Abstract
Genetic engineering of transcription factors is an efficient strategy to improve lignocellulolytic enzyme production in fungi. In this study, the xylanase transcriptional regulators of Trichoderma reesei (Xyr1) and Neurospora crassa (XLR-1), as well as their constitutively active mutants (Xyr1A824V and XLR-1A828V), were heterologously expressed in Penicillium oxalicum. The two heterologous regulators were identified to be able to activate lignocellulolytic enzyme gene expression in P. oxalicum. Particularly, expression of T. reesei Xyr1 resulted in a higher cellulase production level compared with the expression of native xylanase transcriptional regulator XlnR using the same promoter. Xyr1A824V and XLR-1A828V were found to be able to confer P. oxalicum more enhanced lignocellulolytic abilities than wild-type regulators Xyr1 and XLR-1. Furthermore, introduction of regulatory modules containing Xyr1A824V/XLR-1A828V and their target cellulase genes resulted in greater increases in cellulase production than alone expression of transcriptional regulators. Through the cumulative introduction of three regulatory modules containing regulator mutants and their corresponding target cellulase genes from P. oxalicum, T. reesei, and N. crassa, a 2.8-fold increase in cellulase production was achieved in P. oxalicum.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Lignocellulosic materials are sustainable resources for biofuel production due to their vast abundance and renewable nature (Kricka et al. 2015). Filamentous fungi such as Penicillium oxalicum (Liu et al. 2013, 2014), Trichoderma reesei (Kubicek et al. 2009; Peterson and Nevalainen 2012), or Neurospora crassa (Coradetti et al. 2012; Tian et al. 2009) can produce hydrolytic enzymes to synergistically deconstruct lignocellulosic biomass. However, the high cost of lignocellulolytic enzymes is a major bottleneck to the development of an economically viable lignocellulosic ethanol industry (Ellila et al. 2017). Thus, how to increase the production of lignocellulolytic enzyme is an important problem that needs to be solved.
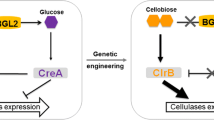
P. oxalicum has a strong cellulolytic ability in saprophytic conditions due to its efficient production of lignocellulolytic enzymes (Fang et al. 2010; Saini et al. 2015). P. oxalicum strain 114-2 has been studied for cellulase production for more than 30 years in China (Gao et al. 2017a). Data from comparative genomics analysis showed that P. oxalicum has a more diverse set of lignocellulolytic enzymes than many other cellulolytic fungi, such as T. reesei (Liu et al. 2013). After genetic engineering of 114-2, a high cellulase productivity of 158.38 U/L/h was reached by recombinant strain RE-10 (Han et al. 2017). In filamentous fungi, the expression of lignocellulolytic enzyme genes is triggered by inducers released from complex plant polysaccharides and tightly regulated by transcription factors (TFs) (Glass et al. 2013; Amore et al. 2013). It has been proved that cellobiose is an inducer of lignocellulolytic enzyme expression in P. oxalicum (Chen et al. 2013). In our previous work, TFs including CreA, ClrB, XlnR, and AmyR were identified as the major regulators of lignocellulolytic enzyme genes in P. oxalicum (Li et al. 2015). The major carbon catabolite repressor CreA plays a negative role in lignocellulolytic enzyme gene expression through carbon catabolite repression (CCR). ClrB was identified to be necessary for efficient cellulase production, and its deletion incurred significantly reduced growth on cellulose (Li et al. 2015). XlnR is the main activator regulating xylanase gene expression and is moderately involved in cellulase production (Li et al. 2015).
XlnR homologs are commonly found in the genomes of filamentous ascomycetes and are essential to the expression of xylanolytic enzyme genes in many species. Nevertheless, the degree of XlnR homologs’ involvement in cellulase regulation differs greatly depending on the species of fungi (Klaubauf et al. 2014). In T. reesei and Aspergillus niger, Xyr1/XlnR was proved to be essential for the expression of both cellulolytic and xylanolytic enzyme genes (Raulo et al. 2016; Stricker et al. 2006). However, in N. crassa, XLR-1 is required for the expression of xylanolytic enzyme genes but is only slightly involved in the expression of cellulase genes (Sun et al. 2012). The genome-wide binding targets of XLR-1 have been identified by the chromatin immunoprecipitation method, which include mainly xylanolytic and xylose-catabolic genes (Craig et al. 2015). The reported consensus binding sequences of T. reesei Xyr1 (5′-GGC(A/T)3-3′) (Rauscher et al. 2006; Furukawa et al. 2009) and N. crassa XLR-1 (5′-GGNTAAA-3′) (Craig et al. 2015) are somewhat different. However, whether this difference results in different regulons between the two TFs remains unclear.
A series of genetic engineering strategies for enhancing lignocellulolytic enzyme production have been performed in lignocellulolytic fungi. Among the strategies, genetic engineering of TFs is efficient due to the simultaneous upregulation of multiple lignocellulolytic enzyme genes (Yao et al. 2015). Deletion of repressor gene creA to release CCR and overexpression of positive regulators (e.g., ClrB and XlnR) to boost transcriptional activation were reported to enhance the production of lignocellulolytic enzymes in several fungal species (Coradetti et al. 2013; Li et al. 2015; Nakari-Setala et al. 2009; Yao et al. 2015). In addition, constitutive activation of transcriptional activators was shown to lead to inducer-independent expression and enhanced induction of lignocellulolytic enzyme genes (Alazi and Ram 2018). For example, a point mutation in T. reesei Xyr1 (A824V) is responsible for constitutively activated production of both endo-xylanases and cellulases (Derntl et al. 2013). The homologous point mutations in XLR-1 of N. crassa and in XlnR of P. oxalicum were also identified to be able to enhance xylanase genes expression (Craig et al. 2015; Gao et al. 2017a). Furthermore, combinatorial genetic manipulation of TF genes and other genes (such as deletion of the intracellular β-glucosidase gene to accumulate inducing molecules or overexpression of cellulase genes) could greatly increase the production level and/or degradation efficiency of lignocellulolytic enzyme mixtures (Chen et al. 2013; Gao et al. 2017a; Qian et al. 2017; Yao et al. 2016).
Although genetic techniques have been well developed, the number of selective markers used for genetic manipulations in P. oxalicum is limited (Li et al. 2015; Yao et al. 2015). Bacterial β serine recombinase (β-rec) can act on six sites flanking a selective marker gene and excises the marker gene sequence between these two six sites, which allows marker reuse in further genetic manipulations (Canosa et al. 1996; Hartmann et al. 2010). The β-rec gene could be controlled by an inducible promoter, which makes the excision step easy to implement (Hartmann et al. 2010). Such β-rec/six self-excising marker recycling system has been established in several fungal species such as N. crassa (Szewczyk et al. 2014), but has not yet been adopted in P. oxalicum.
Considering the overlapping but distinct regulatory target sets of XlnR homologs from different fungal species, it is of interest to test their functions in a heterologous background. On the other hand, expression of heterologous TFs with effective regulatory functions could be used for the engineering of organisms for industrial purposes (Kang et al. 2017; Wang et al. 2007). In this study, we introduced heterologous XlnR homologs from T. reesei and N. crassa, which have relatively clear research backgrounds on regulatory functions and binding sites, into P. oxalicum. The two TFs, as well as their constitutively active mutants (Xyr1A824V and XLR-1A828V), were functionally expressed in P. oxalicum. Also, heterologous regulatory modules containing Xyr1A824V/XLR-1A828V and their target cellulase genes were designed and expressed in P. oxalicum. Besides, the β-rec/six marker recycling system was established in P. oxalicum and used to construct several high-yield lignocellulolytic enzyme-producing strains by integration of combinatory regulatory modules.
Materials and methods
Strains and culture conditions
The mutant M12 (a uracil auxotroph, pyrGQ226*) was derived from P. oxalicum wild-type strain 114-2 (Qin et al. 2013). The strain 114-2 has been deposited at the China General Microbiological Culture Collection Center (CGMCC) under the number of CGMCC 5302. M12 and all mutants constructed from this strain are listed in Table 1. Wheat bran extract agar contained 10% wheat bran extract and 2% agar power. The liquid glucose medium comprised 1× Vogel’s salts (Vogel 1956) and 2% (w/v) glucose. Xylan medium contained 1× Vogel’s salts and 1% xylan (Sigma, St. Louis, USA). Cellulose medium contained 1× Vogel’s salts and 2% microcrystalline cellulose (Sangon Biotech, Shanghai, China). The complex carbon medium was composed of wheat bran (4.66%), corn cob residue (2.00%), soybean cake powder (1.00%), microcrystalline cellulose (0.60%), KH2PO4 (0.30%), NaNO3 (0.28%), (NH4)2SO4 (0.20%), urea (0.10%), and MgSO4 (0.05%).
All the strains used in this study were cultivated on wheat bran extract agar at 30 °C for 3–5 days to harvest conidia. Fresh conidial suspension was added into a 100-mL liquid glucose medium in 300-mL flasks, and cultivated for 20 h to collect mycelia. The mycelia were transferred to a 50-mL cellulose medium for RNA extraction, or transferred to a 50-mL complex carbon medium for enzyme activity assays. All liquid cultures were cultivated in 300-mL flasks at 30 °C and 200 rpm in constant light. Uracil (0.5 g/L) was added to the culture of all uracil auxotrophic strains.
Construction of strains
Construction of strains ΔxlnR, g-PxlnR, g-PxlnRA871V, g-Txyr1, g-Txyr1A824V, g-Nxlr-1, and g-Nxlr-1A828V
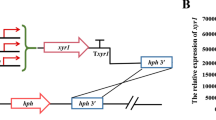
The knockout cassette xlnRU-pyrG-xlnRD was constructed by double-joint PCR (Yu et al. 2004) and introduced to strain M12, and the correct strain was named as ΔxlnR. Then, the strain ΔxlnR was used as a parent strain for the integration of expression cassettes xlnRU-gpdA(p)::PxlnR(or PxlnRA871V/Txyr1/Txyr1A824V/Nxlr-1/Nxlr-1A828V)-hph-xlnRD, respectively. The expression cassettes were introduced into the PxlnR locus of the mutant ΔxlnR by homologous recombination. Accordingly, the strains g-PxlnR, g-PxlnRA871V, g-Txyr1, g-Txyr1A824V, g-Nxlr-1, and g-Nxlr-1A828V were obtained.
Construction of strains DB2-pyrG and DB2
The gene β-rec encoding β-recombinase was synthesized and placed under the control of the promoter of Pbgl2. The selection marker pyrG was obtained from Aspergillus nidulans and was under the control of its native promoter and terminator. Besides, the six sequences were jointed on both sides of pyrG. The knockout cassette Pbgl2(p)::β-rec-pyrG-bgl2D was constructed by a double-joint PCR method, and was transformed into strain M12, which generated mutant DB2-pyrG. To excise the pyrG sequence, strain DB2-pyrG was cultivated in the cellulose medium for 3 days to induce the expression of β-rec. Then, the uracil auxotrophic mutants were screened on solid medium containing 2% glucose, 2% agar power, 0.05% uracil, and 0.20% 5-fluoroorotic acid after cultivation at 30 °C for 4 days. Finally, the resulting strain without pyrG was obtained and named DB2. The schematic representation of the construction of strain DB2 and the establishment of the β-rec/six self-excising marker recycling system is shown in Supplemental Fig. S1.
Construction of strains T-xyr1A824V, N-xlr-1A828V, T-ceT, P-ceT, and P-ceN
The expression cassettes were constructed as indicated in Table 1 and transformed into strain DB2 to generate the expected strains by random insertion.
Construction of strains RE-1-1, RE-1-2, RE-2-1, RE-3-1, RE-3-2, RE-4-1, RE-4-2, RE-5-1, and RE-5-2
The expression cassettes of regulatory modules as indicated in Table 1 were constructed by using the ExoCET direct cloning method (Wang et al. 2018). The strains RE-3-1, RE-4-1, and RE-5-1 were induced by cellulose, respectively, and generated pyrG marker-free strains RE-3-2, RE-4-2, and RE-5-2. All the primers used in this study are provided in Supplemental Table S1.
For all the transformation of P. oxalicum, protoplasts were prepared based on a modified method as described by Gruber et al. (1990). Polyethylene glycol-mediated protoplast transformation was performed according to the method employed by Li et al. (2015).
Phenotype observation
To analyze the phenotypes of mutants on cellulose medium, equal volumes of conidia (107/mL) of strains were spotted on Vogel’s medium plate which contained 1% microcrystalline cellulose as sole carbon source (supplemented with 0.5% Triton X-100) (Dingguo Corp., Beijing, China) at 30 °C for 6 days.
RT-qPCR
Strains were pre-cultured in a glucose medium containing 2% glucose at 30 °C, 200 rpm for 20 h. The mycelia were harvested via vacuum filtration and transferred to 1× Vogel’s salt medium without carbon source for 2 h. The mycelia were collected and then transferred to a 2% cellulose medium for continued cultivation. Total RNA was extracted with the RNAiso™ reagent (TaKaRa, Otsu, Japan). Then, cDNA synthesis was performed using PrimeScript RT Reagent Kit (TaKaRa, Otsu, Japan) according to the manufacturer’s instructions. The RT-qPCR amplification was performed on a LightCycler instrument (Roche, Mannheim, Germany) as previously described (Li et al. 2015). The transcription level of actin was used as an internal standard. All primers used in this study are provided in Supplemental Table S1.
Determination of gene copy numbers by qPCR
To identify the copy numbers of the integrated xlnR, xyr1, or xlr-1 genes in the engineered strains, the genomic DNA was isolated from the mycelia as described by Pentillä et al. (1987) and used as the template for qPCR. The qPCR amplification was performed on a LightCycler instrument (Roche, Mannheim, Germany) as previously described (Li et al. 2015). The actin gene was used to represent a single copy gene. The primers used for qPCR are listed in Supplemental Table S1.
Enzyme activity assays
Filter paper enzyme (FPase) activity was measured using a Whatman No. 1 filter paper (GE Healthcare, Buckinghamshire, UK) as substrate. The reaction system contained a 0.05-g filter paper, a 1.5-mL acetate buffer (pH 4.8), and 0.5 mL of diluted enzyme fraction. The mixtures were maintained at 50 °C for 60 min. CMCase (indicating endoglucanase) and xylanase were assayed using carboxymethylcellulose sodium salt (CMC-Na) (Sigma, St. Louis, USA) and beechwood xylan (Sigma, St. Louis, USA) as the substrates, respectively. The substrate was dissolved in acetate buffer to a final concentration of 1% (w/v). For the assay of CMCase and xylanase activities, 0.5-mL diluted culture supernatants were added to 1.5-mL substrate solutions and incubated at 50 °C for 30 min. The 3,5-dinitrosalicylic acid method was used to determine the amount of released reducing sugars (Miller 1959).
The determination process of pNPCase (indicating cellobiohydrolase) and pNPXase (indicating β-xylosidase) activity was as follows: 50 μL 1 mg/mL pNPC solution (p-nitrophenyl-D-cellobioside, with 10 mg/mL D-glucono-1,5-δ-lactone added in pNPC solution to inhibit the activity of β-glucosidases) or pNPX (p-nitrophenyl-D-xylopyranoside) solution and 100 μL diluted supernatant were maintained at 50 °C for 30 min. Then, 150 μL 10% sodium carbonate was added to stop the reaction. The substrates pNPC/pNPX were purchased from Sigma (St. Louis, USA). Absorbance of the reaction system was measured at 420 nm.
One unit of enzyme activity was defined as the amount of enzyme required to release 1 μmol of product (glucose equivalent/p-nitrophenol) per minute.
Extracellular protein concentration was determined by using Bradford Kit (Sangon Biotech, Shanghai, China).
SDS-PAGE analysis
The mixture of culture supernatants and 5× loading buffers were boiled for 10 min and loaded onto a 12.5% polyacrylamide gel. Coomassie brilliant blue R250 (Sangon, Shanghai, China) was used for staining. Then, the protein gel was washed by a destaining solvent (methanol, acetic acid, and water, 1:1:8, v/v/v) until the background turned clear.
Secretome analysis
The culture broth in complex carbon medium was centrifugated at 12000 rpm for 10 min, and the supernatant was collected. Two hundred micrograms of proteins for each sample were incorporated into SDT buffer (4% SDS, 100 mM DTT, 150 mM Tris-HCl, pH 8.0) (GenView, Florida, USA). The UA buffer (8 M urea, 150 mM Tris-HCl, pH 8.0) was used for repeated ultrafiltration (microcon units, 10 kDa) to remove DTT, detergent, and other low-molecular-weight components. The filters were washed with UA buffer three times and then with 25 mM NH4HCO3 buffer twice. Finally, the protein suspensions were digested overnight with 4 μg trypsin (Sangon, Shanghai, China) in 40 μL 25 mM NH4HCO3 buffer, and the resulting peptides were collected as a filtrate. The peptides of each sample were desalted on C18 cartridges (Empore™ SPE Cartridges C18), concentrated by vacuum centrifugation and reconstituted in 40 μL of 0.1% (v/v) formic acid. LC-MS/MS analysis was performed on a Q Exactive mass spectrometer that was coupled to Easy-nLC (Thermo Fisher Scientific, Waltham, USA). MS data were acquired using a data-dependent top10 method dynamically choosing the most abundant precursor ions from the survey scan (300–1800 m/z) for HCD fragmentation. MS/MS queries were analyzed using Mascot search engine 2.2 (Matrix Science, Ltd., London, UK). The resulting peptide sequences were mapped to an in-house protein sequence database containing the target enzymes (cellobiohydrolase I from T. reesei, GenBank acc. no. XP_006969224.1, endoglucanase I from T. reesei, XP_006965674.1, cellobiohydrolase I from N. crassa, GenBank acc. no. EAA33262.1, and endoglucanase I from N. crassa, EAA29875.2).
Statistical tests
Student’s t tests (one-tailed distribution, homoscedastic) were performed to study the significances of differences between samples by using Microsoft Excel 2013 (Microsoft, Redmond, USA).
Results
Functional expression of heterologous XlnR homologs in P. oxalicum
The putative binding sites of T. reesei Xyr1 and N. crassa XLR-1 were detected in the promoter regions of a major part of lignocellulolytic enzyme genes in P. oxalicum (Supplemental Table S2), implying that the two regulators might play a regulatory role in this fungus. In order to explore the functions of heterologous XlnR homologs in P. oxalicum, a parent strain with the absence of native XlnR was first constructed. M12, a uracil auxotrophic mutant (pyrGQ226*) derived from P. oxalicum wild-type strain 114-2 (Qin et al. 2013), was used as the starting strain in the study. The gene PxlnR (xlnR of P. oxalicum, GenBank acc. no. EPS32714.1) in strain M12 was deleted by knockout and this generated mutant ΔxlnR. Compared with strain M12, ΔxlnR showed lower cellulase activity and faint xylanase activity in a complex carbon medium (Supplemental Fig. S2). The results were similar to the phenotypes of the strain suffering the same modification in the wild-type genetic background (Li et al. 2015). Then, TF-encoding genes Txyr1 (xyr1 from T. reesei, EGR48040.1), Nxlr-1 (xlr-1 from N. crassa, EAA33375.1), and PxlnR were introduced into the PxlnR locus of the mutant ΔxlnR by using the same promoter gpdA(p), respectively. The obtained strains were named g-Txyr1, g-Nxlr-1, and g-PxlnR.
The constructed strains were cultivated in cellulose medium to examine the transcriptional levels of lignocellulolytic enzyme genes. As shown in Fig. 1a, the transcriptional levels of major lignocellulolytic enzyme genes Pcbh1 (cellobiohydrolase gene, EPS32984.1), Pcbh2 (cellobiohydrolase gene, EPS32164.1), Peg1 (endoglucanase gene, EPS32968.1), and particularly Pxyn10A (xylanase gene, EPS33132.1) showed significant increases in all the three mutants when compared with parent strain ΔxlnR (except for cbh2 of g-PxlnR). Interestingly, all the four genes showed the highest transcriptional level in strain g-Txyr1, followed by g-Nxlr-1 and g-PxlnR (Fig. 1a). To further analyze the lignocellulolytic enzyme-producing abilities of strains, the mutants were further cultivated in a complex medium containing wheat bran and corn cob residue. Compared with the parent strain ΔxlnR, all the three mutants showed higher activities of FPase, CMCase, pNPCase, xylanase, and β-xylosidase (Fig. 1b, c, Supplemental Fig. S3). The mutant g-Txyr1 showed the highest cellulase and β-xylosidase activities, while mutant g-PxlnR exhibited the highest xylanase activity, among the strains. The mutant g-Nxlr-1 showed lower cellulase and hemicellulase activities than g-Txyr1 and g-PxlnR. From the above data, we concluded that Xyr1 from T. reesei and XLR-1 from N. crassa were active in P. oxalicum, and both of which could join the regulatory system of lignocellulolytic enzyme genes in P. oxalicum. Particularly, Xyr1 from T. reesei showed superiority relative to native XlnR and XLR-1 from N. crassa.
Transcription level and lignocellulolytic enzyme activity analyses of strains expressing XlnR homologs. a Transcriptional level of lignocellulolytic enzyme genes in strains g-PxlnR, g-Txyr1, and g-Nxlr-1 versus those in ΔxlnR strain (set to one) cultured in cellulose medium at 4 h. b, c FPase and xylanase activities of strains ΔxlnR, g-PxlnR, g-Txyr1, and g-Nxlr-1 cultured in complex carbon medium, respectively. d Transcriptional level of lignocellulolytic enzyme genes in strains g-PxlnRA871V, g-Txyr1A824V, and g-Nxlr-1A828V versus those in ΔxlnR strain (set to one) cultured in cellulose medium at 4 h. e, f FPase and xylanase activities of strains ΔxlnR, g-PxlnRA871V, g-Txyr1A824V, and g-Nxlr-1A828V cultured in complex carbon medium, respectively. Data represent mean ± SD from triplicate cultivations. Statistical significance of the differences between parent strain ΔxlnR and each mutant was calculated for gene transcriptional levels. *p < 0.05, **p < 0.01, ***p < 0.001
Previous studies showed that constitutively active mutations of T. reesei Xyr1 (A824V), N. crassa XLR-1 (A828V), and P. oxalicum XlnR (A871V) could lead to markedly increased production of cellulase and/or xylanase (Craig et al. 2015; Derntl et al. 2013; Gao et al. 2017a). Therefore, it is valuable to determine whether Xyr1A824V and XLR-1A828V exhibit an enhanced activation function relative to wild-type Xyr1 and XLR-1 in P. oxalicum. Subsequently, strains g-Txyr1A824V, g-Nxlr-1A828V, and g-PxlnRA871V were constructed by overexpressing Txyr1A824V, Nxlr-1A828V, and PxlnRA871V in strain ΔxlnR with promoter gpdA(p). The major lignocellulolytic enzyme genes Pcbh1, Pcbh2, Peg1, and Pxyn10A in g-Txyr1A824V, g-Nxlr-1A828V, and g-PxlnRA871V showed higher transcriptional levels than those in g-Txyr1, g-Nxlr-1, and g-PxlnR, respectively, after induction by cellulose (Fig. 1d). Almost all the determined enzyme activities were elevated in g-Txyr1A824V, g-Nxlr-1A828V, and g-PxlnRA871V when compared with those in g-Txyr1, g-Nxlr-1, and g-PxlnR after being cultivated in complex carbon medium (Fig. 1e, f, Supplemental Fig. S3). Again, mutant g-Txyr1A824V exhibited the highest cellulase activity among the three mutants (Fig. 1e). No remarkable differences were observed for xylanase activities among the three strains carrying mutated XlnR homologs (Fig. 1f).
Construction of a cellulase high-producing mutant DB2 using a β-rec/six self-excising marker recycling system
The improvement of lignocellulolytic enzyme expression by introducing heterologous XlnR homologs provided a potential strategy for strain engineering to produce higher levels of these enzymes. To make the recurring use of the screening marker in genetic manipulations possible, we attempted to establish the β-rec/six self-excising marker recycling system in P. oxalicum strain M12. First, the genes β-rec and six-pyrG-six were introduced into strain M12 to replace gene Pbgl2 encoding an intracellular β-glucosidase (GenBank acc. no. EPS25645.1) (Chen et al. 2013), generating strain DB2-pyrG (Supplemental Fig. S1). The strain DB2-pyrG was cultivated in a cellulose medium to induce the excision pyrG gene, which generated the marker-free strain DB2.
Strain DB2 grew well on glucose plates containing uracil, but not on plates without uracil (Supplemental Fig. S4). The result demonstrated that the marker gene pyrG was efficiently excised by β serine recombinase. Through RT-qPCR analysis, we found that strains DB2-pyrG and DB2 both showed higher transcriptional levels of major cellulase genes Pcbh1, Pcbh2, and Peg1 than parent strain M12 (Fig. 2a). The cellulase enzyme activities corresponded to the changes of transcriptional levels in strains DB2-pyrG and DB2. Obvious increases of FPase (2.3-fold), pNPCase (1.8-fold), and CMCase (2.1-fold) activities were observed in DB2 when compared with those of parent strain M12 after 120 h of cultivation (Fig. 2b–d). At the same time, strains DB2-pyrG and DB2 showed no significant differences at transcriptional levels and enzyme activity levels under the same condition (Fig. 2a–d). The results demonstrated that the marker gene pyrG had no effect on the production of lignocellulolytic enzymes. These data implied that the β-rec/six self-excising marker recycling system was successfully implemented in P. oxalicum, providing a viable tool for multi-round transgene manipulations in the following steps.
Transcription level and cellulase activity analyses of strains M12, DB2-pyrG, and DB2. a Transcriptional level analysis of lignocellulolytic enzyme genes of DB2-pyrG and DB2 strains versus those in M12 strain (set to one) cultured in cellulose medium at 4 h. b–d FPase, pNPCase, and CMCase activities of M12, DB2-pyrG, and DB2 strains in complex carbon medium, respectively. Data represent mean ± SD from triplicate cultivations. Statistical significance of the differences between parent strain M12 and each mutant was calculated for gene transcriptional levels. *p < 0.05, **p < 0.01, ***p < 0.001
Comparison of the effects of heterologous regulatory modules on lignocellulolytic gene expression in P. oxalicum
In the above experiments, the heterologous regulators Xyr1 and XLR-1 as well as their constitutively active mutants were proved to be functional in P. oxalicum. To further investigate the regulatory functions of these TFs on their target genes, regulatory modules were designed and introduced into P. oxalicum. Cellulases CBHI and EGI are the main components of the cellulase system of T. reesei, and both of them play crucial roles in the process of cellulose degradation (Kolbe and Kubicek 1990; Stricker et al. 2006). Therefore, a regulatory module, which is composed of Xyr1A824V and its target cellulase genes Tcbh1 (cellobiohydrolase I gene from T. reesei, XP_006969224.1) and Teg1 (endoglucanase I gene from T. reesei, XP_006965674.1), was constructed (Fig. 3a). Within this regulatory module, cellulase genes Tcbh1 and Teg1 were controlled by their native promoters Tcbh1(p) and Teg1(p), respectively. Then, the regulatory module was introduced into strain DB2, generating strain RE-1-1. As controls, strains T-xyr1A824V and T-ceT carrying only Txyr1A824V or Tcbh1-Teg1 were also constructed (Fig. 3a).
Transcription level and cellulase activity analyses of DB2 and various mutants expressing T. reesei or N. crassa genes. a Schematic model of gene expression cassettes (Txyr1* represents Txyr1A824V). b Transcriptional level analysis of heterologous genes Tcbh1 and Teg1 of strains cultured in cellulose medium at 4 h or 18 h. c Transcriptional level analysis of endogenous lignocellulolytic enzyme genes of strains versus those in DB2 strain (set to one) cultured in cellulose medium at 18 h. d–f FPase, pNPCase, and CMCase activities of strains cultured in complex carbon medium, respectively. g Schematic model of gene expression cassettes (Nxlr-1* represents Nxlr-1A828V). h Transcriptional level analysis of heterologous Ncbh1 and Neg1 genes of strains cultured in cellulose medium at 18 h. i Transcriptional level analysis of endogenous lignocellulolytic enzyme genes of strains cultured in cellulose medium at 18 h. j–l FPase, pNPCase, and CMCase activity analysis of strains cultured in complex carbon medium, respectively. Data represent mean ± SD from triplicate cultivations. Statistical significance of the differences between parent strain DB2 and each mutant was calculated transcriptional levels of endogenous genes. *p < 0.05, **p < 0.01, ***p < 0.001
RT-qPCR results showed that the transcripts of heterologous cellulase genes Tcbh1 and Teg1 were detected in RE-1-1, with much higher levels than those in control strain T-ceT (Fig. 3b). Thus, Xyr1A824V improved the transcription of Tcbh1 and Teg1 from their native promoters. The activation of Tcbh1 and Teg1 expression by Xyr1A824V was also supported by the relatively higher cellulase activities of RE-1-1 than strains T-xyr1A824V and T-ceT (Fig. 3d–f, Supplemental Fig. S5). It should be noted that the higher cellulase production in RE-1-1 was not likely due to the increased expression of endogenous cellulases, because no obvious differences in transcriptional levels of genes Pcbh1, Pcbh2, Peg1, and Pxyn10A were observed between RE-1-1 and T-xyr1A824V (Fig. 3c). These results suggested that the heterologous regulatory module was functionally expressed in P. oxalicum, while alone expression of cellulase genes Tcbh1 and Teg1 under their native promoters was much less effective in the absence of their native regulator T. reesei Xyr1.
The above results of enzyme assays suggested that the expression of T. reesei cellulase genes Tcbh1 and Teg1 only slightly enhanced cellulase production in P. oxalicum, even in the presence of their regulator Xyr1A824V. Considering that Xyr1A824V could activate the expression of endogenous lignocellulolytic enzyme genes in P. oxalicum (Fig. 1d), the promoters of Pcbh1 and Peg1 were speculated to be effective targets of Xyr1A824V. Thus, a new regulatory module containing T. reesei genes Tcbh1 and Teg1 controlled by P. oxalicum promoters Pcbh1(p) and Peg1(p) was constructed. This regulatory module was expressed in strain DB2, and the resulted recombinant strain was named RE-1-2 (Fig. 3a). At the same time, the strain containing Tcbh1 and Teg1 genes under promoters Pcbh1(p) and Peg1(p) but lacking Txyr1A824V was constructed and named P-ceT (Fig. 3a).
RT-qPCR data showed that the transcriptional levels of Tcbh1 and Teg1 were significantly higher in RE-1-2 than those in RE-1-1 (Fig. 3b), both containing one copy of Txyr1A824V gene (Supplemental Fig. S6). Actually, the transcriptional levels of Tcbh1 and Teg1 were even higher in the control strain P-ceT than RE-1-2 (Fig. 3b), suggesting that the expression of these two genes from Pcbh1(p) and Peg1(p) were mainly activated by endogenous TFs in P. oxalicum. RE-1-2 showed the highest cellulase activities among the strains (Fig. 3d–f, Supplemental Fig. S5), which should be attributed to the enhanced cellulase expression triggered by Xyr1A824V and efficient expression of Tcbh1 and Teg1 in this strain. Using MS-MS analysis, TCBH1 and TEG1, as the protein products of Tcbh1 and Teg1, were identified in the supernatants of strains RE-1-2 and P-ceT (Supplemental Table S3). These results demonstrated that the new regulatory module containing T. reesei cellulase genes driven by promoters of P. oxalicum cellulase genes was efficient to increase cellulase production in P. oxalicum.
Similar to Xyr1A824V of T. reesei, XLR-1A828V of N. crassa could also effectively enhance lignocellulolytic gene expression in P. oxalicum. Therefore, a strain RE-2-1 expressing a regulatory module containing xlr-1A828V and cellulase genes from N. crassa (Ncbh1, EAA33262.1 and Neg1, EAA29875.2, controlled by P. oxalicum cellulase promoters) was constructed. Another two strains P-ceN and N-xlr-1A828V were also constructed as controls, which contained only the N. crassa cellulase genes or Nxlr-1A828V (Fig. 3g). RT-qPCR data showed the transcription of Ncbh1 and Neg1 in both RE-2-1 and P-ceN (Fig. 3h), and significantly improved transcription of endogenous lignocellulolytic enzyme genes in both RE-2-1 and N-xlr-1A828V (Fig. 3i). Accordingly, RE-2-1 showed the highest cellulase activities among the three strains, especially for pNPCase activity (Fig. 3j–l, Supplemental Fig. S5). The improvements in FPase and CMCase activities in RE-2-1 relative to N-xlr-1A828V were relatively slight (Fig. 3j, l). MS-MS data identified that the protein products of Ncbh1 and Neg1 existed in the supernatants of strains RE-2-1 and P-ceN (Supplemental Table S3). Of note, two copies of the Nxlr-1A828V gene were found in strain RE-2-1 (Supplemental Fig. S6). According to the result of RT-qPCR, the transcriptional levels of endogenous lignocellulolytic enzyme genes were similar between RE-2-1 and N-xlr-1A828V, indicating that the doubled copy number of Nxlr-1A828V is not the reason for the enhanced cellulase activity of RE-2-1.
Improvement of cellulase production via cumulative expression of multiple regulatory modules based on self-excising marker recycling system
Although the regulatory mechanism of lignocellulolytic enzymes is considerably conserved in lignocellulolytic filamentous fungi, the divergence between species may be employed to increase cellulase production. The above regulatory modules were proved to be able to significantly improve the lignocellulolytic enzyme production in P. oxalicum. Thus, the cumulative effects of multiple regulatory modules on lignocellulolytic enzyme expression were expected. To facilitate genetic manipulation, the β-rec/six self-excising marker recycling system was used in the subsequent transformation experiments.
First, P. oxalicum XlnRA871V regulatory module (containing PxlnRA871V, Pcbh1, and Peg1 which were controlled under promoters PDE_02864(p), Pcbh1(p), and Peg1(p), respectively) was constructed, and transformed into strain DB2. The PDE_02864 gene encodes a 40S ribosomal protein S8, and it showed high transcriptional levels in both glucose and cellulose media. In the research of Gao et al. (2017a), PDE_02864(p) was identified as a strong promoter for gene expression. The obtained strain was named RE-3-1. The subsequently obtained marker-free strain was named RE-3-2. Compared with strain DB2, RE-3-2 showed a larger hydrolytic halo on cellulose plates (Fig. 4a), and increased transcriptional levels of genes Pcbh1, Peg1, and Pxyn10A (by 5.4-, 3.7-, and 42.8-folds, respectively) at 18 h (Fig. 4b). Cellulase activities significantly increased compared with those of the parent strain DB2 (Fig. 4d–f). Besides, xylanase activity and extracellular protein concentration of strain RE-3-2 were also higher than those of DB2 (Supplemental Fig. S7).
Lignocellulolytic enzyme producing ability analysis of M12 and its engineered strains. a Phenotype observation of M12 and its engineered strains on 1% cellulose plates after 144 h of cultivation. b Transcriptional levels of endogenous lignocellulolytic enzyme genes in M12 and its engineered strains cultured in cellulose medium at 18 h. c Transcriptional levels of heterologous cellulase genes in DB2, RE-4-2, and RE-5-2 cultured in cellulose medium at 18 h. d–f FPase, pNPCase, and CMCase activities of DB2 and its engineered strains cultured in complex carbon medium, respectively. Data represent mean ± SD from triplicate cultivations. Statistical significance of the differences between strains was calculated for transcriptional levels of endogenous genes. *p < 0.05, **p < 0.01, ***p < 0.001
In order to determine the possibly cumulative effects of co-expression of multiple regulatory modules, the regulatory modules of Xyr1A824V and XLR-1A828V, which contained heterologous cellulase genes under the control of P. oxalicum cellulase promoters (Fig. 3a, g), were transformed into strain RE-3-2, successively. The resulting strains without pyrG were named RE-4-2 and RE-5-2. One copy of PxlnRA871V, Txyr1A824V, and Nxlr-1A828 was detected by qPCR for strains RE-3-2, RE-4-2, and RE-5-2, respectively (Supplemental Fig. S6). Compared with RE-3-2, strain RE-4-2 and RE-5-2 showed larger hydrolytic halos on cellulose plate, with that of the latter strain more obvious (Fig. 4a). The transcriptional levels of endogenous lignocellulolytic enzyme genes of P. oxalicum were significantly enhanced in RE-4-2 and RE-5-2 at 18 h (Fig. 4b). The transcripts of heterologous cellulase genes Tcbh1 and Teg1 were detected in strain RE-4-2 and RE-5-2 (Fig. 4c), and those of Ncbh1 and Neg1 were detected in strain RE-5-2 (Fig. 4c). Strain RE-4-2 showed increases of FPase, pNPCase, CMCase, and xylanase activities by 0.4-, 0.9-, 0.7-, and 0.3-folds at 120 h, respectively, as compared with those of RE-3-2 (Fig. 4d–f, Supplemental Fig. S7). For strain RE-5-2, the activity of pNPCase was also significantly enhanced (p value = 0.031) relative to that of RE-4-2 at 120 h (Fig. 4e, f). As expected, the heterologous cellulases were detected in the culture supernatants of RE-4-2 and RE-5-2 by MS-MS analysis (Supplemental Table S3).
Through the cumulative introduction of three regulatory modules, RE-5-2 showed a 2.8-fold increase in FPase activity when compared with strain DB2 (Fig. 4d). When compared with the original strain M12, strain RE-5-2 showed 5.1-, 10.4-, 5.6-, and 20.0-fold increases in FPase, pNPCase, CMCase, and xylanase activities, respectively (Fig. 4d–f, Supplemental Fig. S7). SDS-PAGE of culture supernatants confirmed the changes increases enzyme production capabilities (Supplemental Fig. S7). The above data demonstrated that the introduction of regulatory modules containing genes encoding XlnR homologs and cellulases from different species could efficiently increase lignocellulolytic enzyme production in P. oxalicum.
The production level of lignocellulolytic enzymes by the engineered strains was also compared in the medium with 1% xylan as sole carbon source. Also, the strains expressing regulatory modules, particularly RE-4-2 and RE-5-2, produced higher cellulase and xylanase activities than the parent strain DB2 (Supplemental Fig. S8). Of note, the advantage of RE-5-2 in cellulase production over RE-4-2 was more remarkable in xylan medium than that in complex carbon medium. Therefore, the N. crassa XLR-1A828V regulatory module was likely to function better in response to xylan-derived signals.
Discussion
The homologs of XlnR are widely present in ascomycete fungi, including species belonging to Dothideomycetes, Eurotiomycetes, and Sordariomycetes classes. While it is clear that these homologous proteins have similar but not the same regulatory functions in different species (Klaubauf et al. 2014), their cross-species regulatory functions have been rarely investigated. In this study, we explored the functions of heterologous Xyr1 and XLR-1 (as well as their constitutively active mutants) in P. oxalicum. Like the native regulator gene PxlnR, expression of both Txyr1 and Nxlr-1 complemented the defective xylanase production in the PxlnR deletion strain (Fig. 1c), suggesting that the role in activating xylanase gene expression is conserved among XlnR homologs. The fold changes of the increase in Pxyn10A expression were higher than those of cellulase genes, suggesting that all the three regulators act more effectively on xylanolytic enzyme genes. Interestingly, the XlnR homologs had different preferences on enhancing the production of cellulase and xylanase (Fig. 1). Specifically, the expression of T. reesei Xyr1 and Xyr1A824V resulted in higher cellulase (but not xylanase) production than the native XlnR and XlnRA871V, respectively (Fig. 1b, e). This could be linked to the critical role of Xyr1 in activating the expression of not only xylanase but cellulase genes in its native background (Stricker et al. 2006). On the other hand, expression of N. crassa XLR-1 led to enhanced cellulase production in P. oxalicum, although it does not bind to cellulase genes in its native background (Craig et al. 2015). Taken together, the results highlighted the partially conserved but also rapidly evolved structure of a XlnR/Xyr1/XLR-1 regulatory network. It was possible that the common ancestor of XlnR homologs regulated the expression of both cellulase and xylanase genes, and some fungal species (e.g., N. crassa) adopted more specialized functions for this regulator during evolution.
The different enhancing effects of XlnR homologs on cellulase expression in P. oxalicum might be due to the different binding abilities of these regulators on cellulase gene promoters. The previous enriched binding motif of T. reesei Xyr1 (5′-GGC(A/T)3-3′) was less stringent than that reported for N. crassa XLR-1 (5′-GGNTAAA-3′) (Craig et al. 2015; Furukawa et al. 2009; Rauscher et al. 2006), although detailed information on their recognition specificities is not clear. For Xyr1, its binding site was found in the promoters of all cellobiohydrolase and endoglucanase genes of P. oxalicum, with an average of six sites in each promoter (Supplemental Table S2). For XLR-1, the binding site was found in 13 of 16 xylanolytic gene promoters, but only in 7 of 14 cellulase gene promoters. Only one or two sites of XLR-1 were present on each cellulase gene promoter. Therefore, T. reesei Xyr1 might bind more efficiently to the cellulase gene promoters of P. oxalicum than to those of N. crassa XLR-1, therefore resulting in higher cellulase activities (Fig. 1b, e). This hypothesis needs to be certified by determining the binding affinities of XlnR homologs on different DNA sites in the future. In addition, indirect activation of cellulase genes by XlnR homologs (e.g., through upregulation of the cellulase gene activator ClrB) could not be excluded.
It was not surprising that the T. reesei cellulase genes with their native promoters were expressed at low levels in P. oxalicum when lacking the activation by T. reesei Xyr1 (Fig. 3b, Supplemental Fig. S5). Co-expression of Xyr1A824V, or replacing the T. reesei promoters by P. oxalicum promoters, could facilitate the expression of T. reesei cellulase genes, suggesting that the two T. reesei promoters need their native activator for efficient transcription. On the other hand, the expression levels of T. reesei cellulase genes by P. oxalicum promoters were not further enhanced by the expression of Xyr1A824V (Fig. 3b), which was different from the cases of endogenous genes (Fig. 3c). This difference might be due to the use of 1500-bp sequences upstream of the coding regions as promoters in our expression module, which might not include the acting site of Xyr1A824V.
The introduction of heterologous XlnR homologs, especially T. reesei Xyr1, is an effective strategy in engineering lignocellulolytic enzyme producing strains from a biotechnological perspective. Due to the relatively specialized functions of TFs, separate engineering of XlnR and cellulase gene activator ClrB was needed to enhance the production of xylanase and cellulase in P. oxalicum (Gao et al. 2017b). Here, single expression of T. reesei Xyr1A824V effectively increased the production of both types of enzymes (Fig. 1e, f). When introduced into the mutant DB2 where the intracellular β-glucosidase gene bgl2 was deleted, the expression of Xyr1A824V could further improve cellulase production by 0.9-fold (Fig. 3d). This suggested that accumulation of inducer and enhancement of transcriptional activator had an additive effect in improving cellulase production. With the aid of the β-rec/six marker recycling system, combination of the expression of Xyr1A824V and manipulation of other targets (e.g., engineering of CreA, Yao et al. 2015) is expected to further improve the production level of lignocellulolytic enzymes.
In summary, we identified that heterologous regulators Xyr1 and XLR-1 could activate lignocellulolytic enzyme gene expression in P. oxalicum. At the same time, regulators Xyr1A824V and XLR-1A828V bearing point mutations were able to confer P. oxalicum higher lignocellulolytic enzyme-producing abilities than wild-type regulators. Furthermore, introduction of regulatory modules containing Xyr1A824V/XLR-1A828V and their target cellulase genes resulted in greater increases in cellulase production than alone expression of transcriptional regulators. These findings support that adoption of regulatory elements from diverse fungal species could be an efficient strategy to genetically engineer lignocellulolytic enzyme-producing strains.
References
Alazi E, Ram AFJ (2018) Modulating transcriptional regulation of plant biomass degrading enzyme networks for rational design of industrial fungal strains. Front Bioeng Biotechnol 6(133). https://doi.org/10.3389/fbioe.2018.00133
Amore A, Giacobbe S, Faraco V (2013) Regulation of cellulase and hemicellulase gene expression in fungi. Curr Genomics 14(4):230–249. https://doi.org/10.2174/1389202911314040002
Canosa I, Rojo F, Alonso JC (1996) Site-specific recombination by the β protein from the streptococcal plasmid pSM19035: minimal recombination sequences and crossing over site. Nucleic Acids Res 24:2712–2717. https://doi.org/10.1093/nar/24.14.2712
Chen M, Qin YQ, Cao Q, Liu GD, Li J, Li ZH, Zhao J, Qu YB (2013) Promotion of extracellular lignocellulolytic enzymes production by restraining the intracellular β-glucosidase in Penicillium decumbens. Bioresour Technol 137:33–40. https://doi.org/10.1016/j.biortech.2013.03.099
Coradetti ST, Craig JP, Xiong Y, Shock T, Tian CG, Glass NL (2012) Conserved and essential transcription factors for cellulase gene expression in ascomycete fungi. Proc Natl Acad Sci U S A 109(19):7397–7402. https://doi.org/10.1073/pnas.1200785109
Coradetti ST, Xiong Y, Glass NL (2013) Analysis of a conserved cellulase transcriptional regulator reveals inducer-independent production of cellulolytic enzymes in Neurospora crassa. Microbiologyopen 2(4):595–609. https://doi.org/10.1002/mbo3.94
Craig JP, Coradetti ST, Starr TL, Glass NL (2015) Direct target network of the Neurospora crassa plant cell wall deconstruction regulators CLR-1, CLR-2, and XLR-1. MBio 6(5):e01452–e01415. https://doi.org/10.1128/mBio.01452-15
Derntl C, Gudynaite-Savitch L, Calixte S, White T, Mach RL, Mach-Aigner AR (2013) Mutation of the xylanase regulator 1 causes a glucose blind hydrolase expressing phenotype in industrially used Trichoderma strains. Biotechnol Biofuels 6(1):62. https://doi.org/10.1186/1754-6834-6-62
Ellila S, Fonseca L, Uchima C, Cota J, Goldman GH, Saloheimo M, Sacon V, Siika-aho M (2017) Development of a low-cost cellulase production process using Trichoderma reesei for Brazilian biorefineries. Biotechnol Biofuels 10(30):30. https://doi.org/10.1186/s13068-017-0717-0
Fang X, Shen Y, Zhao J, Bao XM, Qu YB (2010) Status and prospect of lignocellulosic bioethanol production in China. Bioresour Technol 101(13):4814–4819. https://doi.org/10.1016/j.biortech.2009.11.050
Furukawa T, Shida Y, Kitagami N, Mori K, KatoM KT, Okada H, Ogasawara W, Morikawa Y (2009) Identification of specific binding sites for XYR1, a transcriptional activator of cellulolytic and xylanolytic genes in Trichoderma reesei. Fungal Genet Biol 46(8):564–574. https://doi.org/10.1016/j.fgb.2009.04.001
Gao LW, Li ZH, Xia CQ, Qu YB, Liu M, Yang P, Yu LL, Song X (2017a) Combining manipulation of transcription factors and overexpression of the target genes to enhance lignocellulolytic enzyme production in Penicillium oxalicum. Biotechnol Biofuels 10(100):100. https://doi.org/10.1186/s13068-017-0783-3
Gao LW, Xia CQ, Xu JD, Li ZH, Yu LL, Liu GD, Song X, Qu YB (2017b) Constitutive expression of chimeric transcription factors enables cellulase synthesis under non-inducing conditions in Penicillium oxalicum. Biotechnol J 12(11). https://doi.org/10.1002/biot.201700119
Glass NL, Schmoll M, Cate JHD, Coradetti S (2013) Plant cell wall deconstruction by ascomycete fungi. Annu Rev Microbiol 67:477–498. https://doi.org/10.1146/annurev-micro-092611-150044
Gruber F, Visser J, Kubicek CP, de Graaff LH (1990) The development of a heterologous transformation system for the cellulolytic fungus Trichoderma reesei based on a pyrG-negative mutant strain. Curr Genet 18(1):71–76
Han XL, Song WX, Liu GD, Li ZH, Yang P, Qu YB (2017) Improving cellulase productivity of Penicillium oxalicum RE-10 by repeated fed-batch fermentation strategy. Bioresour Technol 227:155–163. https://doi.org/10.1016/j.biortech.2016.11.079
Hartmann T, Dumig M, Jaber BM, Szewczyk E, Olbermann P, Morschhauser J, Krappmann S (2010) Validation of a self-excising marker in the human pathogen Aspergillus fumigatus by employing the β-rec/six site-specific recombination system. Appl Environ Microbiol 76(18):6313–6317. https://doi.org/10.1128/AEM.00882-10
Kang NK, Kim EK, Kim YU, Lee B, Jeong WJ, Jeong BR, Chang YK (2017) Increased lipid production by heterologous expression of AtWRI1 transcription factor in Nannochloropsis salina. Biotechnol Biofuels 10(231):231. https://doi.org/10.1186/s13068-017-0919-5
Klaubauf S, Narang HM, Post H, Zhou MM, Brunner K, Mach-Aigner AR, Mach RL, Heck AJR, Altelaar AFM, de Vries RP (2014) Similar is not the same: differences in the function of the (hemi-)cellulolytic regulator XlnR (Xlr1/Xyr1) in filamentous fungi. Fungal Genet Biol 72:73–81. https://doi.org/10.1016/j.fgb.2014.07.007
Kolbe J, Kubicek CP (1990) Quantification and identification of the main components of the Trichoderma cellulase complex with monoclonal-antibodies using an enzyme-linked-immunosorbent-assay (ELISA). Appl Microbiol Biotechnol 34(1):26–30. https://doi.org/10.1007/Bf00170918
Kricka W, Fitzpatrick J, Bond U (2015) Challenges for the production of bioethanol from biomass using recombinant yeasts. Adv Appl Microbiol 92:89–125. https://doi.org/10.1016/bs.aambs.2015.02.003
Kubicek CP, Mikus M, Schuster A, Schmoll M, Seiboth B (2009) Metabolic engineering strategies for the improvement of cellulase production by Hypocrea jecorina. Biotechnol Biofuels 2:19. https://doi.org/10.1186/1754-6834-2-19
Li ZH, Yao GS, Wu RM, Gao LW, Kan QB, Liu M, Yang P, Liu GD, Qin YQ, Song X, Zhong YH, Fang X, Qu YB (2015) Synergistic and dose-controlled regulation of cellulase gene expression in Penicillium oxalicum. PLoS Genet 11(9):e1005509. https://doi.org/10.1371/journal.pgen.1005509
Liu GD, Zhang L, Wei XM, Zou G, Qin YQ, Ma L, Li J, Zheng HJ, Wang SY, Wang CS, Xun LY, Zhao GP, Zhou ZH, Qu YB (2013) Genomic and secretomic analyses reveal unique features of the lignocellulolytic enzyme system of Penicillium decumbens. PLoS One 8(2):e55185. https://doi.org/10.1371/journal.pone.0055185
Liu GD, Qin YQ, Li ZH, Qu YB (2014) Improving lignocellulolytic enzyme production with Penicillium: from strain screening to systems biology. Biofuels 4(5):523–534. https://doi.org/10.4155/bfs.13.38
Miller GL (1959) Use of dinitrosalicylic acid reagent for determination of reducing sugar. Anal Chem 31(3):426–428
Nakari-Setala T, Paloheimo M, Kallio J, Vehmaanpera J, Pentillä M, Saloheimo M (2009) Genetic modification of carbon catabolite repression in Trichoderma reesei for improved protein production. Appl Environ Microbiol 75(14):4853–4860. https://doi.org/10.1128/AEM.00282-09
Pentillä M, Nevalainen H, Ratto M, Salminen E, Knowles J (1987) A versatile transformation system for the cellulolytic filamentous fungus Trichoderma reesei. Gene 61(2):155–164. https://doi.org/10.1016/0378-1119(87)90110-7
Peterson R, Nevalainen H (2012) Trichoderma reesei RUT-C30-thirty years of strain improvement. Microbiology 158(Pt 1):58–68. https://doi.org/10.1099/mic.0.054031-0
Qian YC, Zhong LX, Gao J, Sun NN, Wang YF, Sun GY, Qu YB, Zhong YH (2017) Production of highly efficient cellulase mixtures by genetically exploiting the potentials of Trichoderma reesei endogenous cellulases for hydrolysis of corncob residues. Microb Cell Factories 16(1):207. https://doi.org/10.1186/s12934-017-0825-3
Qin YQ, Zheng K, Liu GD, Chen M, Qu YB (2013) Improved cellulolytic efficacy in Penicilium decumbens via heterologous expression of Hypocrea jecorina endoglucanase II. Arch Biol Sci 65(1):305–314. https://doi.org/10.2298/abs1301305q
Raulo R, Kokolski M, Archer DB (2016) The roles of the zinc finger transcription factors XlnR, ClrA and ClrB in the breakdown of lignocellulose by Aspergillus niger. AMB Express 6:5. https://doi.org/10.1186/s13568-016-0177-0
Rauscher R, Würleitner E, Wacenovsky C, Aro N, Stricker AR, Zeilinger S, Kubicek CP, Penttila M, Mach RL (2006) Transcriptional regulation of xyn1, encoding xylanase I, in Hypocrea jecorina. Eukaryot Cell 5(3):447–456. https://doi.org/10.1128/EC.5.3.447-456.2006
Saini R, Saini JK, Adsul M, Patel AK, Mathur A, Tuli D, Singhania RR (2015) Enhanced cellulase production by Penicillium oxalicum for bio-ethanol application. Bioresour Technol 188:240–246. https://doi.org/10.1016/j.biortech.2015.01.048
Stricker AR, Grosstessner-Hain K, Würleitner E, Mach RL (2006) Xyr1 (xylanase regulator 1) regulates both the hydrolytic enzyme system and D-xylose metabolism in Hypocrea jecorina. Eukaryot Cell 5(12):2128–2137. https://doi.org/10.1128/EC.00211-06
Sun JP, Tian CG, Diamond S, Glass NL (2012) Deciphering transcriptional regulatory mechanisms associated with hemicellulose degradation in Neurospora crassa. Eukaryot Cell 11(4):482–493. https://doi.org/10.1128/EC.05327-11
Szewczyk E, Kasuga T, Fan ZL (2014) A new variant of self-excising β-recombinase/six cassette for repetitive gene deletion and homokaryon purification in Neurospora crassa. J Microbiol Methods 100:17–23. https://doi.org/10.1016/j.mimet.2014.02.007
Tian CG, Beeson WT, Iavarone AT, Sun JP, Marletta MA, Cate JHD, Glass NL (2009) Systems analysis of plant cell wall degradation by the model filamentous fungus Neurospora crassa. Proc Natl Acad Sci U S A 106(52):22157–22162. https://doi.org/10.1073/pnas.0906810106
Vogel HJ (1956) A convenient growth medium for Neurospora crassa. Microbiol Genet Bull 13:42–43
Wang HW, Zhang B, Hao YJ, Huang J, Tian AG, Liao Y, Zhang JS, Chen SY (2007) The soybean Dof-type transcription factor genes, GmDof4 and GmDof11, enhance lipid content in the seeds of transgenic Arabidopsis plants. Plant J 52(4):716–729. https://doi.org/10.1111/j.1365-313X.2007.03268.x
Wang HL, Li Z, Jia RN, Yin J, Li AY, Xia LQ, Yin YL, Muller R, Fu J, Stewart AF, Zhang YM (2018) ExoCET: exonuclease in vitro assembly combined with RecET recombination for highly efficient direct DNA cloning from complex genomes. Nucleic Acids Res 46(5):e28. https://doi.org/10.1093/nar/gkx1249
Yao GS, Li ZH, Gao LW, Wu RM, Kan QB, Liu GD, Qu YB (2015) Redesigning the regulatory pathway to enhance cellulase production in Penicillium oxalicum. Biotechnol Biofuels 8:71. https://doi.org/10.1186/s13068-015-0253-8
Yao GS, Wu RM, Kan QB, Gao LW, Liu M, Yang P, Du J, Li ZH, Qu YB (2016) Production of a high-efficiency cellulase complex via β-glucosidase engineering in Penicillium oxalicum. Biotechnol Biofuels 9:78. https://doi.org/10.1186/s13068-016-0491-4
Yu JH, Hamari Z, Han KH, Seo JA, Reyes-Dominguez Y, Scazzocchio C (2004) Double-joint PCR: a PCR-based molecular tool for gene manipulations in filamentous fungi. Fungal Genet Biol 41(11):973–981. https://doi.org/10.1016/j.fgb.2004.08.001
Funding
This study was funded by the National Natural Science Foundation of China (Grant nos. 31030001, 31270089, 31370086, and 31670079), and supported by State Key Laboratory of Microbial Technology Open Projects Fund (No. M2016-07).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This article does not contain any studies with human participants or animals performed by any of the authors.
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(PDF 956 kb)
Rights and permissions
About this article
Cite this article
Xia, C., Li, Z., Xu, Y. et al. Introduction of heterologous transcription factors and their target genes into Penicillium oxalicum leads to increased lignocellulolytic enzyme production. Appl Microbiol Biotechnol 103, 2675–2687 (2019). https://doi.org/10.1007/s00253-018-09612-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-018-09612-y