Abstract
Mitigation of the methane (CH4) emission from ruminants is needed to decrease the environmental impact of ruminant animal production. Different plant materials and chemicals have been tested, but few are both effective and practical. Medicinal herbs contain biological compounds and antimicrobials that may be effective in lowering the CH4 production. However, few studies have systematically evaluated medicinal herbs for their effect on CH4 production or on the rumen microbiota. In this study, extracts from 100 medicinal herbs were assessed for their ability to decrease CH4 production by rumen microbiota in vitro. The extracts of 12 herbs effectively lowered the CH4 production, with the extract of Perilla frutescens seeds being the most effective. The major components of P. frutescens seed extract were identified, and the effects of the extract on the fermentation characteristics and populations of rumen methanogens, fungi, protozoa, and select bacteria were also assessed. The decreased CH4 production induced by the P. frutescens seed extract was accompanied by an increased abundance of Ruminobacter, Selenomonas, Succinivibrio, Shuttleworthis, Pseudobutyrivbrio, Anaerovibrio, and Roseomonas and a decreased abundance of Methanobrevibacter millerae. The abundance of Pedobacter, Anaeroplasma, Paludibacter, Ruminococcus, and unclassified Lachnospiraceae was positively correlated with the CH4 production, with no effects on volatile fatty acids. This study suggests that medicinal herbs may be used to mitigate the CH4 emission from ruminants.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Methane (CH4) emission from ruminant animals accounts for 10–12 % of the global anthropogenic greenhouse gas emissions in carbon dioxide equivalents (IPCC 2007) and wastes 2–15 % of the feed energy depending on the type of diet (Johnson and Johnson 1995). As the demand for meat and milk continues to grow worldwide (especially in developing countries), CH4 emission from ruminants will likely continue to increase in the years to come unless effective and practical mitigation strategies are implemented. Over the past decade, intensive research has been conducted to identify and develop effective and practical means to decrease the CH4 emission from ruminants (Hristov et al. 2013). Phytochemicals, primarily plant secondary metabolites, are particularly attractive because they are naturally produced by plants and can be readily formulated into feed rations. Indeed, several types of phytochemicals have been extensively evaluated both in vitro and in vivo, including saponins (Patra and Yu 2013), tannins (Bhatta et al. 2015), essential oils (Patra and Yu 2014a), and lipids (Benchaar et al. 2015). In most studies, however, effective CH4 suppression was accompanied by a significant decrease in feed digestion and fermentation (Grainger and Beauchemin 2011). Therefore, none of the evaluated CH4-mitigation materials has yet become widely used on diary or beef cattle farms. New research efforts are needed to evaluate various sources to identity those that can effectively decrease CH4 emission from cattle with no or little adverse effects on feed intake, digestion, or fermentation.
Plants have provided the oldest medicines that have been used to treat diseases in both humans and animals. Some medicinal herbs are still being used in many countries, although much less frequently than before chemical and antibiotic medicines became widely available (Liu et al. 2015; Ahmad et al. 2013). The therapeutic efficacy of medicinal herbs is attributed to some of their components that have biological and/or antimicrobial activities. Medicinal herbs may represent a source of natural substances and compounds that can be used to modulate rumen microbial populations to enhance feed digestion and decrease CH4 emission. A number of herbs and combinations of herbs, such as Allium sativum var. pekinense, A. cepa, Artemisia princeps var. orientalis, Zingiber officinale, Citrus unshiu, and Lonicera japonica (Kim et al. 2012a, b); Sambucus nigra, Alchemilla xanthochlora, and Castanea sativa (Jayanegara et al. 2011); Rheum officinale, Frangula alnus, and A. sativum (García-González et al. 2008); Sapindus mukorossi, Terminalia chebula, Populus tremuloides, Syzygium aromaticum, and Psidium guayaba (Kamra et al. 2005); and Sesbania sesban and Acacia angustissima (Zeleke et al. 2005), have been shown to inhibit in vitro methanogenesis. However, most of them are not medicinal herbs, and few studies have comprehensively examined how herbs can affect the composition and structure of rumen microbiota and the population dynamics of rumen methanogens, fungi, protozoa, and select bacteria. The objective of the present study was to evaluate some of the most commonly used medicinal herbs (100 herbs in total) for their ability to modify rumen fermentation and to decrease CH4 production by rumen microbiota. The major components of the most promising medical herbal extracts were identified, and the concentration-dependent responses of these herbal extracts were further examined with respect to rumen fermentation, microbiota populations, and microbiota composition. The findings of this study may help identify potential new plant-derived approaches to mitigate CH4 emission from ruminants and to stimulate future research in achieving sustainable CH4 mitigation in the livestock industry.
Materials and methods
Plant herbs and extract preparation
One hundred common medicinal herbs were purchased from local pharmacy stores and used in this study (Table S1). All of the herbs were in the form of dried raw materials from various parts of herbal plants. They were finely ground to powders (270 to 48 μm) with a portable medicine grinder (HK-02A, Xulang Machinery, China) and then preserved at 4 °C.
Ethanol was used as the solvent to extract the compounds from the herb powders. Each of the powdered herbs (0.25 g) was extracted twice with 70 % ethanol at room temperature using a rotary shaker at 35 rpm (WH-962, Hualida, China) for 1 h. The mixture was then centrifuged at 1660×g for 5 min at room temperature. The supernatants from the two extractions were combined into one pre-weighed glass ampoule, and the solvent was allowed to evaporate overnight under continuous nitrogen gas flux to prevent oxidization. The resultant extract was frozen at −80 °C. The extracts were then freeze-dried (CHRiST, Osterode, Germany), weighed, and stored at −20 °C until use (Kemp and McSweeney 2010).
Preliminary evaluation of herbal extracts
Each of the 100 herbal extracts was evaluated for its effect on rumen fermentation using in vitro cultures with fresh rumen fluid serving as the inoculum (Wang et al. 2015). The Animal Care Committee of Zhejiang University (Hangzhou, China) approved the procedures used to collect rumen fluid from the donor animals. Briefly, rumen fluid was collected from three fistulated lactating cows, which were fed a total mixed ration (roughage/concentrate = 55:45) twice daily, before the morning feeding, and the rumen fluid samples were strained through four layers of cheesecloth under continuous flushing with CO2. The three-rumen fluid samples were combined in equal ratios and mixed as the inoculum. A mixture of an anaerobic medium (Theodorou et al. 1994) and the inoculum (in a 9:1 ratio) was prepared inside of an anaerobic chamber with an atmosphere of 95 % CO2 and 5 % H2. This medium-inoculum mixture was dispensed into 27-mL culture tubes (10 mL/tube), with each containing 100 mg of fermentation substrate consisting of Chinese wild rye grass and corn meal in a 70:30 ratio. To each tube, 4 mg of herbal extract, which was dissolved in dimethyl sulfoxide (DMSO) and adjusted to a concentration of 40 mg/mL, was added, resulting in a final extract concentration of 0.4 mg/mL culture. Each herbal extract was tested in triplicate. Control cultures and blank cultures were also included, and each was tested in triplicate. The control cultures did not receive any herbal extract but did receive the same volume of DMSO (100 μL), while the blank cultures did not receive any herbal extracts or DMSO. All of the tubes were sealed with a butyl rubber stopper that was then secured with an aluminum crimp and incubated at 39 °C with horizontal shaking at 60 rpm.
After 24 h of incubation, the gas pressure in each culture tube was recorded using a pressure sensor (Ruyi, Shanghai, China), and the CH4 concentration in the headspace of each culture tube was measured using a gas chromatograph (GC-2010, Shimadzu, Kyoto, Japan) equipped with a flame ionization detector. The total gas production by each culture was calculated from the corresponding volume of its headspace and gas pressure, while CH4 production was calculated from the CH4 concentration and total gas production. Because this was a preliminary evaluation of a large number of herbal extracts to identify potentially effective ones for further evaluation, no further sampling or analysis was done.
Further evaluation of select effective herbal extracts
Based on the results of the preliminary experiment, 31 herbal extracts (Table 1) that led to the largest decreases in CH4 production were selected for further evaluation using in vitro rumen cultures. The in vitro cultures were prepared and incubated in essentially the same way as described in the preliminary experiment, except for an increased culture volume (50 mL of each extract in 120-mL serum bottles each containing 500 mg of the same fermentation substrate) and the inoculum collected from the rumens of three 2-year-old sheep. The sheep rumen fluid was used as the inoculum to determine if the selected herbal extracts work in sheep as well. Each treatment and the control, which received no herbal extract, had four replicates, and the experiment was repeated once. At the end of the 24-h incubation, total gas production was determined as described above, and the CH4 concentration and volatile fatty acid (VFA) concentrations were determined using gas chromatography (Wang et al. 2015).
Chemical analysis of the fatty acids in Perilla frutescens seed extract
The extract of Perilla frutescens (PF) seeds exhibited the strongest inhibition of CH4 production (Table 1). To identify the major components, this extract was subjected to a chemical analysis for its major fatty acids. Into one glass extraction tube, 0.4 mg of the freeze-dried PF seed extract and 10 mL of a mixture of chloroform/methanol (2:1, v/v) were combined. To prevent oxidation, the headspace of the extraction tube was flushed and then filled with nitrogen gas (99.99 % purity). The tube was sealed with a butyl rubber stopper and shaken at 120 rpm for 1 h at room temperature. The extraction mixture was filtered through one layer of fat-free filter paper (Whatman No. 1), and the retained solid was washed with 5 mL of a chloroform/methanol mixture twice. The filtrate was mixed with 3 mL NaCl (0.73 %, w/v) to precipitate the hydrophilic compounds. After dehydration with anhydrous sodium sulfate and solvent removal by nitrogen sparging, the free fatty acids were converted to their methyl esters (FAMEs) using boron trifluoride-methanol as an esterification reagent (Lopez-Lopez et al. 2001). The FAMEs were analyzed by gas chromatography with a flame-ionization detector. Helium was used as the carrier gas, with a split ratio of 100:1 and a linear gas flow rate at 20 cm/s. After injection, the column temperature was initially maintained at 140 °C for 5 min and then was allowed to ramp up to 240 °C at 4 °C/min. Fatty acids were identified by comparing their retention time to an external standard consisting of 37 known FAMEs (Saint Louis, MO, USA). The quantity of each fatty acid was calculated based on the relative peak area of linolenic acid methyl ester and its quantity in the external standard.
In vitro evaluation of different concentrations of the extract of P. frutescens seeds
The extract from PF seeds was evaluated at five concentrations (0, 0.1, 0.2, 0.3, and 0.4 mg/mL culture) to determine whether it had concentration-dependent effects on in vitro fermentation, CH4 production, and microbiota. The in vitro fermentation was conducted as described above for evaluating the 31 herbal extracts (four replicates per concentrations and one repetition) except for the inoculum being collected from the rumen of three 1-year-old sheep. At the end of the 24-h incubation, the headspace pressure was determined, and the headspace was sampled from each bottle for CH4 analysis. The fermentation bottles were then placed in ice water to stop fermentation. Three subsamples (1 mL each) were collected from each fermentation bottle after mixing. One of the subsamples was immediately stored at −80 °C for determination of the activity of cellulase, carboxymethyl cellulase (CMCase), and xylanase, while the other two subsamples were centrifuged at 13, 000×g for 15 min at 4 °C. The pellets were weighed and immediately stored at −80 °C for analysis of select microbial groups. The supernatants were stored at −80 °C for analysis of the VFAs and ammonia N. The remaining culture in each bottle was filtered through nylon bags (23-μm mesh), and the retained solid was dried in an oven at 65 °C for 48 h to determine the amount of dry matter (DM) (Wang et al. 2015).
Total DNA extraction and real-time quantitative PCR
Total DNA was extracted from the pellet from one of the subsamples of each in vitro culture using the bead-beating method described by Gagen et al. (2010). One standard for each quantative PCR (qPCR) assay was prepared for each group of microbes using PCR (the primers used are listed in Table S2), and cloning of the PCR amplicons was performed using a pGEM®T Easy kit (Promega, Shanghai, China). Purified plasmid preparations containing each amplicon were quantified and serially diluted as the qPCR standard. The abundance of each species or group of the select microbes was determined using qPCR and respective specific primers against each respective standard (Wang et al. 2015). The abundance of each species or group of microbes was expressed as the log10 copies of the 16S rRNA gene (or 18S rRNA gene in the case of protozoa, and ITS1 in the case of fungi) per gram of wet culture pellet.
Enzyme activity assays
The activities of cellulase, CMCase, and xylanase in each culture were determined essentially as described in a previous study (Wang et al. 2015). Briefly, each in vitro fermentation sample (1 mL) was sonicated using a JY92-IIN Ultrasonic Cell Mixer (Ningbo Scientz, Ningbo, China) and was centrifuged at 12,000×g at 4 °C for 10 min. The activities of cellulase, CMCase, and xylanase present in the supernatants were determined using the DNS method (Bailey et al. 1992) with Sigmacell@ cellulose Type 101 (Sigma, Saint Louis, MO, USA), carboxymethyl cellulose sodium (Sigma, Saint Louis, MO, USA), and beechwood xylan (Sigma, Saint Louis, MO, USA) as the substrates, each at 0.01 g/mL. One unit of enzyme activity was defined as the activity that produced 1 μmol of reducing sugars per min.
Analysis of the bacterial and archaeal communities
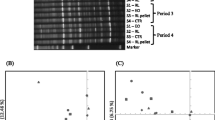
One amplicon library each was prepared for bacteria and for archaea from each of the DNA samples (only the samples from three concentrations 0, 0.2, and 0.4 mg/mL) as described by Caporaso et al. (2011). Briefly, the V4 hypervariable region of the bacterial 16S rRNA gene was amplified using a domain-specific primer set, 515F/806R, while the V1–V3 regions of the archaeal 16S rRNA gene was amplified using the archaea-specific domain primer set, M86F/M448R. Each forward primer had a 6-bp error-correcting barcode unique to each DNA sample at its 5′ end. The amplicon libraries for all of the samples were pooled in equal molar ratios and sequenced on an Illumina MiSeq system using the pair-ended 2 × 250 bp protocol at Novogene Bioinformatics Technology Co., Ltd. (Beijing, China). The paired reads were joined based on overlap regions to form single sequences using FLASH (Magoč and Salzberg 2011). Sequences were demultiplexed and assigned to each sample according to the individual unique barcode and were analyzed using QIIME (Caporaso et al. 2010). Briefly, sequences with a quality score <20 and length <185 bp (for bacteria) and <248 bp (for archaea) were removed. Possible chimeric sequences were identified using the UCHIME algorithm and the Gold database (http://dirve5.com/uchime/uchime_download.html). Then, the UPARSE pipeline was used to pick de novo operational taxonomic units (OTUs) at 97 % sequence similarity. The same number of sequences was used for all of the samples (36,410 sequences/sample for bacteria and 34,130 sequences/sample for archaea) to avoid effects due to different sample sizes. From each OTU, one representative sequence was selected and classified using the Classifier of Ribosomal Database Project (RDP) with a default confidence threshold of 80 %. The community richness was estimated using Chao 1, and the Shannon-Wiener diversity index was calculated using Qiime. The weighted Unifrac distance method was used to conduct a principal coordinates analysis (PCoA). The raw sequence data were deposited in the European Nucleotide Archive under accession no. PRJEB12630 for bacteria and PRJEB12632 for archaea.
Statistical analysis
The data in the preliminary experiment (three replicates, one repetition) and from the further evaluation of the 31 selected herbal extracts (four replicates, one repetition) were analyzed by one-way analysis of variance using SAS 9.1 (SAS Institute, Cary, NC) with each culture bottle used as the experimental unit and the herbal extracts as the main effects. The data from the evaluation of the concentration-dependent effects of PF seed extract were analyzed by orthogonal polynomial contrast to examine the concentration-dependent effects (linear or quadratic). Tukey’s multiple range test was conducted for multiple comparisons of means among treatments. Significance was declared at P < 0.05. Correlations between the fermentation parameters and relative abundance (expressed as the percentage of total bacterial sequences) of bacterial or archaeal genera or groups were calculated as Pearson product-moment coefficients.
Results
Preliminary evaluation to identify effective herbal extracts
The solvent used in this study, dimethyl sulfoxide (DMSO), can accept electrons, so its potential effect on in vitro fermentation was determined by comparing the control cultures that contained DMSO with the blank cultures that contained no DMSO. The added DMSO had no effects on GP but decreased the CH4 from 0.98 to 0.78 mmol/g of dry matter in the incubated substrate. Compared to the control culture, a further decrease by ≥10 % in either the CH4 concentration or CH4 production was considered to be a significant effect (P < 0.05). Based on this criterion, 63 of the 100 herbal extracts significantly decreased the CH4 concentration in the culture headspace, while 31 of the herbal extracts decreased CH4 production (Fig. 1). The 10 most CH4-inhibiting herbal extracts were from the pericarp of Garcinia mangostana; stem, root, and twig peels of Magnolia officinalis; resin of Boswellia carterii; heartwood of Caesalpinia sappan; rhizomes of Curcuma longa; velamen of Acanthopanax gracilistylus; seeds of Cnidium monnieri; roots of Polygala tenuifolia; seeds of P. frutescens; and flower buds of Magnolia biondii. These 10 extracts each decreased CH4 production by at least 28 % (Fig. 1 and Table S1).
The relative changes (increase or decrease compared to the control value, in %) in methane production (mmol/g DM incubated) and gas production (mL/g DM incubated) in response to the addition of the herbal extracts. Only the 31 herbal extracts that significantly decreased methane production are shown. Each diamond represents the source of one herbal extract. (1) Garcinia magostana (pericarp), (2) Magnolia officinalis (stem, root, and twig peels), (3) Boswellia carterii (resin), (4) Caesalpinia sappan (heartwood), (5) Curcuma longa (rhizomes), (6) Acanthopanax gracilistylus (velamen), (7) Cnidium monnieri (seeds), (8) Polygala tenuifolia (roots), (9) Perilla frutescens (seeds), (10) Magnolia biondii (flower buds), (11) Notopterygiumincisum (rhizomes), (12) Lindera aggregate (earthnuts), (13) Cinnamomum cassia (bark), (14) Zingiber officinal (rhizomes), (15) Albizia julibrissin (barks), (16) Zingiber officinale (rhizomes), (17) Cynomorium songaricum (succulent stems), (18) Amomum villosum (fruit), (19) Cirsium japonicum (aerial parts), (20) Alpinia oxyphylla (fruit), (21) Rhapontisum unflorum (roots), (22) Vitex trifolia (seeds), (23) Paeonia lactiflora (roots), (24) Illicium verum (fruit), (25) Erodium stephanianum (aerial parts), (26) Citrus aurantium (fruitlets), (27) Equisetum hiemale (aerial parts), (28) Litchi chinensis (seeds), (29) Ziziphus jujube (seeds), (30) Perilla frutescens (leaves), and (31) Amygdalus persica (seeds)
Effects of select effective herbal extracts on CH4 production and other fermentation characteristics
The preliminary experiment identified 31 CH4-inhibiting herbal extracts (Table 1). When these herbal extracts were further evaluated, they displayed varying effects on the total gas production, CH4 production, total VFA concentration, and acetate/propionate (A/P) ratio (Table 1). Ten of these herbal extracts decreased total gas production, whereas another six increased total gas production. Twelve of these herbal extracts significantly decreased CH4 production per gram of incubated substrate, but one (the extract from Amygdalus persica) increased CH4 production. With the exception of the extract from P. tenuifolia, the other most CH4-inhibiting herbal extracts were confirmed to significantly suppress CH4 production (P < 0.05). Two (M. officinalis and Ziziphus jujube) of the 12 CH4-inhibiting herbal extracts did not decrease (P > 0.05) gas production. Only one herbal extract (from M. biondii) decreased (P < 0.05) total VFA production, while nine of them decreased (P < 0.05) the A/P ratio, including seven of the 12 CH4-inhibiting extracts.
Concentration-dependent effects of P. frutescens seed extract on rumen fermentation and CH4 production
The PF seed extract was the most potent at decreasing CH4 production in vitro (Table 1). All of the measured fermentation characteristics, including gas production (both total and effective), CH4 production, total VFAs, the A/P ratio, and dry matter degradation (DMD), were decreased linearly (P < 0.05) with increasing concentrations of the PF seed extract starting from 0.2 mg/L (Table 2). No effect on ammonia N or pH was noted at any of the concentrations. Methane production was decreased by the extract at concentrations of 0.1 mg/mL and higher, with the greatest decrease (by 32.6 %) being achieved at 0.4 mg/mL.
The PF seed extract did not decrease the population of total bacteria at any of the concentrations tested. The population of R albus and fungi was slightly decreased (P < 0.05) by the PF extract at 0.4 mg/mL. The activities of cellulase, CMCase, and xylanse were not decreased by the treatment (P > 0.05). These effects on microbial populations and enzyme activity corroborate the minimal DMD decrease at the two highest extract concentrations. The population of total methanogens was not affected (P > 0.05), irrespective of the concentration, while that of protozoa decreased (P < 0.05).
To identify the major compounds of the PF seed extract, its chemical composition was determined. Long chain fatty acids (LCFAs) were the major compounds in this herbal extract, accounting for >70 % of the total extract. Unsaturated C18 fatty acids amounted to 675.19 mg/g of the PF seed extract, including 452 mg of α-linolenic acid (18:3 cis,cis,cis-9,12,15), 116.61 mg of linoleic acid (18:2 cis,cis-9,12), and 106.58 mg of oleic acid (18:1 cis-9). Palmitic acid (C16) and stearic acid (C18) were detected as the major saturated LCFAs and were present at 15.47 and 14.59 mg/g, respectively, in the PF seed extract. Polyunsaturated fatty acids accounted for nearly 57 % of the PF seed extract.
Changes in the bacterial communities induced by the extract of P. frutescens seeds
After filtering and quality control, 481,610 quality-checked bacterial 16S rRNA gene sequences were obtained from a total of 520,632 joined sequences. For each sample, 36,410 sequences were used for a bacterial community analysis (Fig. S1A). The Chao 1 estimate was not affected, although a numerical decrease was observed at 0.4 mg/mL (P > 0.05, Fig. 2a). The Shannon-Wiener diversity index was decreased (P < 0.05) in a concentration-dependent manner (Fig. 2b). Collectively, the bacterial sequences represented 40 bacterial phyla, with Proteobacteria, Bacteroidetes, and Firmicutes together accounting for >89.6 % of the total bacterial sequences. The relative abundance of Proteobacteria linearly increased (P < 0.05), whereas that of Bacteroidetes, Fibrobacteres, Cyanobeteria, and Tenericutes linearly decreased (P < 0.05) in a concentration-dependent manner.
Box-and-whisker plots of the diversity indices of the bacterial (a) and (b) and the archaeal (c) and (d) communities affected by the extract of Perilla frutescens (PF) seeds at different concentrations. a and c Chao 1 estimates; b and d Shannon-Wiener diversity indices. Error bars indicate maximum and minimum values, while horizontal lines indicate median values, and boxes indicate values between the 25th and 75th percentiles. Boxes with different letters indicate significant differences (P < 0.05)
At the genus level, 235 taxa were identified, with the most predominant genera being Ruminobacter (24.6 % of all classified sequences), Prevotella (20.1 %), and Succinivibrio (5.2 %). A number of genus-level taxa were significantly affected by the PF seed extract (Fig. 3b). The relative abundance of Anaerovibrio, Pseudobutyrivibrio, Roseomonas, Ruminobacter, Selenomonas, Shuttleworthia, Succinivibrio, unclassified Sinobacteraceae, unclassified R4-41B, unclassified Succinivibrionaceae, and unclassified Burkholderiaceae linearly increased (Table S3) with increasing concentrations of the PF seed extract. However, the proportions of Anaeroplasma, unclassified [Prevotellaceae], Clostridium, Fibrobacter, Luteimonas, Paludibacter, Pedobacter, unclassified RFN20, Ruminococcus, Sphaerochaeta, unclassified YRC22, unclassified Anaeroplasmataceae, unclassified BS11, unclassified Caldilineaceae, unclassified Christensenellaceae, unclassified Lachnospiraceae, unclassified JdFBGBact, unclassified [Paraprevotellaceae], unclassified VC21_Bac22, unclassified Sphingobacteriaceae, and unclassified [Acidimicrobineae] linearly decreased.
Effects of increasing concentrations of the extract of Perilla frutescens seeds on the microbiota, at the phylum level (a) and at the genus level (b) of bacteria, and at the genus level of archaea (c). For bacterial genera, only those that were linearly (P < 0.05) changed are shown, and the heatmap shows the standardized relative abundance of each genus ([relative abundance – average relative abundance of all samples]/standard deviation)
The number of observed OTUs decreased with increasing concentrations of the PF seed extract: these values were 2668 at 0 mg/mL, 2465 at 0.2 mg/mL, and 2252 at 0.4 mg/mL. At the OTU level, the PCoA based on the weighted UniFrac distances (Fig. 4a) also showed that the PF seed extract affected the bacterial community at 0.4 mg/mL but not at 0.2 mg/mL. The Venn diagram (Fig. 4b) revealed that the control culture had the greatest number of unique OTUs (311 OTUs), followed by the cultures supplemented with 0.2 mg/mL (187 OTUs) and 0.4 mg/mL (165 OTUs) of the extract. The three concentrations shared 1680 OTUs (approximately 52.8 % of total OTUs), and these common OTUs accounted for 95.3 to 98.0 % of the total bacterial sequences across all of the samples. Of the shared OTUs, the collective relative abundance of the proteobacterial OTU increased primarily at the expense of that of Bacteroides, with that of Firmicutes remaining unchanged as the concentration of the PF seed extract increased (Fig. 4e).
Effects of increasing concentrations of the extract of Perilla frutescens seeds on bacterial and archaeal communities. Principal coordinate analysis (PCoA) plots of bacterial (a) and archaeal (b) communities; Venn diagrams showing the numbers of OTUs of bacteria (c) and archaea (d) unique to each treatment or common to all treatments; the relative abundance of the bacterial phyla (e) or archaeal genera (f) shared among the three treatments
Effect of the P. frutescens seed extract on archaeal communities
A total of 600,217 joined sequences were obtained for the archaeal 16S rRNA genes, and 509,087 quality-checked sequences remained after filtering and quality control. For each sample, 34,130 sequences were used for community analysis (Fig. S1B). The Shannon-Wiener diversity index decreased linearly with the increasing concentrations of the herbal extract (Fig. 2d), but no significant difference in Chao 1 estimate was noted (Fig. 2c). Among all of the samples, the archaeal sequences represented nine genera (Fig. 3c), with the genus Methanobrevibacter accounting for >82.9 % of the total archaeal sequences, followed by unclassified WCHD3-02 (>8.21 %), unclassified Methanobacteriaceae (>2.37 %), Methanosphaera (>0.84 %), and Methanoplanus (>0.62 %). The sequencing data revealed an increase in the relative abundance of Methanobrevibacter and Methanosphaera, while that of unclassified Methanobacteriaceae decreased in a concentration-dependent manner (Fig. 3c and Table S3). In the genus Methanobrevibacter, M. ruminantium (>17.4 % of Methanobrevibacter sequences) did not change in relative abundance following treatment with the PF seed extract, but the relative abundance of M. millerae (<0.68 % of Methanobrevibacter sequences) decreased, while that of unclassified Methanobrevibacter (>84.1 % of Methanobrevibacter sequences) increased in a concentration-dependent manner.
Corresponding to 0, 0.2, and 0.4 mg/mL of the PF seed extract, 102, 97, and 110 archaeal OTUs were found. The greatest number (26) of unique OTUs was found following treatment with 0.4 mg/mL PF seed extract, and 63 OTUs (approximately 42.0 % of total OTUs) were shared among the samples treated with the three different concentrations (Fig. 4d). These common OTUs accounted for 99.9 % of the total archaeal sequences. No difference in the relative abundance of any of the genera represented by the common OTUs was apparent among the three concentrations (Fig. 4f). At the community level, the three concentrations were separated along the PC2 axis, which explained <17 % of the total variation, but not along the PC1 axis, which explained >71 % of the total variation (Fig. 4b).
Correlation between the rumen microbial taxa and rumen fermentation characteristics
A correlation was found between the number of genera/group of rumen microbes and several fermentation characteristics (Table 3). Based on the correlation patterns, five groups of microbes were found. Group I was positively (P < 0.05, r > 0.607) correlated with the production of both CH4 and total VFA. Included in this group were Fibrobacter, Sphaerochaeta, eight groups of unclassified bacteria, and one group of unclassified Methanobacteriaceae. Group II, containing 14 genera/groups of bacteria, including Clostridium, Pedobacter, Anaeroplasma, Paludibacter, Ruminococcus, Zymomonas, Luteimonas, and seven unclassified bacterial groups, was positively correlated (P < 0.05, r > 0.602) with CH4 production but not with total VFA production (either positively or negatively). In contrast to group I, group III was negatively (P < 0.05, r > −0.598) correlated with the production of both CH4 and VFA, and this group included Roseomonas, Selenomonas, Shuttleworthia, Pseudobutyrivibrio, Anaerovibrio, Ruminobacter, Succinivibrio, Methanosphaera, and three groups of unclassified bacteria. Group IV included Methanobrevibacter and unclassified Sinobacteraceae, and it only exhibited a negative (P < 0.05, r > −0.650) correlation with CH4 production. Group V included two groups of unclassified bacteria, and it was not correlated with the production of either CH4 or total VFA. In general, more genera/groups of the microbes had a correlation with both CH4 production and the A/P ratio than with both CH4 production and DMD (Table 3).
Discussion
This study evaluated the extracts of a large number (100) of medicinal herbs for their potential to decrease CH4 production and modulate rumen fermentation. The decreased CH4 concentration in the headspace of the cultures supported our hypothesis that medicinal herbs are potentially rich sources of compounds that can be explored to mitigate CH4 emission from ruminants. It should be noted that even though DMSO also decreased the CH4 production, the decrease in CH4 production by the herbal extracts was determined by comparison with a control that contained DMSO. When evaluated further, only one of these selected herbal extracts decreased (P < 0.05) total VFA production slightly, while most of these herbal extracts had little effect on fermentation at the tested concentration (0.4 mg/mL). However, some of these herbal extracts altered the fermentation profiles, suggesting that they had effects on some microbial populations. Interestingly, the extract from one of the herbs, A. persica, increased the production of both CH4 and total gas, but not the ratio of CH4 to total gas, suggesting that it had the ability to enhance fermentation. The extracts of two herbs, M. officinalis and Z. jujube, decreased CH4 production but not the total gas or VFA, suggesting that they can inhibit CH4 production without decreasing feed digestion or fermentation. However, they only decreased CH4 production by a small magnitude. Only four of the herbs evaluated in the present study have been tested in previous studies. A powder (70 mg in 50 mL culture) of Zingiber officinale, Illicium verum, Citrus aurantium, and Trigonella foenum-graecum did not affect CH4 production in vitro (García-González et al. 2008). The results of the present study corroborate the findings of the above study. However, Z. officinale extract (6 mg/mL) decreased CH4 production in vitro by 16 % (Kim et al. 2012a, b). A wide range of concentrations need to be used in future screenings of herbs in order to better assess their potential to decrease CH4 emission.
The PF seed extract resulted in the largest decrease in production of both CH4 and total gas, so it was further evaluated for its concentration-dependent effects on common fermentation characteristics, bacterial and archaeal communities, and the populations of selected rumen microbes. It should be noted that although the CH4 production was decreased to a greater extent at 0.4 mg/mL and at concentrations higher than 0.2 mg/mL, both the DMD and total VFA production were also decreased. Therefore, PF seed extract may be used at ≤0.2 mg/mL to achieve significant CH4 mitigation without decreasing feed digestion or fermentation. The decreased CH4 production was accompanied by a decreased A/P ratio, suggesting that it may be possible to direct hydrogen to alter the production of reduced VFA, including propionate.
The population of total bacteria and other groups of rumen microbes known to play an important role in feed digestion and/or CH4 production was quantified by qPCR to help understand the mode of action of the PF seed extract. The population of total bacteria was not decreased by the PF seed extract at any of the concentrations tested, suggesting that there was a lack of inhibition of microbial growth or microbial protein synthesis. The uncoupling of the methanogen population size and CH4 production is consistent with the findings of previous studies involving short-term in vitro incubation (Patra and Yu 2014b). As suggested by Ohene-Adjei et al. (2008), the methanogen abundance would probably decrease after prolonged inhibition of CH4 synthesis.
The bacterial and archaeal communities in the in vitro cultures were examined to help further explain the effects of the PF seed extract and to identify specific groups of bacteria and methanogens that are potentially associated with the altered CH4 production and fermentation characteristics. Because the Chao 1 richness estimate was not affected, but the Shannon-Wiener diversity index, which reflects both richness and evenness, was decreased linearly, the PF seed extract apparently altered the relative abundance of some bacterial populations, rather than completely eliminating or introducing certain bacterial populations. Indeed, the relative abundance of Proteobacteria increased at the expense of that of Bacteroidetes, while that of Firmicutes was not affected. At the genus level, six genera and several unclassified groups gained predominance, including Anaerovibrio, Pseudobutyrivibrio, Roseomonas, Ruminobacter, Selenomonas, and Succinivibrio, all of which are Gram-negative taxa (Bergey’s Manual Trust 2015). On the other hand, some genera lost predominance, including Anaeroplasma (G−), Clostridium (G+), Luteimonas (G−), Paludibacter (G−), Pedobacter (G−), Ruminococcus (G+), and Sphaerochaeta (G−). These results suggest the PF seed extract does not affect bacteria solely based on their Gram-staining characteristics.
The possible relationships between individual genera or genera-level groups and CH4 production, VFAs, and A/P were examined using correlation analysis. Five groups of bacteria/and or methanogens displayed different types of correlations with the production of methane and VFAs (Table 3). The relative abundance of some bacterial genera, including Roseomonas, Shuttleworthis, Selenomonas, Pseudobutyrivbrio, Anaerovibrio, Ruminobacter, Succinivibrio, and three unclassified groups, was negatively correlated with CH4 production. All of the above genera can produce >C2 VFA, such as butyrate, lactate, propionate, and succinate (Stewart et al. 1997; Bergey’s Manual Trust 2015). Indeed, low CH4 emissions were found to be accompanied by a high abundance of Succinivibrionaceae (containing Ruminobacter and Succinivibrio) (Wallace et al. 2015). On the other hand, a number of genera were positively correlated with CH4 production, including Anaeroplasma, Pedobacter, Paludibacter, Ruminococcus, Zymomonas, Fibrobacter, Lachnospiraceae, and Sphaerochaeta. The data generated in the present study cannot directly prove that the above genera are the main contributors to CH4 production in the in vitro cultures, but functions reported in the literature may help understand their involvement in CH4 production. For example, Pedobacter was found to be one of the hydrogen-producing cellulolytic bacteria (Jiang et al. 2015) with CMCase and cellulase activity (Khamtib and Reungsang 2012). Anaeroplasma can ferment sugars to acetate and formate (Robinson et al. 1975). Paludibacter can utilize various sugars and produce acetate and propionate as major fermentation end-products, with succinate as a minor product (Ueki et al. 2006). Ruminococcus (Latham and Wolin 1977) and Lachnospiraceae are known acetate and H2 producers (Stewart et al. 1997). A strong association between CH4 emissions and the abundance of H2-producing bacteria has been reported in sheep (Kittelmann et al. 2014) and in cows (Wallace et al. 2015). Consistent with the decreased A/P ratio observed in the present study and in a previous report (Russell and Strobel 1989), the PF seed extract might have decreased CH4 production by directing hydrogen to >C2 VFA production. However, the current data do not explain the positive correlation with the relative abundance of Fibrobacter, which produces succinate. However, the absolute abundance (16S rRNA gene copies/g of sample) of F. succinogenes, one of the dominant Fibrobacter species in the rumen, was not affected in the in vitro cultures. Thus, the above correlation between the relative abundance of Fibrobacter and CH4 methane production might have been due to changes in the abundance (both relative and absolute) of other bacterial genera. Mathematically, the relative abundance of one bacterial group may change due to changes in the relative abundance of other bacteria, even though the absolute abundance of that bacterial group remains unchanged. Therefore, any effect on bacterial populations should be interpreted with caution if the relative abundance is used as the measurement.
The PF seed extract only affected the archaeal communities to a limited degree by altering the relative abundance of individual taxa and OTUs. It did not affect the species richness, as indicated by the decreased Shannon-Wiener diversity index but unchanged Chao 1 richness estimates. The qPCR data showed that the total population of archaea was not affected by the PF seed extract. The sequencing data revealed an increase in the relative abundance of Methanobrevibacter and Methanosphaera, while the relative abundance of unclassified Methanobacteriaceae was decreased. Methanobrevibacter species are the dominant methanogens in the rumen (Skillman et al. 2006; Wright et al. 2004), and M. gottschalkii, M. smithii, M. millerae, and M. thaueri were correlated with high CH4 production, while M. ruminantium and M. olleyae were correlated with low CH4 production (Danielsson 2016; Danielsson et al. 2012). Although these Methanobrevibacter species were not quantified by qPCR in the present study, the PF seed extract decreased the proportion of M. millerae, suggesting that the PF seed extract has various effects on individual Methanobrevibacter species.
P. frutescens is a plant with a pleasant aroma. It is widely distributed and used as a medicinal herb in Asia, an ornamental plant in Western countries, and a naturalized or invasive plant in the USA (Boning 2010). In the present study, unsaturated fatty acids, especially linolenic acid (C18:3), were found to be the major compounds in the PF seed extract. Polyunsaturated fatty acids (PUFAs) are well documented as being methane-suppressing (Patra 2013) by directly inhibiting methanogens and protozoa (Dohme et al. 1999) and by serving as hydrogen acceptors (Loor et al. 2002). The PUFAs in the PF seed extract are probably the major components that affect fermentation and CH4 production. However, mixed results were reported for the effects of LCFAs and PUFAs on methanogens (Dohme et al. 1999) and bacteria (Machmüller and Kreuzer 1999). In addition to LCFAs, PF seeds also contain other compounds, including policosanol, phytosterol, and tocopherol (Kim et al. 2012a, b). Perilla seeds are rich in protein (17 % of DM) and fat (51 %) (Mellinas et al. 2016). Therefore, P. frutescen seeds, rather than seed extracts, may be incorporated into ruminant rations for practical application to help mitigate CH4 emission from ruminants while enhancing their nutrition. Future feeding trials are warranted to test this premise.
In conclusion, medicinal herbs are potential sources of compounds that can be used to mitigate CH4 emission from ruminant animals. The ethanol extract of P. frutescens seeds decreased CH4 production without adversely affecting feed fermentation when included at a relatively low concentration. Given the high content of proteins and lipids, P. frutescens seeds may be directly incorporated into rations to both enhance nutrition and mitigate CH4 emission. A number of bacteria and methanogens were found to be correlated with CH4 production, and a better understanding of these bacteria and methanogens may facilitate the development of new strategies to mitigate CH4 emission without adversely affecting rumen fermentation.
References
Ahmad A, Husain A, Mujeeb M, Khan SA, Najmi AK, Siddique NA, Damanhouri ZA, Anwar F (2013) A review on therapeutic potential of Nigella sativa: a miracle herb. Asian Pac J Trop Biomed 3(5):337–352
Bailey MJ, Biely P, Poutanen K (1992) Interlaboratory testing of methods for assay of xylanase activity. J Biotechnol 23:257–270
Benchaar C, Hassanat F, Martineau R, Gervais R (2015) Linseed oil supplementation to dairy cows fed diets based on red clover silage or corn silage: effects on methane production, rumen fermentation, nutrient digestibility, N balance, and milk production. J Dairy Sci 98(11):7993–8008
Bergey’s Manual Trust (2015) Bergey’s manual of systematics of Archaea and Bacteria. Wiley, in association with Bergey’s Manual Trust, New York
Bhatta R, Saravanan M, Baruah L, Prasad CS (2015) Effects of graded levels of tannin-containing tropical tree leaves on in vitro rumen fermentation, total protozoa and methane production. J Appl Microbiol 118(3):557–564
Boning C (2010) Florida’s best herbs and spices. Pineapple Press Inc., Sarasota, Florida, pp. 148–149
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7(5):335–336
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci U S A 108(Suppl 1):4516–4522
Danielsson R (2016) Methane production in dairy cows: Impact of feed and rumen microbiota. Acta Univ Agric Sueciae:45
Danielsson R, Schnürer A, Arthurson V, Bertilsson J (2012) Methanogenic population and CH4 production in Swedish dairy cows fed different levels of forage. Appl Environ Microbiol 78(17):6172–6179
Dohme F, Machmüller A, Estermann BL, Pfister P, Wasserfallen A, Kreuzer M (1999) The role of the rumen ciliate protozoa for methane suppression caused by coconut oil. Lett Appl Microbiol 29:187–192
Gagen EJ, Denman SE, Padmanabha J, Zadbuke S, Al Jassim R, Morrison M, McSweeney CS (2010) Functional gene analysis suggests different acetogen populations in the bovine rumen and tammar wallaby forestomach. Appl Environ Microbiol 76(23):7785–7795
García-González R, López S, Fernández M, Bodas R, González JS (2008) Screening the activity of plants and spices for decreasing ruminal methane production in vitro. Anim Feed Sci Technol 147:36–52
Grainger C, Beauchemin KA (2011) Can enteric methane emissions from ruminants be lowered without lowering their production? Anim Feed Sci Technol 166-167:308–320
Hristov AN, Oh J, Firkins JL, Dijkstra J, Kebreab E, Waghorn G, Makkar HPS, Adesogan AT, Yang W, Lee C, Gerber PJ, Henderson B, Tricarico JM (2013) Special topics-mitigation of methane and nitrous oxide emissions from animal operations: I. A review of enteric methane mitigation options. J Anim Sci 91(11):5045–5069
IPCC (Intergovernmetnal Panel on Climate Change) (2007) Climate change 2007: the physical science basis. Cambridge University Press, Cambridge
Jayanegara A, Marquardt S, Kreuzer M, Leiber F (2011) Nutrient and energy content, in vitro ruminal fermentation characteristics and methanogenic potential of alpine forage plant species during early summer. J Sci Food Agric 91(10):1863–1870
Jiang H, Gadow SI, Tanaka Y, Cheng J, Li YY (2015) Improved cellulose conversion to bio-hydrogen with thermophilic bacteria and characterization of microbial community in continuous bioreactor. Biomass Bioenerg 75:57–64
Johnson KA, Johnson DE (1995) Methane emissions from cattle. J Anim Sci 73(8):2483–2492
Kamra DN, Agarwal N, Chaudhary LC (2005) Inhibition of ruminal methanogenesis by tropical plants containing secondary compunds. Int Cong Ser 1293:156–163
Kemp GW, McSweeney CS (2010) Screening of plants for inhibitory activity against pathogenic microorganisms from the gut of livestock. In: Vercoe PE, Makkar HPS, Schlink AC (eds) In vitro screening of plant resources for extra-nutritional attributes in ruminants: nuclear and related methodologies. Springer Netherlands, Berlin, pp. 145–158
Khamtib S, Reungsang A (2012) Biohydrogen production from xylose by Thermoanaerobacterium thermosaccharolyticum KKU19 isolated from hot spring sediment. Int J Hydrog Energy 37:12219–12228
Kim ET, Kim CH, Min KS, Lee SS (2012a) Effects of plant extracts on microbial population, methane emission and ruminal fermentation characteristics in in vitro. Asian-Aust J Anim Sci 25(6):806–811
Kim JK, Park SY, Na JK, Seong ES, Yu CY (2012b) Metabolite profiling based on lipophilic compounds for quality assessment of perilla (Perilla frutescens) cultivars. J Agric Food Chem 60(9):2257–2263
Kittelmann S, Pinares-Patiño CS, Seedorf H, Kirk MR, Ganesh S, McEwan JC, Janssen PH (2014) Two different bacterial community types are linked with the low-methane emission trait in sheep. PLoS One 9(7):e103171
Latham MJ, Wolin MJ (1977) Fermentation of cellulose by Ruminococcus flavefaciens in the presence and absence of Methanobacterium ruminantium. Appl Environ Microbiol 34(3):297–301
Liu MZ, Zhang YL, Zeng MZ, He FZ, Luo ZY, Luo JQ, Wen JG, Chen XP, Zhou HH, Zhang W (2015) Pharmacogenomics and herb-drug interactions: merge of future and tradition. Evid-Based Compl Alt 321091. doi: 10.1155/2015/321091
Loor JJ, Bandara ABPA, Herbein JH (2002) Characterization of 18:1 and 18:2 isomers produced during microbial biohydrogenaation of unsaturated fatty acids from canola and soya bean oil in the rumen of lactating cows. J Anim Physiol Anim Nutr 86(11–12):422–432
Lopez-Lopez A, Castellote-Bargallo AI, Campoy-Folgoso C, Rivero-Urgel M, Tormo-Carnice R, Infante-Pina D, Lopez-Sabater MC (2001) The influence of dietary palmitic acid triacylglyceride position on the fatty acid, calcium and magnesium contents of at term newborn faeces. Early Hum Dev 65(Suppl2):S83–S94
Machmüller A, Kreuzer M (1999) Methane suppression by coconut oil and associated effects on nutrient and energy balance in sheep. Can J Anim Sci 79:65–72
Magoč T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27(21):2957–2963
Mellinas C, Valdés A, Ramos M, Burgos N, Garrigós MDC, Jiménz A (2016) Active edible films: current state and future trends. J Appl Polym Sci 133(2):42631
Ohene-Adjei S, Chaves AV, McAllister TA, Benchaar C, Teather RM, Forster RJ (2008) Evidence of increased diversity of methanogenic archaea with plant extract supplementation. Microb Ecol 56(2):234–242
Patra AK (2013) The effect of dietary fats on methane emissions, and its other effects on digestibility, rumen fermentation and lactation performance in cattle: a meta-analysis. Livest Sci 155(2–3):244–254
Patra AK, Yu Z (2013) Effective reduction of enteric methane production by a combination of nitrate and saponin without adverse effect on feed degradability, fermentation, or bacterial and archaeal communities of the rumen. Bioresour Technol 148:352–360
Patra AK, Yu ZT (2014a) Effects of vanillin, quillaja saponin, and essential oils on in vitro fermentation and protein-degrading microorganisms of the rumen. Appl Microbiol Biotechnol 98(2):897–905
Patra AK, Yu ZT (2014b) Combinations of nitrate, saponin, and sulfate additively reduce methane production by rumen cultures in vitro while not adversely affecting feed digestion, fermentation or microbial communities. Bioresour Technol 155:129–135
Robinson IM, Allison MJ, Hartman PA (1975) Anaeroplasma abactoclasticum gen.nov., sp.nov.: an obligately anaerobic mycoplasma from the rumen. Int J Syst Evol Microbiol 25(2):173–181
Russell JB, Strobel HJ (1989) Effect of ionophores on ruminal fermentation. Appl Environ Microbiol 55(1):1–6
Skillman LC, Evans PN, Strompl C, Joblin KN (2006) 16S rDNA directed PCR primers and detection of methanogens in the bovine rumen. Lett Appl Microbiol 42(3):222–228
Stewart CS, Flint HJ, Bryant MP (1997) The rumen bacteria. In: Hobson PN, Stewart CS (eds) The rumen microbial ecosystem. Blackie Academic & Professional, London, pp. 10–72
Theodorou MK, Williams BA, Dhanoa MS, McAllan AB, France J (1994) A simple gas production method using a pressure transducer to determine the fermentation kinetics of ruminant feeds. Anim Feed Sci Technol 48(3–4):185–197
Ueki A, Akasaka H, Suzuki D, Ueki K (2006) Paludibacter propioicigenes gen. Nov., sp. nov., a novel strictly anaerobic, gram-negative, propionate-producing bacterium isolated from plant residue in irrigated rice-field soil in Japan. Int J Syst Evol Microbiol 56(Pt 1):39–44
Wallace RJ, Rooke JA, McKain N, Carol-Anne D, Hyslop JJ, Ross DW, Waterhouse A, Watson M, Roeche R (2015) The rumen microbial metagenome associated with high methane production in cattle. BMC Genomics 16:839
Wang JK, He B, Du W, Luo Y, Yu Z, Liu JX (2015) Yeast with surface displayed xylanase as a new dual purpose delivery vehicle of xylanase and yeast. Anim Feed Sci Technol 208:44–52
Wright AD, Williams AJ, Winder B, Christophersen CT, Rodgers SL, Smith KD (2004) Molecular diversity of rumen methanogens from sheep in Western Australia. Appl Environ Microbiol 70(3):1263–1270
Zeleke AB, Clément C, Hess HD, Kreuzer M, Soliva CR (2005) Effect of foliage from multi-purpose trees and a leguminous crop residue on in vitro methanogenesis and ruminal N use. Int Cong Ser 1293:168–171
Acknowledgments
The authors acknowledge financial support from the National Key Technology Program (2015BAC02B05) and partial support from the Beijing Key Laboratory for Dairy Cow Nutrition.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
This study was funded by the National Key Technology Program (2015BAC02B05) and partially by the Beijing Key Laboratory for Dairy Cow Nutrition.
Conflict of interest
The authors declare that they have no competing interests.
Electronic supplementary material
ESM 1
(DOCX 214 KB)
Rights and permissions
About this article
Cite this article
Wang, J., Liu, M., Wu, Y. et al. Medicinal herbs as a potential strategy to decrease methane production by rumen microbiota: a systematic evaluation with a focus on Perilla frutescens seed extract. Appl Microbiol Biotechnol 100, 9757–9771 (2016). https://doi.org/10.1007/s00253-016-7830-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-016-7830-z