Abstract
Ferredoxin-NADP+ reductase (FNR) catalyses the electron transfer from ferredoxin to NADP+ via its flavin FAD cofactor. A molecular dynamics theoretical approach is applied here to visualise the transient catalytically competent interaction of Anabaena FNR with its coenzyme, NADP+. The particular role of some of the residues identified as key in binding and accommodating the 2′P-AMP moiety of the coenzyme is confirmed in molecular terms. Simulations also indicate that the architecture of the active site precisely contributes to the orientation of the N5 of the FAD isoalloxazine ring and the C4 of the coenzyme nicotinamide ring in the conformation of the catalytically competent hydride transfer complex and, therefore, contributes to the efficiency of the process. In particular, the side chain of the C-terminal Y303 in Anabaena FNR appears key to providing the optimum geometry by reducing the stacking probability between the isoalloxazine and nicotinamide rings, thus providing the required co-linearity and distance among the N5 of the flavin cofactor, the C4 of the coenzyme nicotinamide and the hydride that has to be transferred between them. All these factors are highly related to the reaction efficiency, mechanism and reversibility of the process.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
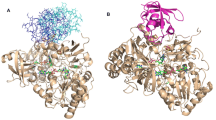
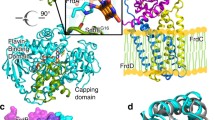
During photosynthesis, ferredoxin-NADP+ reductase (FNR) catalyses the electron transfer from ferredoxin (Fd) to NADP+ via its flavin FAD cofactor (Medina 2009; Medina and Gómez-Moreno 2004). The final hydride transfer (HT) event between WT FNR and the nucleotide is a reversible process, and two different transient charge-transfer complexes (CTC), FNRox-NADPH (CTC-1) and FNRhq-NADP+ (CTC-2), form prior to and upon HT whatever the HT direction (Fig. 1a) (Batie and Kamin 1984a; Lans et al. 2010; Peregrina et al. 2010; Tejero et al. 2007). This suggests that during the HT event the approach of the N5 donor atom of the FADH− isoalloxazine ring of FNR (N5i) to the C4 acceptor of the coenzyme nicotinamide ring (C4n) (or vice versa) occurs through certain stacking (Ortiz-Maldonado et al. 2003; Peregrina et al. 2010). Structures for the FNRox:NADP+ interaction obtained under different experimental conditions suggest a series of conformational rearrangements that might contribute to attaining the catalytically competent complex (Fig. 1b). Recognition of the 2′P-AMP moiety of NADP+ by FNR should be initially expected; this complex is labelled as C-I (PDB 1quf) (Serre et al. 1996). The need for an optimal interaction at this stage has been proposed as critical for the induction of the additional conformational changes (Paladini et al. 2009; Peregrina et al. 2009; Tejero et al. 2007). They must include narrowing the FNR coenzyme binding cavity to fit the 2′P-AMP and the pyrophosphate (PPi), as well as steering the nicotinamide towards the isoalloxazine binding site, as observed in C-II (PDB 1gjr, Fig. 1b) (Carrillo and Ceccarelli 2003; Hermoso et al. 2002; Tejero et al. 2005). The arrangement in the C-II crystal structure will further require the displacement of the C-terminal Tyr, Y303 in Anabaena FNR, and the rotation of the nicotinamide plane for the catalytically competent isoalloxazine-nicotinamide interaction to be achieved. However, there are not experimental data supporting whether such structural changes are produced from the C-II structure, and the final architecture of the catalytically competent complex (herein labelled C-III*) is also thus far unknown (Lans et al. 2010; Paladini et al. 2009; Tejero et al. 2005). Replacement of Y303 by Ser increases the enzyme affinity for NADP+ and produces a close isoalloxazine-nicotinamide stacking (C-III, PDB 2bsa and 1qfy, Fig. 1b), not detected with WT FNR. However, as the result of this strong binding, Y303S FNR shows a much lower catalytic efficiency (Deng et al. 1999; Lans et al. 2010; Piubelli et al. 2000; Tejero et al. 2005). Recent studies have demonstrated that stronger CTCs between the isoalloxazine and the nicotinamide do not relate to faster HT, and that the C-terminal Y303 contributes to providing the geometry of the catalytically competent complex by reducing the stacking probability between the isoalloxazine and nicotinamide (Peregrina et al. 2010).
a Scheme proposed for the reversible HT reaction between FNR and NADP+. MC-1 and MC-2 denote the Michaelis complexes for the non-photosynthetic and photosynthetic reactions, respectively. b Structural conformations proposed for different steps in the interaction of FNR with NADP+ to attain the catalytically competent complex. 1 Crystal structure of FNRox:NADP+ C-I obtained by soaking, PDB 1quf, showing recognition of the 2′P-AMP moiety of NADP+ by FNR (Serre et al. 1996). 2 Crystal structure of FNRox:NADP+ C-II, PDB 1gjr, showing the narrowing of the FNR coenzyme binding cavity to fit the 2′P-AMP and the PPi (Hermoso et al. 2002). 3 Crystal structure of Y303S FNRox:NADP+ C-III, PDB 2bsa, showing a close isoalloxazine-nicotinamide stacking (Tejero et al. 2005). 4 Catalytically competent WT complex C-III*, representing the unknown architecture for the ternary Y303-isoalloxazine-nicotinamide interaction
In the present study, the ability of molecular dynamics (MD) simulations to visualise protein flexibility as well as variations produced by ligand binding (Banci et al. 1992; Gorse and Gready 1997) is exploited to better understand the role of different regions and residues of Anabaena FNR in achieving the transient catalytically competent interaction with the coenzyme. We analyse the role played by the 2′P-AMP binding site in the early stages of coenzyme recognition (Medina et al. 2001; Tejero et al. 2007). Finally, we particularly discuss the putative orientations that the nicotinamide and C-terminal Tyr rings might adopt in the active site of the WT enzyme during catalysis on the basis of the simulations carried out on the complexes with the coenzyme of WT, Y303S and Y303W FNRs, and on the reported experimental HT efficiency for these variants.
Materials and methods
Model building of the biological systems
MD simulations of WT FNR in its complexes with NADP+, determined either by soaking (C-I, 1quf) or co-crystallisation (C-II, 1gjr), of Y303W FNR free and in complex with NADP+ (1w35 and 1w87) and of Y303S FNR free and in complex with NADP+ (1w34 and 2bsa, C-III) were carried out using the Cartesian coordinates of their crystal structures (Hermoso et al. 2002; Serre et al. 1996; Tejero et al. 2005). WT FNRhq:NADP+ and FNRox:NADPH models presenting isoalloxazine-nicotinamide interactions “putatively compatible with HT” through the formation of transient CTCs (C-III*) were generated by replacement of the NADP+ group in the non-productive Anabaena WT FNR:NADP+ C-II (1gjr) with the conformation of NADP+ in Anabaena Y303S FNRox:NADP+ C-III (PDB 2bsa) (Fig. 1 and SP1). Additional models were produced using the structure of the WT FNR:NADP+ C-I (1quf) by in silico replacement with the most conserved rotamer of either a Phe or an Ala for Y235 and an Asp for S223, as well as the NADP+ coenzyme with NAD+. These manipulations were carried out using SpdViewer (Guex and Peitsch 1997). Crystallographic waters were removed from all crystal structures. The FAD cofactor was replaced with FADH− when simulating complexes of FNRhq.
MD simulations
MD simulations were carried out using AMBER (Case et al. 2005), ff94 (Cornell et al. 1995) and the contributed parameters for FAD, FADH−, NADPH and NADP+ (http://www.pharmacy.manchester.ac.uk/bryce/amber#cof) (Antony et al. 2000; Walker et al. 2002). Residues were protonated at pH 7.0 using standard AMBER methods and PROPKA (Bas et al. 2008). In general, simulations have been carried out with cofactors and coenzymes in the redox states expected for the photosynthetic HT process, FADH− and NADP+, with the single exception of WT FNRox:NADPH C-III*, where FAD and NADPH were used. Each molecular system was neutralised by the addition of sodium ions and solvated with a TIP3P water model in a truncated octahedron box centred on the geometric centre of the protein and with the minimum distance of 10 Å from any protein atom.
In order to remove close contacts and highly repulsive orientations of the initial protein-solvent system, solvent molecules and counter ions were relaxed and allowed to redistribute around the restrained protein molecule before energy minimisation of the entire system. The process consisted of 500 steps of steepest descent (SD) and 500 steps of conjugate gradient (CG), in which the protein is restrained with an harmonic potential with force of 500 kcal mol−1 Å−2 centred on each atom to smooth the solvent-protein interaction. A second energy minimisation applied 1,000 steps of SD and 1,500 steps of CG to the whole system. In models generated by in silico replacement of residues or of coenzyme 5,000 steps of SD and 15,000 of CG were applied. The resulting system was heated from 0 to 300 K with the protein atoms weakly constrained with an harmonic potential with force constant of 10 kcal mol−1 Å−2 for 20 ps, to get a solvent pre-equilibration. The whole system was then equilibrated over 20 ps at 300 K and a constant pressure of 1 atm with a weak-coupling pressure algorithm. A Langevine temperature equilibration algorithm, periodic boundary conditions and the Particle Mesh Ewald method for long-range electrostatic interactions were used. All bonds involving hydrogens were constrained with SHAKE. Time step of 2 fs was used for dynamics integration. Each system was simulated for 6–30 ns maintaining the same conditions as for the second equilibration, and the structures were collected every 2 ps. During equilibration and MD productions, the leapfrog Verlet integration scheme was used.
Most of the obtained simulations, as well as some recent crystal structures (Peregrina et al. 2010), show an interaction between the C-terminal carboxylate and the guanidinium of R264 (Fig. 2a), suggesting the position of R264 in C-III* might influence the Y303 one. Additional simulations were conducted following a different protocol that considers such interactions. After allowing Y303 to get relaxed and equilibrated, a 4 ns MD simulation was carried out restraining the FADH− and NADP+ movement (restriction constant of 10 kcal mol−1 Å−2). Then, FADH− and NADP+ were allowed to move freely (6 ns), while R264 was restrained (10 kcal mol−1 Å−2) in one of the most populated conformations interacting with the Y303 carboxylate.
a Overall structure of Anabaena FNR showing the residues discussed in this work as relevant for the interaction with the coenzyme; green sticks represent the carbons. The FAD prosthetic group is represented in sticks with carbons in orange, while a putative position for NADP+ is represented with blue sticks. Panels b, c and d show the relative disposition of the coenzyme (shown as sticks and CPK-coloured) at the equilibrium simulations for WT C-I, Y235A C-I and S223D C-I respectively with regard to the 2′P-AMP binding surface (in pale green). Position of the coenzyme at the end of each simulation is shown with carbons in green, yellow and blue for b, c and d respectively. R224, R233, Y235 (A235 in c), and L263 are shown as sticks with carbons in green
Analysis of MD trajectories
Evolution of three-dimensional conformation was inspected using the programs VMD (Humphrey et al. 1996) and PyMOL (Delano 2002). Interatomic distances and angles, as well as the r.m.s.d. from a given structure, were monitored using the PTRAJ module in AMBER and CARNAL (Case et al. 2005).
Results
The r.m.s.d. of the backbone atoms of the different solvated FNR species along the MD simulations from the initial structure are in general <1.6 Å, and positional r.m.s.d. of α-carbons showed regions of similar flexibility (Fig. SP2, Table SP1). The protein core, formed by the T155-W167 α-helix and flanked by the β-sheets at the N-terminal and C-terminal domains, shows the lowest fluctuation. The G66-P73 and Y104-V113 loops are among the most flexible, corresponding also to the highest B factors in the crystal structures (Hermoso et al. 2002; Serre et al. 1996; Tejero et al. 2005). The K227-R233 and, particularly, the S275-S286 loops are relatively flexible for the C-I of Y235F, Y235A and S223D FNRs. The mobility of these loops was considerably decreased in C-I with NAD+, C-II of Y303W and C-III of Y303S, but flexibility is kept in the simulations representing putative competent C-III*. In general, structures corresponding to C-II and C-III show less flexibility than those corresponding to the enzyme, free, in C-I or in C-III*.
The dynamics around the 2′P-AMP binding site
MD simulations of WT FNRhq:NADP+ C-I (Figs. 1b1 and 2b) maintain the main interactions of the 2′P-AMP portion of NADP+ with the protein shown in the crystal structure (Fig. SP3). R224 and R233 stabilise the 2′P, the adenine plane is stacked between Y235 (angle ~30–40°) and L263 chains, and the S223 and the Q237 chains H-bond the ribose and the adenine respectively (Fig. 2a, b, Table SP2). Additionally, S223 makes polar contacts with R233. These interactions contribute to modulating the conformation of the site from the initial, putatively non-productive, one to one much more similar to that observed in the crystallographic Y303S C-III (Tejero et al. 2005). Simulations did not reproduce subsequent rearrangement of the coenzyme PPi onto the protein surface. This was not a surprise since such strong positional changes are usually not easily produced in the short times simulated.
Replacement of Y235 with either Phe or Ala has an important impact on the coenzyme-enzyme interaction: the adenine of the 2′P-AMP gets out of the enzyme cavity after breaking interactions with Q237 and L263, whereas R224 and R233 accompany the 2′P displacement (Figs. 2c and SP3, Table SP2). Replacement of S223 with Asp disrupts the adenine stacking with Y235 and L263, while D223 ends twisted towards the binding cavity being stabilised by R233 and K227 (Fig. 2d and SP3, Table SP2). Such changes probably prevent the adenine from reaching the final productive position. We also replaced NADP+ with NAD+ in the WT FNRhq:NAD+ C-I MD simulation. Although the adenine interactions with S223, Q237, and L263 were maintained (Fig. SP3, Table SP2), the coenzyme position was hardly modified from the initial one.
MD simulations of the Y303W and Y303S FNRhq:NADP+ complexes, C-II and C-III respectively, maintain most of the interactions of the adenine binding site described in WT C-1 (Fig. SP4). However, in Y303S C-III, the interactions between the protein and the 2′P-AMP of the coenzyme generally show lower levels of occupancy (Fig. SP4, Table SP2). The coenzyme presents an L-shape conformation, with the PPi located at the corner of the two branches accommodating the NMN and 2′-P AMP. L263 contributes with van der Waals contacts to place NMN in Y303W C-II, but in Y303S C-III the contact is reduced to a couple of atoms of the ribose.
The dynamics around the WT Anabaena FNR active site
In the absence of coenzyme, the isoalloxazine stacking with Y79 (on the si-face) and Y303 (on the re-face) remains along the simulation. Y79 keeps a constant distance and angle, ~65°, with regard to the mass centre and plane of the isoalloxazine respectively. However, Y303 moves on the isoalloxazine ring re-face surface, rapidly changing the angles from less than 10° up to 30°, inducing minor distance changes within the atoms of this environment (Figs. 3, 4a and b). The presence of the reduced flavin, FADH−, only produces a minor increase in the N5Hi-HO-S80 distance with regard to the oxidised crystal structure. A water molecule populates the environment of the reacting N5Hi, further contributing to bridging the above-mentioned positions (Table SP3), while S80 is additionally stabilised by CO4i and by an H-bond with the side chain of S59 (Fig. 3). Noticeably, the E301 carboxylate starts the simulations by interacting with the hydroxyls of S80 and S59, but its conformation flips between this one, pointing towards the inside of the protein, and others in which it is solvent exposed and making contact with R264 and T302 (Fig. 4a, b). Water molecules fill the former position of E301 and H-bond the hydroxyls of S80 and S59. One of these conformations with E301 pointing towards the solvent has been recently frozen in an X-ray crystal structure, while its former position is populated by water molecules (Peregrina et al. 2010). Despite rotation of the C261 chain along simulations, its thiol is positioned in the environment of the hydroxyls of S80 and, particularly, Y303. The aromatic ring of Y303 keeps its stacking to the isoalloxazine, while its C-terminal carboxylate points to the surface (Fig. 4a and b, Table SP3). It is also worth noting that the side chain of R264 shows two preferential conformations along the simulations; in one its guanidinium interacts with the C-terminal carboxylate, in the other it points towards the opposite direction and interacts with the carboxylates of E267 and/or E268 (Fig. 4a and b).
Time evolution of some specific interactions at the active site of WT FNRox in C-I along the MD simulation. Atom to atom distance between a the N5i and the oxygen of the hydroxyl group of Ser80, b the oxygens of the hydroxyl groups of Ser80 and Ser59, c the oxygen of the hydroxyl of Ser80 and one of the oxygens of the carboxylate of Glu301. b Angle between the planes of the isoalloxazine and the Tyr303 rings. Planes are defined by at least three atoms of the rings
Position of selected residues directly involved or connected to the active site of free FNR at different moments along the MD simulation. The images correspond to the following times along the simulation: a 0 ns (grey carbons), 0.8 ns (beige carbons) and 3.2 ns (light violet carbons) and b 14.8 ns (light green carbons) and 26.8 ns (pale yellow carbons). c Positions for the dihedral angle of carbons C-α, C-β and C-γ of the side chain of E301 along the MD of WT FNRhq:NADP+ C-III* with fixed R264 (black), FNRox:NADP+ C-III* (red) and free FNRhq (green). d Positions for the dihedral angle for carbons C-α, C-β and C-γ of the side chain of R264 along the MD of WT FNRhq:NADP+ C-III* with fixed R264 and free FNRhq. e Details of the water molecules at the active site of FNRhq:NADP+ C-III* with fixed R264 at 3 ns of the MD simulation. FADH−, NADP+ and Y303 are shown with carbons in orange, blue and cyan sticks, respectively
Replacement of Y303 at the active site
In general, differences found in the dynamics around the active site of the free Y303S and Y303W FNRs are just reduced to interactions of the modified residue. In the Y303S variant the S303 and E301 side chains point towards the protein surface, while the C-terminal carboxylate alternatively interacts either with C261 or R264. The cavity produced by the replacement is filled with water, but the position of the isoalloxazine is not displaced. In Y303W FNR, the introduced Trp is highly stabilised in the active site stacking with the isoalloxazine.
MD simulation of Y303S FNRrd:NADP+, C-III, keeps the A side of the nicotinamide facing the re-face of the central isoalloxazine ring, consistent with the stereochemistry of HT reported for this enzyme. The C4n and the N5Hi are ~3.5–4.0 Å apart and the N5i-hydride-C4n reacting atoms show an angle of ~90–100° (Fig. SP5). The distance is in general reasonable for direct HT, but lack of co-linearity of the reacting atoms has been described as one of the main causes of the hindered HT observed for this variant (Lans et al. 2010; Peregrina et al. 2010). S80 keeps its H-bond interaction with N5Hi but gets separated from C4n. In addition, the carboxamide group of the nicotinamide is stabilised by H-bonds with chains of E301 and S303. In Y303W C-II, the nicotinamide does not replace the C-terminal Trp at any moment, but a displacement of the NMN portion towards the surface compatible with the reorganisation of C-II into C-III, without major alteration of the 2′P-AMP binding site, might be envisaged.
A theoretical model mimicking the interaction between the flavin and the nicotinamide in the active site of WT FNR
Models of WT FNRox/hq complexes with NADP+/H (C-III*) were produced as indicated in the “Materials and methods” section (Fig. SP1). Equilibration of these initial dispositions provided starting structures for MD. In the first simulation of FNRhq:NADP+ C-III*, Y303 prevented any interaction between the isoalloxazine and the nicotinamide, producing N5Hi-C4n distances and angles not compatible with HT. Recent crystallographic data (Peregrina et al. 2010) and the simulations described above (Fig. 4a and b) further support the early hypothesis that R264 might influence the Y303 displacement to allow the nicotinamide approach to the active site. With this idea in mind we carried out a new simulation in which, after equilibration of the system, R264 was restrained in one of the conformations where its guanidinium interacts with the C-terminal carboxylate of Y303. Following this procedure we obtained a WT FNRhq:NADP+ C-III* dynamic structure in which Y303 interacts with both the isoalloxazine and the nicotinamide, placing its hydroxyl between the N10 of the isoalloxazine (N10i) and the N1 of the nicotinamide (N1n). Such organisation allows a close N5Hi-C4n interaction and contributes to increasing the co-linearity of the reacting atoms (Figs. 5a and SP5). Similarly, and without need of any atom restriction, the MD of the WT FNRox:NADPH C-III* arrived at a conformation in which Y303 also situates its hydroxyl between the N10i and the N1n, while keeping a close interaction and favourable orientation among the N5i acceptor, the hydride to be transferred and the C4n donor (Fig. 5b).
Relative disposition of the isoalloxazine and the coenzyme with regard to the protein surface for WT FNRhq:NADP+ C-III* with fixed R264 (green surface) and FNRox:NADPH C-III* (pink surface) at 3 and 6.2 ns of the MD, respectively. a, b Detail of the isoalloxazine-nicotinamide structural relationships; c and d show the corresponding PPi and 2′P-AMP moieties of the coenzyme. Y303, FADH−, FAD, NADP+ and NADPH are shown as sticks and CPK-coloured with carbons in cyan, yellow, orange, pink and blue respectively. e Schemes representing the putative relative dispositions between the flavin (orange) and nicotinamide rings (blue) in the case of the competent complexes of Y303 and WT FNRs with the coenzyme. In the WT complex the putative disposition of the Y303 side chain is indicated by a green plane. α (black) is the average angle formed by the planes of the isoalloxazine central ring and the nicotinamide ring, while β (green) corresponds to the average angle formed by the N5i-hydride-C4n atoms
These WT FNRhq:NADP+ and FNRox:NADPH C-III* structures show the simultaneous allocation in the isoalloxazine active site environment of the nicotinamide and the Y303 (Fig. 5a and b). Y303 breaks the parallelism and decreases the stacking between the isoalloxazine and the nicotinamide by increasing the distance between N10i and N1n, while keeping the reacting atoms, N5i and C4n, at HT distance. Such spatial disposition approaches the two reacting rings in such a way that the N5i-hydride-C4n angle increases, producing a more co-linear disposition (Figs. 4e, 5e and SP5), in agreement with the high efficiency of WT FNR. In general, in both simulations N7n interacts with E301 and S80, whereas C261 and S80 chains stabilise C4n, with these distances being shorter in FNRhq:NADP+ C-III* than in FNRox:NADPH C-III* (Fig. 5a, b and SP5). Simultaneously, S80 gets slightly farther from N5Hi than in Y303S C-III, while the N5i-HO-S80-C4n angle, ~50°, decreases with regard to Y303S C-III (Fig. SP5).
Simulations of WT FNRhq:NADP+ and FNRox:NADPH C-III* also show that the 2′P-AMP moiety of the coenzyme perfectly matches in its protein cavity. In particular, an H-bond is occasionally produced between R233 and Y235-OH, contributing to orienting the aromatic ring of Y235, fixing its stacking to the adenine and reducing its rotation (Fig. SP4), while the mobility of R100 ends considerably reduced with the position of its guanidinium almost unaltered with regard to the PPi. However, simulations also indicate a constant movement of the 2′P-AMP moiety (Figs. 5c, d, and SP4, Table SP2). During this movement the protein adapts its surface to this coenzyme moiety, keeping the particular interactions unaltered, but allowing different relative dispositions between the nicotinamide and the adenine portions of the coenzyme.
Discussion
Our comparative simulations of WT, Y235F, Y235A and S223D C-I confirm that the initial step of recognition and binding of the 2′P-AMP portion of NADP+ by FNR is critical to attain a catalytically competent interaction. Experimental studies showed that binding of NADP+ to FNR is prevented when a negatively charged residue substitutes for S223, an effect related to the electrostatic repulsion between the 2′P and the introduced charge, a strategy commonly used by NADH-dependent members of the FNR family (Medina et al. 2001). Rearrangement of the binding cavity was far from being observed in the simulations of the S223D variant, and interactions of D223 with different cavity residues envisaged blocking of the binding site in the absence of coenzyme. Replacement of Y235 with Phe decreased the FNR HT ability, although this variant is apparently able to induce CTC formation (Medina et al. 2001; Tejero et al. 2007). However, when an Ala was substituted for Y235, formation of CTC and HT were prevented (Medina et al. 2001; Peregrina et al. 2010; Tejero et al. 2007). Simulations confirm S223 is in close interaction with the 2′P and the hydroxyls of the ribose, while the aromatic stacking of Y235 contributes to maintaining the integrity of the AMP binding site, the adenine binding and its rearrangement inside the cavity. Y235 closes the bottom of the cavity, pushing the adenine ring towards L263 and clamping it in the binding site. However, the aromatic character of Y235 is not enough to produce the more optimal stacking contact, since the position of its plane with regard to the adenine tends to rotate (Fig. SP3D). Such rotation is considerably reduced, and the aromatic plane thus stabilised, by the interactions provided by the hydroxyl of Y235 with the 2′P and, through it, with R233. Removal of this Arg has also been reported to have negative effects on the overall HT process (Medina et al. 2001). Therefore, not only contacts between FNR and the coenzyme at the 2′P-AMP binding site but also the ones among protein residues appear efficiently designed to produce the most optimum NMN orientation in the active site for HT. The simulations also indicate that the FNR NADP+-binding domain does not sense the presence of NAD+. This is consistent with binding of NAD+ to FNR not being experimentally observed (Medina et al. 2001) and suggests that the 2′P moiety of the coenzyme is critical for the subsequent conformational changes that both the protein and the coenzyme must undergo to place the NMN moiety in its catalytically competent position.
S80, C261 and E301 of Anabaena FNR are conserved in the active site of all known FNRs, as well as in different members of the FNR family (Ceccarelli et al. 2004; Correll et al. 1992, 1993; Murataliev et al. 2004). These residues modulate the midpoint reduction potential of FAD, the affinity for Fd, and the electron and HT rates (Aliverti et al. 1995, 1998, 1993; Dumit et al. 2010; Medina et al. 1998; Tejero et al. 2005). S80 and C261 maintain stable location and interactions, confirming their contribution to NMN allocation for an efficient flavin-nicotinamide interaction through the production of CTCs during HT (Aliverti et al. 1995, 1993; Musumeci et al. 2008). H-bond interactions between S59 and S80 chains are also populated in our simulations, even when the NMN moiety of the coenzyme is in the active site. Additionally, S59 bridges the E301 side-chain. This latter residue switches positions in and out of the active site (Fig. 4). Experimental and theoretical studies also indicate that E301 contributes to the semiquinone and midpoint potential modulation, acts as a proton donor during FNR reduction by the protein partner and results in key stabilising and destabilising reaction intermediates in the ternary complex (Aliverti et al. 1998; Dumit et al. 2010; Faro et al. 2002; Medina et al. 1998; Peregrina et al. 2010). In fact, in the Anabaena FNR:Fd complex, the carboxylate of E301 is no longer exposed to solvent, and it is one of the residues with the highest propensity for being at the protein–protein interface (Fig. SP6) (Medina et al. 2008; Morales et al. 2000). Such observations reinforce a pathway for proton transfer between the external medium and the isoalloxazine N5i via S80 and the conserved water molecule. This route might also include S59, a residue highly conserved in FNR sequences (Bruns and Karplus 1995; Serre et al. 1996). Additionally, E301 is critical for proper binding of the nicotinamide into the active centre, CTC stabilisation and efficient flavin reduction by NADPH (Aliverti et al. 1998; Medina et al. 1998; Peregrina et al. 2010). These observations are consistent with the WT models derived here for the location of the nicotinamide in the active site; the models show S80, C261 and E301 as having a relevant role in the location of the C4n (hydride acceptor/donor) into the active site.
Y303 has been indicated as a critical residue during the catalytic cycle of FNR. It is expected to get displaced from its stacking position at the re-face of the flavin to allow the nicotinamide to approach the isoalloxazine. In the absence of the coenzyme, although no evidence is presented that Y303 occasionally gets out of its stacking position with the isoalloxazine ring (in agreement with the lack of experimental proof of Y303 leaving the cavity, increasing its solvent accessibility or being directly involved in the HT itself), our simulations clearly envisage displacements of Y303 with regard to the isoalloxazine ring (Figs. 3, 4a and b). Both experimental and theoretical studies have identified Y303 as key for lowering the affinity for NADP+/H to levels compatible with turnover, for stabilising the flavin semiquinone required for electron splitting and for modulating the flavin midpoint potential and, therefore, the electron/hydride transfer rates (Deng et al. 1999; Lans et al. 2010; Nogués et al. 2004; Paladini et al. 2009; Piubelli et al. 2000; Tejero et al. 2005). Therefore, the small movements detected here of Y303 on the surface of the isoalloxazine might contribute to allowing the nicotinamide to find its way into the catalytic site.
Simulations for WT FNRhq:NADP+ and FNRox:NADPH C-III* provide an ensemble of conformations in which Y303 and the nicotinamide are simultaneously allocated in the flavin environment. In these conformations, Y303 contributes to the geometric deformation of the π stacking between the isoalloxazine and the nicotinamide observed in the Y303S FNRhq:NADP+ C-III, brings the N5i-hydride-C4n angle to values compatible with the HT (>160°) and narrows the distance between the hydride donor and acceptor atoms (Fig. 5). These arrangements surely reduce the close contact of the reactant ionic pair in the physiological direction, making it possible and efficient, as experimentally observed. Therefore, although Y303 must get displaced to allow the entrance of the nicotinamide into the active site, it must keep forming part of it during the HT event to allow for the correct alignment among the reacting atoms. Thus, here we provide some putative transient conformations for the isoalloxazine, the nicotinamide and Y303 that might be compatible with the high HT efficiency of plastidic FNRs. These data also suggest a group of orientations among the reacting rings that could all contribute to the HT reaction.
We must keep in mind, however, a key feature that still remains to be explained on molecular terms: how the putative C-I or C-II initial interactions might be converted into C-III*, since the expected molecular movements are not apparent and, despite reversibility, different HT mechanisms might apply in both directions (Carrillo and Ceccarelli 2003). In our simulations, we never detected interactions between the nicotinamide and the aromatic ring of Y303 out of the catalytic site. Therefore we have no information about whether such a possible contribution toward pushing the tyrosine away exists. However, our simulations suggest additional side chains, indirectly connected with the active site, such as R100, T157, R264, E267 and E268, which might be involved in this process (Figs. 2a, 4a and b). Among them, R264 might be particularly critical and changes in the interaction between its guanidinium and the C-terminal carboxylate of Y303 might contribute to displacement of this Tyr and to triggering the entrance of the nicotinamide into the catalytic site. Additionally, the constant movement observed for the adenine and its binding pocket might also contribute to attaining the catalytically competent complex by allowing the nicotinamide to explore the entrance to the catalytic site after the 2′P-AMP moiety has already been anchored to FNR. Finally, during the physiological process, FNR, Fd and NADP+ form a ternary complex (Batie and Kamin 1984a; Carrillo and Ceccarelli 2003; Hermoso et al. 2002; Martinez-Julvez et al. 2009; Velazquez-Campoy et al. 2006). The Fd and coenzyme binding sites on FNR are not completely independent, and the overall reaction is proposed to work in an ordered two-substrate process, with the pyridine nucleotide binding first (Batie and Kamin 1984a, b). Thus, as shown in putative Fd:FNRhq:NADP+ and Fd:FNRox:NADPH ternary models, Fd might also help to place the C-terminal side chain (Fig. SP6). Further work must be done to clarify all these points.
In conclusion, the MD theoretical approach presented here provides new details on the interaction mechanism between FNR and NADP+. The particular role of some of the key residues in the binding and location of the 2′P-AMP portion of the coenzyme and in the FNR active site is described here in molecular terms. In particular, a key role is envisaged for Y303 in providing the co-linearity required among the N5 of the flavin cofactor, the hydride to be transferred and the C4 of the nicotinamide of NADP+ during HT.
Abbreviations
- FNR:
-
Ferredoxin-NADP+ reductase
- FNRox :
-
FNR in the fully oxidised state
- FNRhq :
-
FNR in the anionic hydroquinone (fully reduced) state
- 2′-P:
-
2′-Phosphate group of NADP+/H
- HT:
-
Hydride transfer
- WT:
-
Wild-type
- CTC:
-
Charge-transfer complex
- CTC-1:
-
FNRox-NADPH CTC
- CTC-2:
-
FNRhq-NADP+ CTC
- NMN:
-
Nicotinamide nucleotide moiety of NADP+/H
- 2′-P-AMP:
-
2′-P-AMP moiety of NADP+/H
- PPi:
-
Pyrophosphate
- N5Hi, N5i:
-
N5 hydride donor/acceptor of the FADH−/FAD isoalloxazine ring of FNR
- C4n, C4Hn:
-
C4 hydride acceptor/donor of the NADP+/H nicotinamide ring
References
Aliverti A, Piubelli L, Zanetti G, Lubberstedt T, Herrmann RG, Curti B (1993) The role of cysteine residues of spinach ferredoxin-NADP+ reductase as assessed by site-directed mutagenesis. Biochemistry 32:6374–6380
Aliverti A, Bruns CM, Pandini VE, Karplus PA, Vanoni MA, Curti B, Zanetti G (1995) Involvement of serine 96 in the catalytic mechanism of ferredoxin-NADP+ reductase: structure-function relationship as studied by site-directed mutagenesis and X-ray crystallography. Biochemistry 34:8371–8379
Aliverti A, Deng Z, Ravasi D, Piubelli L, Karplus PA, Zanetti G (1998) Probing the function of the invariant glutamyl residue 312 in spinach ferredoxin-NADP+ reductase. J Biol Chem 273:34008–34015
Antony J, Medvedev DM, Stuchebrukhov AA (2000) Theoretical study of electron transfer between the photolyase catalytic cofactor FADH− and DNA thymine dimer. J Am Chem Soc 122:1057–1065
Banci L, Schroder S, Kollman PA (1992) Molecular dynamics characterization of the active cavity of carboxypeptidase A and some of its inhibitor adducts. Proteins 13:288–305
Bas DC, Rogers DM, Jensen JH (2008) Very fast prediction and rationalization of pKa values for protein-ligand complexes. Proteins 73:765–783
Batie CJ, Kamin H (1984a) Electron transfer by ferredoxin:NADP+ reductase. Rapid-reaction evidence for participation of a ternary complex. J Biol Chem 259:11976–11985
Batie CJ, Kamin H (1984b) Ferredoxin:NADP+ oxidoreductase. Equilibria in binary and ternary complexes with NADP+ and ferredoxin. J Biol Chem 259:8832–8839
Bruns CM, Karplus PA (1995) Refined crystal structure of spinach ferredoxin reductase at 1.7 Å resolution: oxidized, reduced and 2′-phospho-5′-AMP bound states. J Mol Biol 247:125–145
Carrillo N, Ceccarelli EA (2003) Open questions in ferredoxin-NADP+ reductase catalytic mechanism. Eur J Biochem 270:1900–1915
Case DA, Cheatham TE 3rd, Darden T, Gohlke H, Luo R, Merz KM Jr, Onufriev A, Simmerling C, Wang B, Woods RJ (2005) The Amber biomolecular simulation programs. J Comput Chem 26:1668–1688
Ceccarelli EA, Arakaki AK, Cortez N, Carrillo N (2004) Functional plasticity and catalytic efficiency in plant and bacterial ferredoxin-NADP(H) reductases. Biochim Biophys Acta 1698:155–165
Cornell WD, Cieplak P, Bayly CI, Gould IR, Merz KM, Ferguson DM, Spellmeyer DC, Fox T, Caldwell JW, Kollman PA (1995) A second generation force field for the simulation of proteins, nucleic acids, and organic molecules. J Am Chem Soc 117:5179–5197
Correll CC, Batie CJ, Ballou DP, Ludwig ML (1992) Phthalate dioxygenase reductase: a modular structure for electron transfer from pyridine nucleotides to [2Fe-2S]. Science 258:1604–1610
Correll CC, Ludwig ML, Bruns CM, Karplus PA (1993) Structural prototypes for an extended family of flavoprotein reductases: comparison of phthalate dioxygenase reductase with ferredoxin reductase and ferredoxin. Protein Sci 2:2112–2133
Delano WL (2002) They PyMOL molecular graphics system. DeLano Scientific, San Carlos, CA. http://www.pymol.org
Deng Z, Aliverti A, Zanetti G, Arakaki AK, Ottado J, Orellano EG, Calcaterra NB, Ceccarelli EA, Carrillo N, Karplus PA (1999) A productive NADP+ binding mode of ferredoxin-NADP+ reductase revealed by protein engineering and crystallographic studies. Nat Struct Biol 6:847–853
Dumit VI, Essigke T, Cortez N, Ullmann GM (2010) Mechanistic insights into ferredoxin-NADP(H) reductase catalysis involving the conserved glutamate in the active site. J Mol Biol 397:814–825
Faro M, Gómez-Moreno C, Stankovich M, Medina M (2002) Role of critical charged residues in reduction potential modulation of ferredoxin-NADP+ reductase. Eur J Biochem 269:2656–2661
Gorse AD, Gready JE (1997) Molecular dynamics simulations of the docking of substituted N5-deazapterins to dihydrofolate reductase. Protein Eng 10:23–30
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-PdbViewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Hermoso JA, Mayoral T, Faro M, Gomez-Moreno C, Sanz-Aparicio J, Medina M (2002) Mechanism of coenzyme recognition and binding revealed by crystal structure analysis of ferredoxin-NADP+ reductase complexed with NADP+. J Mol Biol 319:1133–1142
Humphrey W, Dalke A, Schulten K (1996) VMD—visual molecular dynamics. J Molec Graph 14:33–38
Lans I, Peregrina JR, Medina M, Garcia-Viloca M, Gonzalez-Lafont A, Lluch JM (2010) Mechanism of the hydride transfer between Anabaena Tyr303Ser FNRrd/FNRox and NADP+/H. A combined pre-steady-state kinetic/ensemble-averaged transition-state theory with multidimensional tunneling study. J Phys Chem B 114:3368–3379
Martinez-Julvez M, Medina M, Velazquez-Campoy A (2009) Binding thermodynamics of ferredoxin:NADP+ reductase: two different protein substrates and one energetics. Biophys J 96:4966–4975
Medina M (2009) Structural and mechanistic aspects of flavoproteins: photosynthetic electron transfer from photosystem I to NADP+. FEBS J 276:3942–3958
Medina M, Gómez-Moreno C (2004) Interaction of ferredoxin-NADP+ reductase with its substrates: optimal interaction for efficient electron transfer. Photosynth Res 79:113–131
Medina M, Martínez-Júlvez M, Hurley JK, Tollin G, Gómez-Moreno C (1998) Involvement of glutamic acid 301 in the catalytic mechanism of ferredoxin-NADP+ reductase from Anabaena PCC 7119. Biochemistry 37:2715–2728
Medina M, Luquita A, Tejero J, Hermoso J, Mayoral T, Sanz-Aparicio J, Grever K, Gómez-Moreno C (2001) Probing the determinants of coenzyme specificity in ferredoxin-NADP+ reductase by site-directed mutagenesis. J Biol Chem 276:11902–11912
Medina M, Abagyan R, Gómez-Moreno C, Fernandez-Recio J (2008) Docking analysis of transient complexes: interaction of ferredoxin-NADP+ reductase with ferredoxin and flavodoxin. Proteins 72:848–862
Morales R, Charon MH, Kachalova G, Serre L, Medina M, Gomez-Moreno C, Frey M (2000) A redox-dependent interaction between two electron-transfer partners involved in photosynthesis. EMBO Rep 1:271–276
Murataliev MB, Feyereisen R, Walker FA (2004) Electron transfer by diflavin reductases. Biochim Biophys Acta 1698:1–26
Musumeci MA, Arakaki AK, Rial DV, Catalano-Dupuy DL, Ceccarelli EA (2008) Modulation of the enzymatic efficiency of ferredoxin-NADP(H) reductase by the amino acid volume around the catalytic site. Febs J 275:1350–1366
Nogués I, Tejero J, Hurley JK, Paladini D, Frago S, Tollin G, Mayhew SG, Gómez-Moreno C, Ceccarelli EA, Carrillo N, Medina M (2004) Role of the C-terminal tyrosine of ferredoxin-nicotinamide adenine dinucleotide phosphate reductase in the electron transfer processes with its protein partners ferredoxin and flavodoxin. Biochemistry 43:6127–6137
Ortiz-Maldonado M, Entsch B, Ballou DP (2003) Conformational changes combined with charge-transfer interactions are essential for reduction in catalysis by p-hydroxybenzoate hydroxylase. Biochemistry 42:11234–11242
Paladini DH, Musumeci MA, Carrillo N, Ceccarelli EA (2009) Induced fit and equilibrium dynamics for high catalytic efficiency in ferredoxin-NADP(H) reductases. Biochemistry 48:5760–5768
Peregrina JR, Herguedas B, Hermoso Dominguez JA, Martínez-Júlvez M, Medina M (2009) Protein motifs involved in coenzyme interaction and enzymatic efficiency in Anabaena ferredoxin-NADP+ reductase. Biochemistry 48:3109–3119
Peregrina JR, Sanchez-Azqueta A, Herguedas B, Martinez-Julvez M, Medina M (2010) Role of specific residues in coenzyme binding, charge transfer complex formation and catalysis in Anabaena ferredoxin NADP+-reductase. Biochim Biophys Acta 1797:1638–1646
Piubelli L, Aliverti A, Arakaki AK, Carrillo N, Ceccarelli EA, Karplus PA, Zanetti G (2000) Competition between C-terminal tyrosine and nicotinamide modulates pyridine nucleotide affinity and specificity in plant ferredoxin-NADP+ reductase. J Biol Chem 275:10472–10476
Serre L, Vellieux FM, Medina M, Gomez-Moreno C, Fontecilla-Camps JC, Frey M (1996) X-ray structure of the ferredoxin:NADP+ reductase from the cyanobacterium Anabaena PCC 7119 at 1.8 Å resolution, and crystallographic studies of NADP+ binding at 2.25 Å resolution. J Mol Biol 263:20–39
Tejero J, Pérez-Dorado I, Maya C, Martínez-Júlvez M, Sanz-Aparicio J, Gómez-Moreno C, Hermoso JA, Medina M (2005) C-terminal tyrosine of ferredoxin-NADP+ reductase in hydride transfer processes with NAD(P)+/H. Biochemistry 44:13477–13490
Tejero J, Peregrina JR, Martínez-Júlvez M, Gutiérrez A, Gómez-Moreno C, Scrutton NS, Medina M (2007) Catalytic mechanism of hydride transfer between NADP+/H and ferredoxin-NADP+ reductase from Anabaena PCC 7119. Arch Biochem Biophys 459:79–90
Velazquez-Campoy A, Goñi G, Peregrina JR, Medina M (2006) Exact analysis of heterotropic interactions in proteins: characterization of cooperative ligand binding by isothermal titration calorimetry. Biophys J 91:1887–1904
Walker RC, de Souza MM, Mercer IP, Gould IR, Klug DR (2002) Large and fast relaxations inside a protein: calculation and measurement of reorganization energies in alcohol dehydrogenase. J Phys Chem B 106:11658–11665
Acknowledgments
This work has been supported by Ministerio de Ciencia e Innovación, Spain (Grant BIO2010-14983 to M.M.), and by CONSI+D, DGA (Grant PM062/2007 to M.M.). We want to thank Dr. K. Teigen and Prof. A. Martínez, from the University of Bergen, for their contribution in the early stages of this study.
Author information
Authors and Affiliations
Corresponding author
Additional information
Special Issue: Transient interactions in biology.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Peregrina, J.R., Lans, I. & Medina, M. The transient catalytically competent coenzyme allocation into the active site of Anabaena ferredoxin NADP+-reductase. Eur Biophys J 41, 117–128 (2012). https://doi.org/10.1007/s00249-011-0704-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00249-011-0704-5