Abstract
The fermentation of xylose is essential for the bioconversion of lignocellulose to fuels and chemicals, but wild-type strains of Saccharomyces cerevisiae do not metabolize xylose, so researchers have engineered xylose metabolism in this yeast. Glucose transporters mediate xylose uptake, but no transporter specific for xylose has yet been identified. Over-expressing genes for aldose (xylose) reductase, xylitol dehydrogenase and moderate levels of xylulokinase enable xylose assimilation and fermentation, but a balanced supply of NAD(P) and NAD(P)H must be maintained to avoid xylitol production. Reducing production of NADPH by blocking the oxidative pentose phosphate cycle can reduce xylitol formation, but this occurs at the expense of xylose assimilation. Respiration is critical for growth on xylose by both native xylose-fermenting yeasts and recombinant S, cerevisiae. Anaerobic growth by recombinant mutants has been reported. Reducing the respiration capacity of xylose-metabolizing yeasts increases ethanol production. Recently, two routes for arabinose metabolism have been engineered in S. cerevisiae and adapted strains of Pichia stipitis have been shown to ferment hydrolysates with ethanol yields of 0.45 g g−1 sugar consumed, so commercialization seems feasible for some applications.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fermentation of xylose to ethanol is driven by political, economic and technical considerations. For example, using biomass as a feedstock for renewable fuel production can substantially reduce the accumulation of greenhouse gasses (McMillan 1997; Claassen et al. 1999; Wyman 1999; Kheshgi et al. 2000), so a recent European Union directive proposed that biofuels should represent 2% of total transportation fuel consumption by 2005 and 5.75% by 2010 (Roca and Olsson 2003). Increased domestic use of agricultural commodities can increase income for farmers, so the agricultural policies of the United States, Brazil and the European Union created nascent ethanol industries for the commercial production of ethanol from grains, sugar cane and other feedstocks. Opportunities for the bioconversion of harvest and processing residues are increasing along with these markets. For example, as grain hull, corn cobs, corn stover and sugar cane bagasse byproducts increase, ethanol production from the waste streams becomes feasible (Saha et al. 1998; Saha and Bothast 1999). Xylose is a major constituent of these and other renewable biomass feedstocks, but its efficient utilization—which is essential for commercial bioconversion (Hinmann et al. 1989; Olsson and Hahn-Hägerdal 1996)—presents a technical barrier. Lignocellulosic crop residues comprise more than half of the world′s agricultural phytomass (Smil 1999) and significant fractions of the total can be recovered without competing with other uses (Lynd 1996; Wyman 1999). Xylose constitutes about 17% of the total dry weight in woody angiosperms and ranges up to 31% in herbaceous angiosperms (Pettersen 1984; Hespell 1998). One source of xylose is the sulfite-pulping of hardwood (Lawford and Rousseau 1993). Depending on the substrate and reaction conditions, dilute acid pretreatments of lignocellulosic residues can recover 80–95% of the xylose from the feedstock (Chen et al. 1998; Kim et al. 2001; Aguilar et al. 2002).
Native strains of Saccharomyces cerevisiae do not use xylose as a carbon source. Candida utilis or ″torula yeast″ will grow on xylose, but this yeast is strictly aerobic and does not produce ethanol. In the early 1980s, following the discovery that S. cerevisiae, Schizosaccharomyces pombe and other yeasts can ferment d-xylulose to ethanol (Wang and Schneider 1980; Wang et al. 1980), intensive screening efforts rapidly revealed that some can convert xylose to ethanol directly under aerobic or oxygen-limiting conditions (Schneider et al. 1981; Jeffries 1982; Slininger et al. 1982. Attention focused on Pachysolen tannophilus, C. shehatae (Du Preez and van der Walt 1983) and Pichia stipitis, which are the best native xylose-fermenting yeasts known (Toivola et al. 1984; Du Preez et al. 1986). Many improvements have been made in the genetic engineering of yeasts and bacteria for the fermentation of xylose and arabinose to ethanol and other products such as lactic acid. However, the bioconversion of pentoses to ethanol still presents a considerable economic and technical challenge (Jeffries and Shi 1999; Aristidou and Penttilä 2000; Hahn-Hägerdal et al. 2001).
The objective of this review is to assess the current state of microbial strain development for the fermentation of pentose sugars. This is a very active field. From January 1999 to June 2003, more than 60 articles on yeast appeared in press. We try to emphasize the latest work and refer the reader to reviews of the earlier literature (Jeffries 1983, 1985; Prior et al. 1989; Jeffries and Kurtzman 1994; Gong et al. 1999; Ho et al. 1999; Jeffries and Shi 1999; Aristidou and Penttilä 2000; Flores et al. 2000; Jeffries and Jin 2000; Ostergaard et al. 2000; Dequin 2001; Hahn-Hägerdal et al. 2001; Galbe and Zacchi 2002).
Metabolic engineering of yeasts
Yeast strain development focuses on the genetic engineering of Saccharomyces cerevisiae, but P. stipitis has also been modified for xylose fermentation. Metabolic engineering can alter sugar transport, assimilation, the pentose phosphate pathway, glycolysis, the terminal steps of fermentation and the relatively complicated interplay between respiration and fermentation that determine the intracellular redox balance. While a few of the changes enable xylose utilization in S. cerevisiae, most have marginal effects. No one factor or enzymatic step is rate-limiting, but some are critical. Even though genes for xylose assimilation are present in S. cerevisiae, they are not expressed at a sufficient level to enable significant sugar assimilation. Engineering of P. stipitis has been more limited, but some principles have been established with this yeast.
Xylose transport
Sugar uptake limits xylose utilization in both S. cerevisiae and P. stipitis. In S. cerevisiae, the HXT family of sugar transporters mediates glucose uptake (Kruckeberg 1996; Boles and Hollenberg 1997). Hxt1–Hxt7 and Gal2 exhibit counter-transport when individually expressed in a hxt1-7 null mutant of S. cerevisiae, which indicates that they function by facilitated diffusion (Maier et al. 2002). Kinetic studies with 14C glucose can distinguish high- and low-affinity uptake systems. Xylose uptake competes with glucose uptake, indicating that they share transport components (Meinander and Hahn-Hägerdal 1997). S. cerevisiae takes up xylose by both low- and high-affinity glucose transport systems (Lee et al. 2002), but after incubation with xylose only the high-affinity system is detected. Cultivation of the xylose-fermenting S. cerevisiae FPL-YSX3 on xylose under aerobic or oxygen-limiting conditions strongly induces (5- to 50-fold) the high-affinity transporters HXT2, HXT6 and HXT7 and the moderate affinity transporter HXT5 (Buziol et al. 2002; Jin 2002). Hxt5 is produced under slow growth conditions (Diderich et al. 2001; Verwaal et al. 2002). With YSX3 cells cultivated in xylose medium, the low-affinity transporters HXT1 and HXT3 are expressed at 2–5% of the level observed with cells grown on glucose (Jin 2002). These results suggest that engineered S. cerevisiae mainly uses the high-affinity system for xylose transport.
Glucose strongly inhibits the transport of xylose by both high- and low-affinity systems (van Zyl et al. 1993). The native S. cerevisiae glucose transporters exhibit a significantly lower affinity for xylose (K m=49–300 mM) than for glucose (K m=1–28 mM; Kötter and Ciriacy 1993; Lagunas 1993; Van Zyl et al. 1999). Therefore, glucose and xylose are consumed simultaneously only under glucose-limited conditions (Meinander and Hahn-Hägerdal 1997). S. cerevisiae (TMB3201)—which is completely deficient in all 18 monosaccharide transporters (Wieczorke et al. 1999) but has a functional xylose utilization pathway (Hamacher et al. 2002)—cannot take up or grow on xylose. When various HXT genes are reintroduced, Hxt4, Hxt5, Hxt7 and Gal2 promote xylose uptake (Hamacher et al. 2002).
Native xylose-metabolizing yeasts have two kinetically distinct xylose transport systems. The low-affinity system is shared with glucose, while the high-affinity system is specific for xylose (Hahn-Hägerdal et al. 2001). Both of these systems are tightly coupled to energy metabolism (Kilian and van Uden 1988; Does and Bisson 1989). Three genes, SUT1, SUT2 and SUT3, encode glucose transporters in P. stipitis (Weierstall et al. 1999). Sut2 and Sut3 are highly similar to the S. cerevisiae glucose transporter family and the Sut2 and Sut3 transporters have a higher affinity for glucose than for xylose. Transcription of SUT1 is induced in P. stipitis independently of the oxygen supply. SUT2 and SUT3 are expressed only under aerobic conditions, but independently of the carbon source. Disruption of SUT1 eliminates the low-affinity xylose transport system in P. stipitis. Xylose uptake in P. stipitis grown on glucose has high-affinity (K m1=3.2 mM) and low-affinity (K m2=80 mM) components. In Δsut1 cells grown under oxygen-limiting conditions—when SUT2 and SUT3 are not expressed—xylose transport is still active, which suggests that specific xylose transporters other than Sut1, Sut2 and Sut3 are present. However, Weierstall et al. (1999) were not able to identify any additional cross-hybridization signals for HXT- or SUT-related genes in P. stipitis.
Xylose isomerase
The initial metabolic engineering of S. cerevisiae for xylose assimilation attempted the heterologous expression of bacterial xylose isomerase (XI). This approach was reasonable, given that S. cerevisiae can grow on and ferment xylulose, but significant XI activity was not attained in the transformed cells (Jeffries and Shi 1999; Hahn-Hägerdal et al. 2001). Probably, this is attributable to improper folding of the protein (Sarthy et al. 1987), but even if small amounts of active XI are formed, xylitol is a competitive inhibitor (Smith et al. 1991). Expression of a XI from the thermophilic bacterium Thermus thermophilus achieved the heterologous production of an active enzyme in S. cerevisiae (Walfridsson et al. 1996) and the genetic background has been modified to reduce xylitol production (Walfridsson et al. 1996; Träff et al. 2001). However, the temperature required for moderate T. thermophilus XI activity is well above the maximum growth temperature of Saccharomyces, so researchers from this same laboratory have used directed evolution to obtain a XI with 9-fold higher activity constants at 60 °C and much higher inhibition constants for xylitol (Lönn et al. 2002). Very recently, Harhangi et al. (2003) introduced a eukaryotic XI, the AraA gene, from the anaerobic fungus species into S. cerevisiae and enabled slow xylose assimilation by this route (Kuyper et al. 2003). Whether the use of XI will prove successful could depend on other factors. At equilibrium, energetics of the isomerization between xylose and xylulose favors xylose formation by 83:17 (Jeffries 1985), so some other driving force is necessary to promote this reaction.
Xylose reductase and xylitol dehydrogenase
As early as 1983, researchers concluded that the key to anaerobic assimilation of xylose by native yeasts such as Pachystolon tannophilus was the presence of an aldose (xylose) reductase (XR) that could accept either NADH or NADPH as a cofactor (Bruinenberg et al. 1983a, 1983b, 1984; Verduyn et al. 1985a, 1985b; Bruinenberg 1986). This hypothesis was based on a study of aerobic xylose utilization by C. utilis. Even though this yeast can rapidly ferment glucose, fermentative activity ceases immediately after transfer to xylose. The XR of C. utilis exclusively uses NADH as a cofactor. In comparison, at least one XR of P. tannophilus can use either NADH or NADPH (Verduyn et al. 1985a). The same is true of Pichia stipitis (Verduyn et al. 1985b) and C. shehatae (Ho et al. 1990); and all of these can metabolize xylose anaerobically—even though they do not grow under those conditions (Wijsman et al. 1985). Because the assimilation of xylose requires two oxidoreductase steps and because all oxidoreductase reactions following these are balanced, researchers hypothesized that, if no transhydrogenase were present to regenerate NAD+ and NADPH, xylose assimilation under anaerobic conditions would quickly halt.
When the XR of P. stipitis is cloned and expressed in S. cerevisiae (Amore et al. 1991; Takuma et al. 1991; Tantirungkij et al. 1993, 1994; Billard et al. 1995; Handumrongkul et al. 1998), the resulting transformants produce xylitol if some other carbon source is present to provide a reductant (Hallborn et al. 1991, 1994). When glucose is used as a co-substrate, xylose assimilation and xylitol production are reduced, presumably because of competition for transport (Thestrup and Hahn-Hägerdal 1995). XYL1 does not limit xylose assimilation in P. stipitis (Dahn et al. 1996). To produce ethanol, it is necessary to have a system that can oxidize xylitol to xylulose while reducing acetaldehyde under oxygen-limiting conditions.
Isolation of the first two genes for xylose assimilation from P. stipitis led to the initial development of S. cerevisiae strains able to metabolize xylose (Kötter et al. 1990; Amore et al. 1991). S. cerevisiae transformed with these two genes can use xylose oxidatively and produce xylitol without the addition of a co-metabolizable carbon source. Increasing xylitol dehydrogenase (XDH) activity relative to XR (XR:XDH=0.6) produces less xylitol and more ethanol than when XR is present in greater abundance (Walfridsson et al. 1997). When XYL2 is strongly expressed, xylulose is secreted—indicating that xylulokinase (XK) activity limits xylose metabolism in these cells (Jin and Jeffries 2003).
Xylulokinase
Much has been reported about S. cerevisiae engineered to express XK. Chang and Ho cloned D-xylulokinase from Pachysolen tannophilus and S. cerevisiae as early as 1988 (Chang and Ho 1988; Deng and Ho 1990) and were the first to report a sequence for the S. cerevisiae XK gene in a patent (Ho and Tsao 1993). The complete S. cerevisiae gene, XKS1, was obtained in the yeast genome project (Rodriguez-Pena et al. 1998). The protein coded for by the original Ho and Tsao sequence has been reported as inactive (Eliasson et al. 2000a). d-xylulokinase activity limits the metabolism of d-xylulose in S. cerevisiae (Chang and Ho 1988; Deng and Ho 1990) when S. cerevisiae xylulokinase is overexpressed along with Pichia stipitis XYL1 and XYL2 in Saccharomyces sp. 1400, which is a fusant product of S. diastaticus and S. uvarum (Moniruzzaman et al. 1997; Ho et al. 1998). Saccharomyces sp. 1400 (pLNH33) can ferment a mixture of 53 g glucose l−1 and 56 g xylose l−1 to give an ethanol concentration of 50 g l−1 within 36 h. (Krishnan et al. 1999). This is the highest ethanol yield and fermentation rate from glucose/xylose mixtures reported for a recombinant Saccharomyces to date.
S. cerevisiae CEN.PK 113-7A, a strain with a well defined genetic background, has also been developed as a xylose-fermenting yeast. XYL1, XYL2 and XKS1 were each integrated into the chromosome under the control of the S. cerevisiae PGK1 promoter to obtain a stable transformant designated S. cerevisiae TMB3001 (Eliasson et al. 2000b) which, like S. cerevisiae sp. 1400, is able to ferment mixtures of glucose and xylose to ethanol, albeit with slightly lower yields (cf. Ho et al. 1999; Krishnan et al. 1999). Overexpression of XKS1 clearly enhances xylose utilization under aerobic conditions, but xylose utilization declines by almost an order of magnitude with decreased aeration.
Intracellular ATP levels and the ratio of ATP:ADP are also dramatically lower in the recombinant yeast (Toivari et al. 2001). Overexpression of XKS1, XYL1 and XYL2 increased ethanol yield in two different strains of S. cerevisiae, but it also decreased total xylose consumption by 50–80% (Johansson et al. 2001; Table 1).
d-xylulokinase is not expressed significantly in native S. cerevisiae (Deng and Ho 1990), but it is essential for xylulose metabolism because xks1 mutants do not grow on xylulose. At the same time, overexpression of XKS1 can decrease growth on xylulose (Rodriguez-Pena et al. 1998). Richard et al. (2000) and later Johansson et al. (2001) suggested that overexpression of XKS combined with unlimited access to xylulose might lead to toxicity, due to ATP depletion in a manner similar to that observed with unregulated glucose uptake in Saccharomyces tps1/ggs1 mutants (Hohmann et al. 1993, 1996; Thevelein and Hohmann 1995; Teusink et al. 1998). Jin et al. (2002) expressed the P. stipitis gene for d-xylulokinase, XYL3, in a S. cerevisiae strain that was engineered for high levels of XYL1 and XYL2 (Jin et al. 2003). The XYL3 XK gene product is more specific for d-xylulose; and it has much lower activity against d-ribulose than the S. cerevisiae XK (Richard et al. 2000). When the P. stipitis XK activity is low, cell growth on xylose is not significantly affected. When P. stipitis XK is strongly overexpressed, aerobic growth is significantly inhibited on xylose. Aeration increases the toxicity of XYL3 or XKS1 overexpression. Xylulokinase activity is higher in cells grown on glucose, but no growth inhibition is observed, which indicates that the inhibitory effect is specific to xylose uptake. Ethanol production and growth are optimal when low levels of XYL3 are expressed from the native promoter. This suggests that the effect of overexpressing XK when d-xylulose is fully accessible is similar to substrate-accelerated cell death.
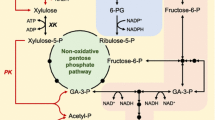
Deletion of XKS1 completely blocks xylitol formation from xylose, but leads to arabitol accumulation and increases ethanol production (Eliasson et al. 2000a). Deletions within the PGI1 promoter that reduce phosphoglucose isomerase activity by one to two orders of magnitude increase the accumulation of fructose-6-phosphate and result in about 15% higher ethanol yield. tps1 and tps2 mutants accumulate sugar phosphates and increase ethanol yield by 20–30%. gnd1 mutants show 30% higher ethanol yield, but rpe1 mutants hardly assimilate xylulose at all—as one might expect, because both ribulose-5-phosphate and xylulose-5-phosphate are necessary to form ribose-5-phosphate and the other intermediates of the pentose phosphate pathway (PPP; Fig. 1).
The pentose phosphate cycle in Saccharomyces cerevisiae engineered for xylose and arabinose assimilation. Reduction of l-arabinose to l-arabitol is mediated by aldose reductase. araA Bacillus subtilis l-arabinose isomerase, araB l-ribulokinase, araD l-ribulose-5-phosphate 4-epimerase, GND1 S. cerevisiae phosphogluconate dehydrogenase, lad1 Trichoderma reesei l-arabinitol 4-dehydrogenase, lxr1 T. reesei l-xylulose reductase, PGI1 S. cerevisiae glucose-6-phosphate isomerase, RKI1 S. cerevisiae ribose-5-phosphate isomerase, RPE1 S. cerevisiae ribulose-phosphate 3-epimerase, TAL1 S. cerevisiae transaldolase, TKL1, S. cerevisiae transketolase, XKS1 S. cerevisiae d-xylulokinase, XYL1 Pichia stipitis xylose (aldose) reductase, XYL2 P. stipitis xylitol dehydrogenase, XYL3, P. stipitis d-xylulokinase, XylA Piromyces xylose isomerase, ZWF1 S. cerevisiae glucose-6-phosphate 1-dehydrogenase
While we can learn a great deal about the mechanics of xylose fermentation by genetically manipulating laboratory strains, commercialization of the xylose fermentation will necessitate the use of strains that can grow vigorously, ferment rapidly and tolerate acids or other inhibitory compounds (Hahn-Hägerdal et al. 2001). Many industrial Saccharomyces strains have been developed that have these characteristics. In fermenting a mixture of glucose and xylose (50 g l−1 each), two industrial yeast strains expressing the xylose assimilation pathway produced more ethanol and consumed more xylose than an engineered laboratory strain (TMB 3001). However, the differences in ethanol production arose almost entirely during glucose consumption rather than during the xylose consumption phase (Zaldivar et al. 2002).
Pentose phosphate pathway
Kötter and Ciracy hypothesized that the excessive production of xylitol by S. cerevisiae genetically engineered with XYL1 and XYL2 was limited by the dual cofactor capacity of the P. stipitis XR, by excessive activity of the oxidative PPP in S. cerevisiae and by insufficient capacity of the non-oxidative PPP (Kötter and Ciriacy 1993). Overexpression of the P. stipitis gene for transketolase (TKL1) in a S. cerevisiae strain expressing heterologous XYL1 and XYL2 greatly reduced growth of the transformant on xylose minimal medium (Metzger and Hollenberg 1994). Strains overexpressing the P. stipitis gene for transaldolase (TAL1), XYL1 and XYL2 grew faster than strains expressing XYL1 and XYL2 alone (Walfridsson et al. 1995). However, the plasmid burden due to the overexpression of TAL1, XYL1 and XYL2 reduced the growth rate of the transformant relative to the host strain (Bao et al. 1997; Meinander et al. 1999).
Redox balance
The production of xylitol by recombinant S. cerevisiae is thought to originate from an overabundance of NADPH relative to NADH for the initial xylose assimilation step. To reduce xylitol production and thereby increase ethanol yield, Jeppson et al. (2002) overexpressed XKS1 in S. cerevisiae gnd1 and zwf1 backgrounds. The Δzwf1 mutant greatly increased ethanol yield by producing 0.41 g ethanol g−1 xylose consumed, as compared with 0.31 g g−1 by the parent strain. The Δgnd1 mutant also showed increased ethanol yield to (0.38 g g−1). However, both mutants showed reduced rates of xylose uptake, which indicates that NADPH production is necessary for xylose assimilation in S. cerevisiae.
Xylitol production by native xylose-metabolizing yeasts varies a great deal with species and aeration conditions. Pachysolen tannophilus, C. shehatae and Pichia stipitis all produce xylitol to varying extents (Du Preez et al. 1984; Sanchez et al. 2002). P. stipitis is notable for its very low xylitol production and high ethanol yields. Ethanol production decreases and xylitol production increases in P. stipitis when the primary alcohol dehydrogenase is deleted (Cho and Jeffries 1998). This suggests that ADH competes with XDH for reductant—presumably NADH—in P. stipitis. In this organism, the main ADH is induced as oxygen availability decreases (Cho and Jeffries 1999; Passoth et al. 2003).
Lowering the cytosolic NADPH concentration while regenerating NAD+ from NADH could reduce xylitol production (Richard et al. 2003). This might be accomplished by expressing a cytosolic transhydrogenase. S. cerevisiae does not possess transhydrogenase activity (Bruinenberg et al. 1985), so Nissen et al. (1997) expressed a transhydrogenase gene from Azotobacter vinlandii in S. cerevisiae and then measured the intracellular concentrations of the NAD(P) and NAD(P)H cofactors. The concentrations of the nicotinamide cofactors in the glucose-grown control cells were as follows:
Expression of the transhydrogenase increased the production of 2-oxoglutarate and glycerol and shifted the intracellular ratio of (NADPH/NADP+):(NADH/NAD+) from 35 to 17 (Nissen et al. 1997). These results indicated that the thermodynamic equilibrium for the transhydrogenase reaction lies in the direction of NADH formation. So it seems unlikely that this approach will be useful in the absence of other energy-consuming reactions.
The XR encoded by P. stipitis XYL1 has K m=3.2 μM for NADPH and K m=40 μM for NADH (Rizzi et al. 1988). The production of NADPH by glucose-6-phosphate dehydrogenase occurs largely on demand in response to the intracellular concentrations of NADPH (Michal 1999). Taken together, these three factors mean that the P. stipitis XR will always favor consumption of NADPH over NADH. Native P. stipitis does not produce significant amounts of xylitol, whereas recombinant S. cerevisiae expressing the P. stipitis XYL1 produces abundant xylitol, so some factor other than XR must be responsible for enabling cofactor balance in P. stipitis.
Respiration
Respiration plays a critical role in the metabolism of xylose by both native and engineered yeasts. The exact nature of the requirement is not fully understood, because oxygen appears to function differently in enabling growth and fermentation. Native xylose-metabolizing yeasts or genetically engineered S. cerevisiae (Eliasson et al. 2000b) can metabolize xylose to ethanol in the absence of oxygen, but oxygen is required for yeasts to grow on xylose (Ligthelm et al. 1988), and for optimal ethanol production, low aeration rates are required (Toivari et al. 2001). Anoxia kills C. shehatae when it is cultivated on xylose, but not when cultivated on glucose. This suggests that there is a fundamental difference in the oxygen requirements for the metabolism of these two sugars (Kastner et al. 1999).
Disruption of CYC1 in P. stipitis blocks much of its capacity for ATP production and results in a petite colony morphology, even though this is a petite-negative yeast. P. stipitis Δcyc1 mutants grow slowly on glucose or xylose, but unlike wild-type P. stipitis, they do not grow on glycerol or xylitol. Volumetric ethanol production rates of P. stipitis Δcyc1 mutants on xylose are similar to the parent, but because cell yields are much lower, the specific fermentation rate is about 50% higher with Δcyc1 mutants (Shi et al. 1999). Loss of CYC1 restricts cell growth but respiration capacity remains high because P. stipitis also possesses an alternative salicylhydroxamic acid-sensitive terminal oxidase, STO1 (Shi et al. 2002). Disruption of STO1 increases ethanol production from xylose but does not affect cell growth, which indicates that electron transfer to Sto1 does not generate ATP.
Anaerobiosis
One of the biggest challenges for commercialization of the yeast xylose fermentation is obtaining growth on xylose under anaerobic conditions. The cost of aerating bioreactors for ethanol production is prohibitive by current practice. As was first noted by Wang and Schneider in 1980, S. cerevisiae and other yeasts can grow on xylulose under aerobic conditions (Wang and Schneider 1980), but even though they can ferment xylulose, they do not grow on this sugar under anaerobic conditions (Maleszka and Schneider 1984). Mitochondrial function appears to be necessary for the growth of yeasts on xylose or xylulose. Ethanol production from xylose, xylitol or xylulose is enhanced by aeration in most yeasts (Maleszka and Schneider 1982); and in some species, such as C. tropicalis, aeration is essential for ethanol production (Jeffries 1981).
Eliasson et al. (2000b) were the first to report the anaerobic production of ethanol from xylose by recombinant S. cerevisiae TMB3001. However, this yeast grew on xylose only in the presence of oxygen; and glucose was included in all fermentations to enable continuous cultivation. Xylose was co-utilized with glucose under anaerobic conditions. Xylose uptake varied inversely with its concentration, but even at the highest xylose concentration, (15 g xylose l−1, 5 g glucose l−1) only 12% of the xylose was consumed. The anaerobic ethanol yield on xylose was estimated at 0.21 g g−1, assuming a constant ethanol yield on glucose. Glucose appears to be required for the anaerobic metabolism of xylose, because these authors were not able to maintain a steady state when cultures were grown on xylose alone (Eliasson et al. 2000b).
To overcome this limitation, Sonderegger and Sauer (2003) maintained S. cerevisiae TMB3001 in continuous culture under progressively more restrictive oxygen limitations. Starting from continuous aerobic cultivation on a mixture of xylose and glucose and progressing to anaerobic cultivation on xylose alone, the authors obtained two cell populations. Clones taken from the larger population grew anaerobically on xylose but showed impaired growth on glucose. Clones taken from the smaller population were incapable of anaerobic growth but produced more ethanol from xylose than the parental strain.
In a separate research effort, Wahlbom et al. (2003) also used continuous cultivation under aerobic, oxygen-limited and anaerobic conditions to obtain improved mutants of recombinant S. cerevisiae with higher capacities for xylose fermentation. However, even the best of these mutants showed only about one-third of the aerobic maximum growth rate and two-thirds of the ethanol productivity obtained with P. stipitis CBS 6054 on xylose (Wahlbom et al. 2003).
C. shehatae can use glucose and xylose simultaneously in a chemostat, but aeration is required for cell growth (Kastner et al. 1998). C. shehatae cells, cultivated aerobically on d-glucose and d-xylose, undergo one doubling or less following a shift to anoxia. Cell viability declines nine times faster in d-xylose than in d-glucose fermentations. Anaerobic growth does not occur on either d-glucose or d-xylose (Kastner et al. 1999).
Heterologous expression of S. cerevisiae URA1 in P. stipitis was reported to confer anaerobic growth, but this apparently required an uncharacterized mutational event in the host cell background, because transformed cells required more than 100 h before initial growth was noted (Shi and Jeffries 1998).
For commercial purposes, it may be possible to cultivate either native or engineered yeasts anaerobically on glucose followed by a respiro-fermentative phase on the residual xylose. Glucose and xylose are almost always obtained as mixtures from lignocellulose hydrolysates, so by properly engineering cell recycle loops, it should be possible to obtain high-yield conversions under anaerobic or oxygen-limited conditions.
Metabolite flux and transcriptome profiling
Metabolic flux analysis and flux estimates based on 13C-labeling experiments (Christensen et al. 2002) have been used to estimate metabolite levels of S. cerevisiae and P. stipitis grown on glucose and xylose. Intracellular metabolite levels are higher in industrial yeast strains than in laboratory strains engineered for xylose utilization (Zaldivar et al. 2002). P. stipitis derives at least 58% of its phospho-enol-pyruvate (PEP) through the non-oxidative PPP, whereas S. cerevisiae uses the non-oxidative PPP for the biosynthesis of less than 4% of its PEP (Fiaux et al. 2003). A flux balance analysis (FBA) showed that the maximum ethanol yield from xylose in yeast is 0.46 g g−1 rather than 0.51 g g−1, because of the cofactor difference between XR and XDH. Metabolic FBA also predicted that there is an optimal aeration level for ethanol production and that xylitol accumulation decreases with higher aeration. Both of these predictions have been confirmed by measuring product yields at various aeration rates (Jin 2002).
Transcriptome profiling methods, such as RT-PCR and microarray experiments, were applied to monitoring the differential expression of genes between glucose and xylose fermentation by recombinant S. cerevisiae. Of the 5,944 genes detected under oxygen-limited culture conditions, 386 (6.6%) showed differential expression when cells were grown on xylose, as compared with glucose. As expected, most of those genes fell in the energy-production category. Expression levels of genes coding for glycolytic, fermentative and pentose phosphate enzymes did not change greatly. However, expression of the genes encoding tricarboxylic acid cycle and respiratory enzymes greatly increased when cells were grown on xylose (Jin 2002).
Protoplast fusion
Protoplast fusion is widely used to improve the fermentative properties of industrial yeasts. By complementing multiple auxotrophic markers, it is possible to obtain stable hybrids between closely related species. In an attempt to increase yeast strains with ethanol tolerance and the ability to ferment xylose, S. cerevisiae has been fused with auxotrophic strains of C. shehatae or P. stipitis (Gupthar 1992), but mononucleate fusants quickly segregate into their parental type strains (Yoon et al. 1996). Other researchers report fusants between S. cerevisiae and P. stipitis that show the capacity for xylose fermentation and an ability to ferment glucose in the presence of 6% ethanol (Kordowska-Wiater and Targonski 2001). Given the instability of the hybrids, this will probably not lead to commercial yeast strains.
Xylitol production
Native xylose-metabolizing yeasts, such as Pachysolen tannophilus, C. shehatae, (Du Preez et al. 1984), C. boidinii (Vandeska et al. 1996; Winkelhausen et al. 1996), Hansenula polymorpha (Sanchez et al. 1998) and C. guillermondii (Rodrigues et al. 2002), all produce xylitol to greater or lesser degrees depending on the pH, oxygen availability and other culture conditions. Pichia stipitis is notable because it produces relatively little xylitol (Sanchez et al. 2002). When its genes for Adh are disrupted, xylitol production increases dramatically (Cho and Jeffries 1998); and when the d-xylulokinase gene is disrupted, P. stipitis produces a mixture of xylitol and arabitol (Jin et al. 2002). S. cerevisiae cells engineered for xylose utilization tend to produce xylitol (Kötter and Ciriacy 1993; Hallborn et al. 1994; Tantirungkij et al. 1994; Walfridsson et al. 1995; Eliasson et al. 2000b). If many copies of XYL1 are integrated into the genome, S. cerevisiae transformants are stable and can produce xylitol in sequential batch or continuous culture (Kim et al. 1999). By increasing the expression of XYL2 relative to XYL1, it is possible to decrease xylitol secretion (Jin et al. 2003).
Arabinose utilization
The utilization of l-arabinose is particularly important in the conversion of corn hulls to ethanol (Saha et al. 1998). Corn fiber consists of about 20% starch, 14% cellulose and 35% hemicellulose; and l-arabinose makes up approximately 28% of the hemicellulosic fraction (Park et al. 2001). An extensive screen of 116 yeasts that can grow on l-arabinose showed that four strains, C. auringiensis, C. succiphila, Ambrosiozyma monospora and Candida sp. (YB-2248) could produce some ethanol (4.1 g l−1 or less) directly from arabinose (Dien et al. 1996). While the production rates were very low, these studies showed for the first time that yeasts can directly convert l-arabinose to ethanol (Fig. 2).
The pathways for l-arabinose and d-arabinose metabolism are distinct in yeasts. P. stipitis will grow very slowly on l-arabinose, but it does not ferment this sugar. A mutant of P. stipitis that was unable to metabolize l-arabinose could grow on d-arabinose (Shi et al. 2000). Complementation of this mutant with XYL2 restored growth on l-arabinose. This showed that the pathway used by P. stipitis for l-arabinose metabolism is similar to that used by Aspergillus niger (Witteveen et al. 1989).
Relatively little is known about l-arabinose uptake by yeasts. In C. shehatae, it appears to be mediated by proton symport (Lucas and van Uden 1986). l-Arabinose is similar in structure to d-galactose (Rees 1977) and its transport in S. cerevisiae is mediated by GAL2 (Kou et al. 1970). In Kluyveromyces lactis, the LAC12 gene codes for an inducible lactose permease that is similar in structure to the Escherichia coli xylose-H+ and arabinose-H+ transporters (Chang and Dickson 1988). d-Arabinitol dehydrogenase (Hallborn et al. 1995) functions in an alternative pathway that connects d-xylulose to d-ribulose (Jin et al. 2002).
Fungi metabolize l-arabinose through five enzymes, aldose (xylose) reductase, l-arabinitol 4-dehydrogenase (lad1; Richard et al. 2001), l-xylulose reductase (lxr1; Richard et al. 2002), xylitol dehydrogenase and d-xylulokinase. Overexpression of lad1 and lxr1 along with XYL1, XYL2 and XKS1 enabled S. cerevisiae to grow on and ferment l-arabinose (Richard et al. 2003). Ethanol production occurred at a very low rate. About 0.1 g of ethanol was formed by 4 g of cells in 70 h under anaerobic conditions. Under aerobic conditions, the ethanol formed from l-arabinose would probably be re-assimilated.
In a different approach to engineer S. cerevisiae for l-arabinose fermentation, Sedlak and Ho (2001) expressed three genes of the araBAD operon from E. coli in S. cerevisiae. They reported activity with all three enzymes, but the transformant did not produce ethanol from l-arabinose (Sedlak and Ho 2001). Becker and Boles (2003) were more successful in this approach. Unlike Sedlak and Ho, they were not able to obtain activity through the heterologous expression of E. coli araA, which codes for l-arabinose isomerase, but they were able to express the araA gene from Bacillus subtilis and this—along with the heterologous expression of E. coli araB and araD plus overexpression of S. cerevisiae GAL2—gave rise to a yeast strain that could grow slowly on l-arabinose. After more than 200 h of cultivation, a transformant arose that could grow on l-arabinose, with a doubling time of about 8 h. They were able to identify two mutational events—one in the bacterial l-ribulokinase that reduced affinity for l-ribulose and one in the yeast genome that increased transaldolase expression. Together, these enabled growth on and fermentation of l-arabinose. The resulting strain produced up to 0.08 g ethanol g−1 biomass h−1. This represents a major breakthrough in the metabolic engineering of arabinose metabolism in yeast.
Cellulase and xylanase expression
Native strains of P. stipitis produce xylanases (Ozcan et al. 1991) that enable the fermentation of xylan directly to ethanol, but the yields are very low (Lee et al. 1986). By increasing xylanase production either through mutation (Basaran et al. 2000) or heterologous expression (Morosoli et al. 1993; Den Haan and Van Zyl 2001), it is possible to enhance the xylan fermentation rate. Yeast β-xylosidase is probably most important for the fermentation of xylobiose and xylotriose, because these are the most conspicuous products of endoxylanase xylanase activity and they are also formed during the acid hydrolysis of xylan. Several researchers have heterologously expressed xylanases in S. cerevisiae (Den Haan and Van Zyl 2001; La Grange et al. 2001).
Spent sulfite and hydrolysate fermentation
In the final analysis, the ability of an organism to ferment sugars in hemicellulosic hydrolysates determines the success of metabolic engineering efforts. Acetic acid and toxic phenolic products from lignocellulose can inhibit growth of yeasts in hydrolysates. However, post-hydrolysis treatments can reduce toxicity and strains can be selected for resistance. The most cost-effective hydrolysate treatment—calcium over-liming—is also one of the oldest. Over-liming is used to prepare sulfite waste liquors (Nigam 2001b) and acid hydrolysates of hardwood (Nigam 2001a). Unfortunately, acid hydrolysis and over-liming produce large amounts of calcium sulfate that must be removed. The toxicity of hydrolysates can be significantly reduced by adding laccase to polymerize the free phenolic compounds (Jönsson et al. 1998). A laccase cloned from Trametes versicolor has been expressed at a high level in S. cerevisiae (Larsson et al. 2001). S. cerevisiae TMB 3001 grew better in laccase-treated hydrolysate than in untreated hydrolysate, but it showed almost no utilization of xylose. In a comparison of P. stipitis with C. shehatae, the latter yeast was better able to ferment hydrolysates (Sreenath and Jeffries 2000). In simultaneous saccharification and fermentation, yields of 0.47 g g−1 substrate were obtained (Sreenath et al. 1999, 2001). Wild-type strains of C. shehatae can ferment rice straw autohydrolysates to ethanol with yields of 0.45 g ethanol g−1 sugar consumed and they can produce 0.37 g g−1 from acid prehydrolysates (Abbi et al. 1996). The ethanol productivity of P. stipitis in wood hydrolysates can be improved by up to 2-fold by selecting for resistant strains (Nigam 2001a).
Prospects for future progress
Metabolic engineering enables S. cerevisiae to ferment d-xylose and l-arabinose to ethanol and it improves the capacity of native xylose-fermenting yeasts, such as P. stipitis. The improvement obtained with any one change has been incremental. Co-production of xylitol, low ethanol production rates, requirements for oxygen and co-metabolizable carbon sources remain problems with recombinant S. cerevisiae. Most trials use haploid laboratory strains rather than industrial yeasts for their genetic backgrounds and trials with mixed sugar hydrolysates are not often reported. Mutagenesis and strain selection has improved xylose utilization in recombinant strains, but most mutations have not been characterized. In some instances, multiple genes have been altered through deletion or overexpression, but rarely have expression levels been manipulated. There are, therefore, many opportunities to obtain further improvements by learning more about the factors that limit xylose utilization under anaerobic conditions, by selecting better genetic backgrounds for heterologous expression, by expressing multiple genes at optimal levels and by combining the various beneficial traits into single strains. While strain improvement will probably continue for several years, ethanol production rates and yields are becoming practicable for some commercial applications.
References
Abbi M, Kuhad RC, Singh A (1996) Fermentation of xylose and rice straw hydrolysate to ethanol by Candida shehatae NCL-3501. J Ind Microbiol 17:20–23
Aguilar R, Ramirez JA, Garrote G, Vazquez M (2002) Kinetic study of the acid hydrolysis of sugar cane bagasse. J Food Eng 55:309–318
Amore R, Kötter P, Kuster C, Ciriacy M, Hollenberg CP (1991) Cloning and expression in Saccharomyces cerevisiae of the NAD(P)H-dependent xylose reductase-encoding gene (XYL1) from the xylose-assimilating yeast Pichia stipitis. Gene 109:89–97
Aristidou A, Penttilä M (2000) Metabolic engineering applications to renewable resource utilization. Curr Opin Biotechnol 11:187–198
Bao X, Gao D, Qu Y, Wang Z, Walfridssion M, Hahn-Hägerdal B (1997) Effect on product formation in recombinant Saccharomyces cerevisiae strains expressing different levels of xylose metabolic genes. Chin J Biotechnol 13:225–231
Basaran P, Basaran N, Hang YD (2000) Isolation and characterization of Pichia stipitis mutants with enhanced xylanase activity. World J Microbiol Biotechnol 16:545–550
Becker J, Boles E (2003) A modified Saccharomyces cerevisiae strain that consumes l-arabinose and produces ethanol. Appl Environ Microbiol 69:4144–4150
Billard P, Menart S, Fleer R, Bolotin-Fukuhara M (1995) Isolation and characterization of the gene encoding xylose reductase from Kluyveromyces lactis. Gene 162:93–97
Boles E, Hollenberg CP (1997) The molecular genetics of hexose transport in yeasts. FEMS Microbiol Rev 21:85–111
Bruinenberg PM (1986) The NADP(H) redox couple in yeast metabolism. Antonie Van Leeuwenhoek 52:411–429
Bruinenberg PM, Debot PHM, Dijken JP van, Scheffers WA (1983a) The role of redox balances in the anaerobic fermentation of xylose by yeasts. Eur J Appl Microbiol Biotechnol 18:287–292
Bruinenberg PM, Dijken JP van, Scheffers WA (1983b) A theoretical analysis of NADPH production and consumption in yeasts. J Gen Microbiol 129:953–964
Bruinenberg PM, Debot PHM, Dijken JP van, Scheffers WA (1984) NADH-linked aldose reductase—the key to anaerobic alcoholic fermentation of xylose by yeasts. Appl Microbiol Biotechnol 19:256–260
Bruinenberg PM, Jonker R, Dijken JP van, Scheffers WA (1985) Utilization of formate as an additional energy source by glucose-limited chemostat cultures of Candida utilis CBS-621 and Saccharomyces cerevisiae CBS-8066—evidence for the absence of transhydrogenase activity in yeasts. Arch Microbiol 142:302–306
Buziol S, et al (2002) Determination of in vivo kinetics of the starvation-induced Hxt5 glucose transporter of Saccharomyces cerevisiae. FEMS Yeast Res 2:283–291
Chang SF, Ho NW (1988) Cloning the yeast xylulokinase gene for the improvement of xylose fermentation. Appl Biochem Biotechnol 17:313–318
Chang YD, Dickson RC (1988) Primary structure of the lactose permease gene from the yeast Kluyveromyces lactis—presence of an unusual transcript structure. J Biol Chem 263:16696–16703
Chen RF, Wu ZW, Lee YY (1998) Shrinking-bed model for percolation process applied to dilute-acid pretreatment hydrolysis of cellulosic biomass. Appl Biochem Biotechnol 70/72:37–49
Cho JY, Jeffries TW (1998) Pichia stipitis genes for alcohol dehydrogenase with fermentative and respiratory functions. Appl Environ Microbiol 64:1350–1358
Cho JY, Jeffries TW (1999) Transcriptional control of ADH genes in the xylose-fermenting yeast Pichia stipitis. Appl Environ Microbiol 65:2363–2368
Christensen B, Gombert AK, Nielsen J (2002) Analysis of flux estimates based on (13)C-labelling experiments. Eur J Biochem 269:2795–2800
Claassen PAM, et al (1999) Utilisation of biomass for the supply of energy carriers. Appl Microbiol Biotechnol 52:741–755
Dahn KM, Davis BP, Pittman PE, Kenealy WR, Jeffries TW (1996) Increased xylose reductase activity in the xylose-fermenting yeast Pichia stipitis by overexpression of XYL1. Appl Biochem Biotechnol 57/58:267–276
Den Haan R, van Zyl WH (2001) Differential expression of the Trichoderma reesei beta-xylanase II (xyn2) gene in the xylose-fermenting yeast Pichia stipitis. Appl Microbiol Biotechnol 57:521–527
Deng XX, Ho NW (1990) Xylulokinase activity in various yeasts including Saccharomyces cerevisiae containing the cloned xylulokinase gene. Appl Biochem Biotechnol 24/25:193–199
Dequin S (2001) The potential of genetic engineering for improving brewing, wine making and baking yeasts. Appl Microbiol Biotechnol 56:577–588
Diderich JA, Schuurmans JM, Van Gaalen MC, Kruckeberg AL, Van Dam K (2001) Functional analysis of the hexose transporter homologue HXT5 in Saccharomyces cerevisiae. Yeast 18:1515–1524
Dien BS, Kurtzman CP, Saha BC, Bothast RJ (1996) Screening for l-arabinose fermenting yeasts. Appl Biochem Biotechnol 57/58:233–242
Does AL, Bisson LF (1989) Characterization of xylose uptake in the yeasts Pichia heedii and Pichia stipitis. Appl Environ Microbiol 55:159–164
Du Preez JC, Walt JP van der (1983) Fermentation of d-xylose to ethanol by a strain of Candida shehatae. Biotechnol Lett 5:357–362
Du Preez JC, Prior BA, Monteiro AMT (1984) The effect of aeration on xylose fermentation by Candida shehatae and Pachysolen tannophilus—a comparative study. Appl Microbiol Biotechnol 19:261–266
Du Preez JC, Bosch M, Prior BA (1986) Xylose fermentation by Candida shehatae and Pichia stipitis—effects of pH, temperature and substrate concentration. Enzyme Microb Technol 8:360–364
Eliasson A, et al (2000a) Xylulose fermentation by mutant and wild-type strains of Zygosaccharomyces and Saccharomyces cerevisiae. Appl Microbiol Biotechnol 53:376–382
Eliasson A, Christensson C, Wahlbom CF, Hahn-Hägerdal B (2000b) Anaerobic xylose fermentation by recombinant Saccharomyces cerevisiae carrying XYL1, XYL2, and XKS1 in mineral medium chemostat cultures. Appl Environ Microbiol 66:3381–3386
Fiaux J, Cakar ZP, Sonderegger M, Wuthrich K, Szyperski T, Sauer U (2003) Metabolic flux profiling of the yeasts Saccharomyces cerevisiae and Pichia stipitis. Eukaryot Cell 2:170–180
Flores CL, Rodriguez C, Petit T, Gancedo C (2000) Carbohydrate and energy-yielding metabolism in non-conventional yeasts. FEMS Microbiol Rev 24:507–529
Galbe M, Zacchi G (2002) A review of the production of ethanol from softwood. Appl Microbiol Biotechnol 59:618–628
Gong CS, Cao NJ, Du J, Tsao GT (1999) Ethanol production from renewable resources. Adv Biochem Eng Biotechnol 65:207–241
Gupthar AS (1992) Segregation of altered parental properties in fusions between Saccharomyces cerevisiae and the d-xylose fermenting yeasts Candida shehatae and Pichia stipitis. Can J Microbiol 38:1233–1237
Hahn-Hägerdal B, Wahlbom CF, Gardonyi M, Zyl WH van, Cordero Otero RR, Jönsson LJ (2001) Metabolic engineering of Saccharomyces cerevisiae for xylose utilization. Adv Biochem Eng Biotechnol 73:53–84
Hallborn J, et al (1991) Xylitol production by recombinant Saccharomyces cerevisiae. Biotechnology 9:1090–1095
Hallborn J, Gorwa MF, Meinander N, Penttilä M, Keranen S, Hahn-Hägerdal B (1994) The influence of cosubstrate and aeration on xylitol formation by recombinant Saccharomyces cerevisiae expressing the XYL1 gene. Appl Microbiol Biotechnol 42:326–333
Hallborn J, Walfridsson M, Penttilä M, Keranen S, Hahn-Hägerdal B (1995) A short-chain dehydrogenase gene from Pichia stipitis having d-arabinitol dehydrogenase activity. Yeast 11:839–847
Hamacher T, Becker J, Gardonyi M, Hahn-Hägerdal B, Boles E (2002) Characterization of the xylose-transporting properties of yeast hexose transporters and their influence on xylose utilization. Microbiology 148:2783–2788
Handumrongkul C, Ma DP, Silva JL (1998) Cloning and expression of Candida guilliermondii xylose reductase gene (xyl1) in Pichia pastoris. Appl Microbiol Biotechnol 49:399–404
Harhangi HR, et al (2003) Xylose metabolism in the anaerobic fungus Piromyces sp. strain E2 follows the bacterial pathway. Arch Microbiol 180:134–141
Hespell RB (1998) Extraction and characterization of hemicellulose from the corn fiber produced by corn wet-milling processes. J Agric Food Chem 46:2615–2619
Hinmann ND, Wright JD, Hoagland W, Wyman CE (1989) Xylose fermentation—an economic analysis. Appl Biochem Biotechnol 20/21:391–401
Ho NW, Tsao GT (1993) Recombinant yeasts for effective fermentation of glucose and xylose. US Patent 5789210
Ho NW, Lin FP, Huang S, Andrews PC, Tsao GT (1990) Purification, characterization, and amino terminal sequence of xylose reductase from Candida shehatae. Enzyme Microb Technol 12:33–39
Ho NW, Chen Z, Brainard AP (1998) Genetically engineered Saccharomyces yeast capable of effective co-fermentation of glucose and xylose. Appl Environ Microbiol 64:1852–1859
Ho NW, Chen Z, Brainard AP, Sedlak M (1999) Successful design and development of genetically engineered Saccharomyces yeasts for effective co-fermentation of glucose and xylose from cellulosic biomass to fuel ethanol. Adv Biochem Eng Biotechnol 65:163–192
Hohmann S, Neves MJ, Koning W de, Alijo R, Ramos J, Thevelein JM (1993) The growth and signalling defects of the ggs1 (fdp1/byp1) deletion mutant on glucose are suppressed by a deletion of the gene encoding hexokinase PII. Curr Genet 23:281–289
Hohmann S, Bell W, Neves MJ, Valckx D, Thevelein JM (1996) Evidence for trehalose-6-phosphate-dependent and -independent mechanisms in the control of sugar influx into yeast glycolysis. Mol Microbiol 20:981–991
Jeffries T (1981) Conversion of xylose to ethanol under aerobic conditions. Biotechnol Lett 3:213–218
Jeffries TW (1982) A comparison of Candida tropicalis and Pachysolen tannophilus for conversion of xylose to ethanol. Biotechnol Bioeng Symp 12:103–110
Jeffries T (1983) Utilization of xylose by bacteria yeasts and fungi. Adv Biochem Eng Biotechnol 27:1–32
Jeffries TW (1985) Emerging technology for fermenting d-xylose. Trends Biotechnol 3:208–212
Jeffries TW, Jin YS (2000) Ethanol and thermotolerance in the bioconversion of xylose by yeasts. Adv Appl Microbiol 2000:221–268
Jeffries TW, Kurtzman CP (1994) Strain selection, taxonomy, and genetics of xylose fermenting yeasts. Enzyme Microb Technol 16:922–932
Jeffries TW, Shi NQ (1999) Genetic engineering for improved xylose fermentation by yeasts. Adv Biochem Eng Biotechnol 65:117–161
Jeppsson M, Johansson B, Hahn-Hägerdal B, Gorwa-Grauslund MF (2002) Reduced oxidative pentose phosphate pathway flux in recombinant xylose-utilizing Saccharomyces cerevisiae strains improves the ethanol yield from xylose. Appl Environ Microbiol 68:1604–1609
Jin YS (2002) Metabolic engineering of xylose fermentation in Saccharomyces cerevisiae. PhD thesis, University of Wisconsin, Madison
Jin YS, Jeffries TW (2003) Changing flux of xylose metabolites by altering expression of xylose reductase and xylitol dehydrogenase in recombinant Saccharomyces cerevisiae. Appl Biochem Biotechnol 105/108:277–285
Jin YS, Jones S, Shi NQ, Jeffries TW (2002) Molecular cloning of XYL3 (d-xylulokinase) from Pichia stipitis and characterization of its physiological function. Appl Environ Microbiol 68:1232–1239
Jin YS, Ni H, Laplaza JM, Jeffries TW (2003) Optimal growth and ethanol production from xylose by recombinant Saccharomyces cerevisiae require moderate d-xylulokinase activity. Appl Environ Microbiol 69:495–503
Johansson B, Christensson C, Hobley T, Hahn-Hägerdal B (2001) Xylulokinase overexpression in two strains of Saccharomyces cerevisiae also expressing xylose reductase and xylitol dehydrogenase and its effect on fermentation of xylose and lignocellulosic hydrolysate. Appl Environ Microbiol 67:4249–4255
Jönsson LJ, Palmqvist E, Nilvebrant NO, Hahn-Hägerdal B (1998) Detoxification of wood hydrolysates with laccase and peroxidase from the white-rot fungus Trametes versicolor. Appl Microbiol Biotechnol 49:691–697
Kastner JR, Jones WJ, Roberts RS (1998) Simultaneous utilization of glucose and d-xylose by Candida shehatae in a chemostat. J Ind Microbiol Biotechnol 20:339–343
Kastner JR, Jones WJ, Roberts RS (1999) Oxygen starvation induces cell death in Candida shehatae fermentations of d-xylose, but not d-glucose. Appl Microbiol Biotechnol 51:780–785
Kheshgi HS, Prince RC, Marland G (2000) The potential of biomass fuels in the context of global climate change: focus on transportation fuels. Annu Rev Energy Environ 25:199–244
Kilian SG, Uden N van (1988) Transport of xylose and glucose in the xylose-fermenting yeast Pichia stipitis. Appl Microbiol Biotechnol 27:545–548
Kim KH, Tucker MP, Keller FA, Aden A, Nguyen QA (2001) Continuous countercurrent extraction of hemicellulose from pretreated wood residues. Appl Biochem Biotechnol 91/93:253–267
Kim YS, Kim SY, Kim JH, Kim SC (1999) Xylitol production using recombinant Saccharomyces cerevisiae containing multiple xylose reductase genes at chromosomal delta-sequences. J Biotechnol 67:159–171
Kordowska-Wiater M, Targonski Z (2001) Application of Saccharomyces cerevisiae and Pichia stipitis karyoductants to the production of ethanol from xylose. Acta Microbiol Pol 50:291–299
Kötter P, Ciriacy M (1993) Xylose fermentation by Saccharomyces cerevisiae. Appl Microbiol Biotechnol 38:776–783
Kötter P, Amore R, Hollenberg CP, Ciriacy M (1990) Isolation and characterization of the Pichia stipitis xylitol dehydrogenase gene, XYL2, and construction of a xylose-utilizing Saccharomyces cerevisiae transformant. Curr Genet 18:493–500
Kou SC, Christensen MS, Cirillo VP (1970) Galactose transport in Saccharomyces cerevisiae. II. Characteristics of galactose uptake and exchange in galactokinaseless cells. J Bacteriol 103:671–678
Krishnan MS, Ho NWY, Tsao GT (1999) Fermentation kinetics of ethanol production from glucose and xylose by recombinant Saccharomyces 1400 (pLNH33). Appl Biochem Biotechnol 77/79:373–388
Kruckeberg AL (1996) The hexose transporter family of Saccharomyces cerevisiae. Arch Microbiol 166:283–292
Kuyper M, et al (2003) High-level functional expression of a fungal xylose isomerase: the key to efficient ethanolic fermentation of xylose by Saccharomyces cerevisiae? FEMS Yeast Res (in press)
La Grange DC, Pretorius IS, Claeyssens M, Zyl WH van (2001) Degradation of xylan to d-xylose by recombinant Saccharomyces cerevisiae coexpressing the Aspergillus niger beta-xylosidase (xlnD) and the Trichoderma reesei xylanase II (xyn2) genes. Appl Environ Microbiol 67:5512–5519
Lagunas R (1993) Sugar transport in Saccharomyces cerevisiae. FEMS Microbiol Rev 104:229–242
Larsson S, Cassland P, Jönsson LJ (2001) Development of a Saccharomyces cerevisiae strain with enhanced resistance to phenolic fermentation inhibitors in lignocellulose hydrolysates by heterologous expression of laccase. Appl Environ Microbiol 67:1163–1170
Lawford HG, Rousseau JD (1993) Production of ethanol from pulp-mill hardwood and softwood spent sulfite liquors by genetically engineered Escherichia coli. Appl Biochem Biotechnol 39:667–685
Lee H, Biely P, Latta RK, Barbosa MFS, Schneider H (1986) Utilization of xylan by yeasts and Its conversion to ethanol by Pichia stipitis strains. Appl Environ Microbiol 52:320–324
Lee WJ, Kim MD, Ryu YW, Bisson LF, Seo JH (2002) Kinetic studies on glucose and xylose transport in Saccharomyces cerevisiae. Appl Microbiol Biotechnol 60:186–191
Ligthelm ME, Prior BA, Du Preez JC (1988) The induction of d-xylose catabolizing enzymes in Pachysolen tannophilus and the relationship to anaerobic d-xylose fermentation. Biotechnol Lett 10:207–212
Lönn A, Gardonyi M, Zyl W van, Hahn-Hägerdal B, Otero RC (2002) Cold adaptation of xylose isomerase from Thermus thermophilus through random PCR mutagenesis. Gene cloning and protein characterization. Eur J Biochem 269:157–163
Lucas C, Uden N van (1986) Transport of hemicellulose monomers in the xylose-fermenting yeast Candida shehatae. Appl Microbiol Biotechnol 23:491–495
Lynd LR (1996) Overview and evaluation of fuel ethanol from cellulosic biomass: technology, economics, the environment, and policy. Annu Rev Energy Environ 21:403–465
Maier A, Volker B, Boles E, Fuhrmann GF (2002) Characterisation of glucose transport in Saccharomyces cerevisiae with plasma membrane vesicles (countertransport) and intact cells (initial uptake) with single Hxt1, Hxt2, Hxt3, Hxt4, Hxt6, Hxt7 or Gal2 transporters. FEMS Yeast Res 2:539–550
Maleszka R, Schneider H (1982) Fermentation of d-xylose, xylitol, and d-xylulose by yeasts. Can J Microbiol 28:360–363
Maleszka R, Schneider H (1984) Involvement of oxygen and mitochondrial function in the metabolism of d-xylulose by Saccharomyces cerevisiae. Arch Biochem Biophys 228. 228:22–30
McMillan JD (1997) Bioethanol production: status and prospects. Renewable Energy 10:295–302
Meinander NQ, Hahn-Hägerdal B (1997) Influence of cosubstrate concentration on xylose conversion by recombinant, XYL1-expressing Saccharomyces cerevisiae: a comparison of different sugars and ethanol as cosubstrates. Appl Environ Microbiol 63:1959–1964
Meinander NQ, Boels I, Hahn-Hägerdal B (1999) Fermentation of xylose/glucose mixtures by metabolically engineered Saccharomyces cerevisiae strains expressing XYL1 and XYL2 from Pichia stipitis with and without overexpression of TAL1. Bioresour Technol 68:79–87
Metzger MH, Hollenberg CP (1994) Isolation and characterization of the Pichia stipitis transketolase gene and expression in a xylose-utilising Saccharomyces cerevisiae transformant. Appl Microbiol Biotechnol 42:319–325
Michal G (1999) Biochemical pathways. Wiley, New York
Moniruzzaman M, et al (1997) Fermentation of corn fibre sugars by an engineered xylose utilizing Saccharomyces yeast strain. World J Microbiol Biotechnol 13:341–346
Morosoli R, Zalce E, Durand S (1993) Secretion of a Cryptococcus albidus xylanase in Pichia stipitis resulting in a xylan fermenting transformant. Curr Genet 24:94–99
Nigam JN (2001a) Development of xylose-fermenting yeast Pichia stipitis for ethanol production through adaptation on hardwood hemicellulose acid prehydrolysate. J Appl Microbiol 90:208–215
Nigam JN (2001b) Ethanol production from hardwood spent sulfite liquor using an adapted strain of Pichia stipitis. J Ind Microbiol Biotechnol 26:145–150
Nissen TL, Schulze U, Nielsen J, Villadsen J (1997) Flux distributions in anaerobic, glucose-limited continuous cultures of Saccharomyces cerevisiae. Microbiology 143:203–218
Olsson L, Hahn-Hägerdal B (1996) Fermentation of lignocellulosic hydrolysates for ethanol production. Enzyme Microb Technol 18:312–331
Ostergaard S, Olsson L, Nielsen J (2000) Metabolic engineering of Saccharomyces cerevisiae. Microbiol Mol Biol Rev 64:34–50
Ozcan S, Kötter P, Ciriacy M (1991) Xylan-hydrolyzing enzymes of the yeast Pichia stipitis. Appl Microbiol Biotechnol 36:190–195
Park NH, Yoshida S, Takakashi A, Kawabata Y, Sun HJ, Kusakabe I (2001) A new method for the preparation of crystalline l-arabinose from arabinoxylan by enzymatic hydrolysis and selective fermentation with yeast. Biotechnol Lett 23:411–416
Passoth V, Cohn M, Schafer B, Hahn-Hagerdal B, Klinner U (2003) Analysis of the hypoxia-induced ADH2 promoter of the respiratory yeast Pichia stipitis reveals a new mechanism for sensing of oxygen limitation in yeast. Yeast 20:39–51
Pettersen RC (1984) The chemical composition of wood. Adv Chem Ser 1984:57–126
Prior BA, Kilian SG, Dupreez JC (1989) Fermentation of d-xylose by the yeasts Candida shehatae and Pichia stipitis—prospects and problems. Process Biochem 24:21–32
Rees DA (1977) Polysaccharide shapes. Wiley, New York
Richard P, Toivari MH, Penttilä M (2000) The role of xylulokinase in Saccharomyces cerevisiae xylulose catabolism. FEMS Microbiol Lett 190:39–43
Richard P, Londesborough J, Putkonen M, Kalkkinen N, Penttilä M (2001) Cloning and expression of a fungal l-arabinitol 4-dehydrogenase gene. J Biol Chem 276:40631–40637
Richard P, Putkonen M, Väänänen R, Londesborough J, Penttilä M (2002) The missing link in the fungal l-arabinose catabolic pathway, identification of the l-xylulose reductase gene. Biochemistry 41:6432–6437
Richard P, Verho R, Putkonen M, Londesborough J, Penttilä M (2003) Production of ethanol from l-arabinose by Saccharomyces cerevisiae containing a fungal l-arabinose pathway. FEMS Yeast Res (in press)
Rizzi M, Erlemann P, Buithanh NA, Dellweg H (1988) Xylose fermentation by yeasts 4. Purification and kinetic studies of xylose reductase from Pichia stipitis. Appl Microbiol Biotechnol 29:148–154
Roca C, Olsson L (2003) Increasing ethanol productivity during xylose fermentation by cell recycling of recombinant Saccharomyces cerevisiae. Appl Microbiol Biotechnol 60:560–563
Rodrigues DC, Da Silva SS, Almeida ESJB, Vitolo M (2002) Xylose reductase activity of Candida guilliermondii during xylitol production by fed-batch fermentation: selection of process variables. Appl Biochem Biotechnol 98/100:875–883
Rodriguez-Pena JM, Cid VJ, Arroyo J, Nombela C (1998) The YGR194c (XKS1) gene encodes the xylulokinase from the budding yeast Saccharomyces cerevisiae. FEMS Microbiol Lett 162:155–160
Saha BC, Bothast RJ (1999) Pretreatment and enzymatic saccharification of corn fiber. Appl Biochem Biotechnol 76:65–77
Saha BC, Dien BS, Bothast RJ (1998) Fuel ethanol production from corn fiber—current status and technical prospects. Appl Biochem Biotechnol 70/72:115–125
Sanchez S, Bravo V, Castro E, Moya AJ, Camacho F (1998) The production of xylitol from d-xylose by fermentation with Hansenula polymorpha. Appl Microbiol Biotechnol 50:608–611
Sanchez S, Bravo V, Castro E, Moya AJ, Camacho F (2002) The fermentation of mixtures Of d-glucose and d-xylose by Candida shehatae, Pichia stipitis or Pachysolen tannophilus to produce ethanol. J Chem Technol Biotechnol 77:641–648
Sarthy AV, McConaughy BL, Lobo Z, Sundstrom JA, Furlong CE, Hall BD (1987) Expression of the Escherichia coli xylose isomerase gene in Saccharomyces cerevisiae. Appl Environ Microbiol 53:1996–2000
Schneider H, Wang PY, Chan YK, Maleszka R (1981) Conversion of d-xylose into ethanol by the yeast Pachysolen tannophilus. Biotechnol Lett 3:89–92
Sedlak M, Ho NWY (2001) Expression of E. coli araBAD operon encoding enzymes for metabolizing l-arabinose in Saccharomyces cerevisiae. Enzyme Microb Technol 28:16–24
Shi NQ, Jeffries TW (1998) Anaerobic growth and improved fermentation of Pichia stipitis bearing a URA1 gene from Saccharomyces cerevisiae. Appl Microbiol Biotechnol 50:339–345
Shi NQ, Davis B, Sherman F, Cruz J, Jeffries TW (1999) Disruption of the cytochrome c gene in xylose-utilizing yeast Pichia stipitis leads to higher ethanol production. Yeast 15:1021–1030
Shi NQ, et al (2000) Characterization and complementation of a Pichia stipitis mutant unable to grow on d-xylose or l-arabinose. Appl Biochem Biotechnol 84-86:201–216
Shi NQ, Cruz J, Sherman F, Jeffries TW (2002) SHAM-sensitive alternative respiration in the xylose-metabolizing yeast Pichia stipitis. Yeast 19:1203–1220
Slininger PJ, Bothast RJ, Van Cauwenberge JE, Kurtzman CP (1982) Conversion of d-xylose to ethanol by the yeast Pachysolen tannophilus. Biotechnol Bioeng 24:371–384
Smil V (1999) Crop residues: agriculture′s largest harvest—crop residues incorporate more than half of the world agricultural phytomass. Bioscience 49:299–308
Smith CA, Rangarajan M, Hartley BS (1991) d-Xylose (d-glucose) isomerase from Arthrobacter strain NRRL B3728. Purification and properties. Biochem J 277:255–261
Sonderegger M, Sauer U (2003) Evolutionary engineering of Saccharomyces cerevisiae for anaerobic growth on xylose. Appl Environ Microbiol 69:1990–1998
Sreenath HK, Jeffries TW (2000) Production of ethanol from wood hydrolyzate by yeasts. Bioresour Technol 72:253–260
Sreenath HK, Koegel RG, Moldes AB, Jeffries TW, Straub RJ (1999) Enzymic saccharification of alfalfa fiber after liquid hot water pretreatment. Process Biochem 35:33–41
Sreenath HK, Koegel RG, Moldes AB, Jeffries TW, Straub RJ (2001) Ethanol production from alfalfa fiber fractions by saccharification and fermentation. Process Biochem 36:1199–1204
Takuma S, et al (1991) Isolation of xylose reductase gene of Pichia stipitis and its expression in Saccharomyces cerevisiae. Appl Biochem Biotechnol 28/29:327–340
Tantirungkij M, Nakashima N, Seki T, Yoshida T (1993) Construction of xylose-assimilating Saccharomyces cerevisiae. J Ferment Bioeng 75:83–88
Tantirungkij M, Seki T, Yoshida T (1994) Genetic improvement of Saccharomyces cerevisiae for ethanol production from xylose. Ann NY Acad Sci 721:138–147
Teusink B, Walsh MC, Dam K van, Westerhoff HV (1998) The danger of metabolic pathways with turbo design. Trends Biochem Sci 23:162–169
Thestrup HN, Hahn-Hägerdal B (1995) Xylitol formation and reduction equivalent generation during anaerobic xylose conversion with glucose as cosubstrate in recombinant Saccharomyces cerevisiae expressing the xyl1 gene. Appl Environ Microbiol 61:2043–2045
Thevelein JM, Hohmann S (1995) Trehalose synthase: guard to the gate of glycolysis in yeast? Trends Biochem Sci 20:3–10
Toivari MH, Aristidou A, Ruohonen L, Penttilä M (2001) Conversion of xylose to ethanol by recombinant Saccharomyces cerevisiae: importance of xylulokinase (XKS1) and oxygen availability. Metab Eng 3:236–249
Toivola A, Yarrow D, Bosch E van den, Dijken JP van, Scheffers WA (1984) Alcoholic fermentation of deuterium-xylose by yeasts. Appl Environ Microbiol 47:1221–1223
Träff KL, Otero Cordero RR, Zyl WH van, Hahn-Hägerdal B (2001) Deletion of the GRE3 aldose reductase gene and its influence on xylose metabolism in recombinant strains of Saccharomyces cerevisiae expressing the xylA and XKS1 genes. Appl Environ Microbiol 67:5668–5674
Vandeska E, Amartey S, Kuzmanova S, Jeffries TW (1996) Fed-batch culture for xylitol production by Candida boidinii. Proc Biochem 31:265–270
Verduyn C, Frank J, Dijken JP van, Scheffers WA (1985a) Multiple forms of xylose reductase in Pachysolen tannophilus CBS 4044. FEMS Microbiol Lett 30:313–317
Verduyn C, Kleef R van, Frank J, Schreuder H, Dijken JP van, Scheffers WA (1985b) Properties of the NAD(P)H-dependent xylose reductase from the xylose-fermenting yeast Pichia stipitis. Biochem J 226:669–677
Verwaal R, Paalman JW, Hogenkamp A, Verkleij AJ, Verrips CT, Boonstra J (2002) HXT5 expression is determined by growth rates in Saccharomyces cerevisiae. Yeast 19:1029–1038
Wahlbom CF, Zyl WH van, Jonsson LJ, Hahn-Hagerdal B, Otero RR (2003) Generation of the improved recombinant xylose-utilizing Saccharomyces cerevisiae TMB 3400 by random mutagenesis and physiological comparison with Pichia stipitis CBS 6054. FEMS Yeast Res 3:319–326
Walfridsson M, Hallborn J, Penttilä M, Keranen S, Hahn-Hägerdal B (1995) Xylose-metabolizing Saccharomyces cerevisiae strains overexpressing the TKL1 and TAL1 genes encoding the pentose phosphate pathway enzymes transketolase and transaldolase. Appl Environ Microbiol 61:4184–4190
Walfridsson M, Bao X, Anderlund M, Lilius G, Bulow L, Hahn-Hägerdal B (1996) Ethanolic fermentation of xylose with Saccharomyces cerevisiae harboring the Thermus thermophilus xylA gene, which expresses an active xylose (glucose) isomerase. Appl Environ Microbiol 62:4648–4651
Walfridsson M, Anderlund M, Bao X, Hahn-Hägerdal B (1997) Expression of different levels of enzymes from the Pichia stipitis XYL1 and XYL2 genes in Saccharomyces cerevisiae and its effects on product formation during xylose utilisation. Appl Microbiol Biotechnol 48:218–224
Wang PY, Schneider H (1980) Growth of yeasts on d-xylulose 1. Can J Microbiol 26:1165–1168
Wang PY, Shopsis C, Schneider H (1980) Fermentation of a pentose by yeasts. Biochem Biophys Res Commun 94:248–254
Weierstall T, Hollenberg CP, Boles E (1999) Cloning and characterization of three genes (SUT1–3) encoding glucose transporters of the yeast Pichia stipitis. Mol Microbiol 31:871–883
Wieczorke R, Krampe S, Weierstall T, Freidel K, Hollenberg CP, Boles E (1999) Concurrent knock-out of at least 20 transporter genes is required to block uptake of hexoses in Saccharomyces cerevisiae. FEBS Lett 464:123–128
Wijsman MR, Bruinenberg PM, Dijken JP van, Scheffers WA (1985) Incapacity for anaerobic growth in xylose-fermenting yeasts. Antonie Van Leeuwenhoek 51:563–564
Winkelhausen E, Pittman P, Kuzmanova S, Jeffries TW (1996) Xylitol formation by Candida boidinii in oxygen limited chemostat culture. Biotechnol Lett 18:753–758
Witteveen CFB, Busink R, Vandevondervoort P, Dijkema C, Swart K, Visser J (1989) l-Arabinose and d-xylose catabolism in Aspergillus niger. J Gen Microbiol 135:2163–2171
Wyman CE (1999) Biomass ethanol: technical progress, opportunities, and commercial challenges. Annu Rev Energy Environ 24:189–226
Yoon G-S, Lee T-S, Kim C, Seo J-H, Ryu Y-W (1996) Characterization of alcohol fermentation and segregation of protoplast fusant of Saccharomyces cerevisiae and Pichia stipitis. J Microbiol Biotechnol 6:286–291
Zaldivar J, et al (2002) Fermentation performance and intracellular metabolite patterns in laboratory and industrial xylose-fermenting Saccharomyces cerevisiae. Appl Microbiol Biotechnol 59:436–442
Zyl C van, Prior BA, Kilian SG, Brandt EV (1993) Role of d-ribose as a cometabolite in d-xylose metabolism by Saccharomyces cerevisiae. Appl Environ Microbiol 59:1487–1494
Zyl WH van, Eliasson A, Hobley T, Hahn-Hägerdal B (1999) Xylose utilisation by recombinant strains of Saccharomyces cerevisiae on different carbon sources. Appl Microbiol Biotechnol 52:829–833
Acknowledgements
The authors thank P. Richard, E. Boles and J. Becker for early access to reviewed manuscripts and to J. Laplaza for useful discussion.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Jeffries, T.W., Jin, YS. Metabolic engineering for improved fermentation of pentoses by yeasts. Appl Microbiol Biotechnol 63, 495–509 (2004). https://doi.org/10.1007/s00253-003-1450-0
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00253-003-1450-0