Abstract
Xylitol production by Pichia stipitis FPL-YS30, a xyl3-Δ1 mutant that metabolizes xylose using an alternative metabolic pathway, was investigated under aerobic and oxygen-limited culture conditions. Under both culture conditions, FPL-YS30 (xyl3-Δ1) produced a negligible amount of ethanol and converted xylose mainly into xylitol with comparable yields (0.30 and 0.27 g xylitol/g xylose). However, xylose consumption increased five-fold under aerobic compared to oxygen-limited conditions. This suggests that the efficiency of the alternative route of xylose assimilation is affected by respiration. As a result, the FPL-YS30 strain produced 26 g/l of xylitol, and exhibited a higher volumetric productivity (0.22 g xylitol l−1 h−1) under aerobic conditions.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Xylose is the second most abundant sugar in nature (Pettersen 1984). Therefore, its degradation into CO2 is important for carbon cycling. Recently, the conversion of xylose into value-added chemicals, such as xylitol, ethanol, and lactic acid, has become attractive to the fermentation industry (Lynd et al. 1999). In particular, the bioconversion process for xylitol production has been intensively studied during the last decade because xylitol can be used as a functional sweetener (Emidi 1978). So far three types of yeasts are known to produce xylitol. First, there are wild type xylose assimilating yeasts, such as Candida boindii (Vandeska et al. 1995), C. guilliermondii (Meyrial et al. 1991), C. tropicalis (Kim et al. 1999), C. parapsilosis (Oh et al. 1998), and Debaryomyces hansenii (Cruz et al. 2002), Second, recombinant Saccharomyces cerevisiae (Hallborn et al. 1991; Lee et al. 2003; Meinander and Hahn-Hagerdal 1997) containing XYL1 from Pichia stipitis is known to convert xylose into xylitol. Third, mutant strains of Pichia stipitis also produce xylitol from xylose. Cho and Jeffries reported the accumulation of xylitol by P. stipitis alcohol dehydrogenase (ADH) disrupted mutants (Cho and Jeffries 1998) and (Kim et al. 2001) reported xylitol production with a xylitol dehydrogenase defective mutant of P. stiptis.
We disrupted the XYL3 gene coding for D-xylulokinase in P. stipitis. The resulting strain, FPL-YS30 (xyl3-Δ1) utilized xylose slowly and accumulated a significant amount of xylitol in the medium (Jin et al. 2002). A metabolic pathway via arabinitol and ribulose-5-phosphate, bypassing the xylulokinase step, was proposed as an alternative pathway mediating xylose assimilation in the xylulokinase mutant. This alternative pathway consists of xylose reductase, xylitol dehydrogenase, arabinitol dehydrogenase, and ribulokinase reactions. This pathway is also involved in L-arabinose assimilation in P. stipitis (Fig. 1). In the present study, we investigated the capability of the alternative pathway for xylitol production by culturing the FPL-YS30 strain under aerobic and oxygen-limited conditions.
Materials and methods
Microorganisms and fermentation conditions
P. stipitis strains FPL-UC7 (ura3-2) (= NRRL Y-21448) (Lu et al. 1998) and P. stipitis FPL-YS30 (xyl3-Δ1) (xyl3::URA3) (=NRRL Y-30785) (Jin et al. 2002) were used in this study. Yeast strains were maintained at 4°C on agar plates of YPD (10 g/l of yeast extract, 20 g/l of Bactopeptone, and 20 g/l of glucose) medium. YPD medium was used for inoculum preparation. YP medium (10 g/l of yeast extract, 20 g/l of Bactopeptone) with 100 g/l of xylose was used for xylose fermentation. Fermentation experiments were carried out at 30°C in 50 ml of medium in a 125 ml Erlenmeyer flask. Cells were grown in two different culture conditions, i.e. oxygen-limited and aerobic conditions by agitating at 100 and 200 rpm, respectively.
Analytical methods
Xylose, xylitol, and ethanol concentrations were determined by HPLC (HP, Wilmington, Del.) using an ION 300 column (Interaction Chromatography, San Jose, Calif.) with 5 mM sulfuric acid solution as a mobile phase at a flow rate of 0.6 ml/min. Cell growth was monitored by optical density at 600 nm (OD600). One OD600 at 600 nm was equivalent to 0.21 g cell/l for P. stipitis.
Results
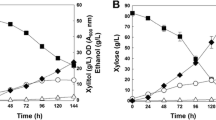
To investigate characteristics of the alternative pathway for xylose assimilation that bypasses xylulokinase, we cultured FPL-YS30 (xyl3-Δ1), and its parental strain, P. stipitis UC7 (ura3), in YP medium with 100 g/l of xylose under two different conditions (100 and 200 rpm). Overall, FPL-YS30 consumed xylose very slowly compared to P. stipitis UC7 at 100 rpm (Fig. 2). During the first 40 h FPL-YS30 consumed xylose faster than P. stipitis UC7 (Fig. 3), but the maximum specific xylose consumption rates of FPL-YS30 were only 17% and 41% of P. stipitis UC7 cultured with aeration at 100 and 200 rpm, respectively. FPL-YS30 consumed only 18 g/l of xylose with aeration at 100 rpm whereas it consumed 99 g/l of xylose with aeration at 200 rpm within the 120 h of the fermentation experiment. P. stipitis UC7 mainly produced ethanol from xylose with yields of 0.40 and 0.32 g ethanol/g xylose with aeration at 100 and 200 rpm, respectively. In contrast, ethanol production from xylose by FPL-YS30 was negligible (less than 2 g/l) under both conditions. FPL-YS30 produced mainly xylitol from xylose with a yield of 0.30 and 0.27 g xylitol/g xylose at aeration at aeration rates of 100 and 200 rpm whereas xylitol production by P. stipitis UC7 under the same conditions was almost negligible (less than 3 g/l). Xylitol accumulation by FPL-YS30 began as soon as 20 h into the fermentation experiments, which can be regarded as fully aerobic conditions (Figs. 2b, 3b). In contrast, ethanol was not produced by P. stipitis UC7 within that same time period under the same conditions (Figs. 2a, 3a).
Discussion
We suggested that FPL-YS30 (xyl3-Δ1) uses an alternative pathway to metabolize xylose because it did not show any detectable D-xyluokinase activity yet it was still able to grow on xylose (Jin et al. 2002). Based on xylose consumption profiles, the metabolic capacity of the alternative pathway was inferior to the main pathway. This suggests that the xylulokinase pathway is mainly responsible for xylose consumption in wild type P. stipitis. This is also supported by previous results that D-xyulokinase activity is three fold higher than D-ribulokinase activity in P. stipitis UC7 grown on xylose (Jin et al. 2002). Although xylose metabolism via either pathway is aeration-dependent, the alternative pathway required a higher aeration rate than the main pathway. The mutant did not produce any ethanol even though it could grow using the alternative pathway. This was similar to our earlier observation that P. stipitis produced very little ethanol from L-arabinose (Shi et al. 2000) even though it could grow slowly on this sugar.
Kim et al. (2001) reported an increase in xylitol production by a P. stipitis xylitol dehydrogenase mutant. Cho and Jeffries (Cho and Jeffries 1998) also reported that P. stipitis PsADH disruptants produce xylitol. Although these mutants exhibit similar phenotypes in terms of xylitol production, metabolic mechanisms underlying these similar phenotypes are distinct. First, the xylitol dehydrogenase mutant requires co-substrates, which support growth and regeneration of cofactor, to produce xylitol from xylose since its xylose metabolic pathway blocked. By comparison, adhΔ and xyl3Δ mutants can produce xylitol from xylose as a sole carbon source. Second, the xylitol dehydrogenase mutant and the xyl3-Δ1 mutant accumulate xylitol because of direct disruption in xylose metabolic pathway, but adhΔ mutants are thought to produce xylitol because of indirect metabolite interactions through cytosolic NADH. Third, adhΔ mutants prefer oxygen-limited conditions for xylitol production, like most xylitol producing yeast strains, but the xyl3-Δ1 mutant exhibited the same xylitol yield independent of aeration. This suggests that xylitol accumulation by adhΔ mutants is mainly caused by redox imbalance in the cytosol under oxygen-limited conditions. In contrast, the YS30 (xyl3-Δ1) mutant produced xylitol under the same conditions that its parent produced ethanol. These results suggest that FPL-YS30 (xyl3-Δ1) produces xylitol mainly because of insufficient glycolytic capacity of the alternative pathway rather than because of a redox imbalance. This also indicates that the oxidative phase of the PPP, which produces NADPH for xylitol accumulation, is sufficient even without activity coded for by XYL3. This is a new class of P. stipitis mutants that accumulate xylitol during xylose fermentation by a different metabolic mechanism compared to mutants reported previously.
References
Cho JY, Jeffries TW (1998) Pichia stipitis genes for alcohol dehydrogenase with fermentative and respiratory functions. Appl Environ Microbiol 64:1350–1358
Cruz JM, Dominguez JM, Dominguez H, Parajo JC (2002) Xylitol production from barley bran hydrolysates by continuous fermentation with Debaryomyces hansenii. Biotechnol Lett 22:1895–1898
Emidi A (1978) Xylitol, its properties and food applications. Food Technol 32:20–32
Hallborn J, Walfridsson M, Airaksinen U, Ojamo H, Hahn-Hägerdal B, Penttila M, Kerasnen S (1991) Xylitol production by recombinant Saccharomyces cerevisiae. Biotechnology 9:1090–1095
Jin YS, Jones S, Shi NQ, Jeffries TW (2002) Molecular cloning of XYL3 (D-xylulokinase) from Pichia stipitis and characterization of its physiological function. Appl Environ Microbiol 68:1232–1239
Kim JH, Ryu YW, Seo JH (1999) Analysis and optimization of a two-substrate fermentation for xylitol production using Candida tropicalis. J Ind Microbiol Biotechnol 22:181–186
Kim MS, Chung YS, Seo JH, Jo DH, Park YH, Ryu YW (2001) High-yield production of xylitol from xylose by a xylitol dehydrogenase defective mutant of Pichia stipitis. J Microbiol Biotechnol 11:564–569
Lee WJ, Kim MD, Yoo MS, Ryu YW, Seo JH (2003) Effects of xylose reductase activity on xylitol production in two-substrate fermentation of recombinant Saccharomyces cerevisiae. J Microbiol Biotechnol 13:725–730
Lu P, Davis BP, Hendrick J, Jeffries TW (1998) Cloning and disruption of the beta-isopropylmalate dehydrogenase gene (LEU2) of Pichia stipitis with URA3 and recovery of the double auxotroph. Appl Microbiol Biotechnol 49:141–146
Lynd LR, Wyman CE, Gerngross TU (1999) Biocommodity Engineering. Biotechnol Prog 15:777–793
Meinander NQ, Hahn-Hägerdal B (1997) Influence of cosubstrate concentration on xylose conversion by recombinant, XYL1-expressing Saccharomyces cerevisiae: a comparison of different sugars and ethanol as cosubstrates. Appl Environ Microbiol 63:1959–1964
Meyrial V, Delgenes JP, Molletta R, Navarro JM (1991) Xylitol production from D-xylose by Candida guilliermondii. Biotechnol Lett 13:281–286
Oh DK, Kim SY, Kim JH (1998) Increase of xylitol production rate by controlling redox potential in Candida parapsilosis. Biotechnol Bioeng 58:440–444
Pettersen RC (1984) The chemical composition of wood. Am Chem Soc Symp Ser 207:57–126
Shi NQ, Prahl K, Hendrick J, Cruz J, Lu P, Cho JY, Jones S, Jeffries T (2000) Characterization and complementation of a Pichia stipitis mutant unable to grow on D-xylose or L-arabinose. Appl Biochem Biotechnol 84–86:201–216
Vandeska E, Amartey S, Kuzmanova S, Jeffries TW (1995) Effects of environmental conditions on production of xylitol by Candida boidinii. World J Microbiol Biotechnol 11:213–218
Acknowledgements
The authors thank Hal Alper for reading the manuscript critically. This research was supported by NRI/CSREES, grant number 2001-35504-10695.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jin, YS., Cruz, J. & Jeffries, T. Xylitol production by a Pichia stipitis D-xylulokinase mutant. Appl Microbiol Biotechnol 68, 42–45 (2005). https://doi.org/10.1007/s00253-004-1854-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-004-1854-5