Abstract
Single-molecule experiments on DNA unzipping are analyzed on the basis of the mobility of nucleic bases in complementary pairs. Two possible scenarios of DNA double-helix unzipping are proposed and studied, using the atom–atom potential function method. According to the first scenario, the base pairs transit into a ‘preopened’ metastable state and then fully open along the ‘stretch’ pathway. In this case, the DNA unzipping takes place slowly and as an equilibrium process, with the opening energies being similar to the energies obtained in thermodynamic experiments on DNA melting. The second scenario is characterized by higher opening forces. In this case, the DNA base pairs open directly along the ‘stretch’ pathway. It follows from our calculations that, in this scenario, the enthalpy difference between the A\(\cdot \)T and G\(\cdot \)C base pairs is much higher than in the first case. The features of the first unzipping scenario show that it can play a key role during the process of DNA genetic information transfer in vivo. It follows from our study that a peculiarity of the second scenario is that it can be used for the development of faster methods for reading genetic information in vitro.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The development of single-molecule micromanipulation techniques (Smith et al. 1992; Simmons et al. 1996; Bockelmann et al. 1998a; Bustamante and Keller 1995) made it possible to investigate biological macromolecules such as isolated systems, allowing one to study the main mechanical processes involved in DNA functioning such as DNA stretching, bending, twisting and the opening of DNA base pairs (Lavery et al. 2002; Bustamante et al. 2000, 2003; Manghi and Destainville 2016). Among them especially interesting to research is the sequential separation of nucleic bases in a DNA double helix under the action of external force (DNA unzipping) (Bockelmann et al. 1998a,b,2002, 2004 Essevaz-Roulet et al. 1997; Thomen et al. 2002), because it plays a key role in genetic information transfer.
In early studies, the separation of nucleic bases in DNA base pairs (bp) was investigated as its denaturation under the influence of temperature, i.e., DNA melting (Owczarzy et al. 2004; Frank-Kamenetskii and Prakash 2014; Wartell and Benight 1985). In this case, the opening of base pairs is not sequential. First, the base pairs in A\(\cdot \)T-rich sites open, and, second, in G\(\cdot \)C-rich sites, due to the lower free opening energy of the A\(\cdot \)T pair as compared with G\(\cdot \)C. In contrast to the melting process, single-molecule micromanipulation methods make it possible to investigate the process of sequential opening of DNA base pairs in the same way, as it takes place in cells in vivo. Thus, with sufficient experimental precision and adequate understanding of the opening of nucleic base pairs, new method for reading nucleotide sequences can be developed.
In a DNA unzipping experiment (Fig. 1), as carried out by single-molecule micromanipulation methods (Bockelmann et al. 1998a, b, 2002, 2004; Essevaz-Roulet et al. 1997; Thomen et al. 2002; Bockelmann and Viasnoff 2008; Huguet et al. 2009, 2010; Danilowicz et al. 2003), the dependence of the opening force on strand separation is measured. For \(\lambda \)-phage DNA at low pulling velocities (\(\le 1~\upmu \text{m}/\text{s}\)), the strand separation begins after some critical force value \((\le 15~\text{pN})\) is reached. As a result, the force-extension curve forms a plateau. Note, that in this case, the unzipping process takes place without significant fluctuations of the opening force. For larger velocities (\(>1~\upmu \text{m}/\text{s}\)), the plateau occurs at a higher force (Thomen et al. 2002). In the last case, the force fluctuates significantly, and the DNA unzipping takes place as non-equilibrium process. To sum up, DNA unzipping depending on the pulling velocity can take place according to two different scenarios.
Scheme of a single-molecule manipulation experiment on DNA unzipping (Essevaz-Roulet et al. 1997; Bockelmann et al. 1998a, b, 2002, 2004; Thomen et al. 2002; Bockelmann and Viasnoff 2008). Force is applied to the polystyrene bead which is manipulated by magnetic or optical tweezers. Here F is the force that acts on a double-stranded DNA in the unzipping fork. Due to the experimental setup geometry, the applied force should have two components (\(F_x\) and \(F_y\))
DNA unzipping has been studied theoretically in Bockelmann et al. (1998a, b), Huguet et al. (2009), Huguet et al. (2010), Lubensky and Nelson (2000) and Volkov and Solovyov (2009). Bockelmann et al. (1998a, b) proposed a model of the unzipping process that considers the opening energies of base pairs, extension energies of single strands, and energy parameters for instruments used in the experiment (elastic energy of the bead in a trap, extension of handles). The force–extension curve calculated within this model describes the experimentally observed plateau value. Later, more accurate selection of the parameters of this model made it possible to describe the unzipping process for shorter DNA fragments with greater accuracy (Huguet et al. 2009, 2010). In papers Lubensky and Nelson (2000, 2002) it was emphasized that the DNA unzipping under physiological conditions can have a randomness and a non-equilibrium nature. The dependence of unzipping process on temperature conditions is considered in Monte Carlo simulations in Voulgarakis et al. (2006), but the model did not accord with experiment. Volkov and Solovyov (2009) propose the mechanism of the unzipping plateau appearance but the difference in unzipping processes for DNA A\(\cdot \)T and G\(\cdot \)C pairs was not considered. The different unzipping scenarios for different velocities of DNA unzipping were discussed in Thomen et al. (2002), but the definite physical mechanism of these processes is not determined yet.
The observed data and the existing theoretical models of DNA unzipping give information on the mesoscopic parameters of DNA unzipping, but cannot give an understanding of the physical mechanism of the unzipping process. It should be noted, that unlike the effect of temperature in the process of DNA melting, the action of force in the experiments on unzipping has a directional nature and requires the consideration of individual pathways of base motion in pairs.
The goal of the present work is to understand DNA unzipping on the scale of the mobility of nucleic bases in a DNA pair. In the next section, we will show that, because of the different possible opening pathways of nucleic base pairs, DNA unzipping can take place according to two different scenarios. In Sect. 3 with the use of atom–atom potential functions, the opening energies for the A\(\cdot \)T and G\(\cdot \)C base pairs within these two scenarios are calculated. It is shown that, during the DNA unzipping process, the base pairs can open not only along the ‘stretch’ pathway, but, under some conditions, they can first transit into a metastable state along the ‘opening’ pathway and then fully open along the ‘stretch’ pathway. In Sect. 4, the critical force that corresponds to the opening of base pairs in the ‘stretch after opening’ scenario is estimated.
Possible scenarios of the DNA unzipping process
It should be noted that, in the unzipping experiment, a part of the DNA macromolecule remains without force tension, so it is in the usual coil-like state in solution (see Fig. 1). As known (Thomen et al. 2002), the effects that are related to the coil influence manifest themselves as a friction drag in DNA unzipping. So, we assume that the force that acts on the unzipping fork due to the presence of the coil has to perform two types of work: to draw out a double-stranded DNA (dsDNA) from the coil and to open the base pairs. Thus, the unzipping force should have two components: \(F_x\) and \(F_y\) (Fig. 1).
Between the unzipping fork and the coil, a closed dsDNA is situated. It experiences a tension from the external force (\(F_y\) component) and the hydrodynamic damping of the coil (so-called effect of drag in solution). Thus, this part of a double-stranded DNA is in a strained state, preparatory to its unzipping. Another part of the DNA consists of two already unzipped DNA strands, which are under the action of external force.
The tension in the DNA chain (\(F_y\) component on Fig. 1) should stimulate the double-helix stretching and be accompanied with losses in the stacking of base pairs. Such a process under natural conditions should lead to the preopening of base pairs and their interactions with surrounding water molecules. That is, base pairs in the region of the unzipping fork can transit into the ‘preopened’ metastable state along the ‘opening’ pathway. The possibility of the occurrence of such a metastable state of complementary base pairs in physiological conditions in the environment of water molecules is shown by quantum mechanical calculations and molecular dynamic simulations of the base pair interaction with water (Kryachko and Volkov 2001; Giudice et al. 2003).
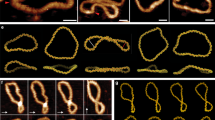
The DNA unzipping process can take place along different pathways, according to the known types of the mobility of nucleic bases in complementary pairs (Diekmann 1989). The most probable for DNA unzipping are the ‘stretch’ (when bases in a pair separate from each other along hydrogen bonds), ‘opening’ (when base pairs open into the major groove), and ‘shear’ (when bases separate in the plane of a pair in a direction perpendicular to the hydrogen bonds) pathways (Fig. 2). All these pathways can induce a large distortions in the hydrogen bonds and lead to base pair opening.
The analysis of the pathways of DNA base pair opening at the level of small amplitude displacements, which are implemented as conformational vibrations, shows that the base displacements in a pair along the ‘stretch’ (Fig. 2a) and ‘opening’ (Fig. 2b) pathways occur together with displacements of the DNA backbone strands (Volkov and Kosevich 1987, 1991; Volkov et al. 1989). But the movements of the bases in a pair along the ‘shear’ pathway occur together with displacements of the strands in the direction perpendicular to the hydrogen bonds that conflicts with the geometry of the DNA unzipping experiment. Thus, displacements of bases in a pair along the ‘shear’ pathway will not be considered in the present work.
It should be noted that the ‘stretch’ and ‘opening’ pathways are considered as the main probable fluctuations of the bases in the double helix under physiological conditions (Frank-Kamenetskii and Prakash 2014). These fluctuations are often found in studying DNA dynamics, and their occurrence is observed in the experiments involving formaldehyde kinetics and hydrogen exchange in DNA molecules (Volkov 1995; Kryachko and Volkov 2001; Giudice et al. 2003).
Due to the construction of the experimental setup and the way, in which the force is applied to a DNA macromolecule, the ‘stretch’ pathway should have a certain advantage. If the external force has a sufficiently high value (and can overcome the hydrodynamic damping of the coil), the opening of DNA base pairs in the unzipping process should always take place along the ‘stretch’ pathway. But if the external force is not sufficient to make the direct separation of bases along the ‘stretch’ pathway, then the unzipping process can proceed via separation of base pairs along the ‘opening’ pathway (as was proposed in Volkov and Solovyov 2009).
Degrees of freedom of nucleic bases in a pair of nucleotides. Picture is taken from Diekmann (1989): a ‘stretch’ pathway; b ‘opening’ pathway; c ‘shear’ pathway
So, we will assume that the unzipping of base pairs can take place according to two possible scenarios: the Watson–Crick pair opens along the ‘stretch’ pathway (‘stretch’ scenario, Fig. 3a) or the Watson–Crick pair first transits into a ‘preopened’ metastable state along the ‘opening’ pathway and then fully opens along the ‘stretch’ pathway (Fig. 3b).
From the analysis of DNA double-helix conformational vibrations, it is known that vibrations of the bases in a pair along the ‘opening’ pathway take place with a much lower frequency than along the ‘stretch’ pathway (Volkov and Kosevich 1987, 1991). This means that base pair opening along the ‘opening’ pathway takes more time than along the ‘stretch’ pathway. Therefore, the ‘stretch after opening’ scenario should be realized with lower unzipping velocities than in the ‘stretch’ scenario.
Therefore, the two considered scenarios can be distinguished by velocity: the slow one (‘stretch after opening’) and the fast one (‘stretch’ scenario).
Methods and results
Let us estimate the energy required for the transition of a base pair from the Watson–Crick or the ‘preopened’ states to the ‘opened’ configuration for DNA base pairs. The interaction energy in the complementary pairs A\(\cdot \)T and G\(\cdot \)C consists of the energies of hydrogen bonds, van der Waals and Coulomb interactions:
The energy of a hydrogen bond between atoms i and j is modeled by the modified Lennard–Jones potential ‘10–12’ (Poltev and Shulyupina 1986):
where \(r_{ij}\) is the distance between atoms i and j. The van der Waals interactions are described by the Lennard–Jones potential ‘6–12’ (Zhurkin et al. 1980):
and the parameters \(A_{ij}, B_{ij}\) and \(A_{ij}^{(10)}, B_{ij}^{(10)}\) are taken from Poltev and Shulyupina (1986) and Zhurkin et al. (1980). The Coulomb interaction is given by the electrostatic potential:
where \(q_i\) and \(q_j\) are the charges of the atoms i and j located at the distance \(r_{ij}\), \(\epsilon _0\) is the vacuum permittivity, and \(\epsilon (r)\) is the dielectric permittivity of a medium.
It should be noted that the calculations performed in the present work for studying DNA unzipping take into account only the interaction energy of nucleic bases in a pair. It is known that an important role in the stability of a DNA base pair is played by stacking interactions (Yakovchuk et al. 2006). But for base pairs that are situated in the unzipping fork, at the end of the closed part of the double helix, the contribution of stacking interaction should be low. This conclusion is in accordance with the known experimental facts.
It is known (Lukashin et al. 1976) that the probability of base pair fluctuations resulting in opening inside the double helix is much less than on the ends. The lower influence of stacking interactions on the opening of base pairs at the end of the double helix is confirmed also by recent data from molecular dynamic simulations (Dans et al. 2016), where it was observed that the bases in a pair at the end of the double helix have significantly larger fluctuational amplitudes and are definitely in a state that differs from the states of the pairs inside the double helix, as well as from the fully opened pairs (Jose et al. 2009). That is these pairs are in an intermediate ‘preopened’ state, where the stacking interaction with neighboring pairs is very small. So we can consider that the contribution of the stacking interaction in the unzipping fork should be much less than for a base pair situated inside the double helix.
In our calculations, we consider that the transition into the ‘preopened’ state takes place through the rotation of bases around the axis passing through the atom \(C_{1}'\) orthogonally to the plane of the base (Fig. 4). As the ‘stretch’ pathway, we consider a motion of bases in a pair relative to each other along the line connecting the \(C_{1}'\) atoms (Fig. 4).
The geometry of the Watson–Crick pairs A\(\cdot \)T and G\(\cdot \)C is taken from Saenger (1984). For the A\(\cdot \)T pair, the distance R between atoms \(C_{1}'\) is taken as 10.44 Å, the angles between the glycosidic bond and R (\(\alpha _1 \) and \(\alpha _2 \)) are calculated from the condition that the Watson–Crick A\(\cdot \)T pair in the closed state has the distance \(\text{N}_6\text{H}\cdots \text{O}_4\) equal to 2.9 Å, and the distance \(\text{N}_1\cdots \text{HN}_3\) is 2.8 Å. For the pair G\(\cdot \)C, the calculations are made in a similar way.
First, we have calculated the energy minima [minima of function (1)] for Watson–Crick base pairs with the value of \(\epsilon (r)\) in (4) set as 1 (see Table 1). It is obvious that the results obtained are very close to the values calculated in Poltev and Shulyupina (1986). Small discrepancies in the range of \(\sim 1\) kcal/mol can be due to uncertainty in the orientation of hydrogen atoms.
Our calculations are sufficient for determining the comparative stability of the A\(\cdot \)T and G\(\cdot \)C pairs in the Watson–Crick configuration. But, for the qualitative analysis of the interaction energies of bases in complementary pairs, the potential functions (4) give an anomalously large contribution to the Coulomb interaction of hydrogen atoms for the G\(\cdot \)C pair (Table 1). As DNA is situated in a water-ionic solution in reality, the atoms of bases are screened by water and other molecules, which results in a weakening of the Coulomb interactions. To take the effect of a physiological medium into account, the dependence of the dielectric permittivity on the distance between interacting atoms in the system is accounted for. Using the approach developed in Hingerty et al. (1985), the dielectric permittivity will be calculated according to the expression
The results of calculations with the use of expression (5) are shown in Table 1 in the last column. As can be seen, these calculations give the consensus result for the A\(\cdot \)T and G\(\cdot \)C pairs. Therefore, the further calculations of energies discussed in the present work are performed accounting for dependence of the dielectric permittivity according to (5). Thus, we will consider that the base pair interaction energy minimum for the Watson–Crick state corresponds to the base pair opening energy in the ‘stretch’ scenario and, accordingly, the energy minimum for the ‘preopened’ state corresponds to the opening energy in the ‘stretch after opening’ scenario. This assumption is proper because both scenarios take place under the same external conditions, i.e., under the action of an external force on the pairs situated in the unzipping fork.
The energies, as well as the geometric parameters, of the A\(\cdot \)T and G\(\cdot \)C Watson–Crick and ‘preopened’ states are shown in Table 2. It is obvious that the geometry of the Watson–Crick minima are in accordance with the standard geometry of Watson–Crick pairs (Saenger 1984).
The transition into the ‘preopened’ state is calculated in two steps. First, we make rotation of bases (‘opening’ pathway) (shown in Fig. 4) up to the angle which corresponds to the rupture of the external hydrogen bond in a pair (\(\text{N}_6\text{H}\cdots \text{O}_4\) for the A\(\cdot \)T and \(\text{O}_6\cdots \text{HN}_4\) to G\(\cdot \)C). It is known (Saenger 1984) that a hydrogen bond comes to be broken when the distance between the heavy atoms (N or O) and the hydrogen atom reaches the value of the sum of their van der Waals radii. As the van der Waals radius of a hydrogen atom is \({\approx } 1.2 \) Å, and those of nitrogen and oxygen are \({\approx } 1.5\) Å (Zefirov and Zorkyi 1974), this distance will be \({\approx } 2.7\) Å. Accordingly, since the length of the covalent bond N–H \({\approx } 1\) Å (Saenger 1984), the hydrogen bond will be considered as broken, when the distance between heavy atoms reaches \(\approx 3.7\) Å. We consider also that the full base pair opening takes place, when the middle hydrogen bond (\(\text{N}_1\cdots \text{N}_3\)) is broken Frank-Kamenetskii and Prakash (2014), i.e., the distance \({\rm N}_{1} {\rm N}_{3} {\approx } 3.7\) Å.
On the second step, we move this rotated configuration along the line \(C_{1}'C_{1}'\) (‘stretch’ pathway) looking for the energy minimum to get the ‘preopened’ state. From Table 2 (‘stretch after opening’, column R), it can be seen that, to achieve the energy minimum, the bases in the A\(\cdot \)T pair should approach each other to \(\approx 0.3\) Å. At the same time, the bases in G\(\cdot \)C pair almost remain in their places after the rotation. We will consider that the obtained ‘preopened’ configurations of the A\(\cdot \)T and G\(\cdot \)C pairs are stable under the action of external force.
The results of our calculations for two possible scenarios of DNA unzipping are shown in Table 3 together with the corresponding data obtained in the mesoscopic approach (Huguet et al. 2010) for low unzipping velocities. Energies in the 4th column of Table 3 are calculated from 10 nearest neighbor energies from Huguet et al. (2010) by averaging over all possible nearest neighbors. Thus, to calculate \(E_{\mathrm{AT}},\) we use averaging: \(E_{\mathrm{AT}}=( E_{\mathrm{AA}/\mathrm{TT}} + E_{\mathrm{TA}/\mathrm{AT}} + E_{\mathrm{AT}/\mathrm{TA}} + E_{\mathrm{AC}/\mathrm{TG}}+ E_{\mathrm{AG}/\mathrm{TC}})/5\). For \(E_{\mathrm{GC}},\) we have: \(E_{\mathrm{GC}}=(E_{\mathrm{CC}/\mathrm{GG}} + E_{\mathrm{CG}/\mathrm{GC}} + E_{\mathrm{GC}/\mathrm{CG}} + E_{\mathrm{CA}/\mathrm{GT}}+ E_{\mathrm{GA}/\mathrm{CT}})/5\). It can be seen that the results of our calculations for the ‘stretch after opening’ scenario fit the data much better than for the ‘stretch’ scenario. Consequently, we can conclude that in the single-molecule micromanipulation experiments performed with low opening velocities (\(\sim 10\) \(\upmu \text{m}/\text{s}\)), the ‘stretch after opening’ scenario is more appropriate.
Critical force estimation
Now, let us consider the implementation of two suggested scenarios of DNA unzipping: ‘stretch after opening’ and ‘stretch’. As follows from the experiment (Sect. 1), their implementation depends on velocities and, correspondingly, on the value of the critical force. We define the critical force as that at which the base pairs start to open at a given velocity. The value of the critical force (as well as the velocity) can be a feature that determines the scenario of the opening of base pairs in the DNA unzipping. In the slow equilibrium process, this value can be calculated by the formula proposed in Bockelmann et al. (1998a)
where \({G}_{\mathrm{pair}}\) is the sum of the free energies of all opened base pairs in a polynucleotide, \({G}_{\mathrm{e}l}\) is the free energy of a single-strand extension, and l is the total extension length of a single strand.
In the estimation, we will use parameters from (Bockelmann et al. 1998a): \(l = 9.5\) \({\AA }\) and \({G}_{\mathrm{e}l}\approx 0.3\) kcal/mol. Doing the same steps as in Bockelmann et al. (1998a) but using our calculated energies, we estimate the average value of opening force for a polynucleotide with a 50\(\%\) A\(\cdot \)T base pair content. We note that \(\lambda -\)phage DNA, which is usually used in unzipping experiments, has approximately the same value of base pair content. The free energy of the base pair unzipping can be calculated by the formula
where E is the enthalpy, S is the entropy, and T is the temperature. The base pair opening entropy averaged over the nearest neighbors is \(S_{\mathrm{AT}}=- 21.3\) cal/(mol K) for an A\(\cdot \)T-containing polynucleotide and is \(S_{\mathrm{GC}}=- 23.83\) cal/(mol K) (\(T=298\) K) for G\(\cdot \)C-containing polynucleotide, according to Huguet et al. (2010). Using the enthalpy values for the ‘stretch after opening’ scenario (Table 3) and formula (7), we calculated the free energies: \(G_{\mathrm{AT}}=- 1.48\) kcal/mol, \(G_{\mathrm{GC}}=-2.32\) kcal/mol. Substituting the average free energy \(G_{\mathrm{av}}=(G_{\mathrm{AT}}+G_{\mathrm{GC}})/2\) in expression (7), we get the average opening force for the 50\(\%\) A\(\cdot \)T-containing polynucleotide: \(F_{\mathrm{av}}\approx \) 13 pN, which coincides with the value calculated in Bockelmann et al. (1998a) and observed in the unzipping experiments (Bockelmann et al. 1998a; Thomen et al. 2002; Bockelmann et al. 2002, 2004; Huguet et al. 2010). It was observed in Thomen et al. (2002) that, with velocities \(\le 1\) \(\upmu \text{m}/\text{s}\) (which correspond to force values \(\le 15\) pN), the DNA unzipping is an equilibrium cooperative process. Consequently, it can be considered that the observed unzipping processes with force values \(\le 15\) pN takes place according to the ‘stretch after opening’ scenario.
We note that, for the ‘stretch after opening’ scenario, the calculated difference \(\Delta G=G_{\mathrm{GC}}-G_{\mathrm{AT}} \approx 0.84\) kcal/mol is in the range of thermal fluctuations in the system. Thus, the unzipping process in this scenario in a heterogeneous DNA takes place as in a homogeneous polymer. It can be considered that, in this scenario, the small difference in the unzipping free energies for the A\(\cdot \)T- and G\(\cdot \)C-rich sequences is the reason for the stable process of DNA opening with low unzipping velocities and, correspondingly, low opening forces.
Unfortunately, the same calculations of the free energy cannot be done for the ‘stretch’ scenario because of the difficulties in accounting for the significant entropy contribution produced by friction torque of a dsDNA coil. Our calculations show that the difference between the A\(\cdot \)T and G\(\cdot \)C-opening enthalpies in the ‘stretch’ scenario is much higher than in the ‘stretch after opening’ scenario (Table 3). It is obvious that this difference together with a significant frictional torque should lead to a greater instability of the DNA unzipping in the ‘stretch’ scenario, which is observed in Thomen et al. (2002).
On the other hand, if the DNA unzipping experiment is performed with short DNA samples, and if the effect of rotational drag is minimized, then the unzipping will be stable enough for any force and velocity. In this case, the DNA unzipping should take place according to the ‘stretch’ scenario without any significant hydrodynamic fluctuations and should be sensitive enough to determine the type of unzipped pair (A\(\cdot \)T or G\(\cdot \)C).
Conclusions
According to our study, the mechanism of DNA double-helix unzipping may follow two different scenarios.
The first scenario (‘stretch after opening’) is slow because of some changes in the base pairs, state before the opening. In this scenario, due to the dsDNA coil tension the base pairs first transit into the ‘preopened’ metastable state along the ‘opening’ pathway (Fig. 3b) and then fully open along the ‘stretch’ pathway. The opening of base pairs in this scenario can take place cooperatively and with low-energy dissipation because the opening energy difference between the A\(\cdot \)T and G\(\cdot \)C pairs (\({\Delta E}\)) is very small in this case (Table 3, ‘stretch after opening’). Therefore, this scenario is more probable in DNA unzipping experiments with low pulling velocities and should take place during the real unzipping processes in vivo such as transcription and translation.
The second scenario is the extension of hydrogen bonds along the ‘stretch’ pathway (Fig. 3a). This scenario is faster, and the opening takes place directly from any conformational form of DNA. In this case, the base pairs do not have time to transit into the ‘preopened’ metastable state. According to our calculations, the difference in enthalpies between the A\(\cdot \)T and G\(\cdot \)C pairs is significantly larger than in the ‘stretch after opening’ scenario. Thus, implementation of the second unzipping scenario can be more appropriate for the reading of DNA sequences by the single-molecule technique.
Change history
06 March 2019
The original article was published with the following errors.
06 March 2019
The original article was published with the following errors.
References
Bockelmann U, Viasnoff V (2008) Theoretical study of sequence-dependent nanopore unzipping of DNA. Biophys J 94:2716–2724
Bockelmann U, Essevaz-Roulet B, Heslot F (1998a) DNA strand separation studied by single molecule force measurements. Phys Rev E 58(2):2386–2394
Bockelmann U, Essevaz-Roulet B, Heslot F (1998b) Molecular stick-slip motion revealed by opening DNA with Piconewton forces. Phys Rev Lett 79(22):4489–4492
Bockelmann U, Thomen Ph, Essevaz-Roulet B, Viasnoff V, Heslot F (2002) Unzipping DNA with optical tweezers: high sequence sensitivity and force flips. Biophys J 82:1537–1553
Bockelmann U, Thomen P, Heslot F (2004) Dynamics of the DNA duplex formation studied by single molecule force measurements. Biophys J 87:3388–3396
Bustamante C, Keller D (1995) Scanning force microscopy in biology. Phys Today 48(12):32–38
Bustamante C, Smith SB, Liphardt J, Smith D (2000) Single-molecule studies of DNA mechanics. Curr Opin Struct Biol 10(3):279–285
Bustamante C, Bryant Z, Smith SB (2003) Ten years of tension: single-molecule DNA mechanics. Nature 421:423–427
Danilowicz C, Coljee VW, Bouzigues C, Lubensky DK, Nelson DR, Prentiss M (2003) DNA unzipped under a constant force exhibits multiple metastable intermediates. Proc Natl Acad Sci USA 100:1694–1699
Dans PD, Danilane L, Ivani I, Drsata T, Lankas F, Hospital A, Walther J, Pujagut RI, Battistini F, Gelpi JL, Lavery R, Orozco M (2016) Long-timescale dynamics of the Drew–Dickerson dodecamer. Nucl Acids Res 44(9):4052–4066
Diekmann RE (1989) Definitions and nomenclature of nucleic acid structure parameters. EMBO 8(1):1–4
Essevaz-Roulet B, Bockelmann U, Heslot F (1997) Mechanical separation of the complementary strands of DNA. Proc Natl Acad Sci USA 94:11935–11940
Frank-Kamenetskii MD, Prakash S (2014) Fluctuations in the DNA double helix: a critical review. Phys Life Rev 11:153–170
Giudice E, Vrnai P, Lavery R (2003) Base pair opening within B-DNA: free energy pathways for GC and AT pairs from umbrella sampling simulations. Nucl Acids Res 31(5):1434–1443
Hingerty BE, Ritchie RH, Ferrell TL, Turner JE (1985) Dielectric effects in biopolymers: the theory of ionic saturation revisited. Biopolymers 24:427–439
Huguet JM, Forns N, Ritort F (2009) Statistical properties of metastable intermediates in DNA unzipping. Phys Rev Lett 103:248106
Huguet JM, Bizarro CV, Forns N, Smith SB, Bustamante C, Ritort F (2010) Single-molecule derivation of salt dependent base-pair free energies in DNA. Proc Natl Acad Sci USA 107(35):15431–15436
Jose D, Datta K, Johnson NP, von Hippel PH (2009) Spectroscopic studies of position-specific DNA ‘breathing’ fluctuations at replication forks and primer-template junctions. PNAS 106(11):4231–4236
Kryachko ES, Volkov SN (2001) Preopening of the DNA base pairs. Int J Quantum Chem 82(4):193–204
Lavery R, Lebrun A, Allemand J-F, Bensimon D, Croquette V (2002) Structure and mechanics of single biomolecules: experiment and simulation. J Phys Condens Matter 14:383–414
Lubensky DK, Nelson DR (2000) Pulling pinned polymers and unzipping DNA. Phys Rev Lett 85:1572
Lubensky DK, Nelson DR (2002) Single molecule statistics and the polynucleotide unzipping transition. Phys Rev E 65:031917
Lukashin AV, Vologodskii AV, Frank-Kamenetskii MD, Lyubchenko YL (1976) Fluctuational opening of the double helix as revealed by theoretical and experimental study of DNA interaction with formaldehyde. J Mol Biol 108:665–682
Manghi M, Destainville N (2016) Physics of base-pairing dynamics in DNA. Phys Rep 631:141
Owczarzy R, You Y, Moreira BG, Manthey JA, Huang L, Behlke MA, Walder JA (2004) Effects of sodium ions on DNA duplex oligomers: improved predictions of melting temperatures. Biochemistry 43:3537–3554
Poltev VI, Shulyupina NV (1986) Simulation of interactions between nucleic acid bases by refined atom–atom potential functions. J Biomol Struct Dyn 3(4):739–765
Saenger W (1984) Principles of nucleic acids structure. Springer, Berlin
Simmons RM, Finer JT, Chu S, Spudich JA (1996) Quantitative measurements of force and displacement using an optical trap. Biophys J 70:1813–1822
Smith SB, Finzi L, Bustamante C (1992) Direct mechanical measurements of the elasticity of single DNA molecules by using magnetic beads. Science 258:1122–1126
Thomen Ph, Bockelmann U, Heslot F (2002) Rotational drag on DNA: a single molecule experiment. Phys Rev Lett 88(24):248102
Volkov SN (1995) Pre-opened state of the DNA duplex. Mol Biol 29(5):1086–1094
Volkov SN, Kosevich AM (1987) About the conformational vibrations of DNA. Mol Biol 21:797–806
Volkov SN, Kosevich AM (1991) Theory of low-frequency vibrations in DNA macromolecules. J Biomol Struct Dyn 8:1069–1083
Volkov SN, Solovyov AV (2009) The mechanism of DNA mechanical unzipping. Eur Phys J D 54:657–666
Volkov SN, Kosevich AM, Weinreb GE (1989) Theoretical study of the low-frequency vibrations of DNA macromolecule. Biopolym Cell 5:32–39
Voulgarakis NK, Redondo A, Bishop AR, Rasmussen K (2006) Probing the mechanical unzipping of DNA. Phys Rev Lett 96:248101
Wartell RM, Benight AS (1985) Thermal denaturation of DNA molecules: a comparison of theory with experiment. Phys Rep 126(2):67–107
Yakovchuk P, Protozanova E, Frank-Kamenetskii MD (2006) Base-stacking and base-pairing contributions into thermal stability of the DNA double helix. Nucl Acids Res 34(2):564–574
Zefirov UV, Zorkyi PM (1974) Van der Waals radii of atoms in crystal chemistry and structure chemistry. Zh Strukt Khimii 15:118–122
Zhurkin VB, Poltev VI, Florent’ev VL (1980) Atom-atom potential functions for conformational calculations of nucleic acids. Mol Biol (USSR) 14:1116–1130
Funding
The present work was partially supported by the Program of Fundamental Research of the Division of Physics and Astronomy of the National Academy of Sciences of Ukraine.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Rights and permissions
About this article
Cite this article
Zdorevskyi, O., Volkov, S.N. Possible scenarios of DNA double-helix unzipping process in single-molecule manipulation experiments. Eur Biophys J 47, 917–924 (2018). https://doi.org/10.1007/s00249-018-1313-3
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-018-1313-3