatases are reported in certain archaea (Kennelly 2014; Iakoucheva et al. 2004; Fig. S1). Alternatively, SaTreH2 may have the potential to hydrolyze other sugars not yet tested. Certain Sulfolobus species and Crenarchaeota Metallosphaera sedula appear to possess two or more GH15 enzymes acting on trehalose or other sugars (Fig. S4).
Recently, Moon et al. (2016) identified a trehalose-hydrolyzing enzyme in the cell-free extract of S. acidocaldarius DSM639. The purified enzyme had a molecular mass of 40 kDa and was optimally active at high temperatures (85 °C) and under acidic conditions (pH 4.5). The KM value for trehalose hydrolysis was 6.6 mM, and the specific activity was 2.71 U/mg under optimal conditions. Our results indicate that S. acidocaldarius has two trehalose-hydrolyzing enzymes with considerably different catalytic properties. Saci1816 and Saci1250 genes are therefore candidates for the unidentified trehalase gene in S. acidocaldarius. The catalytic properties of trehalase reported by Moon et al. (2016) are similar to those of SaTreH2. SaTreH2 (Saci1250 gene product) appears to be constitutively expressed in the thermoacidophilic archaeon S. acidocaldarius and functions as a processed form lacking an N-terminal region. Alternatively, truncated SaTreH1 (Saci1816 gene product), whose activity was partly impaired through processing, may represent the trehalase reported by Moon et al. (2016). In the GH15 family GAs, the N-terminal domain has been reported to be important for enzyme activity (Li et al. 2014; Sakaguchi et al. 2017). However, further studies are needed to clarify this issue.
Trehalose accumulates in archaea, such as Sulfolobus and Thermoplasma species (Martins et al. 1997; Moon et al. 2016). Of the three enzymes implicated in trehalose degradation (Iturriaga et al. 2009), Moon et al. (2016) reported the stationary phase cell extract showed high trehalose-hydrolyzing activity (TreH) but little trehalose glycosyltransferring synthase (TreT) and trehalose phosphorylase (TreP) activities. Most likely, a processed form of either SaTreH1 or SaTreH2, which represent the Sulfolobus trehalase (TreH), is involved in regulating the trehalose content. Although trehalose metabolism in archaea is poorly understood, the discovery of archaeal trehalose-hydrolyzing enzymes that belong to the GH15 family and present distinct characteristics will help in providing insights into trehalose metabolism in archaea and extend our understanding of the structure-function relationships of GH15 family enzymes as well as other carbohydrate-active enzymes.
References
Alblova M, Smidova A, Docekal V, Vesely J, Herman P, Obsilova V, Obsil T (2017) Molecular basis of the 14-3-3 protein-dependent activation of yeast neutral trehalase Nth1. Proc Natl Acad Sci U S A 114:E9811–E9820. https://doi.org/10.1073/pnas.1714491114
Aleshin AE, Feng PH, Honzatko RB, Reilly PJ (2003) Crystal structure and evolution of a prokaryotic glucoamylase. J Mol Biol 327:61–73. https://doi.org/10.1016/S0022-2836(03)00084-6
App H, Holzer H (1989) Purification and characterization of neutral trehalase from the yeast ABYS1 mutant. J Biol Chem 264:17583–17588
Argüelles JC (2000) Physiological roles of trehalose in bacteria and yeasts: a comparative analysis. Arch Microbiol 174:217–124. https://doi.org/10.1007/s002030000192
Barraza A, Sánchez F (2013) Trehalases: a neglected carbon metabolism regulator? Plant Signal Behav 8:e24778. https://doi.org/10.4161/psb.24778
Carroll JD, Pastuszak I, Edavana VK, Pan YT, Elbein AD (2007) A novel trehalase from Mycobacterium smegmatis—purification, properties, requirements. FEBS J 274:1701–1714. https://doi.org/10.1111/j.1742-4658.2007.05715.x
DeBosch BJ, Heitmeier MR, Mayer AL, Higgins CB, Crowley JR, Kraft TE, Chi M, Newberry EP, Chen Z, Finck BN, Davidson NO, Yarasheski KE, Hruz PW, Moley KH (2016) Trehalose inhibits solute carrier 2A (SLC2A) proteins to induce autophagy and prevent hepatic steatosis. Sci Signal 9:ra21. https://doi.org/10.1126/scisignal.aac5472
Elbein AD, Pan YT, Pastuszak I, Carroll D (2003) New insights on trehalose: a multifunctional molecule. Glycobiology 13:17–27. https://doi.org/10.1093/glycob/cwg047
Franco A, Soto T, Vicente-Soler J, Paredes V, Madrid M, Gacto M, Cansado J (2003) A role for calcium in the regulation of neutral trehalase activity in the fission yeast Schizosaccharomyces pombe. Biochem J 376:209–217. https://doi.org/10.1042/bj20030825
Higashiyama T (2002) Novel functions and applications of trehalose. Pure Appl Chem 74:1263–1269. https://doi.org/10.1351/pac200274071263
Horlacher R, Uhland K, Klein W, Ehrmann M, Boos W (1996) Characterization of a cytoplasmic trehalase of Escherichia coli. J Bacteriol 178:6250–6257. https://doi.org/10.1128/jb.178.21.6250-6257.1996
Iakoucheva LM, Radivojac P, Brown CJ, O’Connor TR, Sikes JG, Obradovic Z, Dunker AK (2004) The importance of intrinsic disorder for protein phosphorylation. Nucleic Acids Res 32:1037–1049. https://doi.org/10.1093/nar/gkh253
Iturriaga G, Suárez R, Nova-Franco B (2009) Trehalose metabolism: from osmoprotection to signaling. Int J Mol Sci 10:3793–3810. https://doi.org/10.3390/ijms10093793
Jules M, Beltran G, François J, Parrou JL (2008) New insights into trehalose metabolism by Saccharomyces cerevisiae: NTH2 encodes a functional cytosolic trehalase, and deletion of TPS1 reveals Ath1p-dependent trehalose mobilization. Appl Environ Microbiol 74:605–614. https://doi.org/10.1128/AEM.00557-07
Kennelly PJ (2014) Protein Ser/Thr/Tyr phosphorylation in the archaea. J Biol Chem 289:9480–9487. https://doi.org/10.1074/jbc.R113.529412
Kopecka M, Kosek D, Kukacka Z, Rezabkova L, Man P, Novak P, Obsil T, Obsilova V (2014) Role of the EF-hand-like motif in the 14-3-3 protein-mediated activation of yeast neutral trehalase Nth1. J Biol Chem 289:13948–13961. https://doi.org/10.1074/jbc.M113.544551
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227:680–685. https://doi.org/10.1038/227680a0
Li Z, Wei P, Cheng H, He P, Wang Q, Jiang N (2014) Functional role of β-domain in the Thermoanaerobacter tengcongensis glucoamylase. Appl Microbiol Biotechnol 98:2091–2099. https://doi.org/10.1007/s00253-013-5051-2
Maicas S, Guirao-Abad JP, Argüelles JC (2016) Yeast trehalases: two enzymes, one catalytic mission. Biochim Biophys Acta 1860:2249–2254. https://doi.org/10.1016/j.bbagen.2016.04.020
Martins LO, Huber R, Huber H, Stetter KO, Da Costa MS, Santos H (1997) Organic solutes in hyperthermophilic archaea. Appl Environ Microbiol 63:896–902
Mertens JA, Braker JD, Jordan DB (2010) Catalytic properties of two Rhizopus oryzae 99-880 glucoamylase enzymes without starch binding domains expressed in Pichia pastoris. Appl Biochem Biotechnol 162:2197–2213. https://doi.org/10.1007/s12010-010-8994-0
Mittenbühler K, Holzer H (1988) Purification and characterization of acid trehalase from the yeast suc2 mutant. J Biol Chem 263:8537–8543
Moon JH, Lee W, Park J, Choi KH, Cha J (2016) Characterization of a trehalose-degrading enzyme from the hyperthermophilic archaeon Sulfolobus acidocaldarius. J Biosci Bioeng 122:47–51. https://doi.org/10.1016/j.jbiosc.2015.12.011
Niesen FH, Berglund H, Vedadi M (2007) The use of differential scanning fluorimetry to detect ligand interactions that promote protein stability. Nat Protoc 2:2212–2221. https://doi.org/10.1038/nprot.2007.321
Okawa K, Ohno M, Kashimura A, Kimura M, Kobayashi Y, Sakaguchi M, Sugahara Y, Kamaya M, Kino Y, Bauer PO, Oyama F (2016) Loss and gain of human acidic mammalian chitinase activity by nonsynonymous SNPs. Mol Biol Evol 33:3183–3193. https://doi.org/10.1093/molbev/msw198
Ortiz CH, Maia JC, Tenan MN, Braz-Padrão GR, Mattoon JR, Panek AD (1983) Regulation of yeast trehalase by a monocyclic, cyclic AMP-dependent phosphorylation-dephosphorylation cascade system. J Bacteriol 153:644–651
Panni S, Landgraf C, Volkmer-Engert R, Cesareni G, Castagnoli L (2008) Role of 14-3-3 proteins in the regulation of neutral trehalase in the yeast Saccharomyces cerevisiae. FEMS Yeast Res 8:53–63. https://doi.org/10.1111/j.1567-1364.2007.00312.x
Sakaguchi M, Matsushima Y, Nankumo T, Seino J, Miyakawa S, Honda S, Sugahara Y, Oyama F, Kawakita M (2014) Glucoamylase of Caulobacter crescentus CB15: cloning and expression in Escherichia coli and functional identification. AMB Express 4:5. https://doi.org/10.1186/2191-0855-4-5
Sakaguchi M, Shimodaira S, Ishida S, Amemiya M, Honda S, Sugahara Y, Oyama F, Kawakita M (2015) Identification of GH15 family thermophilic archaeal trehalases that function within a narrow acidic pH range. Appl Environ Microbiol 81:4920–4931. https://doi.org/10.1128/AEM.00956-15
Sakaguchi M, Matsushima Y, Nagamine Y, Matsuhashi T, Honda S, Okuda S, Ohno M, Sugahara Y, Shin Y, Oyama F, Kawakita M (2017) Functional dissection of the N-terminal sequence of Clostridium sp. G0005 glucoamylase: identification of components critical for folding the catalytic domain and for constructing the active site structure. Appl Microbiol Biotechnol 101:2415–2425. https://doi.org/10.1007/s00253-016-8024-4
Sarkar S, Davies JE, Huang Z, Tunnacliffe A, Rubinsztein DC (2007) Trehalose, a novel mTOR independent autophagy enhancer, accelerates the clearance of mutant huntingtin and α-synuclein. J Biol Chem 282:5641–5652. https://doi.org/10.1074/jbc.M609532200
Shirai A, Matsuyama A, Yashiroda Y, Hashimoto A, Kawamura Y, Arai R, Komatsu Y, Sueharu Horinouchi, Minoru Yoshida, (2008) Global analysis of gel mobility of proteins and its use in target identification. J Biol Chem 283(16):10745–10752
Tourinho-dos-Santos CF, Bachinski N, Paschoalin VM, Paiva CL, Silva JT, Panek AD (1994) Periplasmic trehalase from Escherichia coli: characterization and immobilization on spherisorb. Braz J Med Biol Res 27:627–636
Uno I, Matsumoto K, Adachi K, Ishikawa T (1983) Genetic and biochemical evidence that trehalase is a substrate of cAMP-dependent protein kinase in yeast. J Biol Chem 258:10867–10872
Veisova D, Macakova E, Rezabkova L, Sulc M, Vacha P, Sychrova H, Obsil T, Obsilova V (2012) Role of individual phosphorylation sites for the 14-3-3-protein-dependent activation of yeast neutral trehalase Nth1. Biochem J 443:663–670. https://doi.org/10.1042/BJ20111615
Acknowledgements
We are grateful to Yukari Saisaka (High-Tech Research Center, Meiji Pharmaceutical University) for performing the N-terminal sequence analysis and Misa Ohno, Kazuaki Okawa, and Satoshi Wakita for their valuable suggestions.
Funding
This study was supported by the Japan Society for the Promotion of Science (JSPS) KAKENHI (Grant Number 17 K07729); the Strategic Research Foundation Grant-aided Project for Private Universities of the Ministry of Education, Culture, Sport, Science, and Technology, Japan (MEXT) (Grant Number S1411005); the Science Research Promotion Fund from the Promotion and Mutual Aid Corporation for Private Schools of Japan; and the Project Research Grant from the Research Institute of Science and Technology, Kogakuin University.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals.
Electronic supplementary material
ESM 1
(PDF 267 kb)
Rights and permissions
About this article
Cite this article
Yuasa, M., Okamura, T., Kimura, M. et al. Two trehalose-hydrolyzing enzymes from Crenarchaeon Sulfolobus acidocaldarius exhibit distinct activities and affinities toward trehalose. Appl Microbiol Biotechnol 102, 4445–4455 (2018). https://doi.org/10.1007/s00253-018-8915-7
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-8915-7