Abstract
Thermobifida fusca is a moderately thermophilic soil bacterium belonging to Actinobacteria. It has been known for its capability to degrade plant cell wall polymers except lignin and pectin. To know whether it can produce enzymes to facilitate lignin degradation, the extracellular proteins bound to sugarcane bagasse were harvested and identified by liquid chromatography tandem mass spectrometry. Among the identified proteins, a putative copper-containing polyphenol oxidase of 241 amino acids, encoded by the locus Tfu_1114, was thought to presumably play a role in lignin degradation. This protein (Tfu1114) was thus expressed in E. coli and characterized. Similarly to common laccases, Tfu1114 is able to catalyze the oxidation reaction of phenolic and nonphenolic lignin related compounds such as 2,6-dimethoxyphenol and veratryl alcohol. More interestingly, it can significantly enhance the enzymatic hydrolysis of bagasse by xylanase and cellulase. Tfu1114 is stable against heat, with a half-life of 4.7 h at 90 °C, and organic solvents. It is sensitive to ethylenediaminetetraacetic acid and reducing agents but resistant to sodium azide, a potent inhibitor of laccases. Atomic absorption spectroscopy indicated that the ratio of copper to the protein monomer is 1, instead of 4, a feature of classical laccases. All these data suggest that Tfu1114 is a novel oxidase with laccase-like activity, potentially useful in biotechnology application.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
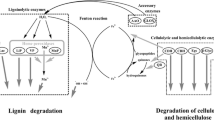
Over the past decades, seeking sustainable energy has gained its momentum due to the continuously increasing price of gasoline. Biofuel produced from lignocellulosic agricultural wastes has been considered to be one of the alternatives if economic and environment-friendly processes can be developed. One of the major hurdles in the way of achieving this goal is the difficulty of removing lignin from the wastes; therefore, the cellulose and hemicellulose within cannot be digested efficiently by the corresponding glycoside hydrolases (Mussatto et al. 2010). Lignin peroxidase, manganese-dependent and manganese-independent peroxidases and laccase, particularly those produced by white-rot fungi, have long been known to be able to promote the lignin degradation (Moreira et al. 1997). Nonetheless, the application of these enzymes in the biofuel production process is still far from practical in the viewpoint of cost.
Laccases (EC 1.10.3.2) are blue multicopper oxidases that catalyze the oxidation of both phenolic and nonphenolic lignin-related compounds as well as highly recalcitrant environmental pollutants, coupled with the reduction of molecular oxygen to water (Madhavi and Lele 2009; Giardina et al. 2010). The broad substrate range makes laccases excellent candidates for many industrial and biotechnological applications, such as decolorization of textile dyes and bioremediation of soils and water (Durán et al. 2002; Rodriguez Couto and Toca Herrera 2006). Laccases have been isolated from many plants, fungi, and bacteria. On sodium dodecyl sulfate polyacrylamide gel electrophoresis (SDS-PAGE), most laccases show mobility corresponding to molecular mass 60–100 kDa, and many, particularly those from eukaryotes, are glycosylated (Baldrian 2006). The plant laccases play a role in the formation of lignin by promoting the oxidative coupling of monolignols (Solomon et al. 1996). Most of the known laccases are of fungal origin, and they participate into a variety of physiological functions such as stress defense and lignin degradation (Giardina et al. 2010; Kües and Rühl 2011). CotA, a component of the endospore coat of Bacillus subtilis, exemplifies the bacterial laccases (Hullo et al. 2001). It was thought to be associated with the biosynthesis of spore pigment, and spore resistance to UV light and H2O2 (Enguita et al. 2003). Fungal and CotA-like bacterial laccases have conserved regions in which histidine residues can bind four copper atoms located at two main sites, the type 1 blue copper center T1 and the T2/T3 trinuclear cluster (Ducros et al. 1998; Enguita et al. 2003; Hakulinen et al. 2002; Hullo et al. 2001; Piontek et al. 2002). These copper atoms are essential for the oxidation activity. Recently, a new family of bacterial laccases (the DUF152 family), including RL5 isolated from bovine rumen microflora, BT4389 from Bacteroides thetaiotaomicron, and YfiH from Escherichia coli, was found to have strong oxidation activities toward various aromatic compounds (Beloqui et al. 2006). Inductively coupled plasma mass spectrometry indicated that RL5 also contains four copper atoms per monomer, although it is quite different from CotA in terms of the length and amino acid sequence of the polypeptides.
Thermobifida fusca is a Gram-positive, moderately thermophilic, filamentous soil bacterium. It is known as an excellent producer of cellulolytic enzymes (Maki et al. 2009). However, no ligninolytic enzyme from T. fusca has been reported in the literature. T. fusca NTU10-1 strain, isolated from local compost, thrives in the medium with sugarcane bagasse as the sole carbon source. This indigenous strain could produce a great quantity of extracellular endoxylanases (Yang et al. 2007). It was postulated that this strain may produce enzymes to help the removal of lignin so that the cellulose and hemicellulose in bagasse could be consumed efficiently. Database mining of the complete genome sequence of T. fusca YX, which was accessible in 2007 (Lykidis et al. 2007), did not find genes that encode fungal laccase-like proteins. However, one open reading frame (Tfu_1114) encodes a hypothetical protein that contains consensus sequences of the protein family DUF152. In this study, the protein, tentatively named Tfu1114, was found in the culture medium of T. fusca NTU10-1 and to be able to bind to sugarcane bagasse. Tfu1114 was thus overexpressed in E. coli and purified to homogeneous state. Activity assay confirmed that Tfu1114 can catalyze the oxidation of a number of aromatic compounds, commonly used in the laccase activity assays. More interestingly, Tfu1114 can help endoxylanase and endocellulase to release reducing sugar equivalents from sugarcane bagasse.
Materials and methods
Bagasse preparation
Sugarcane bagasse, collected from the local market, was washed extensively with running water until the residual soluble sugar was removed. Then, it was dried and smashed with a blender. The small pieces, passed through a 100 mesh screen, were collected and used in this study.
Bacterial strains
T. fusca NTU10-1, cultivated routinely in CYC medium (Czapek-Dox powder, 33 g/L; yeast extract, 2 g/L; casamino acids, 6 g/L; pH 7.2) at 50 °C, was the source of chromosomal DNA and extracellular proteins studied in this study. This strain was deposited in the Bioresource Collection and Research Center (BCRC, Hsinchu, Taiwan) with stock number BCRC 19214. E. coli BL21(DE3) strain (Merck KGaA, Darmstadt, Germany) was used as the host for heterologous protein expression.
Extracellular protein identification
T. fusca NTU10-1 was cultivated in 100 ml CYC medium that also contained 0.125 % dry bagasse at 50 °C, 200 rpm of shaking, for 6 days. The supernatant was collected by centrifugation at 6,000 rpm for 30 min and filtrated through polyethersulfone bottle-top filter unit (pore size, 0.2 μm) (Merck KGaA, Darmstadt, Germany). The clarified sample was concentrated to 5 ml and mixed with approximately 40 ml swollen bagasse in 50 mM phosphate buffer, pH 7.5. After an incubation on a rotary shaker at 4 °C for overnight, the mixture was packed into a column (8 × 2.8 cm) (Bio-Rad Laboratories Inc., Hercules, USA). The unbound proteins in the column were washed away with 400 ml phosphate buffer, and the bound proteins were eluted with 0.2 M succinate buffer, pH 4.0. The proteins in the eluate were precipitated with trichloroacetic acid and redissolved in water. Finally, the protein identity was analyzed by liquid chromatography tandem mass spectrometry (LC-MS/MS) using an Applied Biosystems QStar LC-MS/MS spectrometer (Life Technologies Corp., Carlsbad, USA). The obtained mass spectrometry information was analyzed with Mascot software (Matrix Science Ltd., London, UK) using the NCBInr database. The important parameter settings for Mascot analysis were as follows: mass values, monoisotopic; protein mass, unrestricted; peptide mass tolerance, ±0.5 Da; fragment mass tolerance, ±0.5 Da; and maximal missed cleavages, 2.
Plasmids
Tfu_1114 was amplified from the chromosomal DNA of T. fusca NTU10-1 in PCR using primers 5′-TATGAATTCAGTGACGGGCACCGTGGTCGAG and 5′-TTTGGTACCTCATGGGACCCTCCAGACATATCCGG (the nucleotides in italic are the engineered restriction sites of EcoRI and KpnI, respectively) according to the gene sequence of T. fusca YX (NCBI accession number YP_289175). The PCR-amplified fragment was recovered and digested with EcoRI and KpnI, and then ligated with EcoRI–KpnI-treated pETDuet-1 (Merck KGaA, Darmstadt, Germany) to generate the Tfu1114 expression plasmid. Similarly, Tfu_1213 and Tfu_1074, coding for endoxylanase and endocellulase, respectively, were amplified by PCR and inserted into pETDuet-1 for the production of the mature form of the proteins in E. coli. All the recombinant proteins were fused with a His-tail at the N terminus.
Recombinant protein preparation
The E. coli BL21(DE3) cells, harboring the desired plasmid, were grown in 200 ml ampicillin-containing Luria–Bertani medium at 37 °C. To induce the protein expression, 1 mM isopropyl-β-d-thio-galactopyranoside and 0.5 mM copper sulfate, in the case of producing Tfu1114, were added into the culture when the cell density reached OD600 ≈ 0.8. The cultivation was continued for 4 h. The cell pellet after centrifugation was suspended in 20 ml lysis buffer (50 mM Tris–HCl, pH 8.0, 100 mM NaCl, 10 % glycerol), supplemented with 1 mM phenylmethylsulfonyl fluoride and 5 mM imidazole, and disrupted on ice with Sonicator 3000 (Misonix Inc., Farmingdale, USA) with 5 s pulse and 10 s pause intermittently, power setting at 30 W, for total 15 min. The homogenized cell extract was clarified by centrifugation at 10,000×g for 10 min. The supernatant was loaded onto a 5-ml Ni2+-NTA column (Merck KGaA, Darmstadt, Germany). The column was washed with 100 ml lysis buffer, supplemented with 20 mM imidazole, and the protein bound to the column was finally eluted with the lysis buffer containing 500 mM imidazole. In the case of Tfu1114, the purified enzyme was dialyzed against a 10,000-fold volume of lysis buffer to remove the excess copper ion and imidazole. The protein concentration was determined using the Bradford assay kit (Bio-Rad Laboratories Inc., Hercules, USA) with bovine serum albumin as the standard.
Activity assay
Unless otherwise indicated, the standard laccase activity assay was carried out at 50 °C for 15 min, using 2 mM 2,6-dimethoxyphenol (2,6-DMP) as the substrate in 20 mM phosphate buffer, pH 8.0. The 2,6-DMP oxidation was monitored by the increase in absorbance at 470 nm (ε 470 = 35,645 M−1 cm−1). Alternative substrates for measurement of laccase activity were veratryl alcohol (ε 310 = 9,300 M−1 cm−1), 2,2′-azino-bis(3-ethylbenzthiazoline)-6-sulfonate (ABTS) (ε 420 = 36,000 M−1 cm−1), and guaiacol (ε 470 = 26,600 M−1 cm−1). One unit was defined as the activity to oxidize 1 μmol of the substrate per minute under the indicated reaction condition.
Bagasse hydrolysis assay was carried out by mixing 10 mg dried bagasse with the specified amounts of the recombinant T. fusca endoxylanase (product of Tfu_1213), endocellulase (product of Tfu_1074) or/and Tfu1114, obtained as the aforementioned description in this text, in 1 ml 20 mM phosphate buffer, pH 8.0. The mixture was placed on a rotary shaker, 250 rpm, at 50 °C for 15 h. The reducing sugars released into the supernatant after the incubation was estimated by dinitrosalicylic acid method using glucose as the standard.
Protein binding activity to bagasse was assayed by mixing 0.5 mg of the protein and 10 mg bagasse in 1 ml 20 mM phosphate buffer, pH 8.0. The mixture was shaken at 4 °C for 30 min. After centrifugation at 12,000 rpm for 20 min, the amount of the protein still remained in the supernatant was estimated by SDS–12 % PAGE.
Instrumental analysis
The appearance of bagasse after being treated with the indicated enzymes was observed with a tabletop scanning electron microscope TM-1000 (Hitachi Ltd., Tokyo, Japan). The sample was mounted on a circular aluminum stub with double carbon sticky tape and then examined at an accelerating potential of 1.5 kV. The copper content of Tfu1114 was measured using Pinnacle 900H graphite furnace atomic absorption spectrometer (PerkinElmer Inc., Waltham, USA) equipped with Lumina hollow cathode lamp. Copper pure single-element standard (PerkinElmer Inc., Waltham, USA) was used as the standard.
Results
The reason to study Tfu1114
T. fusca NTU10-1 can efficiently digest sugarcane bagasse (data not shown). To know more about the enzymes involved in bagasse hydrolysis, the extracellular proteins that bound to bagasse were collected as described in the “Materials and methods,” and the protein composition in the collection were analyzed by LC tandem mass spectrometry (Table 1). Details of the spectrometry data can be seen in the supplementary Table S1. As expected, a bunch of cellulases, xylanases, and cellulose-binding proteins were identified. There were also two proteases in the collection. Obviously, these proteins are critical for the hydrolysis of composite polysaccharides and proteins. Another group of the proteins consists of components of the ATP-binding cassette-type transporters and other transport systems. They should participate into active transport of the hydrolyzed soluble products. A couple of oxidoreductases and a hypothetical single-stranded DNA-binding protein were also found in the collection. Their functions in relation to bagasse hydrolysis were unclear. Although all the proteins were eluted from bagasse, some of them might bind to bagasse indirectly through protein–protein interactions.
Interestingly, one of the oxidoreductases found in the collection is Tfu1114, a putative polyphenol oxidase belonging to the protein family DUF152. Would it be possible that Tfu1114 functions as a laccase to promote the enzymatic hydrolysis of bagasse by glycoside hydrolases? To test this hypothesis, Tfu1114 was chosen for functional analysis in this study. The gene Tfu_1114 was amplified from the chromosomal DNA of T. fusca NTU10-1 by PCR. The obtained PCR product has one nucleotide difference from that of T. fusca YX in the total 723 nucleotides, leading to a substitution of lysine for Arg49 in this hypothetical protein. Multiple sequence alignment shows that the percent identities, in amino acid sequences, of Tfu1114 to RL5, YfiH, BT4389, and 1T8H are 26, 21.5 32.6, and 29.8 %, respectively (Fig. 1). 1T8H is an uncharacterized protein from Bacillus stearothermophilus, whose structure was determined by Minasov et al. (2011), available from the RCSB PDB website (http://www.rcsb.org/pdb/home/home.do).
Multiple sequence alignment of some members of the protein family DUF152. The sequences were aligned using the ClustalW program. Tfu1114, a copper oxidase from T. fusca NTU10-1 (this work); RL5, laccase from rumen metagenome (GenBank accession number CAK32504); YfiH, a polyphenol oxidase from E. coli (GenBank accession number AAG57706); BT4389, a polyphenol oxidase from B. thetaiotaomicron (GenBank accession number NP_81330); 1T8H, an uncharacterized protein from B. stearothermophilus. The residues functionally conserved are shown in bold. The assigned residues belonging to the three copper sites in RL5 (Beloqui et al. 2006) are boxed by dashed lines, dash-dotted lines, and solid lines, respectively. The cysteine residues coordinate a zinc atom in 1T8H are boxed by solid lines
Characterization of the oxidation activity of Tfu1114
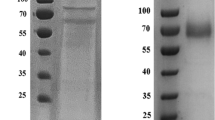
Tfu_1114 was inserted into plasmid pETDuet-1, and the resulting construct was introduced into E. coli BL21(DE3) to produce the recombinant Tfu1114 with a histidine tag fused at the N terminus. Induction with 1 mM IPTG and 0.5 mM copper sulfate at an OD600 of 0.8 was found to be optimal for the protein expression. The recombinant Tfu1114 in the cell extract was purified by immobilized metal affinity chromatography (Fig. 2), followed by dialysis to remove excess copper.
Purification of the recombinant Tfu1114. The E. coli cells were disrupted and the recombinant protein in the crude extract was purified by immobilized metal affinity chromatography as described in the “Materials and methods.” M denotes marker proteins with their molecular masses (kDa) indicated. Lanes 1 and 2 are the crude cell extract and the purified Tfu1114, respectively
The oxidation activity of the recombinant Tfu1114 toward a variety of laccase substrates was assayed at 50 °C in buffer solutions pH 7.0 and pH 8.0 (Fig. 3). The enzyme exhibited the highest activity toward 2,6-DMP at pH 8.0. Veratryl alcohol was the second substrate to be oxidized, followed by guaiacol. The oxidation of ABTS was below the discernible level. Furthermore, the activity for ABTS oxidation was assayed in 20 mM citrate buffer, pH 5, and phosphate buffer, pH 6. The activity was still not detectable. The oxidation rate of 2,6-DMP was determined over a concentration range of 0.5–6 mM at 50 °C, pH 8.0. Based on the dependence of the rate on 2,6-DMP concentration (data not shown), the values of K m and k cat were calculated to be 1 mM and 5 s−1, respectively. To assure Tfu1114 is an authentic copper-containing oxidase, the content of copper in the enzyme was assayed by atomic absorbance spectrometry, and the measurement indicated that there is one copper per Tfu1114 monomer. The UV–visible spectrum of Tfu1114 did not show a peak of absorption at around 608 nm (data not shown), which is typical for the type I Cu(II) and responsible for the deep blue color of laccases (Reinhammar and Malmström 1981).
Substrate specificity of Tfu1114. The reaction was performed at 50 °C for 15 min in 20 mM phosphate buffer that also contained 2 mM substrate as indicated. The oxidation rate was calculated based on the extinction coefficient of the oxidized product of each substrate as described in the “Materials and methods”. These data are representative of three replicate experiments
The optimal working temperature and pH of Tfu1114 were examined. The enzyme was incubated at 50–90 °C for different time periods. The residual activity for 2,6-DMP oxidation was then determined at 50 °C, pH 8.0 (Fig. 4a). Incubation at 50 or 60 °C for up to 4 h did not cause a significant drop in the activity. Inactivation of the enzyme at >70 °C followed the first-order kinetic. From the plot of the log of residual activity versus heating time, the half-lives of Tfu1114 at 70, 80, and 90 °C were calculated to be 15.4, 6.4, and 4.7 h, respectively. In addition, the activity for 2,6-DMP oxidation was examined at various temperatures ranging from 40 to 90 °C, pH 8.0, for 15 min (Fig. 4b). Tfu1114 exhibited the highest activity at 80 °C under the condition. The pH stability of Tfu1114 was examined by incubating the enzyme in buffer solutions pH 6.0–10.0 at 4 °C for 24 h. The residual activity after the treatment was assayed at 50 °C, pH 8.0 (Fig. 4c). Tfu1114 remained alive after 24 h incubation in buffer solutions pH 8.0–9.0, but was quite unstable outside this pH range. The activity at different pH was also examined at 50 °C for 15 min (Fig. 4d). The activity for 2,6-DMP oxidation was higher at alkaline condition. Together, these data indicated that Tfu1114 is a thermophilic and alkaliphilic polyphenol oxidase. Effects of organic solvents on Tfu1114 activity for 2,6-DMP oxidation were examined (Table 2). The activity was moderately tolerant to 10 % (v/v) low-carbon alcohols, acetone, and acetonitrile, but sensitive to dimethyl sulfoxide and formaldehyde.
Effects of temperature and pH on stability and activity of Tfu1114. a The residual activity, measured with 2,6-DMP at 50 °C, was determined after the enzyme had been incubated at 50–90 °C in 20 mM phosphate (pH 8.0) for 0–4 h. b The activity was measured at various temperatures of 40–90 °C in pH 8.0 for 15 min. c The residual activity, measured at 50 °C, pH 8.0, was determined after the enzyme had been incubated in various buffer solutions, pH 6–10, at 4 °C for 24 h. d The activity was measured in various buffer solutions, pH 6.5–9.0, at 50 °C. Tris–HCl buffer (20 mM, pH 6.0–8.0) and phosphate buffer (20 mM, pH 8.0–10.0) were used. These data are representative of three replicate experiments
Several potential laccase inhibitors were included in the reaction solution to get more understanding of Tfu1114 (Table 3). The enzyme was still active in solution containing 1 % sodium azide, a potent inhibitor for most of the studied laccases and heme-containing oxidases. By contrast, the oxidation of 2,6-DMP was very sensitive to ethylenediaminetetraacetic acid, suggesting that the enzyme-bound copper is essential for the catalytic activity. Tfu1114 was also sensitive to dithiothreitol and β-mercaptoethanol. The reducing agents might change the oxidative state of the copper, resulting in the inactivation of the enzyme, or simply re-reduced the oxidized product of 2,6-DMP. Alternatively, disulfide bonds are involved in the formation of the protein structure.
Bagasse hydrolysis
To know whether Tfu1114 is able to help the enzymatic hydrolysis of sugarcane bagasse, it was included in the bagasse digestion mixture that contained the characterized endoxylanase (product of Tfu_1213) or/and endocellulase (product of Tfu_1074) (Ghangas et al. 1989; Cheng et al. 2005; Zhang et al. 2000) (Fig. 5a). No reducing sugar was released from bagasse when endoxylanase (15 mU/ml), endocellulase (75 mU/ml), Tfu1114 (1.25 U/ml), or the combination of endoxylanase and endocellulase was administered. However, the amount of reducing sugar was significantly increased when endoxylanase, endocellulase, and Tfu1114 were applied together. A higher dose of endoxylanase (1.25 U/ml) could accumulate the reducing sugar to a discernible level. Again, addition of Tfu1114 (1.25 U/ml) increased the release of reducing sugar caused by the high dose endoxylanase. The extents of bagasse hydrolysis by different amounts of endoxylanase (15–150 mU/ml) were examined in the presence or absence of Tfu1114 (Fig. 5b). The more endoxylanase, the more reducing sugar was released, and Tfu1114 significantly enhanced the hydrolysis function of endoxylanase. Effects of the enzyme treatment on the appearance of bagasse were examined by scanning electronic microscopy (Fig. 6). Apparently, bagasse was broken into smaller shatters when it was treated simultaneously with endoxylanase and Tfu1114.
Effect of Tfu1114 on the enzymatic hydrolysis of bagasse. Ten milligram bagasse was mixed with specified amounts of enzymes in 1 ml 20 mM phosphate buffer, pH 8.0, and incubated in a shaker incubator (250 rpm) at 50 °C for 15 h. The reducing sugar released from bagasse was measured by dinitrosalicylic acid method. a Single plus symbol denotes 15 mU, 75 mU, and 1.25 U for endoxylanase, endocellulase, and Tfu1114, respectively, while double plus symbol means 1.25 U endoxylanase. b Bagasse was hydrolyzed by various amounts of endoxylanase, 15–150 mU, with (closed circle) or without (open circle) the presence of 5.5 U Tfu1114. One unit of endoxylanase or endocellulase activity was defined as 1 μmol reducing sugar released per min from xylan or carboxymethyl cellulose, respectively. The experiment was performed in triplicate
The appearance of bagasse after being treated with enzymes. Bagasse, 10 mg, was incubated at 50 °C for 15 h in 1 ml 20 mM phosphate buffer, pH 8.0, containing the indicated enzymes. a Buffer control. b 1.25 U endoxylanase. c 1.25 U Tfu1114. d 1.25 U endoxylanase plus 1.25 U Tfu1114. The treated bagasse was observed with a SEM microscope as described in the “Materials and methods”
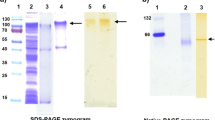
The ability of Tfu1114 to promote the enzymatic hydrolysis of bagasse prompted us to investigate whether it has a direct interaction with bagasse. This was answered by mixing Tfu1114 with bagasse for 30 min and determining the amount of the protein remained in the supernatant (Fig. 7). Tfu1114 and endoxylanase (positive control) disappeared but green fluorescent protein (negative control) remained in the supernatant, indicating the bagasse-binding ability of Tfu1114.
Discussion
In the last decades, considerable efforts have been devoted to find laccases with novel properties, develop laccase-based technologies for various industrial applications, and reduce the enzyme production costs (Rodriguez Couto and Toca Herrera 2006; Kunamneni et al. 2007). From the perspective of cost reduction, laccases of bacterial origin have advantageous properties compared to fungal laccases because they are more suitable for heterologous expression in E. coli (Piscitelli et al. 2010). In view of this, we set out to find enzymes with laccase activity in T. fusca. Groups of proteins involved in the degradation and utilization of the lignocellulosic substrate were identified in the culture medium of T. fusca in this study. Among them, a couple of oxidoreductases, including Tfu1114, were found. It was notable that such oxidoreductases do not contain potential signal peptides for protein secretion. Genome analysis suggested that T. fusca YX mainly use the Sec general secretion system and the two-arginine translocation (TAT) system for the secretion of extracellular proteins (Lykidis et al. 2007). The lack of predicted signal peptide and TAT signal on these proteins implies either the proteins were secreted through other uncharacterized systems or they were simply released into medium due to cell lysis during the cultivation. Considering that we did not observe the phenomenon of cell lysis during the culture and the abundant intracellular house-keeping enzymes were not seen in the collection, the likelihood of the former assumption should be greater than the latter. The fact that many glycoside hydrolases of T. fusca YX also lack the potential signal peptides (Lykidis et al. 2007) supports this argument.
This study demonstrated that Tfu1114 is a copper-containing enzyme able to catalyze the oxidation of phenolic and nonphenolic lignin related compounds. More importantly, it can enhance the catalytic functions of endoxylanase and endocellulase for the hydrolysis of sugarcane bagasse. Therefore, Tfu1114 may be regarded as an enzyme with laccase activity, although it does not contain four copper per monomer, a common feature of classical laccase. In fact, not all laccases hitherto studied possess four copper atoms. For example, the laccase from Phlebia radiata was reported to have two copper and one pyrroloquinoline as the prosthetic group (Karhunen et al. 1990), and a laccase from Pleurotus ostreatus contains one copper, one zinc, and two iron atoms (Palmieri et al. 1997). The stability against heat, with a half-life of 4.7 h at 90 °C, and organic solvents is another notable feature of Tfu1114. With these properties, Tfu1114 may have application potential in biopulping process and biofuel production.
The effects of a number of laccase inhibitors on Tfu1114 activity were examined. Sodium azide has been known as a potent inhibitor of laccases since early studies on the fungal enzymes (Bollg and Leonowicz 1984). Azide was thought to bind types 2 and 3 copper sites, resulting in an interruption of the internal electron transfer and activity inhibition (Ryan et al. 2003). The insensitivity suggests that the configuration of the copper binding center of Tfu1114 differs from that of classical laccases. This result is consistent with the finding that there is only one, rather than four, copper per protein monomer of Tfu1114. RL5, another member of the DUF152 family, contains four copper atoms per monomer (Beloqui et al. 2006). The residues belonging to the copper sites in RL5 were proposed according to protein modeling and site-directed mutagenesis (Beloqui et al. 2006). H73, C75, C118, and H135 coordinate the first copper, while C172, C175, C234, and C237 and N36, Y40, M68, and N114 constitute the other two potential sites, respectively. Among them, only N36, H73, C118, H135, and C234 are conserved (Fig. 1). This raises a probability that different members of the family DUF152 may have different metal binding numbers. This possibility is supported by the fact that only one zinc atom was found in the crystal structure of 1T8H. Alternatively, the relative affinities of the various binding sites vary greatly among different members of the family, and, probably, other copper atoms in Tfu1114 with weaker affinities were removed during the dialysis step of the protein purification. Future studies on Tfu1114 by site-directed mutagenesis and crystallography will be necessary to clear the uncertainty. In addition, questions such as how Tfu1114 interacts with bagasse and oxidizes the phenolic compounds in atomic detail deserve a thorough investigation. It is also interesting to know whether RL5 is able to enhance the enzymatic hydrolysis of bagasse by glycoside hydrolases.
T. fusca does not contain genes encoding classical laccase, lignin peroxidase, and manganese-dependent peroxidase for the destruction of lignin according to the genome analysis. However, it secretes Tfu1114 instead to help cellulolytic enzymes to hydrolyze lignocellulosic materials. Presumably, the oxidation of phenolic subunits of lignin by Tfu1114 may create breaking points in the network structure of lignin, consequently rendering cellulosic fibers within more exposed to glycoside hydrolases. Nonetheless, more uncharacterized oxidoreductases may be produced by T. fusca, and their concerted actions are required to make a sufficient breakdown of lignin.
References
Baldrian P (2006) Fungal laccases—occurrence and properties. FEMS Microbiol Rev 30:215–242
Beloqui A, Pita M, Polaina J, Martinez-Arias A, Golyshina OV, Zumarraga M, Yakimov MM, Garcia-Arellano H, Alcalde M, Fernandez VM, Elborough K, Andreu JM, Ballesteros A, Plou FJ, Timmis KN, Ferrer M, Golyshin PN (2006) Novel polyphenol oxidase mined from a metagenome expression library of bovine rumen: biochemical properties, structural analysis, and phylogenetic relationships. J Biol Chem 281:22933–22942
Bollag JM, Leonowicz A (1984) Comparative studies of extracellular fungal laccases. Appl Environ Microbiol 48:849–854
Cheng YF, Yang CH, Liu WH (2005) Cloning and expression of Thermobifida xylanase gene in the methylotrophic yeast Pichia pastoris. Enzyme Microb Technol 37:541–546
Ducros V, Brzozowski AM, Wilson KS, Brown SH, Østergaard P, Schneider P, Yaver DS, Pedersen AH, Davies GJ (1998) Crystal structure of the type-2 Cu depleted laccase from Coprinus cinereus at 2.2 Å resolution. Nat Struct Biol 5:310–316
Durán N, Rosa MA, D’Annibale A, Gianfreda L (2002) Applications of laccases and tyrosinases (phenoloxidases) immobilized on different supports: a review. Enzyme Microb Technol 31:907–931
Enguita FJ, Martins LO, Henriques AO, Carrondo MA (2003) Crystal structure of a bacterial endospore coat component. A laccase with enhanced thermostability properties. J Biol Chem 278:19416–19425
Ghangas GS, Hu YJ, Wilson DB (1989) Cloning of a Thermomonospora fusca xylanase gene and its expression in Escherichia coli and Streptomyces lividans. J Bacteriol 171:2963–2969
Giardina P, Faraco V, Pezzella C, Piscitelli A, Vanhulle S, Sannia G (2010) Laccases: a never-ending story. Cell Mol Life Sci 67:369–385
Hakulinen N, Kiiskinen LL, Kruus K, Saloheimo M, Paananen A, Koivula A, Rouvinen J (2002) Crystal structure of a laccase from Melanocarpus albomyces with an intact trinuclear copper site. Nat Struct Biol 9:601–605
Hullo MF, Moszer I, Danchin A, Martin-Verstraete I (2001) CotA of Bacillus subtilis is a copper-dependent laccase. J Bacteriol 183:5426–5430
Karhunen E, Niku-Paavola ML, Viikari L, Haltia T, van der Meer RA, Duine JA (1990) A novel combination of prosthetic groups in a fungal laccase; PQQ and two copper atoms. FEBS Lett 267:6–8
Kües U, Rühl M (2011) Multiple multi-copper oxidase gene families in basidiomycetes—what for? Curr Genomics 12:72–94
Kunamneni A, Ballesteros A, Plou FJ, Alcalde M (2007) Fungal laccase—a versatile enzyme for biotechnological applications. In: Méndez-Vilas A (ed) Communicating current research and educational topics and trends in applied microbiology, vol 1, Formatex Research Center. Badajoz, Spain, pp 233–245
Lykidis A, Mavromatis K, Ivanova N, Anderson I, Land M, DiBartolo G, Martinez M, Lapidus A, Lucas S, Copeland A, Richardson P, Wilson DB, Kyrpides N (2007) Genome sequence and analysis of the soil cellulolytic actinomycete Thermobifida fusca YX. J Bacteriol 189:2477–2486
Madhavi V, Lele SS (2009) Laccases: properties and applications. Biol Res 4:1684–1717
Maki M, Leung KT, Qin W (2009) The prospects of cellulase-producing bacteria for the bioconversion of lignocellulosic biomass. Int J Biol Sci 5:500–516
Minasov G, Shuvalova L, Mondragon A, Taneja B, Moy SF, Collart F, Anderson WF (2011) 1.8 Å crystal structure of an uncharacterized B. steaarothermophilus protein. RCSB PDB, Protein Data Bank. doi:10.2210/pdb1t8h/pdb
Moreira MT, Feijoo G, Sierra-Alvarez R, Lema J, Field JA (1997) Biobleaching of oxygen delignified kraft pulp by several white rot fungal strains. J Biotechnol 53:237–251
Mussatto SI, Dragone G, Guimaraes PM, Silva JP, Carneiro LM, Roberto IC, Vicente A, Domingues L, Teixeira JA (2010) Technological trends, global market, and challenges of bio-ethanol production. Biotechnol Adv 28:817–830
Palmieri G, Giardina P, Bianco C, Scaloni A, Capasso A, Sannia G (1997) A novel white laccase from Pleurotus ostreatus. J Biol Chem 272:31301–31307
Piontek K, Antorini M, Choinowski T (2002) Crystal structure of a laccase from the fungus Trametes versicolor at 1.90- Å resolution containing a full complement of coppers. J Biol Chem 277:37663–37669
Piscitelli A, Pezzella C, Giardina P, Faraco V, Giovanni S (2010) Heterologous laccase production and its role in industrial applications. Bioengineered Bugs 1:252–262
Reinhammar B, Malmström BG (1981) “Blue”-copper-containing oxidases. In: Spiro TG (ed) Copper proteins. Wiley-Interscience, New York, pp 109–149
Rodriguez Couto S, Toca Herrera JL (2006) Industrial and biotechnological applications of laccases: a review. Biotechnol Adv 24:500–513
Ryan S, Schnitzhofer W, Tzanov T, Cavaco-Paulo A, Gübitz GM (2003) An acid-stable laccase from Sclerotium rolfsii with potential for wool dye decolourization. Enzyme Microb Technol 33:766–774
Solomon EI, Sundaram UM, Machonkin TE (1996) Multicopper oxidases and oxygenases. Chem Rev 96:2563–2606
Yang C-H, Yang S-F, Liu W-H (2007) Production of xylooligosaccharides from xylans by extracellular xylanases from Thermobifida fusca. J Agric Food Chem 55:3955–3959
Zhang S, Barr BK, Wilson DB (2000) Effects of noncatalytic residue mutations on substrate specificity and ligand binding of Thermobifida fusca endocellulase cel6A. Eur J Biochem 267:244–252
Acknowledgment
We would like to thank the National Science Council, Taiwan, for the financial support (NSC 98-2313-B-005-019-MY3).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Chen, CY., Hsieh, ZS., Cheepudom, J. et al. A 24.7-kDa copper-containing oxidase, secreted by Thermobifida fusca, significantly increasing the xylanase/cellulase-catalyzed hydrolysis of sugarcane bagasse. Appl Microbiol Biotechnol 97, 8977–8986 (2013). https://doi.org/10.1007/s00253-013-4727-y
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-013-4727-y