Abstract
Bacterial members of the coral holobiont play an important role in determining coral fitness. However, most knowledge of the coral microbiome has come from reef-building scleractinian corals, with far less known about the nature and importance of the microbiome of octocorals (subclass Octocorallia), which contribute significantly to reef biodiversity and functional complexity. We examined the diversity and structure of the bacterial component of octocoral microbiomes over summer and winter, with a focus on two temperate (Erythropodium hicksoni, Capnella gaboensis; Sydney Harbour) and two tropical (Sinularia sp., Sarcophyton sp.; Heron Island) species common to reefs in eastern Australia. Bacterial communities associated with these octocorals were also compared to common temperate (Plesiastrea versipora) and tropical (Acropora aspera) hard corals from the same reefs. Using 16S rRNA amplicon sequencing, bacterial diversity was found to be heterogeneous among octocorals, but we observed changes in composition between summer and winter for some species (C. gaboensis and Sinularia sp.), but not for others (E. hicksoni and Sarcophyton sp.). Bacterial community structure differed significantly between all octocoral species within both the temperate and tropical environments. However, on a seasonal basis, those differences were less pronounced. The microbiomes of C. gaboensis and Sinularia sp. were dominated by bacteria belonging to the genus Endozoicomonas, which were a key conserved feature of their core microbiomes. In contrast to previous studies, our analysis revealed that Endozoicomonas phylotypes are shared across different octocoral species, inhabiting different environments. Together, our data demonstrates that octocorals harbour a broad diversity of bacterial partners, some of which comprise ‘core microbiomes’ that potentially impart important functional roles to their hosts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Coral reefs are highly productive marine ecosystems that harbour diverse communities of benthic macroorganisms and microorganisms. Within both tropical and temperate reefs, hard corals (order Scleractinia) and octocorals (subclass Octocorallia) typically represent two of the most important habitat-forming taxa. Octocorals are taxonomically diverse and widely distributed across marine environments, from the shallow tropics to the deep sea [1]. However, unlike hard corals, octocorals form an eightfold symmetry of their polyps and generally do not form calcium carbonate skeletons, but instead consist of fleshy tissue supported by small skeletal elements (sclerites) [2]. While the foundation of tropical reef structures is generally formed from hard coral skeletons, octocorals can contribute to the formation of reefs through the production of sclerites [3], and by also providing structural complexity and biodiversity to reefs [4, 5]. Both hard corals and octocorals host a variety of microorganisms, including symbiotic dinoflagellates, bacteria, viruses and archaea, which together with the coral host comprise a holobiont [6].
The ecological relationships between a coral host and its microbiome have been demonstrated to be central to reef health [7], with bacteria and archaeal partners playing essential roles in biogeochemical cycling [8,9,10], nutrient acquisition [11], protection against pathogens and extending host physiological capacity [12, 13]. Extensive examinations of the microbiome of hard corals have described the substantial diversity of bacterial communities present [14] and the sensitivity of microbial communities to environmental perturbations [15,16,17,18]. Hard corals sustain microbial communities comprising both transient microbial partners and a ‘core microbiome’, which forms a spatially and temporally stable relationship with a host species [19, 20]. While not always present in high abundance [21], members of the core microbiome can be conserved within a coral host even under acute environmental changes [22,23,24]. The presence of these consistent microbial partners points towards potentially important roles in maintaining healthy holobiont function, which may be particularly important during periods of stress [25]. However, while a substantial understanding of the nature and dynamics of hard coral microbiomes has been uncovered over the last two decades [13, 26,27,28,29], much less is known about the microbiome of octocorals.
Recent examinations of octocoral microbiomes have revealed the presence of specific bacterial communities that are distinct from the surrounding environment [30,31,32,33,34,35,36,37], but are generally lower in diversity than the bacterial communities associated with hard corals [38]. The stability of bacterial communities associated with octocorals appears to be species-specific [38]. For example, some studies have shown differences among the microbiomes of octocoral species of the same genus [33], and within an octocoral species inhabiting different locations [30, 39]. In contrast, species among the Gorgoniidae family (subclass Octocorallia) from the Mediterranean were shown to display little variation in bacterial community composition across species, with microbiome structure highly conserved over both space and time [34, 40, 41]. Despite this lack of variation of bacterial communities among some Mediterranean gorgonians, other Mediterranean octocorals such as Corallium rubrum appear to harbour highly distinct bacterial communities [41]. Other recent studies have even begun to explore patterns of phylosymbiosis among octocorals located in the Mediterranean [41] Great Barrier Reef [42] and the Algarve [43], indicating long-term associations between coral hosts and their microbiome.
Despite inconsistencies in the dynamics of octocoral microbiomes, specific groups of bacteria within the class Gammaproteobacteria have regularly been identified as prevalent members of the octocoral microbiome [38]. In particular, the Endozoicomonas genus (Order Oceanospirillales) has been shown to dominate the microbiomes of temperate gorgonians and other octocorals [36, 40, 41, 44,45,46], and has been proposed to play a significant role in octocoral health [47]. These bacteria also frequently associate with many other coral species [24, 48,49,50] and marine invertebrates [51,52,53]. Some other common members of the octocoral microbiome include bacteria belonging to Spirochaetes (Borrelia, Spirochaeta), Mollicutes (Mycoplasma, Hepatoplasma) and Flavobacteriia (Aquimarina) (see review by Van de Water et al. [38]), but the temporal and spatial stability of these octocoral associates is still unclear.
Revealing the structure and stability of the octocoral microbiome is important to aid the identification of potential symbiotic partnerships. In this study, we characterised the bacterial assemblages associated with four temperate and tropical octocoral species that commonly occur in eastern Australian reefs, which were compared to the bacterial communities associated with hard coral species inhabiting the same environments. We additionally examined temporal patterns in octocoral microbiomes over seasonal extremes (summer and winter), revealing that some octocorals (Sarcophyton sp. and Erythropodium hicksoni) host relatively consistent bacterial communities despite pronounced environmental change, whereas others (Sinularia sp. and Capnella gaboensis) appear to undergo changes in the relative abundance of core members seasonally.
Methods
Sample Collection
Specimens of the tropical octocoral species Sinularia sp. and Sarcophyton sp. (family Alcyoniidae) were collected from Heron Island (23°26′39.2″S, 151°54′47.8″E), which is located on the southern Great Barrier Reef (Australia). For comparative purposes, specimens of the hard coral species Acropora aspera were also collected from the same site. Replicate coral samples were collected from separate colonies during low tide at a depth of 1–3 m on the reef flat. The temperate octocoral species, E. hicksoni (family Anthothelidae) and C. gaboensis (family Nephtheidae), and the hard coral species, Plesiastrea versipora, were collected in parallel from Bare Island, Botany Bay, New South Wales (Australia) (33°59′29.5″S, 151°13′57.4″E), from a rocky reef situated in 5–7-m deep water. For each branching coral (Sinularia sp., Sarcophyton sp., C. gaboensis and A. aspera), small fragments (< 5 cm) were removed from the outer edge tips of the colonies, while for encrusting corals (E. hicksoni and P. versipora), < 5-cm fragments were removed from the colony edges. Triplicate samples were collected from each species from both locations and during each season, including the Austral summer (February 2017) and winter (July–August 2017). Coral colonies were unable to be tagged because of permit restrictions, so discrete colonies were sampled between the two seasons. All samples were washed after collection with sterile phosphate buffered saline (PBS) to remove any mucus layers prior to being immediately snap frozen in liquid nitrogen.
Coral Microbiome Characterisation
DNA was extracted from coral tissue by homogenising 200 mg of frozen octocoral fragment (including sclerites) within 1.5 mL of autoclaved PBS, at pH of 7.4, using a TissueRuptor® with sterile probes at full speed (33,000 rpm) for 1 min [54]. For hard corals, frozen tissue from coral fragments (~ 5 cm) was immediately extracted by air-blasting in 5 mL of PBS buffer using an air gun into a small sterile zip lock bag followed by homogenisation [55]. Genomic DNA (gDNA) was extracted directly from coral homogenates using the DNeasy Blood and Tissue Kit (Qiagen, USA), according to the manufacturer’s instructions. Negative extraction controls were also processed in parallel to test for the presence of any kit contaminants [56].
The bacterial component of all coral microbiomes was characterised using 16S rRNA gene amplicon sequencing using the V1–V3 region. For individual PCR reactions, DNA was aliquoted to 10–40 ng−L, with 12.5 µL HotStarTaq Plus Master Mix (Qiagen, USA), 1.5 µL of each 10 M primer: 27F 5′–TCGTCGGCAGCGTCAGATGTGTATAAGAGACAGAGAGTTTGATCMT–3′ and 519R 5′–GTCTCGTGGGCTCGGAGATGTGTATAAGAGACAGWATTACCGCGGCKGCTG–3′ (Illumina adapters underlined). PCR reactions were then performed in triplicate 25-µL reactions under the following thermocycling conditions: 94 °C for 3 min, then 25 cycles each of 94 °C for 30 s, 50 °C for 40 s, 72 °C for 1 min and finally 72 °C for 5 min. Following amplification, PCR products were checked in a 2% agarose gel to determine successful amplifications. Sample triplicates were then pooled and purified using calibrated AMPure XP beads (Beckman Coulter, USA). Barcodes were added to the PCR products using the Nextera XT kit (Illumina, USA) according to the manufacturer’s protocol. 16S rRNA amplicons were sequenced using an Illumina MiSeq following the manufacturer’s guidelines (2 × 300 bp) at the Molecular Research Laboratory (Shallowater, TX, USA).
Sequencing data was analysed using a customised bioinformatics pipeline https://github.com/timkahlke/ampli-tool. Firstly, FLASH (v1.2.11) was used to combine the de-multiplexed paired end reads followed by MOTHUR software (v.2.3.1) to remove any sequences that were shorter or longer than 462–549 bp, and subsequently for quality checks to remove all sequences below an average quality score of 25. The detection and removal of chimeric sequences were performed in vsearch (v2.3.2) [57]. Operational taxonomic units (OTUs) were defined as having 97% similarity using the OTU picking method in vsearch. Taxonomic classifications of OTUs were then assigned using QIIME v1.9 [58] and the RDP classifier [59] against the Silva v138 database [60]. Any sequences found within the sequenced kit negatives (total 112 OTUs), or which corresponded to mitochondria or chloroplast, were discarded from the analysis. To remove the effect of uneven library sizes, sequences were subsampled to 1835 sequences per sample, which equated to the fewest in a single sample after the removal of one exceptionally low read sample (Sa3winter; 1146 reads). Raw data files in FASTQ format were deposited into the NCBI Sequence Read Archive (SRA) and can be accessed under the bioproject number PRJNA656354.
The core microbiome of each coral species was determined using the panbiom python script [61] with the following criteria: An OTU was considered a core microbiome member if it occurred at a minimum abundance 0.001% and was present in at least five out of the six biological replicate samples across the two seasons. A core microbiome was consistently found when an abundance cut off of at least 0.1% was used; however, as outlined by Ainsworth et al. [21], at a threshold of > 0.001%, the core microbiome was considered to better represent the individual variability within replicates of coral samples and to include potentially important rare OTUs.
Statistical Analysis
Alpha diversity of bacterial communities associated with corals was assessed in MOTHUR by calculating observed OTUs and Shannon’s diversity indices. Statistical differences in alpha diversity indices and bacterial composition between coral hosts and seasons were analysed using a permutational multivariate analysis of variance (PERMANOVA) with Bray–Curtis Dissimilarity matrix in the PRIMER-E + PERMANOVA package v1.0.6. Pairwise comparisons with FDR adjustments were performed under type III partial sums of squares with 999 permutations and Monte Carlo simulations, using fixed factors ‘species’ (coral samples) and ‘season’ (summer and winter). Visualisations of beta diversity measurements were conducted using non-metric multidimensional scaling analysis (nMDS) of the bacterial composition between coral hosts and seasons using phyloseq R package [62]. Additionally, to assess differences in OTU dispersion between and among species seasonally, we used a beta-binominal regression model and Wald tests to detect differential abundance and overdispersion simultaneously using the corncob R package [63]. FDR adjustment was applied for multiple comparisons. Finally, seasonal differences among unique Endozoicomonas OTUs for each coral species were assessed using a Mann–Whitney U test in SPSS. Differences were considered significant when p < 0.05.
Results
Sequencing Overview
A total of 1,400,719 joint reads were produced from 38 samples, which included 36 coral fragments from two separate sites and two negative controls. After the removal of low-quality, chimeric and short reads, a total number of 439,343 high-quality sequences were obtained for all coral samples with an average of 13,820 sequences ranging from 1840 to 51,885 per sample analysed. A total of 338,279 and 101,064 sequences were analysed for octocoral and hard coral fragments respectively, and averaged 28,189 for octocorals and 16,844 for hard corals. The sequences were pooled to form a total of 903 OTUs clustered at 97% similarity.
Alpha Diversity of Octocoral-Associated Bacterial Assemblages
The average number of observed OTUs and the diversity of bacterial communities associated with octocorals varied significantly between coral species (Fig. 1; Tables S1 and S2; Shannon’s p < 0.005; observed OTUs p < 0.005). Among the temperate octocoral species, E. hicksoni displayed the highest mean diversity, and harboured more than twice as many OTUs (Shannon’s, 5.1 ± 0.45; observed OTUs, 134 ± 26.1; mean ± SE) than C. gaboensis (Shannon’s, 2.5 ± 0.62; observed OTUs, 47 ± 9.5). For the tropical octocoral species, the microbiome of Sarcophyton sp. was more diverse (Shannon’s, 4.4 ± 0.82; observed OTUs, 63 ± 16) than Sinularia sp. (Shannon’s, 2.8 ± 0.18; observed OTUs, 30 ± 3.9; Tables S1 and S2).
Species diversity measured by Shannon’s diversity index (a) and observed species richness (b), of bacterial operational taxonomic units (OTUs) from temperate corals (Capnella gaboensis, Erythropodium hicksoni and Plesiastrea versipora) and tropical corals (Sinularia sp. Sarcophyton sp. and Acropora aspera). Seasonal samples (summer and winter) were pooled to form n = 6 for all corals except Sarcophyton sp. samples (n = 5). Box plots represent 25th and 75th percentile, lines show medians and the + represents the mean value of the dataset. Asterix above box plots represent significant differences
Differences in bacterial diversity among octocoral and hard coral hosts were also observed. Specifically, the bacterial community associated with the temperate hard coral P. versipora (Fig. 1; Shannon’s, 5.7 ± 0.30; observed OTUs, 143 ± 22) was more diverse than that associated with the temperate octocoral C. gaboensis (Table S1; p < 0.005), but its diversity levels did not differ to E. hicksoni (p > 0.05). Among tropical corals, there was no difference in bacterial diversity between hard coral and octocoral species (p > 0.05). Additional alpha diversity statistics (Table S3), including seasonal differences between octocorals (Tables S4 and S5) and seasonal differences among each coral host (Tables S1 and S2), are available in the supplementary material.
Octocorals Harbour Distinct Bacterial Communities
Comparisons of beta diversity across octocoral microbiomes revealed a seasonal dominance of Endozoicomonas in the temperate octocoral C. gaboensis and the tropical octocoral Sinularia sp. (Fig. 2). In contrast, the microbiomes of the temperate octocoral E. hicksoni and the tropical octocoral Sarcophyton sp. were composed of a more diverse community belonging to a range of different bacterial families including Flavobacteriaceae (E. hicksoni 5.9% ± 3.4, Sarcophyton sp. 3.9% ± 2.4; mean ± SE), Granulosicoccaceae (E. hicksoni 2.5% ± 1) and Spirochaetaceae (Sarcophyton sp. 31.8% ± 16; Fig. 2). Specifically, the two main genera within the Spirochaetaceae family were Spirochaeta (Sarcophyton sp. 20%) and Borrelia (Sarcophyton sp. 11%, E. hicksoni < 1%). Bacterial community structure differed between octocoral species (Fig. 3 and Table S6; p < 0.001). Additionally, there was also a significant interactive effect between species and season (Table S6; p < 0.001). Among the temperate octocorals, differences were observed between bacterial communities of C. gaboensis and E. hicksoni during winter (p < 0.05), but not during summer. In contrast, the bacterial communities of the tropical octocorals Sinularia sp. and Sarcophyton sp. were different in summer (p < 0.05) but not in winter (Fig. 2 and Table S6). Composition of bacterial communities further differed between hard corals and octocorals. Among the tropical species examined, the microbiome of the octocoral Sinularia sp. differed in composition compared to the hard coral A. aspera in both seasons (summer p < 0.05; winter p < 0.05). Additionally, for the temperate site, differences in octocoral versus hard coral microbiome structure were observed between the octocoral C. gaboensis and the hard coral P. versipora in both seasons (summer p < 0.05; winter p < 0.05; Fig. 2 and Table S6).
Bacterial community composition seasonally of temperate (C, Capnella gaboensis; E, Erythropodium hicksoni; P, Plesiastrea versipora) and tropical (Si, Sinularia sp.; Sa, Sarcophyton sp.; A, Acropora aspera) coral replicates expressed at the genus level for (Endozoicomonas and Hepatoplasma) otherwise at the family level (where possible), and displaying taxa that were present in > 1% relative abundance. Each proportion represents taxonomy that matched to one phylogenetic group from the Silva v138 database. Top grey bars represent proportions of low abundance families that represented < 1% of the community. UC, unclassified taxonomic level
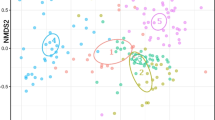
Microbial diversity at the OTU level in winter (w) and summer (s) between the temperate corals: Capnella gaboensis, Erythropodium hicksoni and Plesiastrea versipora, and tropical corals: Sarcophyton sp., Sinularia sp. and Acropora aspera. Presented in non-metric multi-dimensional scaling (nMDS) with Bray–Curtis, stress: 0.20. Triangles indicate winter and circles indicate summer. Coloured ellipses are for illustrative purposes only
Differential Abundance of OTUs Between Coral Species
Between the temperate octocorals, there were seven differentially abundant OTUs, which included one Rhodospirillales OTU with the highest mean relative abundance within the microbiome of C. gabanoesis (OTU951, 79%) in winter (Fig. 2 and Fig. 4a). Among tropical octocorals during summer, there were 17 differentially abundant OTUs (Fig. 4b). Those OTUs representing the highest relative abundance included three from the genera Hepatoplasma (OTU406, 33%; OTU407, 26%; OTU531, 0.29%) and three from Endozoicomonas (OTU137, 2.7%; OTU186, 1%; OTU59, 3.4%), which collectively represented 60% and 7% of the relative abundance within the microbiome of Sinularia sp. respectively.
Significantly differentially abundant OTUs (p < 0.05 following FDR corrections) between coral species seasonally: Erythropodium hicksoni and Capnella gaboensis in winter (a), Sinularia sp. and Sarcophyton sp. in summer (b), Sinularia sp. and Acropora aspera in winter (c) and summer (d), C. gaboensis and Plesiastrea versipora in winter (e) and summer (f), and seasonally within species: C. gaboensis (g) and Sinularia sp. (h). Genus name is provided next to OTU number where possible. Summer samples (s), winter samples (w)
Comparing differentially abundant OTUs between octocorals and hard corals, we found 15 differentially abundant OTUs in summer compared to 11 in winter between the tropical octocoral Sinularia sp. and the tropical hard coral A. aspera (Fig. 4c, d). Among the OTUs significantly higher within Sinularia sp., there were two highly abundant OTUs in winter: (Fig. 4c) Endozoicomonas (OTU129, 44%) and Hepatoplasma (OTU406, 26%), compared to four Endozoicomonas (OTU135, 11%; OTU2, 6%; OTU103, 2.6%; OTU128, 2.6%) and two Hepatoplasma (OTU406, 24%; OTU407, 18%) OTUs in summer (Fig. 4d). Between the temperate octocoral C. gaboensis and the temperate hard coral P. versipora, there were only two differentially abundant OTUs in winter (Fig. 4e) compared to ten in summer (Fig. 4f). During winter, one highly abundant OTU from the order Rhodospirillales (OTU951, 79%) was significantly higher in the octocoral C. gaboensis (Fig. 2), while in summer, the difference between the octocoral and hard coral was largely a result of significantly higher relative abundance of four Endozoicomonas OTUs (OTU100, 20%; OTU124, 4.7%; OTU143, 2.9%; OTU99, 16%) associated with C. gaboensis.
Temporal Shifts in Microbiome Structure Within Coral Hosts
We next assessed differences in microbiome structure for temporal changes within coral hosts. Within both the tropical octocoral Sinularia sp. (p < 0.05) and the temperate octocoral C. gaboensis (p < 0.05), PERMANOVA analysis revealed significant shifts in the bacterial community between summer and winter (Table S6), but no significant seasonal shifts in the bacterial communities occurred in any of the other octocoral species. Within the Sinularia-associated bacterial assemblage, there was a higher relative abundance of Hepatoplasma OTUs in summer (45%) compared to winter (11%). This shift was also coupled with a higher relative abundance of Endozoicomonas OTUs in winter (71%) compared to summer (28%) (Fig. 2). In contrast, Endozoicomonas OTUs associated with the temperate coral C. gaboensis exhibited higher relative abundance in summer (58%) compared to winter (4%) (Fig. 2). Additionally, we tested for significant differential abundant OTUs. Notably, there were five differentially abundant Endozoicomonas OTUs (OTU10, 6.5%; OTU107, 2%; OTU129, 44%; OTU148, 3%; OTU171, 2.9%) which had a higher mean relative abundance in samples of Sinularia sp. collected in winter (Fig. 4g). While among the samples of C. gaboensis collected in summer, there were three highly abundant OTUs, including two Endozoicomonas (OTU122, 9%; OTU100, 20%) and one Rhodospirillales (OTU951, 11%) (Fig. 4h).
Octocoral Core Microbiomes
To identify whether octocoral species host a ‘core microbiome’, we next examined conservation of OTUs within each coral species across summer and winter samples. No universal octocoral core microbiome was observed across all species; however, a core microbiome was revealed in each individual octocoral species (Fig. 5). Notably, at the genus level, Endozoicomonas was found to be present in 100% of coral samples in this study; however, at an OTU level, Endozoicomonas OTUs were not consistently shared between coral species. The core microbiome of the temperate octocoral C. gaboensis was made up of six OTUs, which comprised of three Endozoicomonas OTUs (OTU100, OTU122 and OTU134), two OTUs from the genus Mycoplasma and one OTU that could only be classified to the level of the Rhodospirillales order (Fig. 5). The temperate octocoral E. hicksoni had a core community of five OTUs, comprising one OTU from the genus Lutimonas, two unclassified Acidimicrobiales OTUs, one Rhizobiales OTU and one SAR11 OTU. Among the tropical octocoral species, Sinularia sp. and Sarcophyton sp., one OTU (Hepatoplasma) occurred in the core microbiome of both species, and was in fact the only conserved core microbiome member within Sarcophyton sp. (Fig. 5). In Sinularia sp., the other 6 (of 7) core microbiome members were all Endozoicomonas OTUs (OTU10, OTU56, OTU107, OTU129, OTU171 and OTU193).
Heat map of the coral core microbiome of bacterial operational taxonomic units (OTUs) uniquely present in each coral species classified to the highest taxonomic resolution possible and showing any overlaps between core members in temperate corals: C, Capnella gaboensis (purple); E, Erythropodium hicksoni (red); P, Plesiastrea versipora (orange), and tropical corals: Si, Sinularia sp. (green); Sa, Sarcophyton sp. (yellow) and A, Acropora aspera (blue). Scale on the right represents the relative abundance (%) of each taxa within the microbiome. UC, unclassified taxonomic level
Patterns in Endozoicomonas Diversity
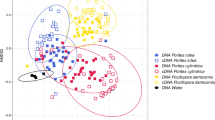
Due to the prominence of Endozoicomonas OTUs in the core microbiome of all tested octocorals, we performed a deeper analysis of patterns in Endozoicomonas diversity. Across octocoral and hard coral samples tested here, a total of 53 unique Endozoicomonas OTUs were observed (Fig. 6), ten of which were associated with the core microbiome of octocoral species (Fig. 5). Of the 53 individual Endozoicomonas OTUs, 28 occurred within the microbiome of all octocoral and hard coral species (in at least one replicate), but their presence often varied with season (Fig. 7). In fact, the diversity of unique Endozoicomonas OTUs significantly decreased from summer (34 ± 3.17–21 ± 2.09; mean ± SE) to winter (17 ± 1.73–5 ± 2.5) across all octocoral and hard coral species (Fig. 7; Mann–Whitney U test, p < 0.001 for all coral species). Endozoicomonas OTU100 dominated the diversity of the temperate octocoral C. gaboensis, while Endozoicomonas OTU129 was consistently associated with both temperate and tropical octocoral species over winter (Fig. 6).
Heat map of operational taxonomic units (OTUs) showing the overall diversity of Endozoicomonas seasonally between temperate (C, Capnella gaboensis; E, Erythropodium hicksoni; P, Plesiastrea versipora) and tropical (Si, Sinularia sp.; Sa, Sarcophyton sp.; A, Acropora aspera) corals. Scale on the right represents the relative abundance (%) of each taxa within the microbiome
Number of unique Endozoicomonas operational taxonomic units (OTUs) from temperate corals (Capnella gaboensis, Erythropodium hicksoni and Plesiastrea versipora) and tropical corals (Sinularia sp. Sarcophyton sp. and Acropora aspera), n = 3 for all corals except samples of Sarcophyton sp. collected in winter (n = 2); error bars represent standard error of the mean. Coloured bars denote summer samples and grey bars denote winter samples. Asterix above bars represent significant differences seasonally
Discussion
The principal goal of this study was to characterise the bacterial communities associated with octocoral species that are common to shallow waters of tropical and temperate reefs in eastern Australia, and to determine to what extent these communities were conserved across species, locations and seasonal extremes. Throughout tropical and temperate reefs, octocorals represent the second most abundant benthic group [2]. Yet, despite substantial evidence that the function of hard corals is strongly governed by their microbiomes [64], comparatively little is known about the microbial associates of octocorals. In this study, we show that octocoral species inhabiting temperate and tropical reefs exhibited diverse and distinct microbiomes that differ from the microbiomes associated with the reference hard coral from the same habitats. We observed significant seasonal differences in over-all microbiome composition in two out of the four octocorals analysed, but several microbiome members were still conserved over time. Notably, despite over-all differences in the microbiome composition of hard corals and octocorals, two out of the four octocoral microbiomes were dominated by the coral symbiont Endozoicomonas [47].
Differences in Coral Microbiome Composition and Seasonal Shifts
Among the octocoral species examined here, the alpha diversity of the bacterial community was comparable to that of hard coral studies [65,66,67], but often higher than previously reported values in other octocorals, such as gorgonians (Shannon’s index 1–3) and deep-sea octocorals (Shannon’s index 1–3) [34, 36, 44, 68,69,70,71]. However, some studies of octocoral microbiome diversity have also reported similar values to our study [69, 72]. In addition, we found that octocoral microbiomes were significantly different from each other, which is consistent with findings among hard coral microbiomes [35, 73, 74] and octocorals [38]. However, previous studies have shown that some octocorals show little differences in microbial community structure between species [41, 75], which has been proposed to be partly due to the incomplete evolutionary divergence between octocoral species of the same genus [41, 76]. Differences in bacterial community composition between octocoral hosts may be a reflection of different host features, such as morphology (e.g. branching vs encrusting), or alternatively may reflect different metabolic requirements among coral hosts, thereby influencing the selection of different microbial communities [77].
One morphological feature of the genus Erythropodium, which may explain the higher bacterial diversity harboured, compared to ‘branching’ octocorals, is the encrusting characteristics that cover rocky reef substrates. Mccaulkley et al. (2016) [46] outline that this feature may potentially allow greater infiltration of bacteria as the coral is both largely exposed to the substrate and the water column. Interestingly, the temperate hard coral P. versipora, which also exhibits encrusting features, was as diverse as E. hicksoni, demonstrating that bacterial diversity may indeed reflect coral morphology (e.g. encrusting vs branching) rather than other characteristics such as soft versus hard structure. Consequently, future studies should include a greater number of coral species with differing morphology to clarify the relationship between octocoral morphology and bacterial diversity, while also incorporating a greater number of replicates to account for the variability observed among samples. We also cannot rule out the possible resemblance of some of the octocoral bacterial communities to those of the surrounding environments as we did not sample for seawater or sediment during this study.
Consistent with previous studies of octocorals, the microbiomes of the tropical octocoral Sarcophyton sp. and the temperate octocoral E. hicksoni were generally stable over time [41, 46], although we did observe seasonal differences among the microbiomes of the tropical octocoral Sinularia sp. and the temperate octocoral C. gaboensis. The seasonal changes observed in the microbiome of both Sinularia sp. and C. gaboensis were predominately driven by changes in the relative abundance of core microbiome members, rather than changes in the presence and absence of bacterial members. Similar observations have also been previously observed among Mediterranean octocorals, where differences were driven by changes in the abundance of core microbiome members [41].
The most notable shift in the microbiome of Sinularia sp. involved changes in the relative abundance of Endozoicomonas OTUs, which were higher in winter than summer. This observation coincides with seasonal shifts previously observed within the temperate gorgonian Paramuricea clavata, where Endozoicomonas dominated the microbiome in summer but completely disappeared in winter [40]. In contrast, Endozoicomonas OTUs associated with the temperate octocoral C. gaboensis were higher in summer compared to winter. Such a pattern may indicate alternate temperature ranges where specific phylotypes of Endozoicomonas thrive. However, to confirm this notion, future studies characterising octocoral microbiomes will need to increase the range of geographical sampling locations and also the range of octocoral taxa characterised. Temporal variability clearly has an impact on some octocoral microbiomes. However, the significant differences observed between all octocoral microbiomes suggest that octocoral host species was the strongest influence on bacterial community composition and plays a more important role in shaping the microbiome than site and changing environmental conditions.
Octocoral Core Microbiomes
The microbiomes of the tropical octocorals Sinularia sp. and Sarcophyton sp. have been previously profiled from similar sites across the Great Barrier Reef [75]. Most notably, Bourne et al. [75] reported high relative abundance (> 50%) of sequences corresponding to Oceanospirillales in the microbiome of Sarcophyton sp., but a much lower proportion of this group (< 2%) in Sinularia sp. Contrary to these findings, we found that members of Oceanospirillales (namely Endozoicomonas) were highly abundant in the microbiome of Sinularia sp., but not in Sarcophyton sp. Such large discrepancies in the abundance of Oceanospirillales between studies could be a result of the different primers used and their affinities to this bacterial taxon. In addition, for the first time, we also profile the microbiomes of the ecologically important temperate octocorals C. gaboensis and E. hicksoni to reveal the presence of their core bacterial partners, which are likely significant for colony health.
The core microbiome of the temperate octocoral C. gaboensis and the tropical octocoral Sinularia sp. was comprised of members of the Rhodospirillales order. Rhodospirillales have previously been associated with healthy hard corals [11]. Members of the Mollicutes genera; Hepatoplasma and Mycoplasma, were also ubiquitous in octocorals, except for the temperate octocoral E. hicksoni where they were not present. While the functional roles of these bacteria are relatively unknown, their consistent association with healthy gorgonians [31, 33, 41] and cold water scleractinians [78] suggests they may contribute to coral function. Notably, the same OTU of Hepatoplasma was shared in the core of both the tropical octocorals Sinularia sp. and Sarcophyton sp., implying this bacterium may fill a similar functional role in both corals.
Endozoicomonas Diversity Among Octocorals
Endozoicomonas OTUs represented a prominent component of the core microbiome of two of the octocorals in this study where they were also among the most relatively abundant bacteria, further supporting the potentially important role of this genus in corals [79]. Additionally, each coral core microbiome contained multiple Endozoicomonas OTUs, suggesting each phylotype may occupy discrete niches [80]. However, whether the functioning of the microbiome has changed or there is functional redundancy remains unclear. The core Endozoicomonas OTUs were not unique to octocorals, as we also observed their presence to be widely spread across both hard and octocorals, and even across geographical locations. In fact, more than half of the individual Endozoicomonas OTUs sequenced in this study appeared in at least one replicate of every coral species examined. The diverse spatial occurrence of these specific phylotypes is in contrast to previous studies of hard corals, where each coral species appears to form its own unique association with specific Endozoicomonas OTUs [49, 81]. Furthermore, a recent meta-analysis revealed strong partitioning of Endozoicomonas phylotypes between hard corals and octocorals [68] inferring different Endozoicomonas OTUs form species-specific relationships within hosts [71, 82]. In our study, however, the data indicate that host specificity of Endozoicomonas, at least at the OTU level, is still somewhat unresolved.
Host Identity May Influence Microbial Community Structure
In addition to the high representation of Endozoicomonas among the microbiomes of Sinularia sp. and C. gaboensis, the tropical octocoral Sarcophyton sp. and the temperate octocoral E. hicksoni exhibited highly diverse microbiomes made up of many more lower abundant bacterial genera (< 5%). Some of the other less dominant bacterial genera found included OTUs from the genus Aquimarina (family Flavobacteriaceae), which were associated with E. hicksoni microbiomes. Aquimarina are commonly associated with both tropical and temperate gorgonians and possess a number of genes involved in nutrient cycling (e.g. carbon, nitrogen and sulphur) [83]; however, their role within the coral holobiont is still unclear. Another interesting feature associated with the E. hicksoni microbiome was the presence of OTUs from the genus Granulosicoccus (family Granulosicoccaceae). Species within this genus have been found in temperate octocorals [41] and are described as obligate chemoheterotrophs [84] with the capacity of sulphur metabolism (e.g. DMSP). Members of the Spirochaetaceae family (Spirochaeta and Borrelia) were also present in the microbiomes of both Sarcophyton sp. and E. hicksoni. Their ubiquity among healthy octocorals [33, 45, 85] suggests a beneficial role in the holobiont; however, this has currently not been determined.
Evidence has been growing that microbial associations are strongly driven by host phylogeny [42, 71, 86]. Additionally, varying strategies of microbial restructuring may exist between coral hosts in response to environmental change. The diverse and somewhat heterogenous microbiomes of the octocorals E. hicksoni and Sarcophyton sp. indicate a level of microbiome flexibility. This concept has been largely explored in hard corals and black corals [72, 81, 87, 88], and suggests corals may adjust their microbial members depending on coral holobiont requirements. Actively changing their microbiome may allow these corals to rapidly respond to environmental change through the selection of beneficial microbes present in the surrounding environment. Alternatively, shifts in coral microbial associates may merely reflect changes in host physiology, thereby creating new ecological niches that are filled by opportunistic transient bacterial communities rather than the host actively seeking beneficial microbial members. Nevertheless, it remains to be determined whether a flexible microbiome will be key to a coral’s ability to survive under future changing conditions caused by climate change.
Conclusion
This study revealed that octocorals harbour unique and specific bacterial assemblages that often show similar diversity levels but different composition to the bacterial communities associated with hard corals. Octocoral species hosted temporally conserved core microbiome members, dominated by Endozoicomonas, indicating a potentially profound role of these bacteria in the octocoral holobiont within both tropical and temperate environments. Seasonal shifts in the relative abundance of core members, in some corals, are indicative of influence from local conditions and may facilitate host acclimation to changing environmental conditions. The stark differences in octocoral bacterial communities locally suggest host specificity is a primary driver in shaping microbial composition rather than environmental influences.
Data availability
‘Methods’ Raw data files in FASTQ format were deposited into the NCBI Sequence Read Archive (SRA) and can be accessed under the bioproject number PRJNA656354.
Code Availability
Not applicable.
References
McFadden CS, Sánchez JA, France SC (2010) Molecular phylogenetic insights into the evolution of octocorallia: a review. Integr comp Biol. pp 389–410. https://doi.org/10.1093/icb/icq056
Fabricius K, Alderslade P (2001) Soft corals and sea fans: a comprehensive guide to the tropical shallow water genera of the central-west Pacific, the Indian Ocean and the Red Sea. Australian Institute of Marine Science, Townsville, p 14
Jeng MS, Huang HD, Dai CF et al (2011) Sclerite calcification and reef-building in the fleshy octocoral genus Sinularia (Octocorallia: Alcyonacea). Coral Reefs 30:925–933. https://doi.org/10.1007/s00338-011-0765-z
Sanchez JAA, Zea S, Diaz JM (1998) Patterns of octocoral and black coral distribution in the oceanic barrier reef-complex of Providencia Island, Southwestern Caribbean. Caribb J Sci 34:250–264
De’Ath G, Fabricius KE, Sweatman H, Puotinen M, (2012) The 27-year decline of coral cover on the Great Barrier Reef and its causes. Proc Natl Acad Sci U S A 109:17995–17999. https://doi.org/10.1073/pnas.1208909109
Rohwer F, Seguritan V, Azam F, Knowlton N (2002) Diversity and distribution of coral-associated bacteria. Mar Ecol Prog Ser 243:1–10. https://doi.org/10.3354/meps243001
Bourne DG, Morrow KM, Webster NS (2016) Insights into the coral microbiome: underpinning the health and resilience of reef ecosystems. Annu Rev Microbiol 70:317–340. https://doi.org/10.1146/annurev-micro-102215-095440
Rädecker N, Pogoreutz C, Voolstra CR et al (2015) Nitrogen cycling in corals: the key to understanding holobiont functioning? Trends Microbiol 23:490–497
Raina JB, Tapiolas D, Willis BL, Bourne DG (2009) Coral-associated bacteria and their role in the biogeochemical cycling of sulfur. Appl Environ Microbiol 75:3492–3501. https://doi.org/10.1128/AEM.02567-08
Siboni N, Ben-Dov E, Sivan A, Kushmaro A (2008) Global distribution and diversity of coral-associated Archaea and their possible role in the coral holobiont nitrogen cycle. Environ Microbiol 10:2979–2990. https://doi.org/10.1111/j.1462-2920.2008.01718.x
Ceh J, Raina JB, Soo RM et al (2012) Coral-bacterial communities before and after a coral mass spawning event on Ningaloo Reef. PLoS ONE 7(5):e36920. https://doi.org/10.1371/journal.pone.0036920
Putnam HM, Barott KL, Ainsworth TD, Gates RD (2017) The vulnerability and resilience of reef-building corals. Curr Biol 27:R528–R540
Peixoto RS, Rosado PM, Leite DCA et al (2017) Beneficial microorganisms for corals (BMC): proposed mechanisms for coral health and resilience. Front. Microbiol 8:341. https://doi.org/10.3389/fmicb.2017.00341
Blackall LL, Wilson B, Van Oppen MJH (2015) Coral-the world’s most diverse symbiotic ecosystem. Mol Ecol 24:5330–5347
Bourne D, Iida Y, Uthicke S, Smith-Keune C (2008) Changes in coral-associated microbial communities during a bleaching event. ISME J 2:350–363. https://doi.org/10.1038/ismej.2007.112
Witt V, Wild C, Anthony KRN et al (2011) Effects of ocean acidification on microbial community composition of, and oxygen fluxes through, biofilms from the Great Barrier Reef. Environ Microbiol 13:2976–2989. https://doi.org/10.1111/j.1462-2920.2011.02571.x
Lee STM, Davy SK, Tang SL et al (2015) Successive shifts in the microbial community of the surface mucus layer and tissues of the coral Acropora muricata under thermal stress. FEMS Microbiol Ecol 91(12):fiv142. https://doi.org/10.1093/femsec/fiv142
Gajigan AP, Diaz LA, Conaco C (2017) Resilience of the prokaryotic microbial community of Acropora digitifera to elevated temperature. Microbiol Open 72:e00478-e00411. https://doi.org/10.1002/mbo3.478
Huggett MJ, Apprill A (2019) Coral microbiome database: integration of sequences reveals high diversity and relatedness of coral-associated microbes. Environ Microbiol Rep 11:372–385. https://doi.org/10.1111/1758-2229.12686
Hernandez-Agreda A, Gates RD, Ainsworth TD (2017) Defining the core microbiome in corals’ microbial soup. Trends Microbiol 25:125–140
Ainsworth TD, Krause L, Bridge T et al (2015) The coral core microbiome identifies rare bacterial taxa as ubiquitous endosymbionts. ISME J 9:2261–2274. https://doi.org/10.1038/ismej.2015.39
Thornhill DJ, Fitt WK, Schmidt GW (2006) Highly stable symbioses among western Atlantic brooding corals. Coral Reefs 25:515–519. https://doi.org/10.1007/s00338-006-0157-y
Lema KA, Willis BL, Bourne DG (2014) Amplicon pyrosequencing reveals spatial and temporal consistency in diazotroph assemblages of the Acropora millepora microbiome. Environ Microbiol 16:3345–3359. https://doi.org/10.1111/1462-2920.12366
Pogoreutz C, Rädecker N, Cárdenas A et al (2018) Dominance of Endozoicomonas bacteria throughout coral bleaching and mortality suggests structural inflexibility of the Pocillopora verrucosa microbiome. Ecol Evol 8:2240–2252. https://doi.org/10.1002/ece3.3830
Epstein HE, Torda G, van Oppen MJH (2019) Relative stability of the Pocillopora acuta microbiome throughout a thermal stress event. Coral Reefs 38:373–386. https://doi.org/10.1007/s00338-019-01783-y
Glasl B, Herndl GJ, Frade PR (2016) The microbiome of coral surface mucus has a key role in mediating holobiont health and survival upon disturbance. ISME J 10:2280–2292. https://doi.org/10.1038/ismej.2016.9
Lema KA, Clode PL, Kilburn MR et al (2016) Imaging the uptake of nitrogen-fixing bacteria into larvae of the coral Acropora millepora. ISME J 10:1804–1808. https://doi.org/10.1038/ismej.2015.229
Thurber RV, Willner-Hall D, Rodriguez-Mueller B et al (2009) Metagenomic analysis of stressed coral holobionts. Environ Microbiol 11:2148–2163. https://doi.org/10.1111/j.1462-2920.2009.01935.x
Reshef L, Koren O, Loya Y et al (2006) The coral probiotic hypothesis. Environ Microbiol 8:2068–2073. https://doi.org/10.1111/j.1462-2920.2006.01148.x
Robertson V, Haltli B, McCauley E et al (2016) Highly variable bacterial communities associated with the octocoral Antillogorgia elisabethae. Microorganisms 4:23. https://doi.org/10.3390/microorganisms4030023
Gray MA, Stone RP, Mclaughlin MR, Kellogg CA (2011) Microbial consortia of gorgonian corals from the Aleutian islands. FEMS Microbiol Ecol 76:109–120. https://doi.org/10.1111/j.1574-6941.2010.01033.x
Penn K, Wu D, Eisen JA, Ward N (2006) Characterization of bacterial communities associated with deep-sea corals on Gulf of Alaska seamounts. Appl Environ Microbiol 72:1680–1683. https://doi.org/10.1128/AEM.72.2.1680-1683.2006
Holm JB, Heidelberg KB (2016) Microbiomes of Muricea californica and M. fruticosa: comparative analyses of two co-occurring Eastern Pacific Octocorals. Front Microbiol 7:917. https://doi.org/10.3389/fmicb.2016.00917
Van de Water JAJM, Melkonian R, Voolstra CR et al (2017) Comparative assessment of Mediterranean gorgonian-associated microbial communities reveals conserved core and locally variant bacteria. Microb Ecol 73:466–478. https://doi.org/10.1007/s00248-016-0858-x
Osman EO, Suggett DJ, Voolstra CR et al (2020) Coral microbiome composition along the northern Red Sea suggests high plasticity of bacterial and specificity of endosymbiotic dinoflagellate communities. Microbiome 8:1–16. https://doi.org/10.1186/s40168-019-0776-5
Bayer T, Arif C, Ferrier-Pagès C et al (2013) Bacteria of the genus Endozoicomonas dominate the microbiome of the Mediterranean gorgonian coral Eunicella cavolini. Mar Ecol Prog Ser 479:75–84. https://doi.org/10.3354/meps10197
Cleary DFR, Polónia & ARM, Reijnen & BT, et al (2020) Prokaryote communities inhabiting endemic and newly discovered sponges and octocorals from the Red Sea. Springer. https://doi.org/10.1007/s00248-019-01465-w
Van De Water JAJM, Allemand D, Ferrier-Pagès C (2018) Host-microbe interactions in octocoral holobionts - recent advances and perspectives. Microbiome 6:1–28. https://doi.org/10.1186/s40168-018-0431-6
Goldsmith DB, Kellogg CA, Morrison CL et al (2018) Comparison of microbiomes of cold-water corals Primnoa pacifica and Primnoa resedaeformis, with possible link between microbiome composition and host genotype. Sci Rep 8:12383. https://doi.org/10.1038/s41598-018-30901-z
La Rivière M, Roumagnac M, Garrabou J, Bally M (2013) Transient shifts in bacterial communities associated with the temperate gorgonian Paramuricea clavata in the Northwestern Mediterranean Sea. PLoS ONE 8(2):e57385. https://doi.org/10.1371/journal.pone.0057385
Van de Water JAJM, Voolstra CR, Rottier C et al (2018) Seasonal stability in the microbiomes of temperate gorgonians and the red coral Corallium rubrum across the Mediterranean Sea. Microb Ecol 75:274–288. https://doi.org/10.1007/s00248-017-1006-y
Pollock FJ, McMinds R, Smith S et al (2018) Coral-associated bacteria demonstrate phylosymbiosis and cophylogeny. Nat Commun 9:1–13. https://doi.org/10.1038/s41467-018-07275-x
Keller-Costa T, Lago-Lestón A, Saraiva JP et al (2021) Metagenomic insights into the taxonomy, function, and dysbiosis of prokaryotic communities in octocorals. Microbiome 9:1–21. https://doi.org/10.1186/S40168-021-01031-Y
Correa H, Haltli B, Duque C, Kerr R (2013) Bacterial communities of the gorgonian octocoral Pseudopterogorgia elisabethae. Microb Ecol 66:972–985. https://doi.org/10.1007/s00248-013-0267-3
Wessels W, Sprungala S, Watson SA, et al (2017) The microbiome of the octocoral Lobophytum pauciflorum: minor differences between sexes and resilience to short-term stress. FEMS Microbiol Ecol 93:fix013. https://doi.org/10.1093/femsec/fix013
McCauley EP, Haltli B, Correa H, Kerr RG (2016) Spatial and temporal investigation of the microbiome of the Caribbean octocoral Erythropodium caribaeorum. FEMS Microbiol Ecol 92:1–10. https://doi.org/10.1093/femsec/fiw147
Neave MJ, Apprill A, Ferrier-Pagès C, Voolstra CR (2016) Diversity and function of prevalent symbiotic marine bacteria in the genus Endozoicomonas. Appl Microbiol Biotechnol 100:8315–8324
Neave MJ, Rachmawati R, Xun L et al (2017) Differential specificity between closely related corals and abundant Endozoicomonas endosymbionts across global scales. ISME J 11:186–200. https://doi.org/10.1038/ismej.2016.95
Bayer T, Neave MJ, Alsheikh-Hussain A et al (2013) The microbiome of the red sea coral stylophora pistillata is dominated by tissue-associated endozoicomonas bacteria. Appl Environ Microbiol 79:4759–4762. https://doi.org/10.1128/AEM.00695-13
Maher RL, Schmeltzer ER, Meiling S et al (2020) Coral microbiomes demonstrate flexibility and resilience through a reduction in community diversity following a thermal stress event. Front Ecol Evol 8:356. https://doi.org/10.3389/FEVO.2020.555698
Dishaw LJ, Flores-Torres J, Lax S et al (2014) The gut of geographically disparate Ciona intestinalis harbors a core microbiota. PLoS ONE 9:e93386. https://doi.org/10.1371/JOURNAL.PONE.0093386
Li Z, Kellogg CA, Gophna U et al (2019) Tissue-specific microbiomes of the Red Sea giant clam Tridacna maxima highlight differential abundance of Endozoicomonadaceae. Front Microbiol 10:2661. https://doi.org/10.3389/fmicb.2019.02661
Forget NL, Juniper SK (2013) Free-living bacterial communities associated with tubeworm (Ridgeia piscesae) aggregations in contrasting diffuse flow hydrothermal vent habitats at the Main Endeavour Field, Juan de Fuca Ridge. Microbiologyopen 2:259–275. https://doi.org/10.1002/MBO3.70
Haydon TD, Seymour JR, Suggett DJ (2018) Soft corals are significant DMSP producers in tropical and temperate reefs. Mar Biol 165:1–7. https://doi.org/10.1007/s00227-018-3367-2
Weber L, DeForce E, Apprill A (2017) Optimization of DNA extraction for advancing coral microbiota investigations. Microbiome 5:18. https://doi.org/10.1186/s40168-017-0229-y
Salter SJ, Cox MJ, Turek EM et al (2014) Reagent and laboratory contamination can critically impact sequence-based microbiome analyses. BMC Biol 12:1–12. https://doi.org/10.1186/s12915-014-0087-z
Rognes T, Flouri T, Nichols B et al (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 18(4):e2584. https://doi.org/10.7717/peerj.2584
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Quast C, Pruesse E, Yilmaz P et al (2013) The SILVA ribosomal RNA gene database project: improved data processing and web-based tools. Nucleic Acids Res 41(Database issue):D590–D596. https://doi.org/10.1093/nar/gks1219
Kahlke T (2018) timkahlke/panbiom 1.0 (Version 1.0). Zenodo. https://doi.org/10.5281/zenodo.1137875
McMurdie PJ, Holmes S (2013) phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS ONE 8:e61217. https://doi.org/10.1371/journal.pone.0061217
Martin BD, Witten D, Willis AD (2020) Modeling microbial abundances and dysbiosis with beta-binominal regression. Ann appl Stat 14:94–115. https://doi.org/10.1214/19-AOAS1283
van Oppen MJH, Blackall LL (2019) Coral microbiome dynamics, functions and design in a changing world. Nat Rev Microbiol 17:557–567. https://doi.org/10.1038/s41579-019-0223-4
Lee OO, Yang J, Bougouffa S et al (2012) Spatial and species variations in bacterial communities associated with corals from the Red Sea as revealed by pyrosequencing. Appl Environ Microbiol 78:7173–7184. https://doi.org/10.1128/AEM.01111-12
Shore-Maggio A, Runyon CM, Ushijima B et al (2015) Differences in bacterial community structure in two color morphs of the Hawaiian reef coral Montipora capitata. Appl Environ Microbiol 81:7312–7318. https://doi.org/10.1128/AEM.01935-15
Sharp KH, Pratte ZA, Kerwin AH et al (2017) Season, but not symbiont state, drives microbiome structure in the temperate coral Astrangia poculata. Microbiome 5:120. https://doi.org/10.1186/s40168-017-0329-8
Kellogg CA (2019) Microbiomes of stony and soft deep-sea corals share rare core bacteria. Microbiome 7:90. https://doi.org/10.1186/s40168-019-0697-3
McCauley M, Jackson CR, Goulet TL (2020) Microbiomes of Caribbean octocorals vary over time but are resistant to environmental change. Front Microbiol 11:1272. https://doi.org/10.3389/fmicb.2020.01272
Woo S, Yang SH, Chen HJ et al (2017) Geographical variations in bacterial communities associated with soft coral Scleronephthya gracillimum. PLoS ONE 12(8):e0183663. https://doi.org/10.1371/journal.pone.0183663
La Rivière M, Garrabou J, Bally M (2015) Evidence for host specificity among dominant bacterial symbionts in temperate gorgonian corals. Coral Reefs 34:1087–1098. https://doi.org/10.1007/s00338-015-1334-7
Van de Water JAJM, Coppari M, Enrichetti F et al (2020) Local conditions influence the prokaryotic communities associated with the mesophotic black coral Antipathella subpinnata. Front Microbiol 11:2423. https://doi.org/10.3389/fmicb.2020.537813
Chu ND, Vollmer SV (2016) Caribbean corals house shared and host-specific microbial symbionts over time and space. Environ Microbiol Rep 8:493–500. https://doi.org/10.1111/1758-2229.12412
Sunagawa S, Woodley CM, Medina M (2010) Threatened corals provide underexplored microbial habitats. PLoS ONE 5(3):e954. https://doi.org/10.1371/journal.pone.0009554
Bourne DG, Dennis PG, Uthicke S et al (2013) Coral reef invertebrate microbiomes correlate with the presence of photosymbionts. ISME J 7:1452–1458. https://doi.org/10.1038/ismej.2012.172
Aurelle D, Pivotto ID, Malfant M et al (2017) Fuzzy species limits in Mediterranean gorgonians (Cnidaria, Octocorallia): inferences on speciation processes. Zool Scr 46:767–778. https://doi.org/10.1111/zsc.12245
Camp EF, Suggett DJ, Pogoreutz C et al (2020) Corals exhibit distinct patterns of microbial reorganisation to thrive in an extreme inshore environment. Coral Reefs 39:701–716. https://doi.org/10.1007/s00338-019-01889-3
Kellogg CA, Lisle JT, Galkiewicz JP (2009) Culture-independent characterization of bacterial communities associated with the cold-water coral Lophelia pertusa in the northeastern Gulf of Mexico. Appl Environ Microbiol 75:2294–2303. https://doi.org/10.1128/AEM.02357-08
Ziegler M, Roik A, Porter A et al (2016) Coral microbial community dynamics in response to anthropogenic impacts near a major city in the central Red Sea. Mar Pollut Bull 105:629–640. https://doi.org/10.1016/j.marpolbul.2015.12.045
Neave MJ, Michell CT, Apprill A, Voolstra CR (2017) Endozoicomonas genomes reveal functional adaptation and plasticity in bacterial strains symbiotically associated with diverse marine hosts. Sci Rep 17(7):40579. https://doi.org/10.1038/srep40579
Ziegler M, Seneca FO, Yum LK et al (2017) Bacterial community dynamics are linked to patterns of coral heat tolerance. Nat Commun 8:14213. https://doi.org/10.1038/ncomms14213
Keller-Costa T, Eriksson D, Gonçalves JMS et al (2017) The gorgonian coral Eunicella labiata hosts a distinct prokaryotic consortium amenable to cultivation. FEMS Microbiol Ecol 93:143. https://doi.org/10.1093/FEMSEC/FIX143
Keller-Costa T, Silva R, Lago-Lestón A (2016) Genomic insights into Aquimarina sp. strain EL33, a bacterial symbiont of the gorgonian coral Eunicella labiata. Genome Announc 4:e00855–16. https://doi.org/10.1128/genomea.00855-16
Lee K, Lee H, Choi T et al (2007) Granulosicoccaceae fam. nov., to include Granulosicoccus antarcticus gen. nov., sp. nov., a non-phototrophic, obligately aerobic chemoheterotroph in the order. J Microbiol Biotechnol 17(9):1483–1490
Van De Water JA, Melkonian R, Junca H, Voolstra CR, Reynaud S, Allemand D, Ferrier-Pagès C (2016) Spirochaetes dominate the microbial community associated with the red coral Corallium rubrum on a broad geographic scale. Sci Rep 6:1–7. https://doi.org/10.1038/srep27277
O’Brien PA, Tan S, Yang C et al (2020) Diverse coral reef invertebrates exhibit patterns of phylosymbiosis. ISME J 14:2211–2222. https://doi.org/10.1038/s41396-020-0671-x
Ziegler M, Grupstra CGB, Barreto MM et al (2019) Coral bacterial community structure responds to environmental change in a host-specific manner. Nat Commun 10(1):3092. https://doi.org/10.1038/s41467-019-10969-5
Voolstra CR, Ziegler M (2020) Adapting with microbial help: microbiome flexibility facilitates rapid responses to environmental change. BioEssays 42:2000004. https://doi.org/10.1002/bies.202000004
Acknowledgements
The authors would like to thank the staff at Heron Island Research Station for their technical support, and Mathieu Pernice, Audrey Commault and Samantha Goyen for their assistance with sampling. We are grateful for the reviewers’ thorough and constructive feedback which has helped improve the quality of this manuscript. Coral samples were collected under the Great Barrier Reef Marine Park permit G17/39149.1 issued to Emma Camp/David Suggett, and Department of Primary Industries permit P15/0042-1.1 issued to Samantha Goyen/David Suggett. Trent Haydon is supported by an Australian Government Research Training Program Scholarship.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Ethics Approval
Not applicable.
Consent to Participate
Not applicable.
Consent for Publication
Not applicable.
Conflict of Interest
The authors declare no competing interests.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Haydon, T.D., Suggett, D.J., Siboni, N. et al. Temporal Variation in the Microbiome of Tropical and Temperate Octocorals. Microb Ecol 83, 1073–1087 (2022). https://doi.org/10.1007/s00248-021-01823-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-021-01823-7