Abstract
Zebrafish (Danio rerio) has become an increasingly important model for immunological study. Its immune system is remarkably similar to that of mammals and includes both the adaptive and innate branches. Zebrafish T cells express functional T cell receptors (TCR), and all four TCR loci are present within the genome. Using 5′-rapid amplification of cDNA ends, we cloned and sequenced zebrafish TCRβ transcripts. TCRβ VDJ coding joints demonstrate conservation of mechanisms used by other vertebrate species to increase junctional diversity. Using the sequences obtained, along with previously published data, we comprehensively annotated the zebrafish TCRβ locus. Overall, organization of the locus resembles that seen in mammals. There are 51 V segments, a single D segment, 27 Jβ1 segments, a single Jβ2 segment, and two constant regions. This description of the zebrafish TCRβ locus has the potential to enhance immunological research in zebrafish and further our understanding of mammalian TCR repertoire generation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Zebrafish (Danio rerio) has emerged as a powerful new vertebrate model of human disease. Initially favored by developmental biologists, zebrafish has been adopted into many research areas and is gaining popularity as an immune disease model (reviewed in Meeker and Trede 2008). Evolution of the adaptive immune system coincided with the emergence of jawed vertebrates, predating the divergence of fish from other vertebrates (reviewed in Traver et al. 2003). Thus, the adaptive and innate branches of the immune system are both intact in zebrafish and remain remarkably similar to their human counterparts (reviewed in Meeker and Trede 2008). T cells with mammalian-like T cell receptors (TCR) are present in all jawed vertebrates (Litman et al. 1999), and T cell and thymic development in zebrafish parallel that of mammals; the primary difference is that the zebrafish thymus remains as two discrete bilateral structures (Willett et al. 1999; Danilova et al. 2004). Like mammals, four TCR loci (α, β, δ, and γ) are found in the zebrafish genome (Haire et al. 2000 (α); Schorpp et al. 2006 (β and δ); Yazawa et al. 2008 (γ)), and Rag-dependent VDJ recombination is also present (Wienholds et al. 2002). The zebrafish genome has been sequenced, and assemblies are continually being refined. Although some sequence data exist for zebrafish TCR α and β loci (Haire et al. 2000; Schorpp et al. 2006), none of the zebrafish TCR loci are fully annotated, and the number and ordering of TCR β segments remain largely unknown.
Here, we describe the genomic organization of the zebrafish TCRβ locus. We employed 5′-rapid amplification of cDNA ends (5′-RACE) to clone and sequence TCRβ transcripts that had undergone VDJ recombination and were expressed as spliced mRNA transcripts. Analysis of VDJ coding joints demonstrate that general locus organization and mechanisms used to generate junctional diversity in zebrafish and mammals are conserved. Using the sequences obtained, together with previously published data, we annotated the TCRβ locus. This description should facilitate the ongoing use of zebrafish as a model for immune development and disease.
Materials and methods
Zebrafish care and maintenance
Fish were housed in a colony at 28.5°C on a 14/10-h circadian cycle. For all procedures, fish were anesthetized with 0.02% tricaine methanesulfonate (MS222). Fish were handled per National Institutes of Health guidelines, under a protocol approved by the University of Utah Animal Care and Use Committee (IACUC number 08-08005).
TCRβ transcript cloning and analysis
Thymi, gut, and other tissues were extracted from 4–6-month-old WIK strain lck::EGFP +/+ zebrafish and RNA isolated with Trizol (Invitrogen, Carlsbad, CA, USA). RNA was reverse transcribed using Moloney murine leukemia virus reverse transcriptase (Clontech, Palo Alto, CA, USA), and RACE was performed using a SMART II RACE kit (Clontech). Gene-specific primers for nested polymerase chain reaction (PCR) were designed for the TCR-Cβ1 and TCR-Cβ2 constant regions (Supplemental Table 1; Schorpp et al. 2006). PCR was performed using kit reagents with cycling conditions for each round as follows: 94°C 30 s, 68°C 30 s, and 72°C 3 min, for 20–25 cycles). PCR products were excised from ethidium bromide-stained agarose gels, and bands were purified using QIAquick Gel Extraction kit (Qiagen, Valencia, CA, USA). DNA was cloned into the TOPO-TA cloning vector (Invitrogen) and transformed into chemically competent DH5α cells. Colonies were screened for inserts using internal primers and colony PCR (Supplemental Table 1). Plasmids from positive clones were purified (Qiagen) and sequenced by MCLAB (San Francisco, CA, USA). Sequences were analyzed using Sequencher 4.8 (Gene Codes Corporation, Ann Arbor, MI, USA) and compared with the Zv6 March 2006 platform in the UCSC Genome Browser using the Blat function (Kent 2002, http://genome.ucsc.edu). These sequence data were produced by the D. rerio Sequencing Group at the Sanger Institute and can be obtained from http://www.sanger.ac.uk/Projects/D_rerio/Zv8_assembly_information.shtml. Zv8 sequence was compared to Zv6 using Vector NTI Advance (DiBiase et al. 2006).
Results and discussion
Expressed TCRβ segments
Clones retrieved by 5′-RACE
Thirty-five unique TCRβ clones were isolated from a total of six WIK strain animals. Thirty-three of these were TCRβ1 transcripts obtained using Cβ1 primers, while primers for Cβ2 yielded only two TCRβ2 cDNAs. One possible explanation for this skewed result is that there is a natural preference for Cβ1 during zebrafish TCRβ rearrangement and/or thymic selection. An alternate possibility is that the 5′-RACE reaction is more robust for Cβ1 as compared to Cβ2. Among the 35 TCRβ isolates, six transcripts contain only D-J recombinations, with no attached V sequence (clones 30–35, Supplemental Fig. 1). In each case, transcription initiated immediately upstream of the D cassette. These cDNAs represent classic sterile transcripts typically generated in immature T cells at the DN1 stage (Porritt et al. 2004). Among these six D-J clones, the single D segment was rearranged with four different J cassettes (Jβ1.18, Jβ1.29, Jβ1.33, and Jβ1.31).
The remaining 29 TCRβ transcripts contained complete VDJ coding joints and are shown aligned in Fig. 1. Two non-functional rearrangements were recorded: recombination in clones 15 and 17 produced out-of-frame transcripts. In addition, clone 15 had a 130-bp truncation at the 3′ end of the V segment. The other 27 clones contained apparently functional VDJ recombinations. Altogether, expression of 22 unique V sequences and 18 different J sequences was identified. Many V and J gene segments were represented in multiple unique recombinations (Fig. 1). The J segment genes most commonly used were Jβ1.04, Jβ1.16, and Jβ1.23, each represented three times. The most common V gene used was Vβ.51, represented in four different transcripts. Notably, Vβ.51 is the V cassette located immediately upstream of the single D segment in the locus, suggesting a preference for usage of D-proximal V gene segments during V-DJ joining. In fact, a closer analysis of Vβ usage supports such a preference. After grouping V segments into three groups based on distance from the D gene, we found that the third of V cassettes farthest away was incorporated into six clones with VDJ recombinations, while the third closest to the D was used in 14. The middle third was used eight times. Regression analysis of this data results in a slope of four additional incorporations per third, a Pearson correlation coefficient of 0.96 and an r-squared value of 0.92. Our finding is reminiscent of the preferential rearrangement of the most DH-proximal heavy chain V segment in the pre-immune repertoire of human B cells (Varade et al. 1993).
Alignment of VDJ coding joints from expressed TCRβ transcripts. Dots indicate nucleotide deletions with respect to the genomic sequence of the V, D, or J gene used. N-nt denotes non-template nucleotide additions. V number (V#) and J number (J#) refer to the designated gene segment, and are bold when a V or J occurred in more than one uniquely cloned VDJ rearrangement. Translation of highly conserved consensus codon sequences is shown in the top line. Single nucleotide polymorphisms, not matching genomic sequences of Zv8, are designated with lowercase. Dashes in clone 3 represent bps that could not be verified by sequence due to limitations of 5′-RACE. Vβ.46 in clone 15 is truncated by 130 bp at its 3′ end
Generation of CDR3 hypervariability
The VDJ coding joints in Fig. 1 provide evidence that mechanisms leading to hypervariability of the complementary determining region 3 (CDR3) loop of TCR are similar in mammals and zebrafish. Endonuclease activity at the joining ends of V, D, and J cassettes is evident. Nucleotide (nt) deletions at coding joints in each TCR ranged from 2 to 18 nt for Cβ1 with a mean of 10, consistent with published data from other teleosts (Fischer et al. 2002). Thus, the CDR3 loop can vary by at least six amino acids in length, augmenting diversity. At the V-D joint, the range of nucleotide loss is 0–10 with a mean of 5.2, and at the D-J joint, 1–13 and 5.7, respectively. In addition to endonuclease activity, definitive non-template-mediated nucleotide (N-nt) additions were detected in 15 of 29 TCR clones with full VDJ rearrangements and five of six D-J recombinations. While each coding joint typically had evidence of at least one or two N-nt additions, this number may be an underestimate as N-nt additions matching pre-existing genomic sequence cannot be distinguished from ends that were left intact.
There is apparent bias for N-nt additions at the D-J joint as compared to the V-D joint. Nineteen of 28 (68%) N-nt additions in the group of full VDJ coding joints were located at the D-J joint. In these transcripts, there is also evidence of bias for G/C versus A/TN-nt additions, as 16 of 28 (57%) N-nt added were G or C. This is also seen in mammals where TdT preferentially adds G nucleotides (Basu et al. 1983; Murray et al. 2006). As noted above, N-nt additions matching deleted genomic sequence cannot be differentiated from non-deletions, and because the D region is particularly G/C rich, we may be underestimating this G/C bias of TdT in zebrafish. Taken together, these findings support that zebrafish generate significant diversity of their TCR CDR3 loop in a similar fashion to mammals.
Genomic annotation of the TCRβ locus
In silico identification of coding sequences
Collectively, via 5′-RACE, we identified 22 unique V gene segments, a single D cassette, 17 unique Jβ1 gene segments, and one Jβ2 gene. Using the Blat function of the UCSC Genome Browser, corresponding sequences were identified on chromosome 17 in the Zv6 release of the zebrafish genome. Schorpp et al. (2006) previously published nine additional Jβ1 sequences for which we could find corresponding genomic sequence, as well as two Jβ2 cassettes. One of these Jβ2 segments was expressed among Cβ1 transcripts in our study, Jβ1.04, and maps to the Jβ1 region. The other Jβ2 sequence of Schorpp et al. (2006) corresponds to Jβ2.01, maps to a genomic position between Cβ1 and Cβ2, and is represented among our Cβ2 transcripts.
To identify other V, D, and J gene segments, we examined chromosome 17 genomic sequences between map positions 45,312,445 and 45,582,949 (Zv8). This 270-kb region includes the TCRβ locus, as well as an additional 30-kb downstream of Cβ2. We searched for putative V, D, and J cassettes and C exons by screening for homology to previously identified TCR genes and for appropriately spaced consensus recombination signal sequences (RSS). An additional putative 30 V cassettes and seven J cassettes were identified. Of these, we classified one V and six Js as pseudogenes based on lack of an open reading frame or absence of a consensus RSS (see below). Thus, only one additional J cassette (Jβ1.22) was identified in silico. Altogether, there are 51 Vβ1 cassettes and one additional Vβ1 pseudogene, 27 Jβ1 cassettes and six additional Jβ1pseudogenes, and one functional Jβ2 cassette. No additional D cassettes or constant region exons were identified. The zebrafish TCRβ locus is depicted schematically in Fig. 2. Our annotation of the TCRβ locus on chromosome 17 is included in the supplemental materials as a text file in GenBank format.
Genomic organization of the zebrafish TCRβ locus. The 51 Vβ (V), a single Dβ (D1), 27 Jβ1 (J1), and a single Jβ2 (J2) cassettes are indicated along with the Cβ1 (C1) and Cβ2 (C2) constant regions. The transcription start site of the Vβ leader exon (L) is marked with a dotted line. The second Vβ exon is denoted V. The internal TATA box (TATA), found in TCR D regions across species, is indicated. Recombination signal sequences characteristic of VDJ rearrangement include heptamers (seven) and nonamers (nine) with spacers following the 12/23 ± 1 rule (12, 23). Genomic splice donors (GT) and acceptors (AG) are indicated. Cβ1 has four exons, and Cβ2 has three exons (not shown; Schorpp et al. 2006). Vβ.52 is found 8.1 kb downstream of Cβ2 on the complementary strand in reverse orientation. Pseudogenes are not shown
Organization of Vβ segments
Similar to mammals and other vertebrates, each zebrafish V cassette consists of two exons that are post-transcriptionally spliced. The first exon contains a 5′-UTR of approximately 100 bp followed by a start codon and an average coding region of 48 bp (range 31–61). The leader exon is followed by the splice donor sequence, GT, with an intron length of 75–145 bp, similar to that seen in the TCRα of teleosts (Fischer et al. 2002). Exon two of the V is preceded by the splice acceptor sequence, AG, and is approximately 300 bp in length. It is followed by a consensus RSS heptamer–nonamer sequence separated by a 22- or 23-bp spacer.
Of the total 51 Vβ1 cassettes, most are separated by 200–9,600 bp of intergenic sequence. The marked exception is 70 kb located between Vβ.31 and Vβ.32. There is also a V cassette (Vβ.52) 8.1 kb downstream of Cβ2 in reverse orientation. Despite this peculiar organization, Vβ.52 is functional, as it was detected in our expression repertoire (clone 11, Fig. 1). In the human TCRβ locus, there is also an inverted Vβ cassette found downstream of Cβ2, and this has been shown to undergo rearrangement by inversion (Malissen et al. 1986). This suggests that the zebrafish Vβ.52 may employ a similar mechanism of recombination.
Single Dβ segment
A single 12-bp D cassette is found in the TCRβ locus with the following genomic organization: nonamer-12 bp spacer-heptamer-D-heptamer-23 bp spacer. It is located between the Vβ group and the Jβ1 group, downstream of Vβ.51 and upstream of the 5′-most Jβ1 gene, Jβ1.02. The D segment map position is 45,339,359 (Zv8).
Organization of Jβ segments
Jβ cassettes are approximately 80 bp long, consist of single exons, and have the following genomic organization: nonamer-12 bp spacer-heptamer-J gene segment. All 27 Jβ1 cassettes are closely spaced within 16 kb and separated quite evenly by approximately 220 bp. Only one Jβ2 cassette was identified located between Cβ1 and Cβ2.
RSS in the TCRβ locus
For all putative zebrafish TCRβ V, D, and J cassettes, the 12/23 ± 1 rule for RSS is completely conserved. Of note, there is a subtle bias for 22 bp gaps (30/51 V segments). This bias has been noted for TCRVα segments in Tetraodon nigroviridis (pufferfish) and TCRVγ in Salmo salar (Atlantic salmon), suggesting that it may be a feature of teleosts (Fischer et al. 2002; Yazawa et al. 2008). We found one instance of a 24-bp spacer in Vβ.13, a V segment found expressed in our analysis (Fig. 1). Heptamers and nonamers demonstrated strong resemblance to consensus sequences for other vertebrates including human (Hesse et al. 1989; Fischer et al. 2002). Sequence logos were created using WebLogo (http://weblogo.berkeley.edu; Crooks et al. 2004) and are depicted in Fig. 3. Conservation is proposed to be most important for the three heptamer bases closest to the recombination crossover site (Akira et al. 1987; Hesse et al. 1989; Akamatsu et al. 1994), in agreement with our observation. Positions 1–3 of Vβ heptamers demonstrate strong consensus of CAC, whereas GTG is conserved at positions 5–7 in Jβ heptamers. Presumably, this allows for strong base-pairing during recombination. Similar consensus sequences of V- and J-associated heptamers have been reported previously for other teleosts (Fischer et al. 2002).
Consensus recombination signal sequences (RSS) for 51 Vβ segments (a) and 27 Jβ segments (b). The first three nucleotides of Vβ heptamers and the last three bp of Jβ heptamers are strongly conserved and complementary. Similarly, central bp of Vβ and Jβ nonamers are highly conserved and complementary with thymidines in V and adenosines in J (b). The 12 nucleotides of the D region (underlined) and the flanking heptamers and nonamers (boxed) are also shown (c). Note the similarity between the D region RSS and the consensus heptamer and nonamer sequences for Vβ and Jβ
For other species, it is reported that nonamer positions five and six are most important (Danilova et al. 2004). In agreement, conservation of zebrafish TCRβ nonamers is most pronounced in central base pairs (Fig. 3). V-associated nonamers and the J-proximal nonamer of the D segment have consensus adenosines in these positions, while J-associated nonamers and the V-proximal nonamer of the D segment show a thymidine bias. This again likely fosters efficient base-pairing during the recombination process and has been seen previously in teleosts (Fischer et al. 2002).
TCRβ promoters
A proposed promoter element for human and murine Vβ segments is a decamer having the consensus sequence, AGTGATG/CATCA. Conservation of the inverted repeat, TGANNTCA, is strongest, and the decamer is typically located 40–75 bp upstream of the transcription start site (Anderson et al. 1988). We recognized similar decanucleotide sequences upstream of many zebrafish Vβ sequences. Twenty-six Vβ genes have easily identifiable upstream promoter elements that conserve the TGANNTCA element. Most of the other Vβ segments have elements that diverge from this consensus by only one base pair. Notably, some consensus motifs were found as far as 1 kb upstream of the Vβ segment start codon and may not represent genuine regulatory elements.
Phylogenetic relationship of TCRβ sequences
V segments
Zebrafish Vβ gene segments encode peptides of about 115 amino acids (range 110–120). Canonical features seen in mammalian and other vertebrates are highly conserved and include the following residues: Gln6, Cys23, Trp34, and Cys92 (Schiffer et al. 1986). Cys23 and Cys92 likely form an intrachain disulfide bond (Fischer et al. 2002; Wermenstam and Pilstrom 2001). Regions corresponding to CDR1, 2, and 3 and framework regions (FR) are evident (Supplemental Fig. 2a). The WY[K/R]Q sequence located in FR2 of TCR from other species is highly conserved in zebrafish TCRβ, as well as the FR3 motif, YFCA (Rast and Litman 1994; Wermenstam and Pilstrom 2001). Like mammals, only one set of Vβ segments contribute to both TCRβ1 and β2 rearrangements.
Beyond these few conserved residues and motifs, there is extensive variation between zebrafish Vβ segments. Vβ segments can be loosely classified into six families (Supplemental Fig. 2b). However, only 16 of the 51 Vβ segments have more than 75% amino acid identity with at least one other Vβ gene to form a family (Supplemental Fig. 3). These 16 genes comprise seven families: one having four members and the remaining six consisting of only pairs. The other 36 Vβ segments are single-member families by this criterion. Such single-member Vβ families are common in mammalian TCRs and suggest rapid independent evolution following gene duplication (Hedrick 1993).
D segment
Only one D gene cassette was previously known to exist in the TCRβ locus, and this was the only segment identified in our studies. Located between the Vβ1 and Jβ1 regions, it is 12 bp in length (Fig. 3c), matches the human Dβ1 segment sequence exactly (NCBI reference sequence: NG_01333.2), and contains the consensus D sequence identified in other teleosts (5′-GGACAGGG-3′; Partula et al. 1995; Litman et al. 1999). The zebrafish TCRβ D segment encodes Gly-Thr-Gly-Gly (Fig. 1), consistent with the observation that a functional CDR3 is typically a Glycine-rich region (Fischer et al. 2002). Also, the TATA box located within the upstream nonamer and 12 bp spacer (Fig. 2) is reported to be biologically important in other species (Tillman et al. 2004). Notably, neither of the two transcripts using the Cβ2 constant region contained a recognizable D segment. Thus, we postulate that TCRβ undergoes only V-J recombination when Cβ2 is used. Supporting this hypothesis, Schorpp et al. (2006) also found that Cβ2 TCRs lacked obvious D segments. Interestingly, we also saw several Cβ1 transcripts apparently lacking a D segment. While we cannot rule out that in these cases the entire D sequence was removed by endonucleases, it is possible that rearrangement at the TCRβ locus can employ both VDJ and V-J recombination.
J segments
Zebrafish J segments show strong conservation of sequence features seen in other vertebrates including the Phenylalanine-Glycine-X-Glycine (FGXG) motif present in nearly all TCR and immunoglobulin light chains (Supplemental Fig. 4a; Moss and Bell 1995). The conserved 6 bp splice site (GTAAGT) at the 3′ ends of J gene segments in other species (Yazawa et al. 2008) is conserved in zebrafish with some variability at the third and sixth position. Jβ gene segments can be loosely classified into seven families, two containing only a single member (Supplemental Fig. 4b). In contrast to Vβ families, Jβ families show higher nucleotide identities, ranging from 59% to 94% within each family (Supplemental Fig. 5).
C regions
The zebrafish TCRβ locus has two alternate C regions. Cβ1 contains four exons and encodes 156 aa, while Cβ2 has three exons encoding 163 aa. They have 36% amino acid identity. Such significant diversity in nucleotide sequence and exon structure has been seen in TCR constant genes of other teleosts (Yazawa et al. 2008). Both zebrafish TCRβ constant regions contain characteristic immunoglobulin, connecting peptide, transmembrane, and cytoplasmic domains along with canonical residues: Cys147, Trp161, and Cys212 (Wilson et al. 1998; Wermenstam and Pilstrom 2001).
Conclusion
In summary, we report expression data and genomic annotation of the zebrafish TCRβ locus. We have comprehensively described the genomic organization of the V, D, and J gene segment repertoire. The structure of the locus is similar to that described in other vertebrates including mammals. Sequence analysis of zebrafish TCRβ transcripts suggests the presence of canonical recombination machinery and the capacity to generate significant diversity within the CDR3 loop composed of the V-D and D-J coding joints. Overall, transcripts show high homology to previously described TCRβ transcripts of other vertebrates, with conservation of crucial sequence motifs and domains. Our description of the zebrafish TCRβ locus is significant with regard to future zebrafish studies as this vertebrate continues to expand its use as an important model of immunological and other human diseases. As an example, these data have recently been instrumental in facilitating an in-depth interrogation of T cell clonality in zebrafish models of T cell leukemia (Frazer et al. 2009).
References
Akamatsu Y, Tsurushita N, Nagawa F, Matsuoka M, Okazaki K, Imai M, Sakano H (1994) Essential residues in V(D)J recombination signals. J Immunol 153:4520–4529
Akira S, Okazaki K, Sakano H (1987) Two pairs of recombination signals are sufficient to cause immunoglobulin V-(D)-J joining. Science 238:1134–1138
Anderson SJ, Chou HS, Loh DY (1988) A conserved sequence in the T-cell receptor beta-chain promoter region. Proc Natl Acad Sci USA 85:3551–3554
Basu M, Hegde MV, Modak MJ (1983) Synthesis of compositionally unique DNA by terminal deoxynucleotidyl transferase. Biochem Biophys Res Commun 111:1105–1112
Crooks GE, Hon G, Chandonia JM, Brenner SE (2004) WebLogo: a sequence logo generator. Genome Res 14:1188–1190
Danilova N, Hohman VS, Sacher F, Ota T, Willett CE, Steiner LA (2004) T cells and the thymus in developing zebrafish. Dev Comp Immunol 28:755–767
DiBiase A, Harte RA, Zhou Y, Zon L, Kent WJ (2006) Piloting the zebrafish genome browser. Dev Dyn 235:747–753
Fischer C, Bouneau L, Ozouf-Costaz C, Crnogorac-Jurcevic T, Weissenbach J, Bernot A (2002) Conservation of the T-cell receptor alpha/delta linkage in the teleost fish Tetraodon nigroviridis. Genomics 79:241–248
Frazer JK, Meeker ND, Rudner L, Bradley DF, Smith AC, Demarest B, Joshi D, Locke EE, Hutchinson SA, Tripp S, Perkins SL, Trede NS (2009) Heritable T-cell malignancy models established in a zebrafish phenotypic screen. Leukemia 23:1825–1835
Haire RN, Rast JP, Litman RT, Litman GW (2000) Characterization of three isotypes of immunoglobulin light chains and T-cell antigen receptor alpha in zebrafish. Immunogenetics 51:915–923
Hedrick SM (1993) Chimeric T cell receptor-immunoglobulin molecules: function and applications. Int Rev Immunol 10:279–290
Hesse JE, Lieber MR, Mizuuchi K, Gellert M (1989) V(D)J recombination: a functional definition of the joining signals. Genes Dev 3:1053–1061
Kent WJ (2002) BLAT–the BLAST-like alignment tool. Genome Res 12:656–664
Litman GW, Anderson MK, Rast JP (1999) Evolution of antigen binding receptors. Annu Rev Immunol 17:109–147
Malissen M, McCoy C, Blanc D, Trucy J, Devaux C, Schmitt-Verhulst AM, Fitch F, Hood L, Malissen B (1986) Direct evidence for chromosomal inversion during T-cell receptor beta-gene rearrangements. Nature 319:28–33
Meeker ND, Trede NS (2008) Immunology and zebrafish: spawning new models of human disease. Dev Comp Immunol 32:745–757
Moss PA, Bell JI (1995) Sequence analysis of the human alpha beta T-cell receptor CDR3 region. Immunogenetics 42:10–18
Murray JM, O’Neill JP, Messier T, Rivers J, Walker VE, McGonagle B, Trombley L, Cowell LG, Kelsoe G, McBlane F, Finette BA (2006) V(D)J recombinase-mediated processing of coding junctions at cryptic recombination signal sequences in peripheral T cells during human development. J Immunol 177:5393–5404
Partula S, de Guerra A, Fellah JS, Charlemagne J (1995) Structure and diversity of the T cell antigen receptor beta-chain in a teleost fish. J Immunol 155:699–706
Porritt HE, Rumfelt LL, Tabrizifard S, Schmitt TM, Zuniga-Pflucker JC, Petrie HT (2004) Heterogeneity among DN1 prothymocytes reveals multiple progenitors with different capacities to generate T cell and non-T cell lineages. Immunity 20:735–745
Rast JP, Litman GW (1994) T-cell receptor gene homologs are present in the most primitive jawed vertebrates. Proc Natl Acad Sci USA 91:9248–9252
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Schiffer M, Wu TT, Kabat EA (1986) Subgroups of variable region genes of beta chains of T-cell receptors for antigen. Proc Natl Acad Sci USA 83:4461–4463
Schorpp M, Bialecki M, Diekhoff D, Walderich B, Odenthal J, Maischein HM, Zapata AG, Boehm T (2006) Conserved functions of Ikaros in vertebrate lymphocyte development: genetic evidence for distinct larval and adult phases of T cell development and two lineages of B cells in zebrafish. J Immunol 177:2463–2476
Tillman RE, Wooley AL, Hughes MM, Khor B, Sleckman BP (2004) Regulation of T-cell receptor beta-chain gene assembly by recombination signals: the beyond 12/23 restriction. Immunol Rev 200:36–43
Traver D, Herbomel P, Patton EE, Murphey RD, Yoder JA, Litman GW, Catic A, Amemiya CT, Zon LI, Trede NS (2003) The zebrafish as a model organism to study development of the immune system. Adv Immunol 81:253–330
Varade WS, Marin E, Kittelberger AM, Insel RA (1993) Use of the most JH-proximal human Ig H chain V region gene, VH6, in the expressed immune repertoire. J Immunol 150:4985–4995
Wermenstam NE, Pilstrom L (2001) T-cell antigen receptors in Atlantic cod (Gadus morhua L.): structure, organisation and expression of TCR alpha and beta genes. Dev Comp Immunol 25:117–135
Wienholds E, Schulte-Merker S, Walderich B, Plasterk RH (2002) Target-selected inactivation of the zebrafish rag1 gene. Science 297:99–102
Willett CE, Cortes A, Zuasti A, Zapata AG (1999) Early hematopoiesis and developing lymphoid organs in the zebrafish. Dev Dyn 214:323–336
Wilson MR, Zhou H, Bengten E, Clem LW, Stuge TB, Warr GW, Miller NW (1998) T-cell receptors in channel catfish: structure and expression of TCR alpha and beta genes. Mol Immunol 35:545–557
Yazawa R, Cooper GA, Beetz-Sargent M, Robb A, McKinnel L, Davidson WS, Koop BF (2008) Functional adaptive diversity of the Atlantic salmon T-cell receptor gamma locus. Mol Immunol 45:2150–2157
Acknowledgments
NM was supported by National Institutes of Health (NIH) NICHD award 2 K12 HD001410 and an Alex’s Lemonade Stand Young Investigator award; JKF was supported by NIH NICHD award 5 K08 HD53350; JKF and NM were also supported by the Children’s Health Research Center at the University of Utah and by Primary Children’s Medical Center Foundation grants; LR was supported by NIH NIDDK T32 award DK007115; NST was supported by NIH NIAID award R21 AI079784 and the Huntsman Cancer Foundation; Huntsman Cancer Institute core facilities supported by grant P30 CA042014 also participated in this work.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
Below is the link to the electronic supplementary material.
Supplemental Fig. 1
Alignment of the DJ coding joints from non-functional TCRβ cDNA transcripts. Dots indicate truncation with respect to the D or J segment used. N-nt denotes non-template nucleotide additions. J number (J#) refers to the gene identification nomenclature and is shown in bold where it occurs in more than one uniquely cloned VDJ rearrangement. Translation of highly conserved sequences is shown in the top line. The 5′-most region of each transcript is identical to genomic sequence upstream of the D cassette (GIF 83 KB)
Supplemental Fig. 2a
Zebrafish TCR Vβ segments. Alignments of 52 zebrafish TCR Vβ amino acid sequence are shown (a). Varying degrees of consensus are indicated (identical =  , block of similar =
, block of similar =  , conservative =
, conservative =  ). Consensus sequence is listed at the bottom, and proposed FR1, FR2, and FR3 boundaries are marked by gray boxes. Arrows point to highly conserved anchor residues (GIF 436 KB)
). Consensus sequence is listed at the bottom, and proposed FR1, FR2, and FR3 boundaries are marked by gray boxes. Arrows point to highly conserved anchor residues (GIF 436 KB)
Supplemental Fig. 2b
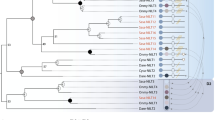
A neighbor-joining tree (b; Saitou and Nei 1987) based on a matrix of distances between all pairs of sequence was constructed with Vector NTI Advance (AlignX). Complete V segment amino acid sequences were included, and the analysis was performed using default settings (GIF 32.4 KB)
Supplemental Fig. 3
Vβ segment identity percentages. The amino acid percent identity of each Vβ pair is shown. Numbers greater than 75% are shaded (GIF 432 KB)
Supplemental Fig. S4
Zebrafish TCR Jβ segments. Alignments of 27 zebrafish TCR Jβ amino acid sequences are shown (a). Translations of Jβ cassettes are approximately 17 aa in length. Consensus regions are indicated (identical =  , block of similar =
, block of similar =  , conservative =
, conservative =  ). A neighbor-joining tree (b; Saitou and Nei 1987) demonstrates seven putative Jβ families (GIF 135 KB)
). A neighbor-joining tree (b; Saitou and Nei 1987) demonstrates seven putative Jβ families (GIF 135 KB)
Supplemental Fig. 5
Jβ segment identity percentages. The amino acid percent identity of each Jβ pair is shown (GIF 153 KB)
Supplemental Table 1
Primer sequences (DOC 56 kb)
Supplementary Material
TCR beta locus annotation (TXT 385 KB)
Rights and permissions
About this article
Cite this article
Meeker, N.D., Smith, A.C.H., Frazer, J.K. et al. Characterization of the zebrafish T cell receptor β locus. Immunogenetics 62, 23–29 (2010). https://doi.org/10.1007/s00251-009-0407-6
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-009-0407-6