Abstract
Defensins comprise an important family of anti-microbial peptides. Among vertebrates, numerous defensin genes have been detected, but their evolutionary background is still discussed. We investigated the molecular evolution and variability of β-defensins of Caprini via sequence analyses of defensin introns. Screening of several domestic and wild species of Caprini revealed a total of 13 discrete β-defensin coding sequences, with three of them described before this study. Phylogenetic analyses revealed that the array of newly described defensin genes is of monophyletic origin and has arisen in numerous independent duplication events after separation of the ancestral defensins. As a result of that scenario, recent defensin genes are distributed in a species-specific manner. Values of synonymous and non-synonymous substitutions demonstrated that both modes of evolutionary pressure, positive as well as negative selection, have acted. In addition, conservation of some β-defensin exons is demonstrated. Discrimination of certain β-defensin genes was possible only due to intron-specific differences. Therefore, sequence analyses restricted to the exons would result in underestimation of the number of β-defensin genes. Our study shows that for reconstruction of the phylogenetic history data of defensin introns are more appropriated. Comparisons among the amino acid sequences show moderate substitutions without changing the net charge of the mature peptides.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cationic anti-microbial peptides are multifunctional peptides of the innate immune system which act directly against host-invading micro-organisms (Boman 1991; Hancock and Lehrer 1998; Lehrer et al. 1993; Martin et al. 1995; Nicholas and Mor 1995). In addition, they trigger the specific immune response via signalling between the early and the specific immune response (Hoover et al. 2002; Yang et al. 1999, 2004). The anti-microbial execution is thought to be caused by the attachment of the cationic peptides (positively charged) at the anionic phospholipids of the microbe's membrane, resulting in its disruption and cell death (Ganz 2003; Kagan et al. 1990; Satchell et al. 2003). One of the most prominent components of that array of anti-microbial peptides is represented by defensins. Defensins are small (2–6 kDa), cationic and contain six cysteine residues at defined positions. Based on their size and the spatial position of the six cysteine residues, defensins of higher vertebrates are classified as α-, β- and θ-defensins. Defensins are translated as unmature precursors and are encoded by two separate exons, of which one encodes the signal sequence and the other the pro- and the mature peptide (reviewed in Ganz 2003; Hughes 1999; Lehrer and Ganz 2002; Schröder 1999; Schutte and McCray 2002; Wang et al. 2003).
The modern era of anti-microbial peptide research began with the description of lysosomal cationic peptides in leucocytes from rabbit and guinea pig (Zeya and Spitznagel 1966), later characterised structurally and classified as α-defensins (Selsted et al. 1985). Up to date α-defensins were also found to be present in human (Gabay et al. 1989; Ganz et al. 1985; Jones and Bevins 1992, 1993; Patil et al. 2004; Territo et al. 1989; Wilde et al. 1989) and non-human primates (Patil et al. 2004; Tanabe et al. 2004; Tang et al. 1999a), mice (Ouellette et al. 1992, 1994, Patil et al. 2004), hamsters and rats (Patil et al. 2004) (reviewed in Raj and Dentino 2002). Because no α-defensins from outside the mammals were detected so far, they seem to be specific to mammals.
β-Defensins were also detected in numerous mammalian species, e.g. primates (Boniotto et al. 2003a,b; Del Pero et al. 2002), human (Bensch et al. 1995; Schutte et al. 2002), cattle (Diamond et al. 1991; Selsted et al. 1993), goat (Zhao et al. 1999), sheep (Huttner et al. 1998) and mice (Bals et al. 1998; Huttner et al. 1997; Jia et al. 2000; Schutte et al. 2002), as well as in birds (Brockus et al. 1998; Harwig et al. 1994), where they are called gallinacins.
θ-Defensins are cyclic defensins that have been identified so far only in primates (Nguyen et al. 2003; Tang et al. 1999b). They are generated by post-translational ligation of two truncated α-defensins (Tang et al. 1999b).
α- and β-defensins seem to be distinct regarding their tissues of expression; α-defensins are mainly expressed by leucocytes and Paneth cells of the small intestine (Ganz 2003; Lehrer and Ganz 2002; Ouellette 2004; Schutte and McCray 2002), vertebrate β-defensins are predominantly produced by epithelial cells lining various organs, e.g. the human skin and bronchial tree, the tongue and the genito-urinary tract (Diamond et al. 1991; Diamond and Bevins 1998; Harder et al. 1997; Schonwetter et al. 1995).
Several studies were carried out to examine the evolutionary relationships of α- and β-defensins detected in several mammalian species (Hughes 1999; Hughes and Yeager 1997; Liu et al. 1997; Morrison et al. 2003). Based on the complete genome sequences of mouse, human and chicken, some studies applied a computational search for defensin coding sequences (Patil et al. 2004; Schutte et al. 2002; Semple et al. 2003; Xiao et al. 2004). In conclusion, the defensin genes were found to be arranged in clusters (Maxwell et al. 2003; Morrison et al. 2003; Patil et al. 2004; Schutte et al. 2002; Semple et al. 2003; Xiao et al. 2004), characterised by a high sequence divergence of discrete genes. This data further indicated rapid duplication and diversifying positive selection of both α- and β-defensins. In addition, the physical linkage of α- and β-defensin clusters in the genome of mouse and human was detected (Liu et al. 1997; Schutte et al. 2002), which may suggest a common ancestry of α- and β-defensins.
However, little is known about the repertoire and evolutionary relatedness of β-defensins belonging to mammalian species outside from the sequenced ones. Studies with numerous mammalian defensins suggested several duplication events following positive selection and diversification within each group studied (Antcheva et al. 2004; Hughes 1999; Hughes and Yeager 1997; Maxwell et al. 2003; Morrison et al. 2003; Semple et al. 2003). In a similar approach, Luenser and Ludwig (2005) estimated the sequence divergence and molecular evolution of bovine β-defensins and found evidence for a common ancestry of different subfamilies of β-defensins as well as lineage-specific evolution. However, all these studies have in common is to be solely based on the defensin's exons, which in several cases were shown to evolve under positive selection. Positively selected genes are characterised by an accelerated evolution likely rendering the development of similar features. Thus, detection of sequence similarity may be explained by a common ancestry as well as by a process of convergent adaptation. It is therefore useful for evolutionary investigations to compare sequences evolving under a neutral mode of evolution. For β-defensins, such sequence is the intron separating exon 1 and exon 2 (Fig. 1). Here we present an array of newly detected β-defensins of members of the tribe Caprini, obtained via a PCR-based genomic search, and explain their evolutionary history, accentuating their importance for the vertebrate immune defence.
Genetic structure of a β-defensin gene. Lengths of both exons and of the intron as well as the splicing sites of the mRNA are shown. The primary translation product is an unmature precursor consisting of a signalpeptide at the N-terminus, a mature peptide at the C-terminus and an intervening propiece
Materials and methods
The following species of the genera Ammotragus, Pseudois, Capra and Ovis, all belonging to the tribe Caprini, were used for molecular investigation: Capra hircus (domestic goat), Ovis aries (domestic sheep), Ovis ammon (argali sheep), Pseudois nayaur (blue sheep), Ammotragus lervia (barbary sheep) and Ovis orientalis (urials and mouflons), represented by two subspecies: O. orientalis severtzovi (Severtzov's urial) and O. orientalis cycloceros (Afghan urial). Nomenclature and common names are based on the geographic origin of samples following the recommendations of the International Union for Conservation of Nature and Natural Resources (IUCN) Wild Sheep and Goat Specialist Group (Shackleton 1997). Total DNA was purified from blood and tissue samples according to standard protocols (QIAGEN Inc., The Netherlands). For amplification of the complete defensin genes, we designed the primers P1 5′-ATCAGCTGCAGAGCTCGTGA-3′ and P2 5′-CAACCTCAATGACCAGTGG-3′. The forward primer (P1) annealed 27 bp upstream of the first exon, and the reverse primer (P2) annealed 132 bp downstream of the second exon. Amplification was performed using standard conditions, with annealing at 58°C. PCR products, 1,860 bp in length, were cloned with the TA-TOPO cloning kit (Invitrogen, The Netherlands). The cloned PCR products were sequenced with several internal primers. Sequencing was carried out on an automated ABI 3100 DNA sequencer (Applied Biosystem, USA). Coding sequences composed of exons 1 and 2 were aligned with published β-defensin genes retrieved from GenBank (AJ009877, Y17679, U75250, U75251) using molecular evolutionary genetics analysis (MEGA) 2.0 (Kumar et al. 1993). The assignment of the ovine and caprine β-defensins obtained in this study to the different defensins was performed using nucleotide Basic Local Alignment Search Tool (BLAST) search in the National Center for Biotechnology Information (NCBI) GenBank. The phylogenetic trees were constructed by neighbor-joining (NJ) algorithms implemented in phylogenetic analysis using parsimony (*and other methods) (PAUP*) 4.0b10 (Swofford 2002) and MEGA 2.0 (Kumar et al. 1993) based on the proportion of nucleotide sites at which the sequences compared were different. Their reliability was assessed by 1,000 bootstrap replications. The significance of branch lengths in the NJ tree was also examined by a standard error test using the confidence probability (CP) program of MEGA. To test if sequences followed the neutral model of molecular evolution (Kimura 1969), the following tests, implemented in the software package DnaSP (v. 4.0; Rozas et al. 2003), were carried out: Hudson–Kreitman–Aguade (HKA) test (Hudson et al. 1987), Tajima's D test (Tajima 1989), Fu and Li's D* test (Fu and Li 1993) and McDonald–Kreitman test (McDonald and Kreitman 1991). The model of evolution was determined using MODELTEST 3.6 (Posada and Crandall 1998). Calculation of the number of synonymous (dS) substitutions per synonymous site and non-synonymous (dN) substitutions per non-synonymous site was done by the method of Nei and Gojobori (1986), likewise using the program DnaSP4.0 (Rozas et al. 2003).
Results
Genomic DNA of specimens belonging to the tribe Caprini, representing two domestic species (O. aries and C. hircus) and four related wild species (O. ammon, P. nayaur, A. lervia and O. orientalis, represented by the two subspecies, O. orientalis cycloceros and O. orientalis severtzovi), was screened for β-defensin genes. Defensin genes of Caprini are composed of two exons with 58 bp (exon 1) and 137 bp (exon 2), respectively, which enclose a single intron of 1,506 bp (Fig. 1). Sequences of exons 1 and 2 are reported in Table 1.
A total of 13 discrete β-defensin coding sequences was detected (Table 2), including two sequences already known from O. aries: sheep β-defensin 1 [sbd1 (U75250)] and sheep β-defensin 2 [sbd2 (U75251)]. Interestingly, in this study, sbd1 was detected in the related wild sheep, O. orientalis cycloceros. Newly detected sequences identified as variants of previously published defensins from C. hircus (Y17679 and AJ009877) and O. aries (U75250 and U75251) were numbered consecutively (Table 2). With the exception of defensin genes from the wild and the highly endangered species P. nayaur, all defensin genes can be designated as variants of either C. hircus β-defensins 1 [chbd1 (Y17679)] and 2 [chbd2 (AJ009877)], or sheep β-defensins 1 [sbd1 (U75250)] and 2 [sbd2 (U75251)].
Comparison of all variants demonstrated high substitution rates, both inter- and intraspecifically. To understand these substitution patterns, we analysed the degree of phylogenetic relatedness of the detected caprine and ovine β-defensins. Concerning our aim to reconstruct the evolutionary history of newly identified defensins, we regarded the non-coding intron sequences to be most suitable for such analysis.
The NJ tree, based on the intron sequences of ovine and caprine defensins (Fig. 2), shows that β-defensins of the tribe Caprini form four distinct species-specific lineages: (1) ovine defensins harbouring all detected defensins of Ovis, (2) caprine defensins harbouring defensins of Capra, (3) defensins of Pseudois and (4) defensins of Ammotragus. Obviously, β-defensins of Ovis, Capra, Pseudois and Ammotragus are homologues derived from a common ancestor. Species-specific clustering indicates lineage-specific evolution. Interestingly, defensins of Capra and Pseudois form a monophyletic group, indicating a closer relation between defensins from Pseudois and Capra than between those of Pseudois and Ovis. Separation of these clusters is supported by high bootstrap values (Fig. 2).
NJ tree based on the proportion difference of nucleotides, using exclusively intron sequence information. Sequences identical in coding regions but differing in introns are termed as intron variants of the corresponding gene (sbd2/I1, sbd2/I2 etc.). The reliability of each branch was tested by 1,000 bootstrap replications. Sequences cluster in four homologous groups according to the phylogenetic position of their hosts
The defensin gene tree, based on the exon regions (Fig. 3), shows that sequences of Pseudois, Ammotragus and Capra grouped together separated from the wild sheep defensins. Thus, the exon tree contradicts the presented intron tree.
Using the hierarchical likelihood ratio tests (hLRTs) implemented in MODELTEST (Posada and Crandall 1998), the JC+Γ model of evolution was determined to fit the data set best. It consists of the equal-base frequency model of Jukes and Cantor (1969) combined with among-site rate variation reflected by a gamma (Γ) shape distribution parameter of α=0.1456. The likelihood of the JC+Γ model was −lnL=584.93. None of the tests of neutral evolution (HKA, D, D* and McDonald–Kreitman) found significant deviations from the null hypotheses of non-neutral evolution (Table 3). Thus, we tested whether positive or negative selection acted on the defensin genes by estimating dS and dN substitutions. By grouping homologous sequences as indicated in the NJ tree (Fig. 2), we computed dN and dS for all pairwise comparisons within the four species-specific lineages: (1) Ovis defensins, (2) Capra defensins, (3) Pseudois defensins and (4) Ammotragus defensins (Table 4 and Fig. 4). In the case of Ammotragus defensins, we found no substitutions, although differences within the intron revealed the existence of at least three sequences. Substitution patterns of Pseudois defensins indicated negative selection due to dN values being lower than those of dS. Comparison of substitutions within ovine defensins showed positive selection if the comparison included all ovine sequences (Table 4). Analysis of substitutions referring to single genes indicated that some ovine defensins are under negative selection (Table 4).
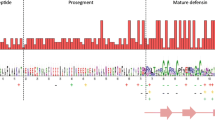
Finally, we examined the extent of constraint at the amino acid level of the presented defensins. Alignment of deduced amino acid sequences (Fig. 5) showed conservation of the signal peptide and the six-cysteine array within the mature peptide. Signal peptides were found to be hydrophobic and rich in leucines, with six and seven leucines respectively, distributed over a length of 19 amino acids. The signalpeptides of all defensins were identical with the exception of chbd2, chbd2_1 and chbd1, differing at one, respectively two sites carrying substituents with similar features (Fig. 5). The mature defensins are positively charged due to the presence of up to five lysines and seven arginines within a range of 38 amino acids. In addition, the six-cysteine motif, as well as the residue positions 47–53, were conserved. All together, we found 22 out of the 38 sites of the mature peptide to be conserved. Three additional sites of the mature peptide show moderate conservation (substitution of residues with similar physico-chemical features). The remaining 13 sites are characterised by radical non-synonymous changes (Fig. 5).
Multiple sequence alignment of the deduced amino acid sequences of caprine and ovine β-defensins. Sites found to be strictly conserved among Caprini are marked by black boxes. Sites with moderate conservation are marked by grey boxes. Changes resulting in substituents with different features are indicated by stars below the alignment. Conserved sites also identical within, human and bovine defensins are also labelled by black boxes. Locations of the signal peptide, the mature peptide and the intervening region of the propiece are depicted
Discussion
In this study, we investigated the variability and evolutionary relatedness of β-defensins of the tribe Caprini. Due to the lack of genome sequence data bank information which would allow a computational search, we screened the genome via PCR. Our genomic search revealed 13 discrete sequences, including two already known from O. aries: sbd1 (U75250) and sbd2 (U75251). Comparison of newly detected sequences with GenBank entries revealed a high sequence similarity to previously known defensins from O. aries [sbd1 (U75259) and sbd2 (U75251)] and C. hircus [chbd1 (Y17679) and chbd2 (AJ009877)]. For this reason, we classified all sequences detected in species belonging to the genera Ovis and Capra as variants of either sbd1 and 2 or of chbd1 and 2. Defensins from P. nayaur showed also remarkable sequence similarity, but due to their distinct origin, we classified them as a new group and named them according to their host, P. nayaur defensin 1, P. nayaur defensin 2 and P. nayaur defensin 3 (pn_1, pn_2 and pn_3).
Interestingly, although the tribe Caprini shows a broad spectrum of discrete and diversified defensins, we identified three orthologues showing sequence identity. Orthologues are genes derived from a speciation event, thus they are present in discrete species and have a common ancestor (in contrast to paralogues which are derived by duplication within single individuals). The first one was sbd1, which, previously known from O. aries, was detected in this study in O. orientalis severzovi, the second was sbd2_2, detected in O. ammon and O. orientalis severtzovi and the third one was sbd1_1, found in O. aries and A. lervia (Table 2). The existence of orthologous β-defensin genes, being identical in their coding regions, was already demonstrated for three bovine species (Bos taurus, Bos frontalis and Bos javanicus) (Luenser and Ludwig 2005). Highly similar or even identical genes, distributed over different species, may provide evidence for a common ancestry of these genes.
However, to understand the relationships of the presented defensin genes, we calculated a NJ tree based on their intron sequences. The resulting gene tree groups all sequences into four clearly distinct clusters, with each cluster composed of numerous genes belonging to one taxon (Ovis, Capra, Pseudois and Ammotragus). According to the branching order, we propose the existence of a common ancestral β-defensin gene, which became subject to lineage-specific evolution, leading to the ancestral defensins of the genus Ammotragus as well as an ancestral defensin common for the genera Ovis, Capra and Pseudois. Subsequent speciation resulted in the divergence of defensins of Ovis from Capra and Pseudois, followed by an additional split into the lineages of Capra and Pseudois. After diversification of these distinct lineages, the ancestral defensins duplicated repeatedly and independently in each lineage/genus. As a result of this scenario, the detected defensins were distributed species-specifically in numerous copies. As mentioned above, our phylogenetic analyses were based on the intron sequences. Comparison restricted to the coding regions will result in an underestimation of the number of β-defensin genes due to the existence of strongly conserved orthologues and paralogues. In detail, we detected conserved paralogous defensins in P. nayaur (pn_1/I1–I6), C. hircus (chbd1_1/I1–I4; chbd2_1/I1 and I2), O. orientalis severtzovi (sbd2_2/I1 and I3; sbd2_3/I1–I3), O. orientalis cycloceros (sbd2_1/I1–I3) and O. ammon (sbd2_4/I1 and I2). Therefore, we postulate numerous duplication events within certain species; for instance, duplication of an ancestral C. hircus defensin resulted in chbd1 and chbd2. Both genes duplicated independently, leading to four variants of chbd1 (chbd1_1/I1, chbd1_1/I2, chbd1_1/I3 and chbd1_1/I4) and two variants of chbd2 (chbd2_1/I1 and chbd2_1/I2), only distinguished by their introns. Thus, we assume that at least four duplication events occurred within C. hircus. Genesis of the whole array of β-defensins within the genera Ammotragus, Pseudois and Ovis occurred most likely in a similar way.
Interestingly, defensin genes of Pseudois are more closely related to those of Capra than to those of Ovis. The phylogenetic position of Pseudois has long been discussed controversially; because of the existence of both goat- and sheep-like features, Pseudois was often referred to as an intermediate. Comparison of the 12S rDNA sequences grouped Pseudois together with Capra, separated from Ovis (Ludwig and Fischer 1998). Resembling these findings, our gene tree is in agreement with the current opinion of the Caprini phylogeny.
In conclusion, the gene tree clearly indicates a homology of all caprine defensins and provides evidence for a lineage-specific evolution. Homologous genes were duplicated repeatedly and independently within each lineage. Redundant gene copies enabled accumulation of mutations by chance; such mutations may endow this gene with a new function (Ohno 1973). Diversification may reflect species-specific adaptation to the local environment or ecological niches. In addition, certain duplicates seem to be strongly conserved (sbd2_1, sbd2_2, sbd2_3 and sbd2_4; chbd2_1, chbd1_1, sbd1_1 and pn1_1).
Gene duplication is regarded to be a key step for the creation of new genes and genes with a new function (Ohno 1970). Due to the prevalence of duplicate genes in genomes of all three domains of life (archaebacteria, bacteria and eucaryotes), the idea of gene duplication has risen to a general principle in evolutionary biology. Examples for duplicated defensins were provided by numerous studies (Hughes 1999; Lynn et al. 2004; Maxwell et al. 2003).
However, while in Ohno's (1970) view, the duplicate may gain a new function (neofunctionalisation), balanced by the other which is retaining the ancestral function, our study provides evidence not only for neofunctionalisation but also for maintaining identical duplicates. Redundancy may directly be advantageous as a mechanism for compensation of deleterious mutations (Clark 1994) and/or developmental accidents (Nowak et al. 1997). In addition, redundancy enables the generation of large amounts of gene products (Zhang 2003). This idea of redundancy being beneficial by providing large amounts of transcripts is supported by the fact that β-defensins are synthesised de novo upon stimulation and not stored like α-defensins (Brockus et al. 1998; Russell et al. 1996; Schonwetter et al. 1995). Yet there is a problem: if a gene is truly redundant, it would not be protected against an accumulation of deleterious mutations, which would lead to a purifying selection of that copy. Therefore, unless the presence of an extra amount of gene product may be advantageous, two genes with identical functions are unlikely to be stably maintained in the genome (Nowak et al. 1997). Theoretical population genetics predicts that both duplicates can be maintained when they differ in some aspects of their function (Nowak et al. 1997). Other than Ohno's (1970) model of neofunctionalisation, differing function can also be reached by division of gene expression after gene duplication (subfunctionalisation) (Force et al. 1999). Detection of two isoforms of a goat defensin expressed specifically in digestive and respiratory tissues (Zhao et al. 1999) may represent such a process of subfunctionalisation.
Comparison of both trees (intron tree and exon tree) depicts the poorly resolved deep branches of the exon tree. The distinct phylogenetic origin of certain defensins, as retrieved from the intron tree, is only poorly reflected and lost. For example, all variants of sbd1_1 representing defensins of the two distinct genera Ovis and Ammotragus are grouped together and close to the ovine defensins sbd1 from domestic sheep and sbd1/I1 from argali sheep, although early speciation separated ancestral Ammotragus defensins (sbd1_1/I2, sbd1_1/I3 and sbd1_1/I4) from all the others (Fig. 2). In addition, certain species-specific duplication events are not reflected in the tree due to exon identity (pn_1, chbd1_1, chbd2_1, sbd2_1, sbd2_2, sbd2_3 and sbd2_4). Obviously, certain defensins are strongly conserved in their coding regions, both intra- as well as interspecifically; therefore, detection of lineage-specific separation of an ancestral defensin and their independent duplication events within distinct lineages requires the information stored in the introns.
However, resuming the branching order of the gene trees, we found evidence for diversifying positive selection, as indicated by the broad array of discrete defensins within a common species, as well as conservation indicated by the existence of identical exons distributed inter- and intraspecifically. To understand the underlying mode of evolution, we first tested for the neutral mode of evolution. Several synonymous nucleotide changes were detected within caprine and ovine defensin genes. The fixation of synonymous substitutions conforms to the neutral theory of molecular evolution (Kimura 1977) and is expected for most genes (Nei 1987). However, the application of a statistical test checking for the neutral mode of evolution clearly demonstrated that the observed variations did not fit that model (Table 3). Therefore, the presence of selection pressure was assumed. Subsequently, we tested for positive and negative selection via a comparison of synonymous and non-synonymous substitutions. In some gene comparisons, the rate of synonymous substitutions exceeded the rate of non-synonymous substitutions. This was the case for all defensins of C. hircus and P. nayaur, indicating that these genes did not evolve under positive-selection pressure. A. lervia defensins showed neither synonymous nor non-synonymous substitutions, a situation which may be regarded as a special case of negative selection. Comparison of ovine defensins revealed positive selection if all sequences were included. Analysis of discrete genes also indicated that some genes were obviously negatively selected (Table 4). In conclusion, comparison of caprine β-defensins showed both positively as well as negatively selected genes (Table 4 and Fig. 4). Such examples were recently also described by others for defensins evolving under positive (e.g. Hughes 1999; Lynn et al. 2004; Luenser and Ludwig 2005; Maxwell et al. 2003) and negative selections (e.g. Del Pero et al. 2002; Luenser and Ludwig 2005).
Comparison of the deduced amino acid sequences of the mature peptides showed moderate changes, which did not affect the net charge of the peptides. The mature peptide consists of 38 sites, out of which 22 sites were found to be conserved within the Caprini (Fig. 5). These sites included the six cysteines, the region from position 47–52 and some single positions. Thirteen positions within the mature peptide were found to be substituted by amino acids with different features (Fig. 5). It is likely that these changes affected the biological activity, but in which way is unclear. It is known from other defensins that small changes in primary structure can have an enormous effect on their potency. Two human neutrophil peptides (HNP1 and HNP3), which differ by only one amino acid, show drastic differences regarding their potency against Candida albicans: While the addition of HNP1 to C. albicans results in the yeast's death, that power is totally lost in HNP3 (Lehrer et al. 1988; Raj et al. 2000). A similar observation was made for two mouse α-defensins (cryptdin 4 and cryptdin 6). Variants of cryptdin 4 and cryptdin 6, truncated at the N-terminus for one and two sites, respectively, showed markedly less anti-microbial activity (Ouellette et al. 2000). However, we can only speculate which changes of toxicity or specificity may result from the 13 positions found to be variable within the mature caprine β-defensins. To gain more information, studies about the expression of these newly detected genes need to be carried out. Perhaps, there are differences regarding the tissues where they are expressed, resembling the findings of Zhao et al. (1999), who detected two isoforms of goat defensins expressed specifically in distinct tissues.
In conclusion, we detected a broad array of defensin genes which was shown to be monophyletic and which we suggest to be the result of several rounds of gene duplication events. Comparison of the coding sequences suggested distinct fates of the duplicates. We found evidence of neofunctionalisation, indicated by sequence diversification, as well as evidence for subfunctionalisation, which may explain the conservation of exons. Numerous duplicates may have been lost due to deleterious mutations and purifying selection. However, our data accentuate how important and informative the inclusion of the defensin introns is for the reconstruction of phylogenetic history. Obviously, evolutionarily stably maintained duplicates on one side and their diversification on the other provide evidence for rapid evolution and confirm the importance of the innate immunity despite the existence of the adaptive immunity.
References
Antcheva N, Boniotto M, Zelezetsky I, Pacor S, Falzacappa MV, Crovella S, Tossi A (2004) Effects of positively selected sequence variations in human and Macaca fascicularis beta-defensins 2 on antimicrobial activity. Antimicrob Agents Chemother 48:685–688
Bals R, Goldman MJ, Wilson JM (1998) Mouse beta-defensin 1 is a salt-sensitive antimicrobial peptide present in epithelia of the lung and urogenital tract. Infect Immun 66:1225–1232
Bensch KW, Raida M, Magert HJ, Schulz-Knappe P, Forssmann WG (1995) hBD-1: a novel beta-defensin from human plasma. FEBS Lett 368:331–335
Boman HG (1991) Antibacterial peptides: key components needed in immunity. Cell 65:205–207
Boniotto M, Antcheva N, Zelezetsky I, Tossi A, Palumbo V, Verga Falzacappa MV, Sgubin S, Braida L, Amoroso A, Crovella S (2003a) A study of host defence peptide beta-defensin 3 in primates. Biochem J 374:707–714
Boniotto M, Tossi A, DelPero M, Sgubin S, Antcheva N, Santon D, Masters J, Crovella S (2003b) Evolution of the beta defensin 2 gene in primates. Genes Immun 4:251–257
Brockus CW, Jackwood MW, Harmon BG (1998) Characterization of beta-defensin prepropeptide mRNA from chicken and turkey bone marrow. Anim Genet 29:283–289
Clark AG (1994) Invasion and maintenance of a gene duplication. Proc Natl Acad Sci U S A 91:2950–2954
Del Pero M, Boniotto M, Zuccon D, Cervella P, Spano A, Amoroso A, Crovella S (2002) Beta-defensin 1 gene variability among non-human primates. Immunogenetics 53:907–913
Diamond G, Bevins CL (1998) beta-Defensins: endogenous antibiotics of the innate host defense response. Clin Immunol Immunopathol 88:221–225
Diamond G, Zasloff M, Eck H, Brasseur M, Maloy WL, Bevins CL (1991) Tracheal antimicrobial peptide, a cysteine-rich peptide from mammalian tracheal mucosa: peptide isolation and cloning of a cDNA. Proc Natl Acad Sci U S A 88:3952–3956
Force A, Lynch M, Pickett FB, Amores A, Yan YL, Postlethwait J (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531–1545
Fu YX, Li WH (1993) Statistical tests of neutrality of mutations. Genetics 133:693–709
Gabay JE, Scott RW, Campanelli D, Griffith J, Wilde C, Marra MN, Seeger M, Nathan CF (1989) Antibiotic proteins of human polymorphonuclear leukocytes. Proc Natl Acad Sci U S A 86:5610–5614
Ganz T (2003) Defensins: antimicrobial peptides of innate immunity. Nat Rev Immunol 3:710–720
Ganz T, Selsted ME, Szklarek D, Harwig SS, Daher K, Bainton DF, Lehrer RI (1985) Defensins. Natural peptide antibiotics of human neutrophils. J Clin Invest 76:1427–1435
Hancock RE, Lehrer RI (1998) Cationic peptides: a new source of antibiotics. Trends Biotechnol 16:82–88
Harder J, Bartels J, Christophers E, Schroder JM (1997) A peptide antibiotic from human skin. Nature 387:861
Harwig SS, Swiderek KM, Kokryakov VN, Tan L, Lee TD, Panyutich EA, Aleshina GM, Shamova OV, Lehrer RI (1994) Gallinacins: cysteine-rich antimicrobial peptides of chicken leukocytes. FEBS Lett 342:281–285
Hoover DM, Boulegue C, Yang D, Oppenheim JJ, Tucker K, Lu W, Lubkowski J (2002) The structure of human macrophage inflammatory protein-3alpha /CCL20. Linking antimicrobial and CC chemokine receptor-6-binding activities with human beta-defensins. J Biol Chem 40:37647–37654
Hudson RR, Kreitman M, Aguade M (1987) A test of neutral molecular evolution based on nucleotide data. Genetics 116:153–159
Hughes AL (1999) Evolutionary diversification of the mammalian defensins. Cell Mol Life Sci 56:94–103
Hughes AL, Yeager M (1997) Coordinated amino acid changes in the evolution of mammalian defensins. J Mol Evol 44:675–682
Huttner KM, Kozak CA, Bevins CL (1997) The mouse genome encodes a single homolog of the antimicrobial peptide human beta-defensin 1. FEBS Lett 413:45–49
Huttner KM, Lambeth MR, Burkin HR, Burkin DJ, Broad TE (1998) Localization and genomic organization of sheep antimicrobial peptide genes. Gene 206:85–91
Jia HP, Wowk SA, Schutte BC, Lee SK, Vivado A, Tack BF, Bevins CL, McCray PB Jr (2000) A novel murine beta-defensin expressed in tongue, esophagus, and trachea. J Biol Chem 275:33314–33320
Jones DE, Bevins CL (1992) Paneth cells of the human small intestine express an antimicrobial peptide gene. J Biol Chem 267:23216–23225
Jones DE, Bevins CL (1993) Defensin-6 mRNA in human Paneth cells: implications for antimicrobial peptides in host defense of the human bowel. FEBS Lett 315:187–192
Jukes T, Cantor C (1969) Evolution of protein molecules. In: Munro HN (ed) Mammalian protein metabolism. Academic, New York, pp 21–132
Kagan BL, Selsted ME, Ganz T, Lehrer RI (1990) Antimicrobial defensin peptides form voltage-dependent ion-permeable channels in planar lipid bilayer membranes. Proc Natl Acad Sci U S A 87:210–214
Kimura M (1969) The number of heterozygous nucleotide sites maintained in a finite population due to steady flux of mutations. Genetics 61:893–903
Kimura M (1977) Preponderance of synonymous changes as evidence for the neutral theory of molecular evolution. Nature 267:275–276
Kumar S, Tamura K, Nei M (1993) MEGA: Molecular Evolutionary Genetics Analysis, version 1.01. Pennsylvania State University, University Park, PA
Lehrer RI, Ganz T (2002) Defensins of vertebrate animals. Curr Opin Immunol 14:96–102
Lehrer RI, Ganz T, Szklarek D, Selsted ME (1988) Modulation of the in vitro candidacidal activity of human neutrophil defensins by target cell metabolism and divalent cations. J Clin Invest 8:1829–1835
Lehrer RI, Lichtenstein AK, Ganz T (1993) Defensins: antimicrobial and cytotoxic peptides of mammalian cells. Annu Rev Immunol 11:105–128
Liu L, Zhao C, Heng HH, Ganz T (1997) The human beta-defensin-1 and alpha-defensins are encoded by adjacent genes: two peptide families with differing disulfide topology share a common ancestry. Genomics 43:316–320
Ludwig A, Fischer S (1998) New aspects of an old discussion—phylogenetic relationships of Ammotragus and Pseudois within the subfamily Caprinae based on comparison of the 12S rDNA sequences. J Zoolog Syst Evol Res 36:173–178
Luenser K, Ludwig A (2005) Variability and evolution of bovine beta-defensin genes. Genes Immun 6:115–122
Lynn DJ, Lloyd AT, Fares MA, O'Farrelly C (2004) Evidence of positively selected sites in mammalian alpha-defensins. Mol Biol Evol 21:819–827
Martin E, Ganz T, Lehrer RI (1995) Defensins and other endogenous peptide antibiotics of vertebrates. J Leukoc Biol 58:128–136
Maxwell AI, Morrison GM, Dorin JR (2003) Rapid sequence divergence in mammalian beta-defensins by adaptive evolution. Mol Immunol 40:413–421
McDonald JH, Kreitman M (1991) Adaptive protein evolution at the Adh locus in Drosophila. Nature 351:652–654
Morrison GM, Semple CA, Kilanowski FM, Hill RE, Dorin JR (2003) Signal sequence conservation and mature peptide divergence within subgroups of the murine beta-defensin gene family. Mol Biol Evol 20:460–470
Nei M (1987) Molecular evolutionary genetics. Columbia University Press, New York, pp 79–83
Nei M, Gojobori T (1986) Simple methods for estimating the numbers of synonymous and nonsynonymous nucleotide substitutions. Mol Biol Evol 3:418–426
Nguyen TX, Cole AM, Lehrer RI (2003) Evolution of primate theta-defensins: a serpentine path to a sweet tooth. Peptides 24:1647–1654
Nicholas P, Mor A (1995) Peptides as weapons against microorganisms in the chemical defense system of vertebrates. Annu Rev Microbiol 49:227–304
Nowak MA, Boerlijst MC, Cooke J, Smith JM (1997) Evolution of genetic redundancy. Nature 388:167–171
Ohno S (1970) Evolution by gene duplication. Springer, Berlin Heidelberg New York
Ohno S (1973) Ancient linkage groups and frozen accidents. Nature 244:259–262
Ouellette AJ (2004) Defensin-mediated innate immunity in the small intestine. Best Pract Res Clin Gastroenterol 18:405–419
Ouellette AJ, Miller SI, Henschen AH, Selsted ME (1992) Purification and primary structure of murine cryptdin-1, a Paneth cell defensin. FEBS Lett 304:146–148
Ouellette AJ, Hsieh MM, Nosek MT, Cano-Gauci DF, Huttner KM, Buick RN, Selsted ME (1994) Mouse Paneth cell defensins: primary structures and antibacterial activities of numerous cryptdin isoforms. Infect Immun 62:5040–5047
Ouellette AJ, Satchell DP, Hsieh MM, Hagen SJ, Selsted ME (2000) Characterization of luminal Paneth cell alpha-defensins in mouse small intestine. Attenuated antimicrobial activities of peptides with truncated amino termini. J Biol Chem 275:33969–33973
Patil A, Hughes AL, Zhang G (2004) Rapid evolution and diversification of mammalian alpha-defensins as revealed by comparative analysis of rodent and primate genes. Physiol Genomics 20:1–11
Posada D, Crandall KA (1998) MODELTEST: testing the model of DNA substitution. Bioinformatics 14:817–818
Raj PA, Dentino AR (2002) Current status of defensins and their role in innate and adaptive immunity. FEMS Microbiol Lett 206:9–18
Raj PA, Antonyraj KJ, Karunakaran T (2000) Large-scale synthesis and functional elements for the antimicrobial activity of defensins. Biochem J 347:633–641
Rozas J, Sanchez-DelBarrio JC, Messeguer X, Rozas R (2003) DnaSP, DNA polymorphism analyses by the coalescent and other methods. Bioinformatics 19:2496–2497
Russell JP, Diamond G, Tarver AP, Scanlin TF, Bevins CL (1996) Coordinate induction of two antibiotic genes in tracheal epithelial cells exposed to the inflammatory mediators lipopolysaccharide and tumor necrosis factor alpha. Infect Immun 64:1565–1568
Satchell DP, Sheynis T, Shirafuji Y, Kolusheva S, Ouellette AJ, Jelinek R (2003) Interactions of mouse Paneth cell alpha-defensins and alpha-defensin precursors with membranes. Prosegment inhibition of peptide association with biomimetic membranes. J Biol Chem 278:13838–13846
Schonwetter BS, Stolzenberg ED, Zasloff MA (1995) Epithelial antibiotics induced at sites of inflammation. Science 267:1645–1648
Schröder JM (1999) Epithelial antimicrobial peptides: innate local host response elements. Cell Mol Life Sci 56:32–46
Schutte BC, McCray PB Jr (2002) [beta]-Defensins in lung host defense. Annu Rev Physiol 64:709–748
Schutte BC, Mitros JP, Bartlett JA, Walters JD, Jia HP, Welsh MJ, Casavant TL, McCray PB Jr (2002) Discovery of five conserved beta-defensin gene clusters using a computational search strategy. Proc Natl Acad Sci U S A 99:2129–2133
Selsted ME, Brown DM, DeLange RJ, Harwig SS, Lehrer RI (1985) Primary structures of six antimicrobial peptides of rabbit peritoneal neutrophils. J Biol Chem 260:4579–4584
Selsted ME, Tang YQ, Morris WL, McGuire PA, Novotny MJ, Smith W, Henschen AH, Cullor JS (1993) Purification, primary structures, and antibacterial activities of beta-defensins, a new family of antimicrobial peptides from bovine neutrophils. J Biol Chem 268:6641–6648
Semple CA, Rolfe M, Dorin JR (2003) Duplication and selection in the evolution of primate beta-defensin genes. Genome Biol 4:R31. DOI 10.1186/gb-2003-4-5-r31
Shackleton DM, IUCN/SSC Caprinae Specialist Group (1997) Wild sheep and goats and their relatives. Status survey and conservation action plan for Caprinae. IUCN, Gland, Switzerland
Swofford DL (2002) PAUP*: phylogenetic analysis using parsimony (*and other methods). Sinauer Associates, Sunderland, MA
Tajima F (1989) Statistical method for testing the neutral mutation hypothesis by DNA polymorphism. Genetics 123:585–595
Tanabe H, Yuan J, Zaragoza MM, Dandekar S, Henschen-Edman A, Selsted ME, Ouellette AJ (2004) Paneth cell alpha-defensins from rhesus macaque small intestine. Infect Immun 72:1470–1478
Tang YQ, Yuan J, Miller CJ, Selsted ME (1999a) Isolation, characterization, cDNA cloning, and antimicrobial properties of two distinct subfamilies of alpha-defensins from rhesus macaque leukocytes. Infect Immun 67:6139–6144
Tang YQ, Yuan J, Osapay G, Osapay K, Tran D, Miller CJ, Ouellette AJ, Selsted ME (1999b) A cyclic antimicrobial peptide produced in primate leukocytes by the ligation of two truncated alpha-defensins. Science 286:498–502
Territo MC, Ganz T, Selsted ME, Lehrer RI (1989) Monocyte-chemotactic activity of defensins from human neutrophils. J Clin Invest 84:2017–2020
Wang W, Cole AM, Hong T, Waring AJ, Lehrer RI (2003) Retrocyclin, an antiretroviral theta-defensin, is a lectin. J Immunol 170:4708–4716
Wilde CG, Griffith JE, Marra MN, Snable JL, Scott RW (1989) Purification and characterization of human neutrophil peptide 4, a novel member of the defensin family. J Biol Chem 264:11200–11203
Xiao Y, Hughes AL, Ando J, Matsuda Y, Cheng JF, Skinner-Noble D, Zhang G (2004) A genome-wide screen identifies a single beta-defensin gene cluster in the chicken: implications for the origin and evolution of mammalian defensins. BMC Genomics 5:56
Yang D, Chertov O, Bykovskaia SN, Chen Q, Buffo MJ, Shogan J, Anderson M, Schroder JM, Wang JM, Howard OM, Oppenheim JJ (1999) Beta-defensins: linking innate and adaptive immunity through dendritic and T cell CCR6. Science 286:525–528
Yang D, Biragyn A, Hoover DM, Lubkowski J, Oppenheim JJ (2004) Multiple roles of antimicrobial defensins, cathelicidins, and eosinophil-derived neurotoxin in host defense. Annu Rev Immunol 22:181–215
Zeya HI, Spitznagel JK (1966) Antimicrobial specificity of leukocyte lysosomal cationic proteins. Science 154:1049–1051
Zhang J (2003) Evolution by gene duplication. Trends Ecol Evol 18:292–298
Zhao C, Nguyen T, Liu L, Shamova O, Brogden K, Lehrer RI (1999) Differential expression of caprine beta-defensins in digestive and respiratory tissues. Infect Immun 67:6221–6224
Acknowledgements
We thank A.K. Hett and C. Pitra for helpful comments and discussions. Furthermore, the authors recognise the following veterinarians and biologists: M. Stöck, A. Pauly, G. Straub and G. Wibbelt, for providing samples and information on the health status of feral sheep.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Fig6
Luenser et al. 2005; Evolution of caprine and ovine β-defensin genes Multiple sequence alignment of complete caprine and ovine defensin genes. Vertical numbers indicate nucleotide positions. Dots indicate identical sites referring to sbd2, and dashes indicate gaps introduced by the alignment. Nucleotides of exon1 (1–57) and 2 (1564–1621) are in bold letters. Sequences solely differing in intron regions are subsequently termed as intron variants of that gene (e.g. sbd2_1/I1, sbd2_1/I2 etc.)
Rights and permissions
About this article
Cite this article
Luenser, K., Fickel, J. & Ludwig, A. Evolution of caprine and ovine β-defensin genes. Immunogenetics 57, 487–498 (2005). https://doi.org/10.1007/s00251-005-0001-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-005-0001-5