Abstract
Host-specific Bacteroides–Prevotella 16S rRNA genetic markers are promising alternative indicators for identifying the sources of fecal pollution because of their high abundance in the feces of warm-blooded animals and high host specificity. However, little is known about the persistence of these genetic markers in environments after being released into environmental waters. The persistence of feces-derived four different host-specific Bacteroides–Prevotella 16S rRNA genetic makers (total, human-, cow-, and pig-specific) in environmental waters was therefore investigated at different incubation temperatures (4, 10, 20, and 30°C) and salinities (0, 10, 20, and 30 ppt) and then compared with the survival of conventional fecal-indicator organisms. The host-specific genetic markers were monitored by using real-time polymerase chain reaction (PCR) assays with specific primer sets. Each host-specific genetic marker showed similar responses in non-filtered river water and seawater: They persisted longer at lower temperatures and higher salinities. In addition, these markers did not increase in all conditions tested. Decay rates for indicator organisms were lower than those for host-specific genetic markers at temperature above 10°C. Furthermore, we investigated whether the PCR-detectable 16S rRNA genetic markers reflect the presence of live target cells or dead target cells in environmental waters. The result revealed that the detection of the Bacteroides–Prevotella 16S rRNA genetic markers in environmental waters mainly reflected the presence of ‘viable but non-culturable’ Bacteroides–Prevotella cells. These findings indicate that seasonal and geographical variations in persistence of these host-specific Bacteroides–Prevotella 16S rRNA genetic markers must be considered when we use them as alternative fecal indicators in environmental waters.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Contamination with animal and human feces limits the use of surface waters for recreation, agriculture, and aquiculture. The potential sources of fecal contaminations are numerous and can be classified into point sources (e.g., wastewater treatment plants, sewer overflows, slaughterhouses, and animal feedlots) and non-point sources (e.g., wildlife, septic systems, livestock, land application of manures, landfills, and pastures) (Kim et al. 2005). Regardless of efforts to reduce or eliminate fecal pollution, a large percentage of watersheds in Japan continue to be impaired mainly due to failure to control and identify non-point fecal pollution.
The most widely used method for measuring fecal pollution is to count viable fecal-indicator bacteria [e.g., total coliform (TC), fecal coliforms (FC), fecal enterococci and Escherichia coli]. Despite widespread implementation of fecal-indicator bacteria monitoring programs, there are numerous limitations associated with their application including detection of non-fecal bacteria that inherently exist in natural environment (Meays et al. 2004; Scott et al. 2002; Simpson et al. 2002), ability to multiply after releasing into water column (Byappanahalli and Fujioka 2004; Desmarais et al. 2002; Solo-Gabriele et al. 2000), inability to identify the source of fecal contamination (Field et al. 2003). Therefore, it is extremely important to develop a reliable method for promptly identifying and quantifying the sources of non-point fecal contaminations, which does not rely on culturing fecal-indicator bacteria.
For this reason, several researchers have proposed that members of the genera Bacteroides and Prevotella (Bacteroides–Prevotella) can be used as alternative fecal-indicator organisms (Allsop and Stickler 1985; Fiksdal et al. 1985). Members of these genera are strict anaerobes and restricted to warm-blooded animals. They are also present at high abundance in the gastrointestinal tract and have host-specific distributions (Bernhard and Field 2000; Simpson et al. 2004). These notable ecophysiological features of the genera Bacteroides–Prevotella highlight that the detection of these organisms is more probable than fecal coliforms and more specific to host animals (Bower et al. 2005) because they have little potential for growth or persist in natural environment (Kreader 1998).
For the detection of these anaerobic organisms, molecular-based methods are generally superior to culture-dependent methods. polymerase chain reaction (PCR)-based assays were commonly used to detect host-specific 16S rRNA genetic markers of the genera Bacteroides and Prevotella (Bernhard and Field 2000; Carson et al. 2005; Dick et al. 2005). More recently, real-time PCR assays were successfully developed for simultaneous identification and quantification of the host-specific 16S rRNA genetic markers (Dick and Field 2004; Griffith et al. 2003; Layton et al. 2006; Okabe et al. 2007; Seurinck et al. 2005). We have also previously developed a real-time PCR assay for quantification of genetic markers for total, human-, cow-, and pig-specific Bacteroides–Prevotella 16S rRNA genes as an alternative fecal indicator and evaluated their host-specificity and sensitivity (Okabe et al. 2007).
For practical applications, it is essential to understand the temporal (seasonal) and geographical stability of Bacteoides–Prevotella 16S rRNA genetic markers in environmental waters. We want to test the hypothesis that different host-specific species or genetic markers (e.g., total, human-, pig-, and cow-specific markers) may have different survival abilities in different environmental waters. If decay rates of each genetic marker under different water conditions were understood, we would be able to estimate the extent and contribution of each fecal source from field measurements. However, little is known about the persistence of PCR-detectable Bacteroides–Prevotella 16Sr RNA genetic markers in different environmental waters.
The goal of this study was therefore to investigate the persistence of feces-derived host-specific Bacteroides–Prevotella 16S rRNA genetic markers, which were developed previously in our laboratory, in river and seawater at different temperatures and salinities. The behaviors of the genetic markers were compared with those of the conventional fecal-indicator organisms. Furthermore, we investigated whether the PCR-detectable 16S rRNA genetic markers during the aerobic incubation in environmental waters were originated from live target cells or dead target cells because PCR assay can detect 16S rRNA genes in live cells, severely stressed cells, and even dead cells (Josephson et al. 1993).
Materials and methods
Samples collections
River water samples were collected from the Atsubetsu River, and seawater samples were collected from the Ishikari Bay in Hokkaido, Japan. Both filtered and non-filtered river water and seawater were used in following experiments. Filtered water samples were prepared by passing through 0.2-μm-pore-size Supor-200 filters (Ann Arbor, MI, USA) to remove the indigenous microorganisms and suspended particles. Human fecal samples were collected from healthy adults in Sapporo City. Fresh cow and pig feces were collected from farms in Hokkaido University and subjected to the experiments within a few hours.
Experimental design
Quantification limits of real-time PCR assays
Quantification limits of the real-time PCR assays were determined using suspended feces in non-filtered river water. The quantification limit was defined as the lowest concentration of the Bacteroides–Prevotella 16S rRNA genetic marker that is within the linear range of quantification of the real-time PCR assay in serially diluted suspensions.
Fresh human, cow, and pig feces were suspended into non-filtered river water (0.1 g total feces/100 ml). Two milliliters of the suspension (100) was concentrated to 200 μl by centrifugation at 8,000×g for 5 min (101). Tenfold dilution series (10−1–10−7) of the original suspension (100) were prepared with the non-filtered river water. Two hundred microliters of the ten times concentrated samples (101) and 100–10−2 diluted samples were used for DNA extraction. At the same time, 2 ml of 10−3 dilution and 20 ml of 10−4–10−7 dilutions were concentrated to 200 μl by centrifugation and used for DNA extraction. Real-time PCR assay was performed with a primer sets specific for total, human-, cow-, and pig-specific Bacteroides–Prevotella group (Bac-Pre1, Human-Bac1, Cow-Bac2, and Pig-Bac2), which was designed previously in our laboratory (Okabe et al. 2007). In addition, enumeration of conventional fecal coliforms (TC and FC) was conducted by membrane filter technique (APHA, AWWA, and WEF 1995) to compare the quantification limits of Bac-Pre1 marker and conventional indicator microorganisms.
Studies using pure culture of B. fragilis
As mentioned above, detection of target 16S rRNA genes with real-time PCR do not mean the presence of intact living Bacteroides cells. We therefore observed the change in numbers of Bac-Pre1 markers, culturable cells, and live cells simultaneously to elucidate the entity of PCR-detectable 16S rRNA genetic markers in aerobic environmental waters. In this experiment, a pure culture of Bacteroides fragilis strain (ATCC 25285) was used instead of fecal samples. B. fragilis strain was pre-cultured anaerobically in 50 ml of Gifu anaerobic media (GAM) broth (Nissui Pharmaceutical, Tokyo, Japan) at 37°C for 15 h. The cells were centrifuged at 8,000×g for 10 min and washed three times with sterile 0.85% w/v NaCl. The cell pellets were suspended in both non-filtered and filtered river and seawater, respectively, resulting in the final cell concentrations of 107–108 cells/ml. The original non-filtered river and seawater contained total bacteria (1.4 × 105–6.0 × 105 cells/ml) and Bac-Pre1 marker (1.2 × 103–2.8 × 104 copies/ml), which was less than 0.1% of the inoculated B. fragilis population. Thus, the natural bacterial flora could be ignored in this experiment. The suspended samples were then incubated aerobically at 10°C under the dark condition in the laboratory. Subsamples were taken at regular intervals for real-time PCR assay using Bac-Pre1 primer set, culturable cell counting, and live/dead staining up to 16 days. Agar plate cultivation and live/dead staining were conducted within 4 h after sampling. Subsamples for real-time PCR were stored at −70°C until further use. Each set of the experiment was conducted in duplicate, and the averages were reported.
Studies using fecal samples
To simulate natural environmental conditions, 1 g of each fresh human, cow, and pig feces were suspended separately in 50 ml of non-filtered river water and combined into one beaker using extra non-filtered river water for washing (3 g total feces/250 ml). The suspended fecal samples were divided equally (50 ml each) into five Erlenmeyer flasks and further diluted to 500 ml with non-filtered river water, respectively (a final concentration of 1.2 mg total feces/ml). Finally the diluted fecal sample solutions were incubated aerobically at different temperatures (4, 10, 20, and 30°C) in the dark. Subsamples (0.2–20 ml) were taken at regular intervals for real-time PCR assays and enumeration of TC and fecal coliforms FC up to 35 days. Subsamples for real-time PCR were stored at −70°C until DNA extraction. The diluted fecal solution was incubated anaerobically at 10°C and used as a control.
In addition, the effect of salinity was determined in sample solutions adjusted to the salinity of 0, 10, 20, and 30 ppt by mixing non-filtered river water and non-filtered seawater. The sample solutions with different salinity were all incubated similarly at 10°C. Each set of the experiment was also conducted in duplicate, and the averages were reported.
Analytical methods
DNA extraction
Subsamples (the maximum volume was set to 20 ml) for real-time PCR assays were concentrated to 200 μl by centrifugation at 8,000×g for 10 min. Total DNA was extracted using the QIAamp DNA Stool Mini Kit (Qiagen, Hilden, Germany) as described in the manufacturer’s instructions. The extracted DNA solution (50 μl) was stored at −20°C until real-time PCR analyses.
Real-time PCR assay
Four primer sets specific for total, human-, cow-, and pig-specific Bacteroides–Prevotella group (Bac-Pre1, Human-Bac1, Cow-Bac2, and Pig-Bac2), which were developed previously in our laboratory, were used for detection of each host-specific 16S rRNA genetic marker. Compositions of PCR mixture and conditions for each real-time PCR assay were described elsewhere (Okabe et al. 2007). According to standard curves with B. fragilis genomic DNA and plasmid DNA, the lowest detection limit was 4.4 × 101 copies per reaction for Bac-Pre1 and Human-Bac1 primer sets and 6.2 × 100 copies per reaction for Cow-Bac2 and Pig-Bac2 primer sets, respectively. Within each real-time PCR run, all samples were analyzed in triplicate.
Enumeration of total and fecal coliforms
Enumeration of TC and FC was carried out using membrane filter technique (APHA, AWWA, and WEF 1995). Samples were diluted with sterile 0.85% w/v NaCl and filtered through 0.45-μm pore-size mixed cellulose ester filters (Advantec, Tokyo, Japan), and the filters were placed on m-End or m-FC agar. Triplicate plates were then incubated at 35°C for 48 h and 44.5°C for 24 h for total and fecal coliforms, respectively. The indole, methyl red, Voges–Proskauer, and Simons’ Citrate tests were conducted for identification of suspected colonies on the agar (APHA, AWWA, and WEF 1995). Average numbers of colony forming units (CFU) per 100 ml were reported.
Enumeration of B. fragilis
Culturable B. fragilis cells were enumerated by spreading 100 μl of tenfold diluted samples with sterile 0.85% w/v NaCl on triplicate Bacteroides Bile Esculin (BBE) agar plates (Livingston et al. 1978), which were then incubated for 2 days at 37°C under anaerobic conditions in GasPak anaerobic jars (Becton Dickinson Microbiology Systems, Cockeysville, MD, USA). Average counts were reported as CFU per ml.
Live B. fragilis cells were enumerated using Live/Dead® BacLight™ Bacterial Viability Kits (Molecular Probes, Oregon, USA). After live/dead staining, the samples (1 ml) were diluted with pure water or concentrated at 5,000×g for 10 min to obtain adequate cell numbers and then filtered with a 25 mm, 0.2-μm-pore-size black nucleopore polycarbonate filter (Costar Scientific). Duplicate filters were examined with an epifluorescence mode of a Carl Zeiss LSM 510 confocal scanning laser microscope (CSLM; Carl Zeiss, Oberkochen, Germany). Total cells and live cells (cells/ml) were enumerated by direct counting at least 20 randomly chosen microscopic fields for each filter.
Decay rate calculations
Decay rates of each 16S rRNA genetic markers and culturable indicator organisms (TC and FC) were calculated using the following equation:
where r = decay rate, N t = log10 (copies or CFU/100 ml) at time t, N 0 = log10 (copies or CFU/100 ml) at time zero, and t = time (in days). Time (t) was determined by the days between the beginning of incubation and after 7 days for temperature effect experiments and after 11 days for salinity effect experiments, respectively.
Results
Quantification limit of real-time PCR assay
Linear decrease in copy numbers of Bac-Pre1 marker as a function of tenfold dilution series of suspended feces in non-filtered river water was observed down to 1.2 × 104 copies per 100 ml (120 copies per reaction; Fig. 1a). Similarly, the detection limits of Human-Bac1, Cow-Bac2, and Pig-Bac2 were determined to be 2.1 × 103 (21 copies per reaction), 4.2 × 102 (4.2 copies per reaction), and 1.1 × 103 copies per 100 ml (11 copies per reaction), respectively (Fig. 1b–d). On the other hand, fecal coliforms were countable for 10−3 and 10−4 g total feces per 100 ml of suspended fecal samples, when we directly filtered 1 ml of the sample solution (Fig. 1a).
Determination of quantification limits. Up to 20 ml of the samples were concentrated to 200 μl by centrifugation for DNA extraction. a Comparison of copy numbers of Bac-Pre1 markers with total fecal coliforms (CFU); b Human-Bac1 markers; c Cow-Bac2 marker; d Pig-Bac2 marker in tenfold dilutions of suspended feces mixture (human, cows, and pigs) in non-filtered river water. ND*, Not detected. Dotted lines in the figures indicated the detection limits determined previously in standard curves generated with B. fragilis genomic DNA or target specific plasmid DNA (Okabe et al. 2007). Error bars indicate the standard deviation of duplicate reactions. In some cases, the error bars were too small to illustrate
Persistence of B. fragilis strain and its survivability
Time-dependent changes in numbers of Bac-Pre1 markers, live cells, and culturable cells of B. fragilis strain were monitored in filtered and non-filtered river and seawater by real-time PCR with Bac-Pre1 primer, live/dead staining, and agar plate counting, respectively. The results in filtered river water showed no decrease in both numbers of Bac-Pre1 markers and live cell counts within 16-day incubation. In contrast, culturable cell counts began to decrease from the beginning and decreased for about 4 logs CFU/ml within 16 days (Fig. 2a). In non-filtered river water, the copy numbers and live cell counts started to decrease after 5 days. The culturable cell counts decreased from the beginning and could not be detected after 11 days (Fig. 2b). In filtered seawater, the numbers of Bac-Pre1 markers and live cells appeared to remain constant throughout 16-day incubation, while the culturable cells decreased rapidly from the beginning (Fig. 2c). In non-filtered seawater, the numbers of Bac-Pre1 markers and live cell counts started to decrease after 10 days and the culturable cell counts decreased rapidly and reached the detection limit after 7 days (Fig. 2d).
Persistence of culturable Bacteroides fragilis in CFU (circle), live cell counts (square) and copy numbers of Bac-Pre1 markers (diamond) incubated at 10°C in filtered river water (a), non-filtered river water (b), filtered seawater (c), and non-filtered seawater (d). Error bars indicate the standard deviation of duplicate runs. In some cases, the error bars were too small to illustrate
Persistence of feces-derived Bacteroides–Prevotella 16S rRNA genetic markers
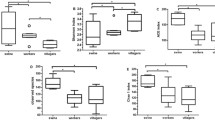
Time-dependent changes in copy numbers of feces-derived host-specific Bacteroides–Prevotella 16S rRNA genetic markers were monitored by real-time PCR with primer sets (Bac-Pre1, Human-Bac1, Cow-Bac2, and Pig-Bac2). At the same time, enumerations of total and fecal coliforms were conducted. The decay pattern of each host-specific Bacteroides–Prevotella 16S rRNA genetic marker was essentially the same at a given incubation temperature and salinity (Figs. 3 and 4). In control samples (incubated at 10°C under anaerobic condition), no significant decay or proliferation was observed in all experiments (data not shown).
Persistence of feces-derived Bac-Pre1 markers (a), Human-Bac1 marker (b), Cow-Bac2 marker (c), Pig-Bac2 marker (d), total coliforms, (e) and fecal coliforms (f) in non-filtered river water incubated at 4°C (circle), 10°C (square), 20°C (diamond) and 30°C (triangle), respectively. Dotted lines in the figures indicated the quantification limits of each independent marker as shown in Fig. 1. Means±SD were reported. In some cases, the error bars were too small to illustrate
Persistence of feces-derived Bac-Pre1 markers (a), Human-Bac1 marker (b), Cow-Bac2 marker (c), Pig-Bac2 marker (d), total coliforms (e), and fecal coliforms (f) in mixture of non-filtered river water and non-filtered seawater at 10°C, where the salinity is 0 ppt (circle), 10 ppt (square), 20 ppt (diamond), and 30 ppt (triangle), respectively. Dotted lines in the figures indicated the quantification limits of each independent marker as shown in Fig. 1. Means±SD were reported. In some cases, the error bars were too small to illustrate
The effect of temperatures
Temperature had influence on the decay of both Bacteroides–Prevotella 16S rRNA genetic markers and total and fecal coliforms in non-filtered river water. Decay rates for the copy numbers of Bac-Pre1, Human-Bac1, Cow-Pac2, and Pig-Bac2 markers were high at higher temperatures (Table 1). The copy numbers decreased about 1–3 logs within the first 2 days at 20 and 30°C, 7 days at 10°C and 11 days at 4°C (Fig. 3a–d). The copy numbers eventually reached each individual quantification limit of genetic markers (102–104 copies per 100 ml as shown in Fig. 1). On the other hand, TC and FC counts remained relatively unchanged for the first 4 days and then gradually decreased in all temperatures tested. The decay rates of TC and FC were faster at 4°C. The lowest decay rates were observed at 10°C for both TC and FC.
The effect of salinity
Each host-specific Bacteroides–Prevotella 16S rRNA genetic marker behaved similarly: The copy numbers of Bac-Pre1, Human-Bac1, Cow-Pac2, and Pig-Bac2 markers started to decrease after 4 days (Fig. 4a–d). The number of each genetic marker tended to decrease faster at lower salinities (Table 1). The decay rate of TC counts was greatly enhanced with salinity (Fig. 4e and Table 1). TC counts increased about 2 logs CFU/100 ml during the first 4 days and then gradually decreased in low salinity samples (0 and 10 ppt), giving positive decay rates (Table 1). The TC counts after 35 days of incubation were still at the same levels as the initial TC counts. In contrast, the TC counts in high salinity samples (20 and 30 ppt) were constant for the first 4 days and then started to decrease. No significant effect of salinity was observed for the FC (Fig. 4f). The FC counts started to decrease rapidly after 4 days and reached the quantification limit after 10-day incubation in all salinities tested.
Discussion
A quantification limit of our real-time PCR with Bac-Pre1 primer set specific to 16S rRNA genes of total Bacteroides–Prevotella spp. was about 1.2 × 104 copies per 100 ml (120 copies per reaction). In addition, the quantification limits of other markers (Human-Bac1, Cow-Bac2, and Pig-Bac2) were slightly lower than the Bac-Pre1 primer sets. These quantification limits were comparable to those for feces-derived Bacteroides 16S rRNA genetic marker (ten copies per reaction; Dick and Field 2004) and human-specific HF183 Bacteroides 16S rRNA genetic marker (9.4 × 103 copies per 100 ml; Seurinck et al. 2005). When we used the real-time PCR assay to investigate the persistence of feces-derived markers at high temperatures, the copy numbers reached the quantification limit (102–104 copies per 100 ml) of each individual marker (Fig. 3). Quantification limit is, however, complicated by a series of factors including the cell collection steps, the efficiency of target DNA recovery, and the efficiency of PCR amplification. All of these factors vary between each sample and study, which prevents exact direct comparison of the quantification limits determined in different studies. We anticipate that further refinements in bacteria collection from samples by using, for example, filtration and DNA extraction will improve the quantification limit of our real-time PCR assay.
In this study, persistence of host-specific Bacteroides–Prevotella 16S rRNA genetic makers in environmental waters was investigated using a pure culture strain of B. fragilis and mixed fecal samples. In the experiments with the pure cultured B. fragilis, the number of culturable cells started to decrease right after initiating incubation, while the numbers of Bac-Pre1 markers and live cells of B. fragilis still remained constant in filtered river water and seawater. The results indicated the presence of ‘viable but non-culturable’ cells, which is a proportion of the viable population that cannot be recovered by normal culturing techniques (Xu et al. 1982; Roszak et al. 1984). The ‘viable but non-culturable’ phenomenon is thought to be a survival strategy adopted by some Gram-negative species at low nutrient concentrations (Kogure et al. 1979). The copy numbers of Bac-Pre1 markers were, however, always smaller than live cell counts in filtered river water and seawater. As each cell of B. fragilis is carrying six operons of the 16S rRNA genes, theoretically, the numbers of Bac-Pre1 marker should be higher than the numbers of live B. fragilis cells. This phenomenon could be attributed to insufficient efficiency of cell collection, DNA extraction, or PCR amplification. In non-filtered river water and seawater that contained natural microbial flora (only less than 0.1% of the inoculated B. fragilis population), the growth of natural bacteria may occur and contribute to increases in the gaps between the numbers of Bac-Pre1 marker and live cells in the later phases of the incubation.
Our experimental results revealed that four different host-specific genetic markers showed similar persistence at each given condition (Figs. 3a–d and 4a–d). This finding is important to evaluate the source and extent of fecal contaminations in natural environmental waters. Kreader (1998) has examined the effects of temperature and predators on the persistence of Bacteroides distasonis derived from human feces by using a dot blot hybridization assay. He found that the PCR target was preserved for at least a week at 24°C in filtered or cyclohexamide (a compound expected to inhibit eukaryotic organisms) treated river water but only 1–2 days in non-filtered river water. He, therefore, concluded that the predator and degradation processes were less active at lower temperatures and seasonal variations in persistence must be considered. Seurinck et al. (2005) have also reported that feces-derived human-specific Bacteroides 16S rRNA genetic marker, which was quantified by real-time PCR assay, persisted longer (more than 24 days) at lower temperatures (4 and 12°C). The results in this study also showed longer persistence of all host-specific Bacteroides–Prevotella 16S rRNA genetic markers at lower temperatures in filtered river and seawater (data not shown) as reported in previous studies. This is probably because the activity of predators and the degradation rate of DNA are expected to be low at lower temperatures.
The effect of salinity was less obvious than that of temperature. The host-specific Bacteroides–Prevotella 16S rRNA genetic markers tended to decrease faster at low salinity (Fig. 4). In the experiment with B. fragilis, the Bac-Pre1 markers remained constant for 11 days in non-filtered seawater as compared with the non-filtered river water (5 days). In addition, no decay of the 16S rRNA genetic markers was observed in both filtered river water and filtered seawater. Based on these results, we speculated that the salinity has indirect influences on the persistence of host-specific Bacteroides–Prevotella 16S rRNA genetic markers, for example, by controlling the activity of predators or other indigenous microorganism, which would not be virtually affected by the salinity.
Considering all the results together, we could suggest that the presence of indigenous microorganisms or protozoa plays a major role in the decay of Bacteroides–Prevotella 16S rRNA genetic markers in environmental waters. We speculated that rapid declines in copy numbers of the genetic markers in non-filtered environmental waters coincided with an increase in densities of predators as reported in previous survival studies of E. coli in surface water (Enzinger and Cooper 1975; McCambridge and McMeekin 1979, 1980; Anderson et al. 1983). However, enumeration and identification of predators was not conducted in our study, and further examination is definitely required to confirm the speculation.
For E. coli, the effects of environmental factors, including sedimentation (Craig et al. 2004; Marino and Gannon 1991), predation (Enzinger and Cooper 1975; McCambridge and McMeekin 1980) sunlight, temperature, salinity, and nutrient deficiencies on their survival in fresh and marine environments have been studied extensively (Rozen and Belkin 2001). In this study, survivals of culturable TC and FC under different temperatures and salinities were investigated by using real fecal samples and compared with the persistence of host-specific Bacteroides–Prevotella 16S rRNA genetic markers. Total and fecal coliforms tended to decay faster at 4°C (Table 1). Similar results were observed in the survival studies of E. coli conducted in estuarine in situ over a variety of seasonal temperatures (5.9–28.2°C; Rhodes and Kator 1988). In addition, our observations support the previous findings that E. coli die-off (Anderson et al. 1983) and sublethal stress (Rhodes et al. 1983) in filtered water were inversely related to temperature. The significant effect of salinity was only observed for TC, but not for FC, in our study. Decay rate of TC was greatly enhanced by salinity, while FC decreased similarly in all salinity levels. Several previous studies have indicated that increasing salinity decreased their survival (Anderson et al. 1979; Anderson et al. 2005; Bordalo et al. 2002; Craig et al. 2004). In general, our results exhibited prolonged survival potential of total and fecal coliforms as compared to most of the previous studies. This is probably because the previous studies used a pure culture of E. coli, while our studies used fresh fecal samples that contain various phylotypes of enteric bacteria and relatively high concentrations of organic carbon and other nutrients. In addition, the dark incubation conditions might contribute to prolonged persistence of total and fecal coliforms in our studies because several researchers have reported that light appears to be a critical factor affecting the survival of enteric bacteria (Davies and Evison 1991; Rozen and Belkin 2001; Troussellier et al. 1998).
Conclusions
Based on the results of this study, the following conclusions can be drawn:
-
1.
The detection of the Bacteroides–Prevotella 16S rRNA genetic markers in environmental waters by real time PCR assay mainly reflected the presence of ‘viable but non-culturable’ Bacteroides–Prevotella cells.
-
2.
Four different host-specific genetic markers (total, human-, cow-, and pig-specific Bacteroides–Prevotella) showed similar persistence patterns in non-filtered river and seawater: The genetic markers persisted longer at lower temperatures and higher salinities. In addition, these markers did not increase in all incubation conditions tested.
-
3
Decay rates for TC and FC counts were lower than those for host-specific genetic markers at temperature above 10°C. The decay rates for TC were greatly enhanced by increasing salinity, whereas the decay rates for FC were relatively constant, regardless of salinity.
These findings indicate that seasonal and geographical variations in persistence of these host-specific Bacteroides–Prevotella 16S rRNA genetic markers must be considered when we use them as alternative fecal indicators in environmental waters.
References
Allsop K, Stickler JD (1985) An assessment of Bacteroides fragilis group organisms as indicators of human faecal pollution. J Appl Bacteriol 58:95–99
Anderson IC, Rhodes MW, Kator HI (1979) Sublethal stress in Escherichia coli: a function of salinity. Appl Environ Microbiol 38:1147–1152
Anderson IC, Rhodes MW, Kator HI (1983) Seasonal variation in survival of Escherichia coli exposed in situ in membrane diffusion chamber water. Appl Environ Microbiol 45:1877–1883
Anderson KL, Whitlock JE, Harwood VJ (2005) Persistence and differential survival of fecal indicator bacteria in subtropical water and sediments. Appl Environ Microbiol 71:3041–3048
APHA, AWWA, WEF (1995) Standard method for the examination of water and wastewater (19th edn.). American Public Health Association, Washington, DC
Bernhard AE, Field LG (2000) Identification of nonpoint sources of fecal pollution in coastal waters by using host-specific 16S ribosomal DNA genetic markers from fecal anaerobes. Appl Environ Microbiol 66:1587–1594
Bordalo AA, Onrassami R, Dechsakulwatana C (2002) Survival of faecal indicator bacteria in tropical estuarine water. J Appl Microbiol 93:864–887
Bower PA, Scopel CO, Jensen ET, Depas MM, McLellan SL (2005) Detection of genetic markers of fecal indicator bacteria in Lake Michigan and determination of their relationship to Escherichia coli densities using standard microbiological methods. Appl Environ Microbiol 71:8305–8313
Byappanahalli M, Fujioka R (2004) Indigenous soil bacteria and low moisture may limit but allow faecal bacteria to multiply and become a minor population in tropical soils. Water Sci Technol 50:27–32
Carson CA, Christiansen JM, Yampara-Iquise H, Benson VW, Baffaut C, Davis JV, Broz RR, Kurtz WB, Rogers WM, Fales WH (2005) Specificity of a Bacteroides thetaiotaomicron marker for human feces. Appl Environ Microbiol 71:4945–4949
Craig DL, Fallowfield HJ, Cromar NJ (2004) Use of microcosms to determine persistence of Escherichia coli in recreational coastal water and sediment and validation with in situ measurements. J Appl Microbiol 96:922–930
Davies CM, Evison LM (1991) Sunlight and the survival of enteric bacteria in natural waters. J Appl Bacteriol 70:265–274
Desmarais TR, Solo-Gabriele HM, Palmer CJ (2002) Influence of soil on fecal indicator organisms in tidally influenced subtropical environment. Appl Environ Microbiol 68:1165–1172
Dick LK, Bernhard AE, Brodeur TJ, Domingo WS, Simpson JM, Walters SP, Field KG (2005) Host distributions of uncultivated fecal Bacteroidales bacteria reveal genetic markers for fecal source identification. Appl Environ Microbiol 71:3184–3191
Dick LK, Field KG (2004) Rapid estimation of numbers of fecal Bacteroides by use of a quantitative PCR assay for 16S rRNA genes. Appl Environ Microbiol 70:5696–5697
Enzinger RM, Cooper RC (1975) Role of bacteria and protozoa in the removal of Escherichia coli from estuarine waters. Appl Environ Microbiol 31:758–763
Field KG, Bernhard AE, Brodeur TJ (2003) Molecular approaches to microbiological monitoring: fecal source detection. Environ Monit Assess 81:313–326
Fiksdal L, Maki JS, Lacroix SJ, Staley JT (1985) Survival and detection of Bacteroides spp., prospective indicator bacteria. Appl Environ Microbiol 49:148–150
Griffith JF, Weinsberg SB, Mcgee CD (2003) Evaluation of microbial source tracking methods using mixed fecal sources in aqueous test samples. J Water Health 1:141–151
Josephson KL, Gerba CP, Pepper IL (1993) Polymerase chain reaction detection of nonviable bacteria pathogens. Appl Environ Microbiol 59:3513–3515
Kim G, Choi E, Lee D (2005) Diffuse and point pollution impacts on the pathogen indicator organism level in the Geum River, Korea. Sci Total Environ 340:94–105
Kogure K, Shimidu U, Taga N (1979) A tentative direct microscopic method for counting living marine bacteria. Can J Microbiol 25:415–420
Kreader C A (1998) Persistence of PCR-detectable Bacteroides distasonis from human feces in river water. Appl Environ Microbiol 64:4103–4105
Layton A, McKay L, Williams D, Garrett V, Genry R, Sayler G (2006) Development of Bacteroides 16S rRNA gene TaqMan-based real-time PCR assays of total, human, and bovine fecal pollution in water. Appl Environ Microbiol 72:214–4224
Livingston SJ, Kominos SD, Yee RB (1978) New medium for selection and presumptive identification of the Bacteroides fragilis group. J Clin Microbiol 7:448–453
Marino RP, Gannon JJ (1991) Survival of fecal coliforms and fecal streptococci in storm drain sediment. Water Res 25:1089–1098
McCambridge J, McMeekin TA (1979) Protozoan predation of Escherichia coli in estuarine waters. Water Res 13:659–663
McCambridge J, McMeekin TA (1980) Relative effects of bacterial and protozoan predators on survival of Escherichia coli in estuarine water samples. Appl Environ Microbiol 40:907–911
Meays CL, Broersma K, Nordin R, Mazumder A (2004) Source tracking fecal bacteria in water: a critical review of current methods. J Environ Manag 73:71–79
Okabe S, Okayama N, Savichtcheva O, Ito T (2007) Identification and quantification of host-specific Bacteroides-Prevotella 16S rRNA genetic markers for assessment of fecal pollution in freshwaters. Appl Microbiol Biotechnol 74:890–901
Rhodes MW, Kator H (1988) Survival of Escherichia coli and Salmonella spp. in estuarine environments. Appl Environ Microbiol 54:2902–2907
Rhodes MW, Anderson IC, Kator HI (1983) In situ development of sublethal stress in Escherichia coli: effects on enumeration. Appl Environ Microbiol 45:1870–1876
Roszak DB, Grimes DJ, Colwell RR (1984) Viable but no recoverable stage of Salmonella enteritidis in aquatic systems. Can J Microbiol 30:332–338
Rozen Y, Belkin S (2001) Survival of enteric bacteria in seawater. FEMS Microbiol Rev 25:513–559
Scott TM, Rose JB, Jenkins TM, Farrah SR, Lukasik J (2002) Microbial source tracking: current methodology and future direction. Appl Environ Microbiol 68:5796–5803
Seurinck S, Defoirdt T, Verstraete W, Siciliano SD (2005) Detection and quantification of the human-specific HF183 Bacteroides 16S rRNA genetic marker with real-time PCR for assessment of human faecal pollution in freshwater. Environ Microbiol 7:249–259
Simpson JM, Santo Domingo JW, Reasoner DJ (2002) Microbial source trackings of science. Environ Sci Technol 36:5279–5288
Simpson JM, Santo Domingo JW, Reasoner DJ (2004) Assessment of equine fecal contamination: the search for alternative bacterial source-tracking targets. FEMS Microbial Ecology 47:65–75
Solo-Gabriele HM, Wolfert MA, Desmarais TR, Palmer CJ (2000) Sources of Escherichia coli in a coastal subtropical environment. Appl Environ Microbiol 66:230–237
Troussellier M, Bonnefont JL, Courties C, Derrien A, Dupray E, Gauthier M, Gourmelon M, Joux F, Lebaron P, Martin Y, Pommepuy M (1998) Responses of enteric bacteria to environmental stress in seawater. Oceanol Acta 21:965–981
Xu HS, Roberts N, Singleton FL, Attwell RW, Grimes DJ, Colwell RR (1982) Survival and viability of nonculturable Escherichia coli and Vibrio cholera in the estuarine and marine environment. Microbiol Ecol 8:313–323
Acknowledgment
We gratefully appreciate Koji Kawata, Department of Urban and Environmental Engineering, Hokkaido University, for valuable comments. This research was partly supported by the 21st Century Center Of Excellence (COE) program “Sustainable Metabolic System of Water and Waste for Area-Based Society” and grant-in-aid for developmental science research (No.15360283) from the ministry of Education, Science, and Culture of Japan.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Okabe, S., Shimazu, Y. Persistence of host-specific Bacteroides–Prevotella 16S rRNA genetic markers in environmental waters: effects of temperature and salinity. Appl Microbiol Biotechnol 76, 935–944 (2007). https://doi.org/10.1007/s00253-007-1048-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-007-1048-z