Abstract
IL-10 inhibits the production of many pro-inflammatory cytokines. Polymorphisms in the IL10 gene promoter at positions −1082G→A, −819C→T and −592C→A occur as three haplotypes, ATA, GCC and ACC. These influence several infectious and inflammatory diseases including community-acquired pneumonia, where a role for IL-10 is suggested by fluctuations in plasma levels of the cytokine. However, the effects of the haplotypes on IL-10 production are unclear. We stimulated peripheral blood mononuclear cells (PBMC) from at least five individuals homozygous for each of the three haplotypes with lipopolysaccharide (LPS, 10 μg/ml) or heat-killed Streptococcus pneumoniae (107cfu/ml) and measured IL-10 mRNA by RT-PCR. Following S. pneumoniae stimulation, PBMC with the ATA haplotype had higher IL-10 mRNA levels than those with the GCC haplotype at 4 h (independent t-test; P=0.024), or the ACC haplotype at 4 h (P<0.0001) and 8 h (P=0.007). Following LPS stimulation, IL-10 mRNA levels were not significantly influenced by the IL10 haplotype, but similar trends were observed, consistent with the variable outcome of published studies. The results suggest that the −819 and/or −592 alleles affect transcription.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Interleukin-10 [IL-10; cytokine synthesis inhibitory factor (CSIF)] is a multifunctional anti-inflammatory cytokine produced by monocytes, macrophages, B cells, T cells and mast cells (de Waal Malefyt and Moore 1998). IL-10 inhibits the production of pro-inflammatory cytokines including TNFα, IL-1, IL-6 and IL-8 (de Waal Malefyt et al. 1991). In sepsis, septic shock and community-acquired pneumonia (CAP), IL-10 serum levels correlate with the intensity of the inflammatory response, the severity of injury and the clinical outcome (Marchant et al. 1994; Gomez-Jimenez et al. 1995; Glynn et al. 1999). This suggests that dysregulation of IL-10 may affect clinical outcome.
Family studies attribute up to 75% of differences in IL-10 production to genetic factors (Westendorp et al. 1997). The IL10 gene is located on Chromosome 1, with multiple single nucleotide polymorphisms (SNP) in the distal and proximal promoter regions (Eskdale et al. 1998). SNP at positions –1082G→A, −819C→T and –592C→A relative to the transcription start site occur as three haplotypes, GCC, ACC and ATA (Turner et al. 1997). These SNP are associated with various inflammatory diseases including CAP (Gallagher et al. 2003; Schaaf et al. 2003), inflammatory bowel disease (Tagore et al. 1999), rheumatoid arthritis (Hajeer et al. 1998), primary Sjogren’s syndrome (Hulkkonen et al. 2001), multiple organ dysfunction in patients with sepsis (Reid et al. 2002) and meningococcal sepsis (van der Pol et al. 2001).
The effects of the IL10 promoter haplotypes on IL-10 production are unclear. Crawley et al. (1999) and Koss et al. (2000) exposed whole blood to LPS and found that individuals with the ATA genotype produced significantly less IL-10 than non-ATA carriers. However, Schaaf et al. (2003) associated the –1082A allele with decreased IL-10 protein levels in similar whole blood assays, while Keijsers et al. (1998) associated low levels with the −1082G allele. Carriage of the ATA haplotype was associated with high IL-10 plasma levels in neonates (Helminen et al. 2001), but this was not apparent in adults with or without injections of LPS (Fijen et al. 2001; Hulkkonen et al. 2001).
A limitation of these studies as models of CAP is that cells were stimulated with the Gram-negative cell wall component, LPS. While Gram-negative bacteria can cause CAP, Streptococcus pneumoniae is more often responsible (Marrie 2000). Moreover many studies have analyzed IL-10 protein in plasma or culture supernatants, rather than mRNA production where the effects of promoter polymorphisms are likely to be clearest. IL-10 protein levels may be affected by factors that modulate translation, secretion and utilization of the cytokine, independent of the IL-10 promoter haplotype. Here we investigated whether IL-10 mRNA levels in cultured peripheral blood mononuclear cells (PBMC) are influenced by SNP in the proximal promoter of IL10 after stimulation with S. pneumoniae or LPS.
Ninety-three healthy Caucasian volunteers from Perth (Western Australia) were genotyped (Table 1), and 16 individuals homozygous for the 3 known haplotypes (ATA, GCC, ACC) were selected for further study (Table 2). These were hospital and laboratory staff, had no chronic illness and were not on any medications or affected by any acute medical condition at the time of this study. They gave written informed consent and the Medical Ethics Committee of the Royal Perth Hospital approved the study. Typing for the −1082, −819 and −592 polymorphisms was performed by a single base extension in multiplexed reactions using Sequenom’s Mass ARRAY technology (Ross et al. 1998). The –819 and –592 polymorphisms were found to be in complete linkage disequilibrium (Table 1).
PBMC from the five or six individuals with each IL10 haplotype were then stimulated for 4 and 8 h with S. pneumoniae or LPS. PBMC were isolated by density gradient centrifugation of heparinized blood on Ficoll-Hypaque and stored in liquid nitrogen in 10% DMSO/90% heat-inactivated (HI) fetal calf serum (FCS). Thawed PBMC were washed once and resuspended at 0.5×106 cells/ml in RPMI/10% HI FCS. Aliquots of 500 μl were placed into 6 ml round-bottom polypropylene tubes with 500 μl RPMI/10% HI FCS with or without 10 μg/ml LPS (from Escherichia coli 0111:B4; Sigma) or 107cfu/ml heat-killed S. pneumoniae, prepared as described previously (Temple et al. 2003), and incubated for 4 and 8 h (37°C, 5% CO2).
Expression of IL-10 mRNA was measured by real time RT-PCR. RNA was extracted from PBMC using the RNeasy Mini Kit (Qiagen, Clifton Hill, Victoria, Australia). RNA (6 μl) was used for first-strand cDNA synthesis using Omniscript reverse transcriptase (Qiagen) and oligo (dT15) primers (Promega, Australia) in a final volume of 20 μl. IL-10 mRNA was quantitated with a forward primer spanning the exon 2–3 junction (5′-TTAAGGGTTACCTGGGTTGC-3′) and a reverse primer spanning the exon 3–4 junction (5′-GGGAAGAAATCGATGACAGC-3′), yielding a 191 bp product. β-Actin was amplified with a forward primer spanning the exon 2–3 junction (5′-GATGACCCAGATCATGTTTGA-3′) and a reverse primer spanning the exon 3–4 junction (5′-GACTCCATGCCCAGGAAGGAA-3′), yielding a 459 bp product. For amplification using the LightCycler (Roche, Indianapolis, Ind., USA), the reaction mixture contained 5 μl cDNA, 20 ng/μl primers (Geneworks, Adelaide, South Australia), 0.25 μg/μl bovine serum albumin (CSL Biosciences, Parkville, Victoria, Australia), 0.08 mm dNTP, 2.0 mm MgCl2 pH8.9 (IL-10) or 2.5 mm MgCl2 pH8.9 (β-actin), 2 μl SYBRGreen 1 (Sigma, St Louis, Mo., USA), 1.5 U Platinum Taq DNA polymerase (Invitrogen Life Technologies, Mount Waverley, Victoria, Australia) in 20 μl. Cycling conditions for IL-10 were 95°C for 5 min; 45 cycles of 96°C for 1 s, 65°C for 5 s, 72°C for 15 s; 96°C for 1 min, 60°C for 5 s, 99°C for 1 min and 40°C for 30 s. For β-actin, the annealing temperature was 64°C and 35 cycles were run. 2% agarose gels containing ethidium bromide were used to confirm amplification of a single product of the correct size. Negative controls were run without cDNA template. Serial 10-fold dilutions of pGEMT-Easy plasmids (Promega, Annandale, NSW, Australia) containing the amplified region of each gene were used to establish standard curves. The method of least squares was used to calculate the initial template concentration using the Lightcycler software. The inter-assay correlation coefficient was 0.97±0.02.
Levels of IL-10 mRNA from the 16 individuals assayed were elevated at 4 h, and continued to increase at 8 h (S. pneumoniae, P=0.01, LPS, P=0.036; compared to unstimulated cells) (Fig. 1). There was no significant difference between cells stimulated with S. pneumoniae or LPS.
Following S. pneumoniae stimulation, PBMC with the ATA haplotype had higher IL-10 mRNA levels than those with the GCC haplotype at 4 h (P=0.024), or the ACC haplotype at 4 h (P<0.0001) and 8 h (P=0.007) (Fig. 2a). A similar trend was evident after LPS stimulation (Fig. 2b), but the differences were not significant (P>0.05).
IL10 mRNA levels after stimulation with a S. pneumoniae (107cfu/ml) and b LPS (10 μg/ml) for 4 and 8 h. Results are presented as the geometric means and 95% confidence intervals for duplicate assays of 5–6 samples per group. *P<0.05, **P<0.01, ***P<0.001, compared with the ATA haplotype (independent t-tests using SPSS v10.1)
After stimulation with S. pneumoniae PBMC with the IL10−819TT or IL10−592AA haplotype had higher IL-10 mRNA levels (4 h, P=0.001; 8 h, P=0.004) than those with IL10−819CC or IL10−592CC (Table 3). In contrast, no significant associations were observed with LPS stimulation. IL-10 mRNA levels were not affected by IL10–1082 (Table 3).
These results establish that IL10 promoter haplotypes influence IL-10 mRNA levels in stimulated PBMC from healthy subjects. S. pneumoniae-stimulated PBMC with the ATA haplotype produced more IL-10 mRNA than those with the ACC and GCC haplotypes after 4 and 8 h. LPS stimulation generated similar trends but no differences were significant. This may account for some variation between published studies.
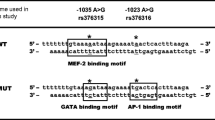
The haplotypic association shown here can be explained by the SNP at −819 and/or –592. These are in complete linkage disequilibrium and lie within positive and negative regulatory regions, respectively (Kube et al. 1995). The −819T and −592A may increase stimulated transcriptional activity by influencing transcription factor binding. However this has not been demonstrated experimentally and other loci should also be considered. For example, nine extended haplotypes, formed by the distal promoter SNP −3575T→A, −2849G→A, and −2763C→A and the proximal IL10 SNP (−1082G→A, −819C→T, and −592C→A, including −1330G→A) have been demonstrated in Caucasians (Gibson et al. 2001). In addition, two microsatellites (IL-10.R/IL-10.G) associated with differential IL-10 protein secretion form four haplotype families with the IL10 haplotypes at positions −1082, −819 and –592 (Eskdale et al. 1998; Eskdale et al. 1999).
Although we have established the IL10−819TT:IL10−592AA haplotype as a determinant of high IL-10 transcription, it is still not clear whether levels of IL-10 protein are affected. Previous studies have obtained variable results using LPS as the stimuli and measuring protein levels of IL-10 (Crawley et al. 1999; Hulkkonen et al. 2001). Our results concur with those of Helminen et al. (2001), who found high plasma IL-10 from neonates with the ATA haplotype. More comprehensive studies are required.
References
Crawley E, Kay R, Sillibourne J, Patel P, Hutchinson I, Woo P (1999) Polymorphic haplotypes of the interleukin-10 5′ flanking region determine variable interleukin-10 transcription and are associated with particular phenotypes of juvenile rheumatoid arthritis. Arthritis Rheum 42:1101–1108
Eskdale J, Gallagher G, Verweij CL, Keijsers V, Westendorp RG, Huizinga TW (1998) Interleukin 10 secretion in relation to human IL-10 locus haplotypes. Proc Natl Acad Sci USA 95:9465–9470
Eskdale J, Keijsers V, Huizinga T, Gallagher G (1999) Microsatellite alleles and single nucleotide polymorphisms (SNP) combine to form four major haplotype families at the human interleukin-10 (IL-10) locus. Genes Immun 1:151–155
Fijen JW, Tulleken JE, Hepkema BG, van der Werf TS, Ligtenberg JJ, Zijlstra JG (2001) The influence of tumor necrosis factor-alpha and interleukin-10 gene promoter polymorphism on the inflammatory response in experimental human endotoxemia. Clin Infect Dis 33:1601–1603
Gallagher PM, Lowe G, Fitzgerald T, Bella A, Greene CM, McElvaney NG, O’Neill SJ (2003) Association of IL-10 polymorphism with severity of illness in community acquired pneumonia. Thorax 58:154–156
Gibson AW, Edberg JC, Wu J, Westendorp RG, Huizinga TW, Kimberly RP (2001) Novel single nucleotide polymorphisms in the distal IL-10 promoter affect IL-10 production and enhance the risk of systemic lupus erythematosus. J Immunol 166:3915–3922
Glynn P, Coakley R, Kilgallen I, Murphy N, O’Neill S (1999) Circulating interleukin 6 and interleukin 10 in community acquired pneumonia. Thorax 54:51–55
Gomez-Jimenez J, Martin MC, Sauri R, Segura RM, Esteban F, Ruiz JC, Nuvials X, Boveda JL, Peracaula R, Salgado A (1995) Interleukin-10 and the monocyte/macrophage-induced inflammatory response in septic shock. J Infect Dis 171:472–475
Hajeer AH, Lazarus M, Turner D, Mageed RA, Vencovsky J, Sinnott P, Hutchinson IV, Ollier WE (1998) IL-10 gene promoter polymorphisms in rheumatoid arthritis. Scand J Rheumatol 27:142–145
Helminen ME, Kilpinen S, Virta M, Hurme M (2001) Susceptibility to primary Epstein-Barr virus infection is associated with interleukin-10 gene promoter polymorphism. J Infect Dis 184:777–780
Hulkkonen J, Pertovaara M, Antonen J, Lahdenpohja N, Pasternack A, Hurme M (2001) Genetic association between interleukin-10 promoter region polymorphisms and primary Sjogren’s syndrome. Arthritis Rheum 44:176–179
Keijsers V, Verweij CL, Hazes M, Westendorp RGJ, Breedveld FC, Huizinga TWJ (1998) Innate differences in IL-10 production are present a the level of transcription and associated with haplotypes: association of IL-10 haplotypes and rheumatoid arthritis. Clin Exp Rheumatol 16:200
Koss K, Satsangi J, Fanning GC, Welsh KI, Jewell DP (2000) Cytokine (TNF alpha, LT alpha and IL-10) polymorphisms in inflammatory bowel diseases and normal controls: differential effects on production and allele frequencies. Genes Immun 1:185–190
Kube D, Platzer C, von Knethen A, Straub H, Bohlen H, Hafner N, Tesch H (1995) Isolation of the human interleukin 10 promoter: characterization of the promoter activity in Burkitt’s lymphoma cell lines. Cytokine 7:1–7
Marchant A, Deviere J, Byl B, De Groote D, Vincent JL, Goldman M (1994) Interleukin-10 production during septicaemia. Lancet 343:707–708
Marrie TJ (2000) Community-acquired pneumonia in the elderly. Clin Infect Dis 31:1066–1078
Pol WL van der, Huizinga TW, Vidarsson G, van der Linden MW, Jansen MD, Keijsers V, de Straat FG, Westerdaal NA, de Winkel JG, Westendorp RG (2001) Relevance of Fc gamma receptor and interleukin-10 polymorphisms for meningococcal disease. J Infect Dis 184:1548–1555
Reid CL, Perrey C, Pravica V, Hutchinson IV, Campbell IT (2002) Genetic variation in pro-inflammatory and anti-inflammatory cytokine production in multiple organ dysfunction syndrome. Crit Care Med 30:2216–2221
Ross P, Hall L, Smirnov I, Haff L (1998) High level multiplex genotyping by MALDI-TOF mass spectrometry. Nat Biotechnol 16:1347–1351
Schaaf BM, Boehmke F, Esnaashari H, Seitzer U, Kothe H, Maass M, Zabel P, Dalhoff K (2003) Pneumococcal septic shock is associated with the interleukin-10 −1082 gene promoter polymorphism. Am J Respir Crit Care Med 168:476–480
Tagore A, Gonsalkorale WM, Pravica V, Hajeer AH, McMahon R, Whorwell PJ, Sinnott PJ, Hutchinson IV (1999) Interleukin-10 (IL-10) genotypes in inflammatory bowel disease. Tissue Antigens 54:386–390
Temple SE, Cheong KY, Almeida CM, Price P, Waterer GW (2003) Polymorphisms in lymphotoxin alpha and CD14 genes influence TNF alpha production induced by Gram-positive and Gram-negative bacteria. Genes Immun 4:283–288
Turner DM, Williams DM, Sankaran D, Lazarus M, Sinnott PJ, Hutchinson IV (1997) An investigation of polymorphism in the interleukin-10 gene promoter. Eur J Immunogenet 24:1–8
Waal Malefyt R de, Abrams J, Bennett B, Figdor CG, de Vries JE (1991) Interleukin 10 (IL-10) inhibits cytokine synthesis by human monocytes: an autoregulatory role of IL-10 produced by monocytes. J Exp Med 174:1209–20
Waal Malefyt R de, Moore KW. (1998) Interleukin-10, 3rd edn. Academic Press, San Diego
Westendorp RG, Langermans JA, Huizinga TW, Verweij CL, Sturk A (1997) Genetic influence on cytokine production in meningococcal disease. Lancet 349:1912–1913
Acknowledgements
This work was funded by Genomics Collaborative Inc, USA, and the Raine Medical Research Foundation of Western Australia.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Temple, S.E.L., Lim, E., Cheong, K.Y. et al. Alleles carried at positions −819 and −592 of the IL10 promoter affect transcription following stimulation of peripheral blood cells with Streptococcus pneumoniae . Immunogenetics 55, 629–632 (2003). https://doi.org/10.1007/s00251-003-0621-6
Received:
Revised:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00251-003-0621-6