Abstract
NADP+-dependent aminoalcohol dehydrogenase (AADH) of Rhodococcus erythropolis MAK154 catalyzes the reduction of (S)-1-phenyl-1-keto-2-methylaminopropane ((S)-MAK) to d-pseudoephedrine, which is used as a pharmaceutical. AADH is suggested to participate in aminoalcohol or aminoketone metabolism in this organism because it is induced by the addition of several aminoalcohols, such as 1-amino-2-propanol. Genetic analysis of around the aadh gene showed that some open reading frames (ORFs) are involved in this metabolic pathway. Four of these ORFs might form a carboxysome-like polyhedral organelle, and others are predicted to encode aminotransferase, aldehyde dehydrogenase, phosphotransferase, and regulator protein. OrfE, a homologous ORF of the FadR subfamily of GntR transcriptional regulators, lies downstream from aadh. To investigate whether or not orfE plays a role in the regulation of aadh expression, the gene disruption mutant of R. erythropolis MAK154 was constructed. The ΔorfE strain showed higher AADH activity than wild-type strain. In addition, a transformed strain, which harbored multi-orfE, showed no AADH activity even in the induced condition with 1-amino-2-propanol. These results suggest that OrfE is a negative regulator that represses aadh expression in the absence of 1-amino-2-propanol.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
NAD(P)+-dependent l-1-amino-2-propanol dehydrogenase (EC 1.1.1.75) is found in several microorganisms, such as Escherichia coli, Pseudomonas sp., and Bacillus subtilis (Turner 1967; Turner and Willetts 1967; Pickard et al. 1968; Willetts and Turner 1971; Faulkner and Turner 1974). These enzymes might be involved in the metabolism of aminoacetone, which is formed from threonine in the case of B. subtilis (Willetts and Turner 1971). On the other hand, NAD+-dependent d-1-amino-2-propanol dehydrogenase, which is also involved in threonine metabolism via aminoacetone, was found in E. coli cells (Dekker and Swain 1968; Lowe and Turner 1970; Campbell and Dekker 1973; Campbell et al. 1978; Kelley and Dekker 1984, 1985). These enzymes might play important roles in the metabolism of 1-amino-2-propanol, aminoacetone, or threonine, but biochemical and enzymological analyses of their enzyme properties have been insufficient, especially for l-1-amino-2-propanol dehydrogenase.
We have recently discovered NADP+-dependent l-1-amino-2-propanol dehydrogenase activity in Rhodococcus erythropolis MAK154 and cloned the gene encoding this enzyme (hereafter referred to as aminoalcohol dehydrogenase (AADH)) (Kataoka et al. 2006, 2008). AADH also catalyzes the reduction of (S)-1-phenyl-1-keto-2-methylaminopropane ((S)-MAK) to d-pseudoephedrine with an optical purity of >99% e.e., accompanied by stereo-position recognition of the methylamino group (α-carbon) of (S)-MAK in racemic MAK (Kataoka et al. 2006). Many enzymes have been found to catalyze the asymmetric reduction of prochiral carbonyl compounds, but AADH exhibits a unique stereospecificity: the formation of a double chiral compound. Because of its unique and high stereoselectivity, AADH might have strong potential for practical application to chiral aminoalcohol production.
This enzyme activity was induced by the addition of any of several aminoalcohols, such as 1-amino-2-propanol, during the growth of R. erythropolis MAK154. However, the expression level of this enzyme was quite low even in induced condition. Therefore, mutant strains showing higher AADH activity are needed when this microorganism is used as a biocatalyst of d-pseudoephedrine production. In R. erythropolis PR4, the whole genomic sequence has been determined (GenBank accession no. AP008957), and no gene showed high homology with aadh. Therefore, AADH seemed to be a unique enzyme in R. erythropolis MAK154, and it is interesting what role AADH plays in this bacterium. Here, we report the genetic analysis of the gene around aadh and the construction of a mutant R. erythropolis MAK154 strain that has higher AADH activity with the disruption of the putative regulator gene.
Materials and methods
Chemicals and reagents
(RS)-1-Phenyl-1-keto-2-ethylaminopropane (EAM) and 2-ethylamino-1-phenyl-1-propanol (EPE) were donated from Daiichi Fine Chemical (Toyama, Japan). Glucose dehydrogenase was purchased from Amano Enzyme (Aichi, Japan). Rhodococcus–Escherichia shuttle vector pK4 (Hashimoto et al. 1992) was kindly gifted from Mitsubishi Rayon (Tokyo, Japan). All restriction enzymes and DNA modification enzymes were purchased from Takara-Bio (Shiga, Japan). All other chemicals used in this study were of analytical grade, commercially available, and used without further purification.
Bacterial strains and culture conditions
R. erythropolis MAK154 (FERM BP-7451; International Patent Organism Depositary, National Institute of Advanced Industrial Science and Technology, Japan) was cultured at 28°C in medium (pH 7.0) containing 10 mg/ml glucose, 15 mg/ml peptone, 3 mg/ml KH2PO4, 2 mg/ml NaCl, 1 mg/ml yeast extract, and 0.2 mg/ml MgSO4·7H2O. dl-1-Amino-2-propanol and/or kanamycin was added into the medium for final concentrations of 0.1% (v/v) and/or 75 μg/ml, respectively. E. coli JM109 was employed as a regular cloning host. E. coli XL1-Blue MRF' was used as a host for propagating cosmid clones. All E. coli transformants were grown in Luria–Bertani (LB) medium (10 mg/ml tryptone, 5 mg/ml yeast extract and 10 mg/ml NaCl). Antibiotics were included as appropriate at the following concentrations: ampicillin, 50 μg/ml; kanamycin 25 μg/ml.
General DNA technique and DNA sequence analysis
All basic DNA procedures, such as genomic or plasmid DNA preparation, restriction enzyme digestion, ligation of DNA, and transformation of E. coli, were followed as standard protocols (Sambrook and Russell 2001). PCR was performed using a PTC-200 thermal cycler (MJ Research, Watertown, MA, USA) and Ex-Taq DNA polymerase (Takara-Bio). The nucleotide sequence was determined by the dideoxy chain termination method (Sanger et al. 1977) using a CEQ dye terminator cycle sequencing kit (Beckman Coulter, Fullerton, CA, USA) with an automated sequencer DNA analysis system, CEQ2000XL (Beckman Coulter).
Construction and screening of genomic library
The total DNA of R. erythropolis MAK154 was partially digested with MboI and ligated with BamHI/XbaI-treated SuperCos I (Stratagene, La Jolla, CA, USA). The in vitro packaging process was carried out using a Gigapack III XL packaging extract (Stratagene). The packaging phages were transfected into E. coli XL1-Blue MRF'. A library of cosmid clones was screened by the colony hybridization method using the AlkPhos Direct Labeling and Detection System (GE Healthcare Bio-Sciences, Buckinghamshire, UK), and a full-length aadh gene was used as a hybridization probe, which was amplified with PCR using primers AADH-N (5′-GAATTCATGTTCAACTCCATTGAAGGTCGTT-3′) and AADH-C (5′-CCCGGGTTACAGTTCGCCGAGCGC-3′). A cosmid vector obtained from a positive clone was named pMAK4000 and used for further analysis.
Construction of disruption mutant of orfE
A 3-kb SacII-digested fragment that was derived from pMAK4000 and that contained partial aadh and orfF and the entire orfE was blunt-ended and then ligated with SmaI-digested pBluescript II SK+ (Stratagene). The resulting plasmid pBMAK300 was digested with SmaI. A larger fragment, from which most of orfE had been removed, was recovered. This fragment was then ligated with an EcoRV-digested PCR fragment containing the kanamycin resistance gene (Km r) obtained by PCR using pK4 as template and primers Km-f (5′-GGGATATCTAGGCGGTGCTACAGAGTTC-3′: EcoRV site was underlined) and Km-r (5′-GGGATATCCAGCTGGCGTAATAGCGAAG-3′: EcoRV site was underlined). The resulting plasmid, pBMAK360, was used for the construction of an orfE disruption mutant (see Fig. 2).
The plasmid pBMAK360 was introduced into R. erythropolis MAK154 by electroporation, as follows. A mid-exponential culture of R. erythropolis MAK154 cells was harvested and washed with cold demineralized water. Cells were then concentrated 20-fold in cold demineralized water (compared with culture volume) and kept on ice. Ice-cold cells (200 μl) were mixed with 200 ng of pBMAK360 in TE buffer (10 mM Tris–HCl and 1 mM EDTA, pH 8.0) in a 2-mm gapped electrocuvette, and subjected to a 2.5-kV electric pulse from a Gene Pulser (Bio-Rad Laboratories, Hercules, CA, USA) connected to a pulse controller (25 μF capacitor; external resistance, 400 Ω). Pulsed cells were diluted immediately with 1 ml of MYP medium (Hashimoto et al. 1992) and incubated for 2 h at 28°C. They were then spread on LB medium containing 75 μg/ml kanamycin. Kanamycin resistance clones were cultured, and total the DNA was isolated. The homologous recombination was checked with PCR using primers orfE-N (5′-GGGATATCTATGGGTGCGCAAACTGAGG-3′) and orfE-C (5′-GGGATATCTCAGGATTCGGAGTCGGTGC-3′), with which the entire region of orfE was amplified.
Construction of strain bearing multi-copy numbers of orfE
A 2.2-kb EcoRV/ScaI fragment of pBMAK300, which contained 70-bp upstream to 1,200-bp downstream of orfE (Fig. 2) was ligated with FspI-digested Rhodococcus–Escherichia shuttle vector pK4. R. erythropolis MAK154 was transformed with the resulting plasmid pKMAK309 using the electroporation method as described above.
AADH activity analysis
AADH activity was measured with a bioconversion rate of (RS)-EAM to EPE. EAM and EPE are corresponding analogs of MAK and d-pseudoephedrine, and EAM is a good substrate for AADH. The wild-type or ΔorfE strain of R. erythropolis MAK154 was cultured in medium with or without 0.1% (v/v) 1-amino-2-propanol for 24 h. Cells were harvested, frozen at −80°C, and lyophilized. The reaction was carried out in 500 μl of reaction mixture comprised of 2 mg/ml each of lyophilized cells, 100 mM sodium phosphate (pH 8.0), 0.6 mg/ml NADP+, 125 mM glucose, 20 mg/ml (RS)-EAM, and 15 units/ml glucose dehydrogenase. Incubation was performed at 30°C with shaking for 24 h. After centrifugation, the resulting supernatant was analyzed by HPLC on an Inertsil Ph-3 column (4.6 × 75 mm; GL Science, Tokyo, Japan) at 40°C. Then, 3% (v/v) acetonitrile containing 50 mM NaH2PO4 was used as a mobile phase, and the flow rate was 1.2 ml/min. The absorbance of the eluent was monitored at 220 nm.
RT-PCR
After the wild-type or ΔorfE strain of R. erythropolis MAK154 was cultured, 1-amino-2-propanol was added for a final concentration of 0.1% (v/v) when OD660 reached 0.4, and the cells were further cultivated for 2 h. Total RNA was extracted from these cells using RNeasy Mini Kit (Qiagen, Valencia, CA, USA). The relative amount of aadh RNA was quantified with real-time PCR using the LightCycler Quick System 350S (Roche Diagnostics, Mannheim, Germany), LightCycler RNA Amplification Kit SYBR Green I (Roche Diagnostics) and primers aadh-f (5′-GAGGACATGGACAGCG-3′) and aadh-r (5′-GCAAAGAACAGTGCGG-3′). mRNA of the glyceraldehyde-3-phosphate dehydrogenase gene was used as an internal standard amplified with primers gapdh-f (5′-CTGCCCTCAACATCGT-3′) and gapdh-r (5′-GTTGGAGTAGCCCCAC-3′).
Results
Sequence analysis of the gene around aadh
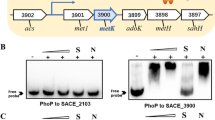
A genomic cosmid library of R. erythropolis MAK154 was constructed in SuperCos I. About 2,000 cosmid clones were screened using the colony hybridization method. With the resulting positive clone of pMAK4000, the nucleotide sequence of an approximately 14-kb DNA region around the aadh gene was determined (GenBank accession no. AB573180). Overall, the ORFs were identified using the BLAST program. The computer-aided analysis of the 14-kb sequence revealed the presence of 14 complete ORFs (Fig. 1). Features of the sequence and selected alignments with other genes are listed in Table 1. Recently, whole or draft genomic sequences of some Mycobacterium strains, i.e., Mycobacterium sp. MCS, Mycobacterium sp. KMS, Mycobacterium sp. JLS, Mycobacterium flavenscens PYR-GCK, and Mycobacterium vanbaalenii PYR-1, each of which can degrade pyridine or polycyclic aromatic hydrocarbon, were determined at the US DOE Joint Genome Institute. In most of these microorganisms, the homologous genes of 11 ORFs (orfV, -W, -X, -Y, -Z, -A, -B, -C, -D, -E, and aadh) exist in a cluster and in the same order with R. erythropolis MAK154 (gene ID Mmcs_0199 to 0189, in the case of Mycobacterium sp. MCS, Fig. 1). In R. erythropolis PR4, there are no ORFs showing significant identities (over 40% in amino acid identity) with these ORFs.
ORF map of the 14-kb region around aadh isolated from R. erythropolis MAK154 and its homologous region of Mycobacterium sp. MCS. Open arrows indicate the ORFs and their direction of transcription. The aadh and orfE are shown in solid gray. The similarities of deduced amino acids from each corresponding ORF are also shown in figure. NotI, SacII, and SmaI restriction enzyme sites are shown with No, Sa, and Sm, respectively. The putative OrfE binding sequences are shown with the text. Arrows under the text indicate the palindromic regions of the OrfE binding sequences
Construction of orfE disruption mutant
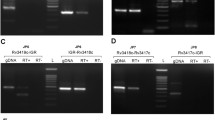
OrfE showed homology with many GntR family transcriptional regulator proteins of several microorganisms. Because regulator proteins of prokaryotic cells are frequently located adjacent to genes under their control, we hypothesized that OrfE had some relationship with the expression of aadh. Thus, the orfE disruption mutant of R. erythropolis MAK154 (ΔorfE strain) was constructed as described in “Materials and methods” (Fig. 2). The gene disruption of mutant R. erythropolis MAK154 was confirmed with PCR analysis that amplified the entire region of orfE (data not shown). In the ΔorfE strain, an approximately 1,700-bp fragment (derived from ΔorfE::Km r) was amplified, and no fragment of about 770 bp (derived from wild-type orfE) was detected. Southern hybridization analysis also indicated that the homologous recombination occurred in the correct position (data not shown). The ΔorfE strain showed a similar growth rate and morphology to the wild-type strain (data not shown).
Construction of orfE-disrupted mutant of R. erythropolis MAK154. Plasmid pBMAK360 was introduced into R. erythropolis MAK154 by electroporation, and gene disruption via a double-crossover homologous recombination was performed. SacII, SmaI, and EcoRV restriction enzyme sites are shown with Sac, Sma, and RV, respectively. The aadh* and orfF* show respectively the partial genes of aadh and orfF. Open triangles are PCR primers to amplify the entire region of orfE. RV and ScaI are EcoRV and ScaI restriction enzyme sites, respectively, used for construction of pKMAK309
Functional analysis of OrfE
To investigate the influence of orfE disruption, AADH activities of the wild-type and ΔorfE strains were analyzed using lyophilized cells grown with or without an inducer (1-amino-2-propanol) (Fig. 3). The non-induced wild-type cells showed little EPE-producing (AADH) activity, whereas the induced wild-type cells produced about 1 mg/ml of EPE. On the other hand, regardless of whether or not 1-amino-2-propanol was added, ΔorfE strain cells showed higher levels of AADH activity than the wild-type and produced about 7 mg/ml EPE. Non-induced ΔorfE strain cells showed almost the same AADH activity level as induced cells. The strain transformed with pK4-derived plasmid pKMAK309, a multi-copy plasmid that contained orfE, showed no AADH activity regardless of induction (Fig. 3).
AADH activities of the wild-type (WT) strain, ΔorfE strain (ΔorfE), and wild-type strain bearing pKMAK309 (WT [pKMAK309]) of R. erythropolis MAK154 under the conditions with (+) or without (−) 1-amino-2-propanol addition. AADH activities were analyzed by the bioconversion rate of (RS)-EAM to EPE using lyophilized cells. Error bars indicate standard deviation (SD) from three independent experiments
The aadh expression level of each strain was also analyzed using quantitative RT-PCR (Fig. 4). In the wild-type strain, aadh expression was about 100-fold higher when 1-amino-2-propanol was added to the medium than when it was not added. On the other hand, in the ΔorfE strain, aadh expression was not affected by the addition of 1-amino-2-propanol. Furthermore, regardless of the addition of 1-amino-2-propanol, the ΔorfE strain showed almost the same expression levels as the induced wild-type strain. These results suggested that OrfE was a negative regulator of the expression of aadh.
Relative expression level of aadh in R. erythropolis MAK154 wild-type (WT) strain and ΔorfE strain (ΔorfE) under the conditions with (+) or without (−) 1-amino-2-propanol addition. Quantitative RT-PCR was done using LightCycler. The relative amounts of mRNA against non-induced wild-type strain are shown in a semi-logarithmic graph. Error bars indicate SD from five independent experiments
Discussion
AADH of R. erythropolis MAK154 is induced with the addition of several aminoalcohols, such as 1-amino-2-propanol, during growth, and catalyzes the reversible conversion of l-1-amino-2-propanol to aminoacetone (Kataoka et al. 2006). These facts suggest that AADH might participate in the metabolic pathway of aminoalcohols or aminoketones, but little is known about the biological role of this enzyme. To elucidate the physiological role of this enzyme, genetic analysis of the region around aadh was performed.
An approximately 14-kb sequence around aadh of R. erythropolis MAK154 was determined. In this region, the existence of 14 putative complete ORFs was indicated. Most of the homologous genes in this region (from orfV to orfE) are found in several Mycobacterium strains. Considering that each non-coding region between orfW and aadh is quite short, that orfV is coded complementary strand in Mycobacterium and that OrfE is a putative regulator gene (see Fig. 1 and Table 1), it seems that at least the genes from orfW to aadh constitute an operon and form a metabolic pathway of some compound.
OrfX, OrfA, and OrfB showed homology with EutE, EutN, and EutM, respectively. EutE, EutN, and EutM together form an ethanolamine utilization (eut) operon with the other 14 eut genes in Salmonella enterica. The biochemistry underpinning ethanolamine catabolism in S. enterica is simple. The initial step in this process is catalyzed by an adenosylcobalamin (also known as a coenzyme form of vitamin B12)-dependent ethanolamine ammonia lyase encoded by the eutBC genes; this catalysis converts ethanolamine to acetaldehyde and free ammonia (Faust et al. 1990). Subsequently, acetaldehyde is converted to acetyl coenzyme A (acetyl-CoA) in a single step presumably catalyzed by the acetaldehyde dehydrogenase (EutE) enzyme (Stojiljkovic et al. 1995). Several eut genes (i.e., eutS, -M, -N, -L, and -K) encode proteins with homology to cyanobacterial carboxysomal shell proteins, and these proteins are suggested to form a carboxysome-like polyhedral organelle that is sometimes called a metabolosome (Stojiljkovic et al. 1995; Kofoid et al. 1999). The physiological role of this organelle is poorly understood, but recent studies suggest that the eut organelle might have the function of protecting cells against aldehyde toxicity; moreover, it has been suggested that these structures function to concentrate both metabolic enzymes and their substrates to allow high catalytic efficiency while maintaining CoA balance (Brinsmade et al. 2005). OrfA and OrfB showed similarities to EutN and EutM, and OrfY and OrfC contain a BMC (bacterial microcompartment) domain, which is found in the shell protein that form microcompartments such as carboxysomes or eut organelles (Kerfeld et al. 2005; Cheng et al. 2008). These observations suggest that in R. erythropolis MAK154, the metabolism of aminoalcohol or aminoketone is carried out in carboxysome-like polyhedral organelle.
In Erwina carotovora N.C.P.P.B.1280, which can grow with 1-amino-2-propanol as a sole nitrogen source, 1-amino-2-propanol is metabolized with phosphorylation and produces 1-amino-2-propanol O-phosphate (Jones and Turner 1971; Jones et al. 1973). In R. erythropolis MAK154, a similar reaction seemed to occur because OrfD showed homology with homoserine kinases and aminoglycoside phosphotransferases. In E. carotovora, 1-amino-2-propanol O-phosphate is converted to propionaldehyde and free ammonia catalyzed by aminoalcohol O-phosphate phospho-lyase (Jones and Turner 1971; Jones et al. 1973). However, no ORF seemed to catalyze such a reaction around aadh. Therefore, a different metabolic pathway might exist in R. erythropolis MAK154. OrfW and OrfX bear similarities with various transaminases (aminotransferases) and aldehyde dehydrogenases, respectively. Tramsaminases catalyze the conversion of amine to aldehyde accompanied by amino transfer to an amino acceptor, and aldehyde dehydrogenases might catalyze the exchange of aldehyde to acyl-CoA. Since the details on this metabolic pathway remain unclear, more detailed examination is required.
OrfE shows similarity with many GntR family transcriptional regulator proteins. The GntR family was first described by Haydon and Guest (1991). This protein family is defined by a 64-amino acid N-terminal DNA-binding helix-turn-helix motif and represses transcription by binding to operator sequences existing in the promoters of target genes. DNA-binding activity is modulated by an inducer molecule that binds to the C-terminal domain of the regulator protein. In general, when the inducer is present, DNA binding is inhibited, and the repression of target genes is relieved (Rigali et al. 2002).The GntR family proteins can be divided into four subfamilies based on the homology of the C-terminal effector-binding domain (Rigali et al. 2002). A sequence comparison of OrfE with several GntR family regulators showed that OrfE is a member of the FadR subfamily and is most closely related to the archetypal family member, the B. subtilis gluconate operon regulator, GntR. FadR subfamily members typically bind metabolites that are substrates of the target enzymes. For example, GntR binds gluconate, thereby relieving repression of the gluconate utilization operon (Miwa and Fujita 1988), and FadR binds fatty acyl-CoA, resulting in the derepression of genes involved in fatty acid degradation (Dirusso and Black 2004). Our results suggested that OrfE was a negative regulator of the expression of aadh. The proposed expression mechanism of aadh is as follows: When 1-amino-2-propanol does not exist, OrfE binds to operator sequences of aadh with N-terminal helix-turn-helix motif and represses transcription of aadh. On the other hand, when 1-amino-2-propanol exists, OrfE presumably binds to 1-amino-2-propanol with C-terminal effector-binding domain, then loses DNA-binding activity and leaves operator sequence of aadh; thus, the transcription of aadh is permitted. In the ΔorfE strain, the differences of AADH activity or aadh expression level were not observed between induced and non-induced cells (Figs. 3 and 4). It might be due to the lack of transcriptional repressor of aadh (i.e., OrfE). In the wild-type strain bearing pKMAK309, AADH activity was not detected regardless of induction (Fig. 3). This recombinant strain might produce larger amount of OrfE in the cells as compared with wild-type strain; consequently, aadh transcription might be strongly repressed by OrfE. The amount of 1-amino-2-propanol added in the medium might be insufficient to take all OrfE away from operator sequence, and transcription might be restricted completely even on the induced condition. Among FadR subfamily proteins, the deduced sequence (palindromic 5′-Nx-T-N-G-T-N3-A-C-N-A-Nx-3′) is conserved in their binding sites (Rigali et al. 2002). In R. erythropolis MAK154, a putative OrfE binding sequence (5′-CTTTATTGGTCCGACCATTAATG-3′) exists about 80-bp upstream from the orfW start codon (Fig. 1). The R. erythropolis MAK154 mutant showing high AADH activity was isolated using a traditional mutagenesis method (data not shown). The genomic region around aadh in this mutant strain was also sequenced. In this region, two positions of base substitution were found, one in orfD and the other in orfE, and both base substitutions caused amino acid substitutions. Because the amino acid substitution in OrfE occurred in the N-terminal helix-turn-helix motif, this mutant OrfE might lose DNA-binding ability. This result strongly suggests that OrfE is the negative regulator of aadh.
The disruption mutant of orfE shows higher AADH activity than that of the wild-type strain. AADH might have high potential for practical application to chiral aminoalcohol production by virtue of its unique and high stereoselectivity (Kataoka et al. 2006, 2008). We expect that the mutant obtained in this study will be useful in the biocatalytic process of chiral aminoalcohols and that the obtained insights will be helpful in understanding the aminoalcohol metabolic pathway.
References
Brinsmade SR, Paldon T, Escalante-Semerena JC (2005) Minimal functions and physiological conditions required for growth of Salmonella enterica on ethanolamine in the absence of the metabolosome. J Bacteriol 187:8039–8046
Campbell RL, Dekker EE (1973) Formation of D-1-amino-2-propanol from L-threonine by enzymes from Escherichia coli K-12. Biochem Biophys Res Commun 53:432–438
Campbell RL, Swain RR, Dekker EE (1978) Purification, separation, and characterization of two molecular forms of D-1-amino-2-propanol:NAD+ oxidoreductase activity from extracts of Escherichia coli K-12. J Biol Chem 253:7282–7288
Cheng S, Liu Y, Crowley CS, Yeates TO, Bobik TA (2008) Bacterial microcompartments: their properties and paradoxes. BioEssays 30:1084–1095
Dekker EE, Swain RR (1968) Formation of Dg-1-amino-2-propanol by a highly purified enzyme from Escherichia coli. Biochim Biophys Acta 158:306–307
Dirusso CC, Black PN (2004) Bacterial long chain fatty acid transport: gateway to a fatty acid-responsive signaling system. J Biol Chem 279:49563–49566
Faulkner A, Turner JM (1974) Microbial metabolism of amino alcohols. Aminoacetone metabolism via 1-aminopropan-2-ol in Pseudomonas sp. N.C.I.B. 8858. Biochem J 138:263–276
Faust LR, Connor JA, Roof DM, Hoch JA, Babior BM (1990) Cloning, sequencing, and expression of the genes encoding the adenosylcobalamin-dependent ethanolamine ammonia-lyase of Salmonella typhimurium. J Biol Chem 265:12462–12466
Hashimoto Y, Nishiyama M, Yu F, Watanabe I, Horinouchi S, Beppu T (1992) Development of a host-vector system in a Rhodococcus strain and its use for expression of the cloned nitrile hydratase gene cluster. J Gen Microbiol 138:1003–1010
Haydon DJ, Guest JR (1991) A new family of bacterial regulatory proteins. FEMS Microbiol Lett 79:291–296
Jones A, Turner JM (1971) Microbial metabolism of amino alcohols via aldehydes. J Gen Microbiol 67:379–381
Jones A, Faulkner A, Turner JM (1973) Microbial metabolism of amino alcohols. Metabolism of ethanolamine and 1-aminopropan-2-ol in species of Erwinia and the roles of amino alcohol kinase and amino alcohol O-phosphate phospho-lyase in aldehyde formation. Biochem J 134:959–968
Kataoka M, Nakamura Y, Urano N, Ishige T, Shi G, Kita S, Sakamoto K, Shimizu S (2006) A novel NADP+-dependent L-1-amino-2-propanol dehydrogenase from Rhodococcus erythropolis MAK154: a promising enzyme for the production of double chiral aminoalcohols. Lett Appl Microbiol 43:430–435
Kataoka M, Ishige T, Urano N, Nakamura Y, Sakuradani E, Fukui S, Kita S, Sakamoto K, Shimizu S (2008) Cloning and expression of the L-1-amino-2-propanol dehydrogenase gene from Rhodococcus erythropolis, and its application to double chiral compound production. Appl Microbiol Biotechnol 80:597–604
Kelley JJ, Dekker EE (1984) D-1-amino-2-propanol: NAD+ oxidoreductase. Purification and general properties of the large molecular form of the enzyme from Escherichia coli K-12. J Biol Chem 259:2124–2129
Kelley JJ, Dekker EE (1985) Identity of Escherichia coli D-1-amino-2-propanol:NAD+ oxidoreductase with E. coli glycerol dehydrogenase but not with Neisseria gonorrhoeae 1, 2-propanediol:NAD+ oxidoreductase. J Bacteriol 162:170–175
Kerfeld CA, Sawaya MR, Tanaka S, Nguyen CV, Phillips M, Beeby M, Yeates TO (2005) Protein structures forming the shell of primitive bacterial organelles. Science 309:936–938
Kofoid E, Rappleye C, Stojiljkovic I, Roth J (1999) The 17-gene ethanolamine (eut) operon of Salmonella typhimurium encodes five homologues of carboxysome shell proteins. J Bacteriol 181:5317–5329
Lowe DA, Turner JM (1970) Microbial metabolism of amino ketones: D-1-aminopropan-2-ol and aminoacetone metabolism in Escherichia coli. J Gen Microbiol 63:49–61
Miwa Y, Fujita Y (1988) Purification and characterization of a repressor for the Bacillus subtilis gnt operon. J Biol Chem 263:13252–13257
Pickard MA, Higgins IJ, Turner JM (1968) Purification and properties of L-1-aminopropan-2-ol: NAD oxidoreductase from a pseudomonad grown on DL-1-aminopropan-2-ol. J Gen Microbiol 54:115–126
Rigali S, Derouaux A, Giannotta F, Dusart J (2002) Subdivision of the helix-turn-helix GntR family of bacterial regulators in the FadR, HutC, MocR, and YtrA subfamilies. J Biol Chem 277:12507–12515
Sambrook J, Russell DW (2001) Molecular cloning: a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, New York
Sanger F, Nicklen S, Coulson AR (1977) DNA sequencing with chain-terminating inhibitors. Proc Natl Acad Sci U S A 74:5463–5467
Stojiljkovic I, Baumler AJ, Heffron F (1995) Ethanolamine utilization in Salmonella typhimurium: nucleotide sequence, protein expression, and mutational analysis of the cchA cchB eutE eutJ eutG eutH gene cluster. J Bacteriol 177:1357–1366
Turner JM (1967) Microbial metabolism of amino ketones. L-1-aminopropan-2-ol dehydrogenase and L-threonine dehydrogenase in Escherichia coli. Biochem J 104:112–121
Turner JM, Willetts AJ (1967) Amino ketone formation and aminopropanol-dehydrogenase activity in rat-liver preparations. Biochem J 102:511–519
Willetts AJ, Turner JM (1971) Threonine metabolism in a strain of Bacillus subtilis enzymic oxidation of 1-aminopropan-2-ol and aminoacetone. Biochim Biophys Acta 252:98–104
Acknowledgements
This work was supported in part by a Grant-in-Aid for Scientific Research, No. 20380051 (to MK), from the Japan Society for the Promotion of Science (JSPS), and by the Targeted Proteins Research Program (TPRP) of the Ministry of Education, Culture, Sports, Science and Technology (MEXT), Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Urano, N., Kataoka, M., Ishige, T. et al. Genetic analysis around aminoalcohol dehydrogenase gene of Rhodococcus erythropolis MAK154: a putative GntR transcription factor in transcriptional regulation. Appl Microbiol Biotechnol 89, 739–746 (2011). https://doi.org/10.1007/s00253-010-2924-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-010-2924-5