Abstract
HLA alleles are representative of ethnicities and may play important roles in predisposition to hematological disorders. We analyzed DNA samples for HLA-A, -B, -C, -DRB1, and -DQB1 loci, from 1550 patients and 4450 potential related donors by PCR-SSO (Polymerase chain reaction sequence-specific oligonucleotides) and estimated allele frequencies in donors and patients from 1550 families who underwent bone marrow transplantation (BMT) in Egypt. We also studied the association between HLA allele frequencies and incidence of acute myeloid leukemia, acute lymphoblastic leukemia, and severe aplastic anemia. The most frequently observed HLA class I alleles were HLA- A*01:01 (16.9%), A*02:01 (16.1%), B*41:01 (8.7%), B*49:01 (7.3%), C*06:02 (25.1%), and C*07:01 (25.1%), and the most frequently observed class II alleles were HLA-DRB1*11:01 (11.8%), DRB1*03:01 (11.6%), DQB1*03:01 (27.5%), and DQB1*05:01 (18.9%). The most frequently observed haplotypes were A*33:01~B*14:02 ~ DRB1*01:02 (2.35%) and A*01:01~B*52:01~DRB1*15:01 (2.11%). HLA-DRB1*07:01 was associated with higher AML odds (OR, 1.26; 95% CI, 1.02–1.55; p = 0.030). Only HLA-B38 antigen showed a trend towards increased odds of ALL (OR, 1.52; 95% CI, 1.00–2.30; p = 0.049) HLA-A*02:01, -B*14:02, and -DRB1*15:01 were associated with higher odds of SAA (A*02:01: OR, 1.35; 95% CI, 1.07–1.70; p = 0.010; B*14:02: OR, 1.43; 95% CI, 1.06–1.93; p = 0.020; DRB1*15:01: OR, 1.32; 95% CI, 1.07–1.64; p = 0.011). This study provides estimates of HLA allele and haplotype frequencies and their association with hematological disorders in an Egyptian population.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human leukocyte antigens (HLA) are encoded by the major histocompatibility complex (MHC) on the short arm of chromosome 6 (p21). The human MHC loci are classified into Class I (HLA-A, -B, and -C) and Class II (HLA-DR, -DP, and -DQ) (Gfeller and Bassani-Sternberg 2018). Donor-recipient histocompatibility is critical for the success of allogeneic stem cell transplantation (allo-SCT) (Simpson and Dazzi 2019).
Allo-SCT is the standard of care for many hematological disorders. HLA allele matching plays a key role in the etiology of allogeneic immune responses following allo-SCT, such as graft-versus-host disease (GVHD) (Park and Seo 2012), and significantly impacts the transplant outcome (Petersdorf 2017). The most used source of hematopoietic stem cells is bone marrow (BM). However, peripheral blood stem cell harvesting is the common practice in our BMT unit due to the relatively easier and safer procedure compared to BM harvesting (Mahmoud et al. 2008).
Since there is no donor registry in Egypt, therefore, allo-SCT transplants are only performed from related donors -chiefly siblings and parents. Unfortunately, about 50% of our patients do not find a fully matched HLA donor. Haploidentical stem cell transplants (Haplo-SCT) were recently introduced into practice (Chang et al. 2021). However; matched unrelated donor (MUD) transplants have better outcomes and are usually the preferred alternative to Haplo-SCT (Gooptu et al. 2021).
HLA polymorphic regions have functional significance for disease susceptibility due to their association with peptide-binding grooves. Certain HLA antigens and alleles are linked to the pathophysiology and outcomes of hematological disorders (Kazemi et al. 2021) and could be critical factors for disease predisposition. Abnormal HLA expression is associated with the occurrence of several hematological disorders such as lymphomas, acute and chronic leukemias, severe aplastic anemia (SAA), hemoglobinopathies (e.g., B-thalassemia, sickle cell disease), and congenital immunodeficiencies (Villemagne et al. 2005; Huang et al. 2012; Gragert et al. 2014; Zaimoku et al. 2017).
The frequencies of HLA alleles and haplotypes are unique to each ethnic group and their diversity reflects the extent of variation between different populations. The information about the frequencies and distributions of HLA alleles and haplotypes among Egyptians is currently limited (Elshakankiry et al. 2017). In this study, we retrospectively collected HLA typing data performed on HLA- loci A, B, C, DRB1, and DQB1 for patients and donors at the Histocompatibility Laboratory of Nasser Institute for Health and Research (NIH) and estimated HLA allele and haplotype frequencies in Egyptians in an attempt to establish a National stem cell donor registry, which will help transplant candidates to find a compatible stem cell donor. Furthermore, we studied the association between HLA alleles and incidence of the most frequently allografted hematological disorders in our BMT Unit; mainly acute myeloid leukemia (AML), acute lymphoblastic leukemia (ALL), and severe aplastic anemia (SAA) to unravel risky HLA alleles for each hematological disorder.
Patients and methods
Patients and donors’ selection
Between 2016 and 2022, we received peripheral blood samples from 1550 allo-SCT candidates with various hematological disorders -including hematological malignancies, bone marrow failure syndromes, primary immunodeficiencies, and hemoglobinopathies (Table 1). We also typed blood samples from 4450 potential donors.
DNA extraction and HLA typing method
DNA was extracted using the QiAamp Blood Mini Kit® (Qiagen, Hilden, Germany) according to the manufacturer's protocol. HLA-A, -B, -C, -DRB1, and -DQB1 were typed using the polymerase chain reaction sequence-specific oligonucleotide probes (PCR-SSO). The detection method is based on PCR amplification principles (exons 1–4 of HLA-A locus, exons 2–4 of HLA -B locus, exon 2–3 of HLA-C locus, exon 2 of HLA-DRB1 locus, and exon 2–3 of HLA-DQB1 locus). Amplified DNA was then hybridized with specific oligonucleotide probes immobilized as parallel lines on membrane-based strips. HLA allele polymorphisms were genotyped using INNO-LIPA (FUJIREBIO Europe N.V. Gent, Belgium). The INNO-LIPA kit detects the HLA allele groups listed in IMGT/HLA version reports 3.52. The results were analyzed by using the LiPA interpretation software LIRAS.
Statistical analysis and modeling
For each locus, only fully typed subjects were included in the analysis. Antigen and allele frequencies were estimated by direct counting and were estimated for each locus and the studied subgroups (i.e. HLA-A, -B, -C, -DRB1, and -DQB1 in patients with ALL, AML, SAA, and donors independently). Standard errors were calculated as the square root of AF(1-AF)/na, where AF is the allele frequency, and na is the number of alleles given the locus and the subgrou: Antigen and allele frequencies were calculated using R software (R Core Team 2022) version 4.2.0.
Haplotype frequencies were estimated in subjects with full 4-digit typing of HLA-A, -B, and -DRB1. Haplotype phasing and haplotype frequency were estimated using the expectation–maximization algorithm implemented in Python for Population Genomics (PyPOP) software (Lancaster et al. 2007) version 1.0.0b7, -which was installed using the package installer for Python package management system (pip) version 23.2.1 and executed in Python (Van Rossum and Drake 2009) version 3.11.5-. Haplotype frequencies standard errors were calculated as the square root of HF(1-HF)/2N, where HF is the haplotype frequency, and N is the number of subjects with complete haplotype data. Haplotype frequencies were estimated in donors and patients and the frequencies were compared using Fisher’s exact test.
A univariate multinomial regression analysis was performed to determine the association between HLA antigens, alleles, and the odds of ALL, AML, and SAA, taking donors as the reference control grou: All additive, dominant, and recessive genetic models were considered, and all the odds ratios were age- and sex-adjusted. The regression analysis was performed using the ‘nnet’ package (Venables and Ripley 2002) version 7.3–17 in R software. The alleles included in the regression analysis satisfied the following conditions: (1) A frequency of at least 0.03 in all the patient groups and donors, (2) A count of at least 5 in all of the selected patient subgroups and 5 in donors, (3) For the dominant and recessive models, for each antigen group/allele, there must be at least 5 heterozygous patients and 5 heterozygous donors for the dominant model, 5 homozygous patients and 5 homozygous donors for the recessive model (the latter condition was not applied for the additive model, as the genotypes are coded and analyzed as numerical variables). P-values were adjusted for false-detection rate (FDR) using Benjamini-Hochberg’s procedure. Missing data were omitted from the analysis.
Tests for deviations from the Hardy-Weinberg equilibrium (HWE) were performed using Guo and Thomson’s Chi-squared test (Guo and Thompson 1992), and testing for selective neutrality was performed using Slatkin’s test (Slatkin 1994). Both tests were performed for each locus in donors using PyPO: Two-locus pairwise linkage disequilibrium (LD) analysis was performed using the same package. Haplotypes were compared between donors and recipients in each disease using Fisher’s exact test, and p-values were also adjusted for FDR.
Allele frequencies of Egyptian donors were compared to other populations using the package ‘midasHLA’ (Hammer and Migdał, 2022) version 1.4.0 in R, which fetches allele frequency data from the Allele frequency net database (AFND, www.allelefrequencies.net). We included only alleles of frequencies > 0.03 and populations with missing frequency data of no more than 5%. We used agglomerative clustering to examine the ancestral relation of the included populations, with correlation as a measure of genetic distance, and Ward’s method for clustering. Allele frequencies were transformed using the arcsine square root transformation before clustering. Heatmaps were plotted using the package ‘pheatmap’ (Kolde 2019) version 1.0.12.
Results
Demographics, antigen, and allele frequencies
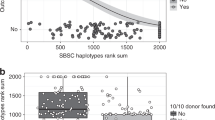
Patients represented 26% of the whole cohort (1550/6000). The median age of donors was 30 years, and the median age of patients was 27 years. The most frequent hematological conditions eligible for allogeneic stem cell transplantation (allo-SCT) were AML (30%), SAA (28%), and ALL (17%). Fifty-three percent of patients had at least one fully matched donor per family, and 1.9% had more than one fully matched donor per family. Siblings represented 87% of potential donors while only 7.2% were parents (Table 1).
Patients born to consanguineous parents represented 23.4% of study population. The highest rates of consanguinity were observed among families of patients with Fanconi anemia (83%) and Beta-Thalassemia (67%). MDS patients were the most likely to find a fully matched donor (66.1% match probability), while patients with inherited immunodeficiency syndromes were the least likely (15.4%) (Table 2).
HLA antigen and allele frequencies in donors
For class I loci, we observed 24 unique HLA-A antigens, 37 -B antigens, and 14 -C antigens. The most frequent HLA-A antigens were A2 (23.2%), A1 (18.2%), and A30 (9.28%). For HLA-B, the most frequent were B41 (10.7%), B35 (9.47%), and B14 (7.87%). For HLA-C the most frequent antigens were Cw7 (35.7%), Cw6 (20.7%), and Cw4 (16.6%). For HLA class II loci, we observed 13 unique DR antigens and 5 unique DQ antigens; the most frequent were DR13 (16.9%), DR4 (16.9%), and DR3 (14.2%). For HLA-DQ, the most frequent antigens were DQ3 (36.2%), DQ6 (25.8%), and DQ5 (20.6%) (ST1).
For HLA class I alleles, we observed 52 unique -A alleles, 76 -B alleles, and 24 -C alleles; the most frequent A alleles were A*01:01 (16.9%), A*02:01 (16.1%), and A*03:01 (7.31%); the most frequent B alleles were B*41:01 (8.71%), B*49:01 (7.31%), and B*14:02 (6.81%), and the most frequent HLA-C alleles were C*06:02 (25.1%), C*07:01 (25.1%), and C*04:01 (17.1%). For HLA class II alleles, we observed 37 DRB1 alleles and 14 DQB1 alleles; the most frequent DRB1 alleles were DRB1*11:01 (11.8%), DRB1*03:01 (11.6%), and DRB1*13:01 (11.4%), and the most frequent DQB1 alleles were DQB1*03:01 (27.5%), DQB1*05:01 (18.9%), and DQB1*06:01 (13.8%) (Table 3). Agglomerative hierarchical clustering suggested a distant ancestry between Egyptian donors and the common ancestor of Tunisians and Saudis (Fig. 1).
Hardy-Weinberg equilibrium and selective neutrality testing in donors
Except for HLA-DQ, there was a significant deviation from expected genotype frequencies under HWE in all loci. Additionally, excessive homozygosity was observed in all loci. Slatkin's test showed that there was a significant departure from the infinite allele model in the HLA-B and HLA-DRB1 loci, with negative Fnd values suggesting balancing selection. Furthermore, the expected homozygosity under HWE was significantly lower than expected under the infinite allele model (Table 4).
HLA antigen and allele frequencies in patients
The most common HLA class I antigens in ALL patients were A2 (22.2%), B41 (10.7%), and Cw7 (37.7%). The most common alleles were A*02:01 (17.5%), B*41:01 (8.3%), C*04:01 (19.2%) and C*07:01 (19.2%). The most common HLA class II antigens were DR13 (20.2%), DQ3 (25%), and DQ6 (25%), while the most common alleles were DRB1*13:01 (13.6%) and DQB1*04:02 (18.9%) (ST2 and Table 5).
In AML patients, the most common HLA class I antigens were A2 (22.8%), B41 (11.3%), and Cw7 (32.4%). The most common alleles were A*01:01 (19.5%), B*41:01 (9.6%), C*04:01 (23%), and C*06:02 (23%). The most common HLA class II antigens were DR13 (18.2%), DQ2 (30%), DQ3 (30%), and DQ4 (30%), while the most common alleles were DRB1*07:01 (12.9%), DQB1*02:01 (30%) and DQB1*04:01 (30%) (ST2 and Table 5).
In SAA patients, the most common HLA class I antigens were A2 (29.6%), B41 (11.3%), and Cw7 (40%). The most common alleles were A*02:01 (19.8%), B*41:01 (9.76%), B*14:02 (9.76%), and C*04:01 (18.5%). The most common HLA class II antigens were DR4 (17.9%) and DQ3 (30%), while the most common alleles were DRB1*03:01 (13.2%), DRB1 *15:01 (13.1%), and DQB1*03:01 (25%) (ST2 and Table 5).
Haplotype frequencies and association with hematological conditions
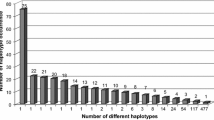
The EM algorithm identified 1230 unique haplotypes; the most frequent were HLA-A*33:01~B*14:02~DRB1*01:02 (2.35%), HLA-A*01:01~B*52:01 ~ DRB1*15:01 (2.11%), and HLA-A*01:01~B*08:01~DRB1*03:01 (0.89%) (Table 6). The most common haplotypes in ALL patients were A*01:01~B*52:01~DRB1*15:01 (2.45%), A*33:01~B*14:02~DRB1*01:02 (2.45%), and A*32:01~B*41:01~DRB1*03:01 (1.84%). The most common haplotypes in AML and myeloid sarcoma (MS) were A*01:01~B*52:01~DRB1*15:01 (2.62%), A*33:01~B*14:02~DRB1*01:02(2.27%), and A*02:01~B*41:01~DRB1*07:01 (1.92%). The most common haplotypes in SAA were A*01:01~B*52:01~DRB1*15:01 (2.32%), A*33:01~B*14:02~DRB1*01:02 (2.14%), and A*33:01~B*14:02~DRB1*11:01 (1.25%). The haplotype A*02:01~B*18:01~DRB1*03:01 frequency was significantly higher in patients with ALL than donors (1.22% vs. 0.13%). Additionally, the frequencies of the haplotypes A*33:01~B*14:02~DRB1*11:01 and A*02:01~B*14:02~DRB1*01:02 (1.25% and 1.07% respectively) were significantly higher than donors (0.17% and 0.26% respectively) (Table 7).
Pairwise linkage disequilibrium (LD) in donors
The haplotypes A*31:01~C*06:02, B*13:01~C*06:02, and B*44:02~C*05:01, were in complete linkage disequilibrium. A strong LD was found between B*50:01~C*06:02, B*52:01~C*12:02, B*14:02~C*08:02, C*12:02~DRB1*15:01, DRB1*11:01~DQB1*03:01, and DRB1*10:01~DQB1*05:01 (ST3).
HLA antigens and alleles association with ALL
Only HLA-B38 showed a trend towards increased odds of ALL in the additive (OR, 1.52; 95% CI, 1.00–2.30; p = 0.049; q = 0.729) and the dominant models (OR, 1.64; 95% CI, 1.06–2.54; p = 0.026, q = 0.392) (ST4).
HLA antigens and alleles association with AML
HLA-DR1 showed a trend towards lower odds of AML in the additive (OR, 0.67; 95% CI, 0.48–0.92; p = 0.013, q = 0.157) and dominant models (OR, 0.66; 95% CI, 0.47–0.93; p = 0.018, q-value = 0.214). On the other hand, DR7 showed a trend towards higher AML odds (OR, 1.25; 95% CI, 1.02–1.54; p = 0.033, q = 0.200). HLA-DRB1*01:02 showed a trend towards lower AML odds (OR, 0.64; 95% CI, 0.45–0.91; p = 0.014, q = 0.214) in the additive model. HLA-DRB1*07:01 was associated with higher AML odds in both the additive (OR, 1.26; 95% CI, 1.02–1.55; p = 0.030, q = 0.296) and dominant models (OR, 1.27; 95% CI, 1.00–1.62; p = 0.045, q = 0.454) (ST4 and Table 8).
HLA antigens and alleles association with SAA
HLA-A2 showed a significant association with higher SAA odds in the additive (OR, 1.38; 95% CI, 1.16–1.64; p < 0.001, q = 0.007), dominant (OR, 1.44; 95% CI, 1.15–1.81; p = 0.002, q = 0.041), and recessive models (OR, 1.71; 95% CI, 1.16–2.51; p = 0.006, q = 0.037). Furthermore, HLA-B14 and -DR15 showed a trend towards higher SAA odds in the additive (B14: OR, 1.41; 95% CI, 1.09–1.83; p = 0.008, q = 0.250; DR15: OR, 1.31; 95% CI, 1.06–1.62; p = 0.012, q = 0.157) and dominant models (B14: OR, 1.43; 95% CI, 1.06–1.92; p = 0.018, q = 0.392; DR15: OR, 1.38; 95% CI, 1.08–1.75; p = 0.009, q = 0.210). On the other hand, HLA-A30 and -DR13 showed a trend toward lower odds of SAA in both the additive (A30: OR, 0.72; 95% CI, 0.53–0.98; p = 0.036, q = 0.349; DR13: OR, 0.79; 95% CI, 0.65–0.97; p = 0.025, q = 0.200) and recessive models (A30: OR, 0.70; 95% CI, 0.50–0.97; p = 0.034, q = 0.413; DR13: OR, 0.79; 95% CI, 0.63–1.00; p = 0.047, q = 0.281). Finally, Cw6 showed a trend towards lower SAA odds in the additive model (OR, 0.67; 95% CI, 0.46–0.98; p = 0.039, q = 0.530) (ST4).
HLA-A*02:01, -B*14:02, and -DRB1*15:01 alleles were associated with higher odds of SAA in the additive (A*02:01: OR, 1.35; 95% CI, 1.07–1.70; p = 0.010, q = 0.242; B*14:02: OR, 1.43; 95% CI, 1.06–1.93; p = 0.020, q = 0.425; DRB1*15:01: OR, 1.32; 95% CI, 1.07–1.64; p = 0.011, q = 0.214) and dominant models (A*02:01: OR, 1.36; 95% CI, 1.03–1.80; p = 0.032, q = 0.411; B*14:02: OR, 1.44; 95% CI, 1.01–2.07; p = 0.046, q = 0.818; DRB1*15:01: OR, 1.38; 95% CI, 1.09–1.76; p = 0.008, q = 0.246). Furthermore, A*02:01 showed a trend towards increased SAA odds in the recessive model (OR, 1.90; 95% CI, 1.07–3.38; p = 0.029, q = 0.176). On the other hand, A*24:02 showed a trend towards decreased SAA odds (OR, 0.59; 95% CI, 0.36–0.96; p = 0.035, q = 0.373) (Table 8).
Discussion
The BMT unit at NIH, Cairo became operational in 1997, and the transplant rates increased dramatically since that time. The challenges that face the clinical hematopoietic stem cell transplantation program in Egypt are financial, technological, and administrative (Mahmoud et al. 2008; Mahmoud et al. 2020). However, this is our first report on HLA-A, -B, -C, -DRB1, and -DQB1 allele and haplotype frequencies from HLA data of 6000 individuals from different regions within Egypt. The history of Egypt reflects its geographical location at the crossroads of several cultures. In addition, Egypt took part in many regional empires throughout its history, which explains the complexity and heterogeneity of its genetic makeu: Allele and haplotype frequencies encountered in this study were similar to those reported by AFND (Gonzalez-Galarza et al. 2020).
In this study, 52, 76, 24, 37, and 14 HLA-A, -B, -C, -DRB1, and -DQB1 alleles were observed in Egyptian donors. Among the HLA-A alleles, A*01:01 (16.9%) and A*02:01 (16.1%) showed the highest frequency and represented 33% of HLA -A allelic diversity. HLA-B*41:01 (8.7%), B*49:01 (7.4%), B*14:02 (6.8%), B*52:01 (5.8%), and B*50:01 (5.1%) accounted for more than 33% of genetic diversity at the B locus. Although HLA-B*50:01 was reported to be frequent in Arabs (Jawdat et al. 2014) Caucasians, North Africans, and West-South Asians (Gonzalez-Galarza et al. 2020), B*41:01 was the most frequent B allele observed in Egyptians.
The most frequent HLA-C alleles were C*06:02 (25.1%), C*07:01 (25.1%), and C*04:01 (17.1%), accounting for 63% of HLA-C diversity. For HLA-DRB1, DRB1*11:01 (11.8%), DRB1*03:01 (11.6%), and DRB1*13:01 (11.4%) showed a frequency of more than 33%, followed by DRB1*07:01 (10.5%), and DRB1*15:01 (10.2%), which collectively accounted for 55% of HLA-DRB1 diversity in this study. Both HLA-DRB1*07:01:01G and DRB1*03:01:01G are common in Arabs of Central and Eastern regions of Saudi (Jawdat et al. 2020), and are frequent in Arabs from Tunisia and Jordan (Gonzalez-Galarza et al. 2020). For HLA-DQB1, five alleles showed a frequency of almost 70%. These alleles include DQB1*03:01 (27.5%), DQB1*05:01(18.9%), DQB1*06:01 (13.8%), DQB1*04:02 (6.9%), and DQB1*02:02 (5.1%). HLA-DQB1*02:01:01 and DQB1*03:01:01 are frequent in Arabs, while HLA-DQB1*03:02:01 is frequent in Arabs, Chinese, Indians, and Caucasians (Gonzalez-Galarza et al. 2020).
The most frequent HLA alleles in this study were A*01:01, DRB1*11:01, DRB1*03:01, DRB1*13:01, DRB1*07:01, DQB1*03:01, DQB1*05:01, DQB1*06:01, and DQB1*04:02. These alleles are listed in the catalog from the European Federation for Immunogenetics (EFI) as common “COM” alleles within all European sub-regions (Sanchez-Mazas et al. 2017). The frequencies of A*02:01, A*01:01, A*03:01, C*07:01, C*04:01, DRB1*15:01, DRB1*07:01, and DRB1*03:01 in the European American population (Creary et al. 2019) were similar in their frequencies to Egyptians.
Regarding allelic frequencies, HLA-A*01:01 (16.9%) frequency was similar to Caucasians of Australia (18.7%), Belgians (15.5%), Austrians (14.6%), peoples of Northwestern England (20.8%) and Southeastern France (15%), Czechs (16%), Palestinian Arabs of Gaza (17.8%), French BM donors (13%), and Germans (15.3%). HLA-A*02:01 frequency was 16.6%, which is similar to Brazilians (19.2%) and Chinese (18.7%), but lower than Argentinians (20–40%), Austrians (29.4%), Belgians (26.6%), German donors (28.3%), and the French BM donor registry (29%).
HLA-B*41:01 frequency was 8.7% in this analysis, which is higher than that of the Western population (1–2%). HLA-B*49:01 (7.1%) was similar in frequency to Armenians (9.5%), lower than Israeli Ethiopian Jews (18.6%), and higher than Western population (3%). HLA-B*14:02 (6.9%) was similar in frequency to Armenians (6%), lower than Ashkenazi and Polish Jews (10%), and higher than French and Germans (2–3%).
HLA-C*07:01 (25.1%) was similar in frequency to Cameroonian Baka Pygmies (25%), Southern Irish (21%), Southern Italians (20.6%), and Germans of Essen (20.9%), but higher than Burkina Faso Fulani (17.4%), peoples of Northwestern England (19%) and Southeastern France (17.3%), Brazilians (15.6%), and Australia New South Wales Caucasians (17.3%). HLA-C*06:02 (25.1%) was higher in frequency than the populations of the USA (2–9%), Burkinabes (5.3%), South-Asian Indians (13.9%), Cameroonians (17%), Czechs (16%), Germans (11%), and Southeastern French (10.6%), but similar to Indians (22.2%), Kenyans (21.7%), Moroccans (23.6%), Pakistanis (21.4%), Saudi Arabians (21.6%), South Africans (20.1%), and Tanzanians (19.2%).
HLA-DRB1*11:01 frequency (11.8%) was similar to Brazil's South Caucasians (11.1%), Chinese (11.7%), Gabonese (13.5%), Georgians (13%), German-Italian minority (10.4%), Greece (11.4%), African-Americans (11.3%), Swedes (11.6%), Indians (12%), Israel-Arabs (10.7%), Italy central (12.8%), but lower than Irani Parsis (21.9%), Lebanese (30.2%), Russian Siberians (16.7%), and Ukrainians (13.5%). HLA-DRB1*03:01 frequency (11.6%) was similar to Algerians (13.9%), Argentinians (10.9%), Austrians (11.8%), Belgians (15.7%), mixed-race Brazilians (11.6%), Chinese (13.1%), Colombians (15%), Czechs (11%), Croatians (12%), Danes (10.2%), British (12.7%), French (15.2%), German donors (10.6%), Israel Ethiopia Jews (12.5%) but higher in Italy Sardinians (55.7%). DRB1*13:01 (11.4%) was similar in frequency to Cameroonians and central Africans, Georgians, Indians, Iranians, Portuguese, Saudi Arabians, and Spanish (10–15%), but higher in Russian Siberians (23.8%). HLA-DQB1*03:01 (27.5%) was slightly higher in frequency than Algerians, Austrians, Belgians, and French (16–21%), but lower than in Brazilians (60%). HLA-DQB1*05:01 (18.9%) was similar in frequency to Austrians, Belgians, and the people of South France and Europe (12–25%) (Gonzalez-Galarza et al. 2020).
The most prevalent HLA-A~B~DRB1 haplotype in our donors were A*33:01~B*14:02~DRB1*01:02 (2.35%), A*01:01~B*52:01~DRB1*15:01 (2.12%), and A*01:01~B*08:01~DRB1*03:01 (0.89%). None of them included the A*02:01:01 allele unlike the Saudi population (Jawdat et al. 2020). HLA-A*33:01~B*14:02~DRB1*01:02 frequency was similar to Israelis Americans, Ashkenazi Jews, Italians, and Poles (1.7–2.3%), but is higher in Americans, Europeans, Russians, Saudis, and Croatians (41–44%). HLA-A*01:01~B*52:01~DRB1*15:01 was found at a similar frequency in Germans, Turks, and Poles (1.2%), but this haplotype frequency was higher in Israelis, Iraqi Jews (40.3%), and Indians (28.7%) (Gonzalez-Galarza et al. 2020). The haplotype A*01:01~B*08:01~DRB1*03:01 (0.89%) was the third most common HLA haplotype in our study; this haplotype was also seen in Caucasian and Hispanic ethnicities reported in the 16th IHIW, with frequencies of 0.07, 0.01, and 0.10. This haplotype ranked first in Europeans, Hispanics, and Dutch (Askar et al. 2013).
The allele frequencies that we reported in our study are consistent with a previous study on 286 Egyptian donors (Elshakankiry et al. 2017), the most common HLA class I antigens in their study were HLA-A2 (19.2%), A1 (15.8%), B41 (10.9%), and B35 (10.2%). However, they reported different frequencies of HLA-DRB1*13:01 (18.7%), DRB1*04:01 (15%), DRB1*11:01 (14.7%), and DRB1*03:01 (13.2%). These differences might be attributed to the small sample size.
The deviations from the HWE were expected a priori. Consanguinity rates among Egyptians were estimated to be between 29 and 39% in 2012 (Temtamy and Aglan 2012). In 2017, it was estimated to be over 40% depending on the region and urbanization (Ahmed 2017). Inbreeding increases with consanguinity, resulting in higher homozygosity. However, we cannot attribute the observed deviation in HWE solely to inbreeding given the small sample size of HLA-C and HLA-DQ alleles and the evidence of balancing selection in HLA-B and -DR loci.
We studied the association between HLA antigen groups and alleles with the ALL, AML, and SAA since they were the most prevalent. Most of our findings were trends toward significance, and generally, no remarkable antigen groups or alleles correlated with the odds of ALL or AML. However, only HLA-A2 showed a statistically significant and positive association with SAA. Our findings are consistent with a report by Gluckman et al. where they found an excess of homozygous HLA-A2 in SAA patients compared to healthy controls (Gluckman et al. 1981). In our study population, the frequency of HLA-A2 was 23.3% in donors and 29.6% in SAA patients. HLA-A2 correlated with higher odds of SAA in all the genetic models we conducted. The exact allele that is correlated with SAA could not be identified precisely in our study. However, we think that it is most likely to be HLA-A*02:01 since it showed a strong trend towards higher SAA odds.
We also found unprecedented significant associations between the haplotypes A*33:01~B*14:02~DRB1*11:01 and A*02:01~B*14:02~DRB1*01:02 with SAA. It is noteworthy that HLA-A*33:01, -B*14:02, and -A*02:01 showed strong trends toward increased SAA odds. The noted association with haplotype could be a result of the higher power of haplotype analysis compared to analysis of the individual antigens or alleles. Also, we found an unprecedented association between A*02:01~B*18:01~DRB1*03:01 and the odds of ALL.
The main limitation of this study is that we used PCR-SSO instead of sequencing due to the limited budget provided by the National Insurance for HLA typing. We perform HLA typing by screening patients and their family members for HLA-DRB1. We only proceed to class I (AB) typing for identical patients and donors on class II or for patients capable of affording the cost of a haploidentical transplant, which until now is not fully covered by insurance, and explains why the number of HLA-DRB1 typed samples is more than twice the number of HLA class I (AB) in our study.
Knowledge of population-specific allele and haplotype frequencies increases the chance of finding matched donors. Unfortunately, less than 3% of donors listed in international registries are from oriental origins, which makes it difficult to find a match for a patient with no sibling donor indicated for BMT in Egypt (Besse et al. 2016).
Conclusion
The results of this study present valuable information about HLA genes in Egyptians that can be used for hematopoietic stem cell unrelated donor recruitment, and selection, and as a helpful resource for population genetic studies and HLA disease associations.
Data availability
Data available on request from the authors.
References
Ahmed SM (2017) Consanguineous marriage among Egyptian youth: secondary analysis of survey of young people in Egypt, 2014. Egypt J Commun Med 35(2):85–94. https://doi.org/10.21608/EJCM.2017.3571
Askar M et al (2013) 16(th) IHIW: global distribution of extended HLA haplotypes. Int J Immunogenet 40(1):31–38. https://doi.org/10.1111/IJI.12029
Besse K et al (2016) On modeling human leukocyte antigen-identical sibling match probability for allogeneic hematopoietic cell transplantation: estimating the need for an unrelated donor source, biology of blood and marrow transplantation. J Am Soc Blood Marrow Trans 22(3):410–417. https://doi.org/10.1016/J.BBMT.2015.09.012
Chang YJ, Zhao XY, Huang XJ (2021) Haploidentical stem cell transplantation for acute myeloid leukemia: current therapies challenges and future prospective. Front Oncol. https://doi.org/10.3389/FONC.2021.758512
Creary LE et al (2019) Next-generation sequencing reveals new information about HLA allele and haplotype diversity in a large European American population. Hum Immunol 80(10):807–822. https://doi.org/10.1016/J.HUMIMM.2019.07.275
Elshakankiry NH et al (2017) P227 determination of HLA -A, -B and - DRB1 alleles and HLA-A -B haplotype frequencies in Egyptians based on family study. Hum Immunol 78:222. https://doi.org/10.1016/J.HUMIMM.2017.06.287
Gfeller D, Bassani-Sternberg M (2018) Predicting antigen presentation-what could we learn from a million peptides? Front Immunol 9:1716. https://doi.org/10.3389/FIMMU.2018.01716/BIBTEX
Gluckman E et al (1981) HLA markers in patients suffering from aplastic anaemia. Haematologia 14(2):165–172
Gonzalez-Galarza FF et al (2020) Allele frequency net database (AFND) 2020 update: gold-standard data classification, open access genotype data and new query tools. Nucleic Acids Res 48(D1):D783–D788. https://doi.org/10.1093/NAR/GKZ1029
Gooptu M et al (2021) HLA-haploidentical vs matched unrelated donor transplants with posttransplant cyclophosphamide-based prophylaxis. Blood 138(3):273–282. https://doi.org/10.1182/BLOOD.2021011281
Gragert L et al (2014) Fine-mapping of HLA associations with chronic lymphocytic leukemia in US populations. Blood 124(17):2657–2665. https://doi.org/10.1182/BLOOD-2014-02-558767
Guo SW, Thompson EA (1992) Performing the exact test of Hardy-Weinberg proportion for multiple alleles. Biometrics 48(2):361. https://doi.org/10.2307/2532296
Hammer C, Migdał M (2022) midasHLA: R package for immunogenomics data handling and association analysis
Huang X et al (2012) HLA associations in classical Hodgkin lymphoma: EBV status matters. PLoS ONE 7(7):39986. https://doi.org/10.1371/JOURNAL.PONE.0039986
Jawdat D et al (2014) HLA-B50 polymorphism in the Saudi population. Int J Immunogenet 41(2):95–97. https://doi.org/10.1111/IJI.12096
Jawdat D et al (2020) HLA-A, -B, -C, -DRB1, -DQB1, and -DPB1 Allele and Haplotype Frequencies of 28,927 Saudi Stem Cell Donors Typed by Next-Generation Sequencing. Front Immunol. https://doi.org/10.3389/FIMMU.2020.544768
Kazemi MH et al (2021) Association of HLA alleles with hematologic malignancies. Gene Rep 25:101346. https://doi.org/10.1016/J.GENREP.2021.101346
Kolde R (2019) pheatmap: pretty heatmaps
Lancaster AK et al (2007) PyPop update – a software pipeline for large-scale multilocus population genomics. Tissue Antigens 69(SUPPL. 1):192–197. https://doi.org/10.1111/J.1399-0039.2006.00769.X
Mahmoud HK et al (2008) Hematopoietic stem cell transplantation in Egypt. Bone Marrow Transplant. https://doi.org/10.1038/BMT.2008.136
Mahmoud HK et al (2020) Hematopoietic stem cell transplantation in Egypt: challenges and opportunities. Mediterr J Hematol Infect Dis 12(1):2020023. https://doi.org/10.4084/MJHID.2020.023
Park M, Seo JJ (2012) Role of HLA in hematopoietic stem cell transplantation. Bone Marrow Res 2012:1–7. https://doi.org/10.1155/2012/680841
Petersdorf EW (2017) Which factors influence the development of GVHD in HLA-matched or mismatched transplants? Best Pract Res Clin Haematol 30(4):333–335. https://doi.org/10.1016/J.BEHA.2017.09.003
R Core Team (2022) R: A Language and Environment for Statistical Computing. R Foundation for Statistical Computing, Vienna, Austria. https://www.R-project.org/
Sanchez-Mazas A et al (2017) Common and well-documented HLA alleles over all of Europe and within European sub-regions: a catalogue from the European Federation for Immunogenetics. HLA 89(2):104–113. https://doi.org/10.1111/TAN.12956
Simpson E, Dazzi F (2019) Bone marrow transplantation 1957–2019. Front Immunol. https://doi.org/10.3389/FIMMU.2019.01246
Slatkin M (1994) An exact test for neutrality based on the Ewens sampling distribution. Genet Res 64(1):71–74. https://doi.org/10.1017/S0016672300032560
Temtamy S, Aglan M (2012) Consanguinity and genetic disorders in Egypt. Middle East J Med Genet 1(1):12–17. https://doi.org/10.1097/01.MXE.0000407744.14663.D8
Van Rossum G, Drake FL (2009) Python 3 reference manual. CreateSpace, Scotts Valley, CA
Venables WN, Ripley BD (2002) Modern applied statistics with S. Fourth. Springer, New York
Villemagne B et al (2005) Two new cases of familial hairy cell leukemia associated with HLA haplotypes A2, B7, Bw4, Bw6. Leuk Lymphoma 46(2):243–245. https://doi.org/10.1080/10428190400013589
Zaimoku Y et al (2017) Identification of an HLA class I allele closely involved in the autoantigen presentation in acquired aplastic anemia. Blood 129(21):2908–2916. https://doi.org/10.1182/BLOOD-2016-11-752378
Funding
This research did not receive any specific grant from funding agencies in the public, commercial, or non-profit sectors, and no funding was received to assist with the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
Yasser ElNahass, Nabil M Abdelfattah and Nourhan Mekky formulated the research question, study design, and collected the data. Yasser ElNahass and Nabil M Abdelfattah conducted the statistical analysis and wrote the manuscript. All authors reviewed, edited, and provided guidance throughout all the phases of this study and writing the manuscript.
Corresponding author
Ethics declarations
Ethical approval
All procedures performed in studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments. All patients and donors gave written informed consent and the study was approved by the National Cancer Institutional Review Board number 2301-302-053.
Competing interests
The authors have no competing interests to declare that are relevant to the content of this article.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
ElNahass, Y., Mekky, N., Abdelfattah, N.M. et al. HLA alleles, haplotypes frequencies, and their association with hematological disorders: a report from 1550 families whose patients underwent allogeneic bone marrow transplantation in Egypt. Immunogenetics 76, 243–260 (2024). https://doi.org/10.1007/s00251-024-01343-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-024-01343-x