Abstract
There is evidence that shows that phosphorus (P) fertilization has a moderate effect on the rhizosphere microbial composition of cultivated crops. But how this effect is manifested on wild species of the same crop is not clear. This study compares the impact of phosphorus fertilization with rhizosphere bacterial community composition and its predicted functions, related to P-cycling genes, in both cultivated and non-cultivated potato (Solanum sp.) plants. It was found that the biomass of non-cultivated potatoes was more responsive to P fertilization as compared with cultivated plants. Differences in general bacterial community composition patterns under increasing P amendments were subtle for both potato groups. However, potato genotype significantly influenced community composition with several bacterial families being more abundant in the cultivated plants. In addition, the predicted phosphatases had lower abundances in modern cultivars compared with non-cultivated potatoes. In summary, despite higher accumulation of differentially abundant bacteria in the rhizosphere of cultivated plants, the responsiveness of these plants to increase P levels was lower than in non-cultivated plants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The microbial community associated with the rhizosphere can have a profound effect on nutrition, growth, and health of plants in agro-ecosystems [1]. Root symbionts and free-living microbes are responsible for the acquisition of nutrients that are otherwise inaccessible to plants [2]. The effectiveness of plant root and microbial synergies is influenced by management practices impacting soil biological communities [3]. Previous studies have shown that the applications of herbicides, pesticides, and tillage can lead to a shift of the microbial community composition and function [4,5,6]. Despite this, our understanding of the effects of excessive fertilization and its impact in rhizosphere microbiome composition and crop health remain sparse.
Phosphorus (P) is an essential element in plant nutrition and a major limiting nutrient in agriculture [7]. After being applied to soils, P can be fixed by soil sorption or precipitated by free aluminum (Al+3), iron (Fe+3), or calcium (Ca+2) depending on soil pH, thus becoming unavailable to plants [8]. Additionally, P uptake in most crops is generally restricted to 10–20% of the P applied [9]; thus, farmers are continually encouraged to increase P fertilization and any excess P tends to accumulate in soils [10]. This residual P fertilizer is also known as “P legacy” and is defined as the difference of cumulative inputs including mineral P fertilizer, atmospheric deposition, and weathering and cumulative outputs, such as P removal during harvest, erosion, leaching, and run off [11]. Currently, P fertilization inputs into arable lands are nearly double the rate of actual plant uptake capacity. For instance, during the period 1965–2007, the input of P fertilizer and manure in North America was 500 kg ha−1, while cumulative P uptake was approximately 250 kg ha−1: meaning that only half of the applied P was utilized by crops [12]. The excess of synthetic fertilization in soil is becoming more apparent [13]; recent evidence shows that phosphatase demand in plant influences root microbiome community [14], however, the evidence on the potential detrimental effect of P on microbial communities is scarce. Hence, there is a need to understand the impact of P legacy and its potential interaction with microbial communities in plant rhizospheres.
Cultivated potatoes (Solanum tuberosum L.) were domesticated between 8000 and 10,000 years ago from wild Solanum species native to the Andes of southern Peru [15]. In its natural habitat, wild potato species grow in nutrient-limited (and P deficient) soils, suggesting that there may be a rich interaction between plants and beneficial microbes to support adequate plant nutrition. Over the last 150 years, potatoes have been bred mainly for below-ground traits, such as tuber development to increase yield [16, 17]. In conjunction, many cultivars have been bred within a context of frequent P fertilization during domestication. As a consequence, high fertility conditions used during plant breeding and selection may have hampered the ability of plants to initiate beneficial interactions with microbes related to nutrient availability and solubilization [18, 19]. In natural environments, numerous microorganisms including species of fungi and bacteria are effective at releasing P from organic and inorganic soil P pools through mineralization and solubilization, respectively [20]. Short- and long-term studies have shown that this symbiosis can be altered when high levels of P are added to soils. A short-term greenhouse experiment, in blueberries, showed that high levels of P (192 kg ha−1) reduced the abundance of genes associated with phosphatases production in soil, accompanied by a shift in the microbial community associated of the rhizosphere [21]. In a different study, incremental addition of soil phosphate in a long-term P fertilization study significantly affected arbuscular mycorrhizal fungal community composition [22]. This microbial shift caused by fertilization over time seems to be more prevalent in modern crop varieties. For example, the nitrogen-fixing endophyte Azoarcus spp. preferentially colonized wild rice species and old varieties as compared with modern cultivars [23]. A more recent study demonstrated that an indigenous landrace of maize grown using little or no fertilizer showed high levels of nitrogen (N) fixation, deriving up to 82% of the plant N from the atmosphere. This was mainly due to diazotrophic bacteria which were not present in modern maize varieties [24]. A reduced number of studies have addressed the rhizosphere microbiome of potato, particularly focusing on differences between cultivars and plant developmental stages [25, 26]. The genotypic effect on the microbiome of wild and domesticated plants is available for limited number of crops such as Phaseolus vulgaris [27], but not for potato under phosphorus fertilization amendment.
To successfully manage beneficial interactions between plant roots and microbes, we need to better understand the impact of soil management on root-associated microbial communities along a domestication gradient. In this study, it was hypothesized that high P rates would differentially affect the microbial composition particularly of P solubilizing bacteria in modern cultivars compared with non-cultivated varieties. In addition, we anticipated a higher P content and biomass gain for the non-cultivated potatoes compared with modern potato cultivars. Lastly, bacteria associated with the P-cycle (i.e., containing phosphatase enzymes) were expected to be significantly more abundant for non-cultivated varieties.
Materials and Methods
Selection of Potato Accessions and Growth Conditions
This study was performed in a greenhouse at the Plant Growth Facility (PGF) of Colorado State University (CSU), Fort Collins, CO, between the month of March and May 2018. Three commercial cultivars of Solanum tuberosum widely grown in North America were selected: “Yukon Gold” (YG), “Russet Burbank” (RB), and “Red Norland” (RN); one direct progenitor of the three commercial cultivars: Solanum tuberosum subsp. tuberosum (PI 595492) (STT); one landrace: Solanum tuberosum subsp. andigena (PI258907) (STA); and one wild type: Solanum bulbocastanum (PI 275184) (SB) were chosen to evaluate the impact of P fertilization on rhizosphere soil microbial communities along a crop domestication gradient. Certified organic tubers from commercial cultivars were obtained from Grand Teton Organics farm, Idaho, USA. Tubers were pre-sprouted during ten 10 days at room temperature. Botanical seeds from the three non-cultivated accessions were obtained from the Potato Gene Bank, Sturgeon Bay, Wisconsin, USA. Potato seeds were pre-treated with gibberellic acid at 2000 ppm and surface disinfected with 3% of sodium hypochlorite and left for 24 h in distillate water. Seeds were pre-germinated on wet filter paper and moved to Hoagland nutritional solution (4.33 g L−1) for 9 days. Small plants were transplanted to a substrate made of vermiculite, sand, and soil (1:1:1) during 10 days for fast growth. Tubers of the three cultivars and seedlings of three accessions were transplanted to 15-cm pots containing a mix of two parts sand and one-part soil collected (0-–20-cm depth) from a field previously planted with winter squash (field 7 South) from the Agricultural Research, Development and Educational Center–South farm (ARDEC) of Colorado State University. Soil was collected on March 13, 2018, soil texture is sandy clay loam, pH of 8.4, and organic matter of 3.0%. Plants were grown on benches in the greenhouse for approximately 70 days.
Fertilizer super triple phosphate (0-45-0) was applied at 3 rates equivalent to 0, 67, and 133 kg ha−1 every 15 days starting on March 22nd to all six different potato genotypes. Suggested P fertilizer rates are based on preplant application related to soil test levels. Extractable P level in the soil used for this experiment was > 11 ppm, which is considered a high level for the AB-DTPA (ammonium bicarbonate-diethylenetriaminepentaacetic acid) test. The fertilizer rate suggested when P content is above 11 ppm, is 67 kg ha−1, and expected yield of 45 tons of potato per hectare [28].
Plants were irrigated to field capacity with a sprayer on alternate days. The range of average min and max temperatures in the greenhouse during the experiment was 16 to 30 °C. Treatments were arranged in a randomized complete block design with 10 replicates per treatment. Every replicate consisted of a single potato plant per pot. A research randomizer software version 4.0 of open access was used for this purpose (http://www.randomizer.org).
Rhizosphere Soil Collection and Plant Biomass Sampling
For this study, we refer to the ectorhizosphere as rhizosphere soil. The ectorhizosphere is the area surrounding the roots [29]. We sampled the immediate soil adjacent to the ectorhizosphere. Rhizosphere soil from each plant was collected after 70 days of growth by removing the plants from the pots, and gently shaking the whole plant to separate the bulk soil and exposing the plant roots. The remaining soil particles, still attached to the root hairs in the rhizosphere zone, were collected, pooled into bags as individual samples, stored for 24 h at 2 °C, and then the DNA was extracted. Fresh plants, shoots, and roots were weighed and fresh biomass recorded immediately after harvest. Plants were then placed in an oven for 4 days at 80 °C and weighed.
The fact that the three cultivated potatoes were grown from tuber pieces (as opposed to the non-cultivated plants grown from botanical seeds) added an additional belowground weight to the cultivated group. To correct this, we reduced 3 g (i.e., average dry weight of the starting tuber piece) from the total dry weight of each of the three cultivated potatoes.
Plant Nutrient Analysis
All 10 dried replicate plants from each given cultivar per each of the three P rates were mixed and homogenized. One dry sample per each treatment was submitted for analysis to the Soil, Water and Plant Testing Laboratory at CSU to determine total P content.
DNA Extraction, PCR, and 16S rRNA Amplicon Sequencing
Total DNA was extracted from each 0.25 g of rhizosphere soil sample using a DNeasy Power Soil DNA isolation kit (Qiagen, Hilden, Germany) according to the manufacturer’s instruction. The DNA was quantified using a spectrophotometer (Thermo Scientific NanoDrop 2000c, Vernon Hills, IL). All isolated DNA had an absorbance ratio (A260/A280) between 1.8 and 2.0.
Quantitative PCR (qPCR) amplification of the bacterial 16 S ribosomal RNA (rRNA) genes (V1–V3 hypervariable region) was performed with the 27 F and 338 R primers [30, 31]. The reaction mix of 20 μL contained 2.0 μL of each primer (10 μM), 4 μL HPLC water, and 10 μL Maxima SYBRgreen 2 × (cat # K0242, Thermo-Fisher Scientific). The DNA samples were diluted to a concentration of 3 ng μL−1, 2 uL was used per reaction. Amplification was performed as follows: (1) 95 °C for 5 min; (2) 95 °C × 40 s; 55 °C × 120 s, 72 °C × 60 s, repeated 30 times; (3) 72 °C × 7 min. Genomic DNA isolated from Pseudomonas putida KT2440 was used as an external standard in order to calculate 16S rRNA copies per g soil FW extracted assuming a P. putida genome size of 3.174 fg and seven 16S rRNA copies per genome. qPCR efficiency was 90% and could detect as little as 100 P. putida genomes in a single PCR reaction.
Sample pool sequencing libraries were constructed by amplification of the V4 region of the 16S rRNA gene using primers 515F and 806R following the protocol for the Earth Microbiome Project [32]. Briefly, amplicon libraries containing Illumina adaptors and 12-bp Golay barcodes were generated for each sample, cleaned using AmPure beads, quantified with PicoGreen, and pooled in equimolar ratios prior to sequencing at Colorado State University Next-Generation Sequencing (NGS) core facility using a 2 × 250 Miseq flow cell (Illumina, San Diego, CA).
Paired-end sequence reads were concatenated and all combined 16S sequences were filtered, trimmed, and processed with the DADA2 (R bioconductor package) [33]. The DADA2 implementation is included in the open-source bioinformatics tool myPhyloDB version 1.2.1 [34]. All primers were removed from each sequence using the open-source Python program Cutadapt [35] and sequence variants were inferred using the default pipeline in DADA2. Each sequence variant identified in DADA2 was classified to the closest reference sequence contained in the Green Genes reference database (Vers. 13_5_99) using the usearch_global option (minimum identity of 97%) contained in the open-source program VSEARCH [36].
After processing, the 16S library pool data was rarefied to 19,221 sequences reads and one sample (5131 reads from the STA, 133 kg) was removed as result. A rarefaction curve of number of reads and OTUs per sample was generated (Supplementary Fig. 1). Additionally, myPhyloDB version 1.2.1 was used to perform a principal coordinate analysis and differential abundance analysis. The abundance of phosphorus cycle genes, E3.1.3.2, appA, phoA, phoD, phoN, and pqqC (see Table 1 for more detail), in each 16S library was predicted using myPhyloDB’s implementation of PICRUSt [37] and used to calculate the proportion of the microbial community that maps to the gene of interest. Total gene-specific abundance (copies g−1 soil FW) was calculated as the product of the proportion of the community with the specified gene and the total 16S copies (copies g−1 soil FW).
Statistical Analysis
Three-way ANOVA was used to analyze dry weight from cultivated and non-cultivated, root and shoots, and P fertilization rates (Supplementary Table 1). Differences in phosphorus content on above-ground tissues were determined using only one repetition and two-way ANOVA analysis was used to test significance between P rates across the 6 potato genotypes (Supplementary Table 2). To test differences in total biomass between cultivated and non-cultivated, one-way ANOVA was performed. RStudio, Version 1.2.5001 was utilized for all above-mentioned analysis.
The effect of P rate and genotype differences in bacterial community composition (16S rRNA sequences mapped to Green Genes database (Ver. 13_5_99) was visualized by constrained principal coordinates analysis (PCoA) using Bray-Curtis distances, (capscale in R vegan package). PERMANOVA analysis was generated by myPhyloDB for each of the PCoA presented. The Adonis command of the R vegan package was used for this task.
Bacteria phyla with different total abundances (16S copies g−1 soil) at the low (0 kg ha−1 of P) or high (133 kg ha−1) were tested by differential expression analysis based on negative binomial distribution using the edgeR package [38]. Differential expression analysis of total abundances between low (0 kg ha−1 of P) and high (133 kg ha−1 of P) rates was also determined for all OTU (99% genetic distance) that mapped to the selected P-cycle genes. Comparisons of the proportion of sequences per individual predicted phosphatase in response to P level and potato group (cultivated and non-cultivated) were performed by analysis of variance (ANOVA).
Results
Above and Below Potato Plant Biomass Analysis
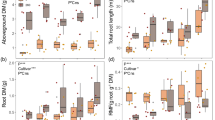
Three-way ANOVA analysis (Supplementary Table 1) showed the effect of P amendments was apparent in all six genotypes. However, cultivated potato genotypes (“Russet Burbank,” “Yukon Gold,” “Red Norland,” and Solanum tuberosum subsp. tuberosum) in the presence of P fertilizer exhibited more vigorous growth compared with non-cultivated potato species (Solanum bulbocastanum and Solanum tuberosum subsp. andigena) (p < 0.001) (Fig. 1A, B). Solanum tuberosum subsp. tuberosum, a direct progenitor of modern potatoes (three commercial cultivars used in this study), showed significantly less biomass compared with the other cultivars. Above-ground dry biomass increased significantly (p < 0.001) with the application of 67 kg and 133 kg ha−1 of P compared with the untreated control (0 kg ha −1) for all genotypes, except for Solanum bulbocastanum (Fig. 1A, B). Belowground dry biomass significantly increased for cultivar “Yukon Gold” and the progenitor Solanum tuberosum subsp. tuberosum when exposed to phosphorus (p < 0.05; Fig. 1B). Cultivated potato genotypes showed a higher biomass in below-ground organs (roots and tubers), contrary to what was observed in non-cultivated species, which concentrated most of their biomass in their above-ground organs (leaves and stems) (p < 0.001). Potato genotypes responded differently in biomass allocation to the same P level addition. For instance, “Russet Burbank” outperformed other cultivated potatoes in above biomass and “Yukon Gold” excelled in below-ground biomass. For non-cultivated potatoes, Solanum tuberosum subsp. andigena accumulated more above-ground biomass than Solanum bulbocastanum and the opposite occurred for below-ground biomass.
Dry biomass of six potato genotypes treated with 0, 67, and 133 kg ha−1 of P. A Above- (leaves and stems) and below (roots and tubers)-ground biomass of four potato accessions. From left to right: “Russet Burbank” (RB), “Red Norland” (RN), “Yukon Gold” (YG), and Solanum tuberosum subsp. tuberosum (STT). B Above- and below-ground biomass of two non-cultivated accessions: Solanum bulbocastanum and Solanum tuberosum subsp. andigena. Three-way ANOVA was used to determine significances between genotypes, P levels, and above- and below-ground biomass
Biomass Differences among Cultivated and Non-cultivated Potatoes
Differences of the total biomass average values (g) between the lowest rate (0 kg ha−1), medium (67 kg ha−1), and the highest rate (133 kg ha−1) of P for cultivated and non-cultivated potatoes were calculated (Supplementary Fig. 2). Solanum tuberosum subsp. tuberosum (STT) is a progenitor used to breed modern cultivars. Consequently, in order to compare biomass gain between cultivated and non-cultivated potato genotypes, STT was removed from further analysis.
The biomass of cultivated potato varieties increased 11% and 22%; whereas for non-cultivated potatoes, biomass increased 47% and 55% from 0 to 67 kg ha−1 of P and from 0 to 133 kg ha−1 of P treatments, respectively. The non-cultivated potato group showed a considerably higher biomass accumulation proportional to P addition compared with the cultivated varieties; however, these differences were not statistically significant.
Phosphorus Content Analysis of Above-Ground Organs
Leaf samples corresponding to each P level (0, 67, 133 kg ha−1) from the 6 genotypes were analyzed (Fig. 2). The two-way ANOVA analysis showed significant differences between P levels (p = 0.002) and the six potato genotypes (p = 0.039) (Supplementary Table 2). Total P concentration gradually increased for cultivated and non-cultivated potato types. For P concentration in above-ground tissue, non-cultivated potatoes had a higher content of P per unit of biomass (mg kg−1) in response to increasing P addition. Further, Solanum bulbocastanum showed twice as much of P concentration values compared with the three potato cultivars, when contrasting the highest P level (133 kg ha−1). In a similar manner, Solanum tuberosum subsp. andigena presented a higher but marginal P content in tissues compared with all three cultivars. The phosphorus content in shoots for cultivated types ranged from 1040 to 6066 (mg kg−1) while the non-cultivated potatoes ranged from 1964 to 8767 (mg kg−1) (Supplementary Table 3). Although the two non-cultivated species showed a different pattern of P uptake, they were superior in P content in above-ground tissues compared with cultivated potatoes.
P concentration (mg kg−1) in dry leaves of six potato accessions. Solanum bulbocastanum (SB), Solanum tuberosum subsp. andigena (STA), Solanum tuberosum subsp. tuberosum (STT), “Yukon Gold” (YG), “Red Norland” (RN), “Russet Burbank” (RB). Each potato accession presents three P treatments (0, 67, 133 kg ha−1). Two-way ANOVA evidenced differences between P levels (p = 0.002) across the 6 genotypes (p = 0.039)
Effect of P Amendment on Potato Rhizosphere Soil Microbial Composition
The effect of P amendment on microbial communities of potato rhizosphere soil was determined by Illumina sequencing analysis. Principal coordinates analysis (PCoA) showed dissimilarities in microbial communities at the potato genotype level, with the 1st two axes explaining 17.5% of variation in the data. A PERMANOVA analysis identified significant variation in the rhizosphere soil microbial communities between genotypes (p < 0.001) and P levels (p = 0.038) and for the interaction between genotype and P level (p = 0.039) (Supplementary Table 4). Furthermore, PCoA ordination of the six potato genotypes and three fertilizer levels showed that all three potato cultivars clustered together while the three non-cultivated potatoes were grouped individually and further apart (Fig. 3). In addition, plant genotype appears to drive microbial community composition to a greater degree than increasing P levels (Fig. 3). Interestingly, when microbial composition of every genotype was analyzed individually, the divergence per P fertilizer level became visually more apparent. However, “Red Norland” was the only genotype showing a significant difference among P levels (p = 0.029) by a PERMANOVA analysis (Supplementary Figs. 3, 4).
Principal coordinate analysis (PCoA) using Bray-Curtis distances, depicting 16S rRNA data from rhizosphere soil samples, amended with three levels of P in six potato genotypes. Potato genotype drives the separation of rhizosphere microbial communities. Confidence ellipses are shown around each potato accession: SB: Solanum bulbocastanum (green), STA: Solanum tuberosum subsp. andigena (purple), STT: Solanum tuberosum subsp. tuberosum (orange), “Russet Burbank” (blue), “Red Norland” (red), “Yukon Gold” (yellow). P levels are 0 kg ha−1 (circle), 67 kg ha−1 (square), and 133 kg ha−1 (Rhomb)
Differential Abundance Analysis of Bacteria for Modern Cultivars and Non-cultivated Potatoes
The differential abundance analysis showed a distinctive bacteria taxon in each group that varied the most in relative abundance from the lowest to the highest P treatment (0 to 133 kg ha −1). A phylum and family-level analysis of the microbiota in the two different potato groups indicated a higher degree of differential abundances (between the low and the high P treatments) for taxa in the cultivated potato over the non-cultivated group. The Rhodospirillaceae, Micromonosporaceae, Nocardioidaceae, Streptomycetacea, and Actinosynemataceae were significantly more abundant (FDR < 0.1; p < 0.01) in the cultivated potatoes. The family taxa Ellin5301, Cystobacteraceae, and unclassified families were less abundant among cultivated varieties (FDR < 0.1; p < 0.01). Within the cultivated group and when comparing low (0 kg ha −1) and high (133 kg ha −1) P levels, one unclassified family from the phylum Gemmatimonadetes was found to decrease in abundance; however, the remaining 10 phyla increased in relative abundance under P addition. In contrast, fewer families were significantly different for the non-cultivated group. The taxa Flavobacteriaceae, Cyclobacteriaceae, Enterobacteriaceae, Aeromonadaceae, and Alteromonadaceae were all present in the non-cultivated potato. Alteromonadaceae was the only taxa that decreased in abundance from the non-cultivated potato group (Fig. 4).
Log fold changes (LogFC) of microbial abundances means from high (133 kg ha−1) and low (0 kg ha−1) phosphorus amendment. A Rhizosphere soil samples from cultivated potatoes. B Rhizosphere samples from non-cultivated potato plants. LogFC was calculated by subtracting the baseMean counts of log ratios of each microbial taxa (at the family level; FDR < 0.1) present at low P (0 kg ha−1) from microbial taxa present at the highest P level. Analysis was performed using the edgeR package, within the bioinformatic software myPhyloDB
Effect of P Rate on Predicted Acid and Alkaline Phosphatase Genes by PICRUSt
In these analyses, we assessed the abundance of predicted phosphatases across the whole bacterial community. Six different phosphatase genes were mapped in the two groups of potatoes of interest (cultivated or non-cultivated) and under two P levels (low or high). Overall, high P amendments increased the abundance of all predicted phosphatases compared with low P rates, for both potato groups (Fig. 5); this increase in abundance was not statistically significant for any of the six phosphatases. When comparing abundances of predicted phosphatases between non-cultivated and cultivated potatoes, irrespective of the P level, the enzymes in the former group were found to be more abundant than in the cultivated potatoes. Four out of the six predicted enzymes were more abundant in the non-cultivated group, including three acid phosphatases (E3.1.3.2, appA, phoN) and one alkaline phosphatase (phoA). Significant differences between cultivated and non-cultivated were found only for appA (p = 0.0369). The remaining two phosphatases (phoD and pqqC) were similarly abundant when comparing cultivated and non-cultivated potato groups. Two-way ANOVA was also performed to evaluate interaction between P level and potato group, but no significance was found (data not shown).
Comparisons of the proportion of sequences per each predicted phosphatase (acid and alkaline), between low and high P treatments (0 and 133 kg ha −1) and between cultivated and non-cultivated potatoes. ANOVA and Tukey HSD test were performed per each phosphatase (low and high, and cultivated and non-cultivated). Letters denote statistical significance. Acid phosphatases: E3.1.3.2 (A), appA (B) (p = 0.0369), and phoN (C). Alkaline phosphatases: phoD (D) and phoA (E). Pyrroloquinoline-quinone synthase: pqqC (F). The proportion of sequences from predicted phosphatases was generated by PICRUSt, using myPhyloDB software
Effect of P Rate on Presence or Absence of Predicted Phosphatase Genes for the Subset of Taxa that Were Differentially Abundant
A subsequent analysis was conducted to postulate the presence or absence of the predicted phosphatase genes that mapped only to those families identified on the differential abundance analysis for each potato group (Supplementary Fig. 5). To be marked as present, the P-cycle gene (per each of the six genes) had to appear in at least one member within a given bacterium family, regardless of the number of members within each family. Contrary, to be considered as absent, each gene had to be not present in each member within a given bacterium family. The relationship between the relative or total abundances of P-cycle genes and bacteria taxa was predicted with the aid of PICRUSt. Potato cultivars had the highest frequency of P-related genes present compared with non-cultivated potatoes for this subset of families. Five out of six different P-cycle genes were present in this group compared with four out of six different genes in non-cultivated potatoes. For both groups, alkaline phosphatases genes were predicted to be more frequent.
Discussion
Non-cultivated Potatoes Assimilate P more Efficiently than Modern Cultivars
Differences in biomass (from 0 to 133 kg ha −1 of P) between the modern cultivars (tuber-bearing potatoes: “Russet Burbank,” “Yukon Gold,” and “Red Norland”) and the wild species and landrace (non-tuber-bearing potatoes: Solanum bulbocastanum and Solanum tuberosum subsp. andigena) showed that the latter group outperformed the former by fourfold, and that the P content in above-ground tissues was twice as much for the non-cultivated group. This may suggest a higher efficiency of certain potato types to uptake P from soils. It is worth noting that there are two conflicting plant traits in our dataset (biomass versus P uptake). From the perspective of decreasing P legacy in soils, which translates into less runoff and eutrophication, it may be that a more efficient uptake of P is more favorable regardless of how it impacts plant growth [11]. In contrast, for the purpose of yield increases, belowground biomass traits (tubers in particular) are more desired even if it does not result in the maximum possible P uptake by the plant [39]. Current plant breeding favors increasing below plant biomass at the expense of increasing total P uptake. However, we suggest a common ground between both traits to enhance potato cultivar selection and resources efficiency traits.
Rhizosphere Microbial Communities Differ Among Modern Cultivars and Non-cultivated Potato Genotypes and Between P Level
At a broader level, our results indicated that the root-associated bacterial community was strongly influenced by the host more so than by the effect of fertilizer addition. This finding supports previous research showing that plants can host their own distinct root microbiota not only at a plant species level, but also at a cultivar level [26, 40,41,42]. Most of the differences in this study were seen at the plant species level. All cultivars were similar in their microbial community composition based on Principal coordinate analysis and differed drastically from non-cultivated potato species. Additionally, a shift in bacteria community composition with the addition of P was observed when individual potato genotypes were analyzed; however, it was only statistically significant for Red Norland cultivar. Recently, Bodenhausen et al. [41] demonstrated a compositional difference in microbial communities of Petunia sp. (Solanaceae) after P supplementation. Similar response has been reported for blueberry, a recently domesticated plant [21]. In both studies, it was observed that incremental P amendments selected certain taxa causing divergence of the whole community in the rhizosphere soil. In a similar manner, [43] Kaminsky et al. showed that soil nutrient amendment can significantly alter the structure and activity of the soil microbiome of Medicago sativa impacting the ability of the microbiome to support crop health.
These observed differences in microbial community composition between potato groups and the differences in biomass gain and P content favoring non-cultivated potatoes might hint at a relationship between plant fitness and specific rhizosphere microbial communities. Namely, this plant response could be linked to distinctive microbial communities inhabiting the rhizosphere of one potato group but absent in the other. An existing hypothesis proposed a decrease in the ability to promote microbial symbiosis in modern cultivars in response to highly fertile conditions used during the plant breeding process [18]. A number of studies on fungi and bacteria among a variety of crops such as maize, wheat, pea, and soybean showed evidence for this hypothesis [44,45,46,47]. In contrast to this hypothesis, our differential abundance analysis exhibited a higher degree of differential abundance and larger number of bacteria taxa at family-level present in modern cultivars compared with non-cultivated potatoes under the same cut off (FDR < 0.1; p < 0.001). Additionally, the Shannon diversity index showed an increase in diversity for cultivated potatoes and the opposite for non-cultivated after P amendment for the overall community (Supplementary Table 6). This could imply a potential lack of plant control in cultivated plants over its associated microbiome leading to the presence of microbes that are more directly linked to soil nutrient levels or any other physiochemical condition in the rhizosphere determined by soil type, root activity, or climate [48]. We acknowledge that the larger number of plants used for the cultivated potato group (3 cultivars) compared with the non-cultivated (2 plant species) could have influenced, at least partially, the power of the observed results favoring the number of the observed taxa in the differential abundance analysis of cultivated potato group. However, the degree of response of the differentially abundance per each taxon for both potato groups should not be affected by the sample size, but by the P fertilization rate (low versus high P levels).
Differential Microbial Abundance and Predicted P-Cycle Genes Suggest a Subset of Opportunistic Bacteria Competing with the Plant for P in Modern Potato Genotypes
Bacterial taxa (differentially abundant under the highest P rate) present in cultivated and non-cultivated potatoes were estimated. Most of the bacteria displayed in the differential abundance analysis for the cultivated potato group belong to Proteobacteria and Actinobacteria phyla. Actinobacteria have been shown to be linked to soil P-cycling and a recent study demonstrated that P fertilization modified microbial community structure to a more copiotroph bacterial community [49, 50]. Moreover, a different study observed that nutrient-rich environments are preferred by bacteria with fast-growing rates as Proteobacteria and Actinobacteria [51, 52]. The higher frequency of phosphatases and the presence of both Actinobacteria and Proteobacteria on the cultivated potatoes may indicate that the occurrence of fast-growing bacteria tends to have increased presence of phosphatase genes when exposed to high P amendments. Based on these observations, we hypothesized two possible scenarios.
First, a case in which bacteria expressing phosphatase activity might be direct competitors with the plant for P fertilizer. Although if there is an excess of P, and P tends to build up, this would suggest that bacteria are not up taking as much P away to impact plant growth. A second and more likely scenario is that fast-growing bacteria are mineralizing P into organic forms making it unavailable for cultivated potato varieties over non-cultivated. Liu et al. [53] showed that inorganic fertilizer amendment increased mineralization of nitrogen. [54] Bardgett et al. reported that soil microbes are capable of competing effectively with plants for organic-nitrogen in temperate grasslands, and recently Fan et al. [55] revealed that long-term fertilization selects against microbial N fixers favoring other microbial groups that support their own growth. Nevertheless, our predicted phosphatases analysis needs to be further tested as potential mechanism of plant-microbial competition for P.
Proportion of Sequences of Predicted Phosphatases for Cultivated and Non-cultivated Potatoes
In order to understand a potential link between phosphatase abundance, plant biomass, and P content, we assessed the abundance of predicted phosphatases across the whole bacterial community (Fig. 5). Overall, most phosphatases increased in abundance under high P amendment whether they were cultivated or non-cultivated. Kaminsky et al. [43] observed that enzyme activity for acid and alkaline phosphatases was the highest (when compared with lower P rate and organic P treatments) for microbiomes conditioned to high P treatment during several generations which agrees with our results.
We expected to see a higher abundance of phosphatase enzymes present in the non-cultivated group, which could help to explain the higher P content and biomass gain in this potato group. This was only true for acid phosphatase appA in non-cultivated plants compared with cultivated groups. When looking at the bacterial community as a whole, most enzymes showed a higher proportion of predicted sequences in non-cultivated groups, but these were not statistically different. It remains speculative to conclude if the increase of predicted phosphatases is related to the ability of the non-cultivated potatoes to uptake P efficiently. Another possible hypothesis is that P fertilizer addition might stimulate distinctive root exudation profiles that fuels microbial growth and enzyme production differentially for cultivated and non-cultivated potatoes [56].
Bacteria with Phosphatase Activity as a Potential Indirect Competitor with the Plant for P Fertilizer
This analysis was conducted to determine the presence or absence of predicted phosphatase genes that mapped to families identified on the differential abundance analysis for cultivated and non-cultivated potatoes. Such taxa drastically changed its abundance in response to P amendment. For this reason, we further investigated the predicted phosphatase genes related to this subset of microbes. It is well established that the addition of inorganic phosphate inhibits microbial phosphatase enzyme activity and synthesis, leading to decreasing rates of mineralization of organic phosphate compounds [51]. In our study, we observed a higher incidence of predicted phosphatases for cultivated potato plants under P addition in a portion of the microbial community (Supplementary Fig. 5). This led us to hypothesize that the excess of P fertilization selects opportunistic microbial members that may preferentially release phosphatases to mineralize the extra P available in the rhizosphere of cultivated potatoes, and as consequence, reducing plant growth. In our study, the extra P fertilizer added to the soil was in inorganic form (i.e., orthophosphate) readily available for the plant. Recently, inorganic phosphate-solubilizing genes have been reported [57]; but studies linking inorganic P addition and expression of those genes have not been conducted. Despite this situation, upon P fertilization and based on the chemical conditions of the soil (pH, organic matter content, CEC), a considerable part of the newly applied P can become immobile by soil particles in a process known as adsorption and/or converted to organic forms such as phytic acid by soil microbes [58, 59]. This soil P conversion can prevent the cultivated potato plants from accessing P from the soil. In support of this, it has been shown that phosphate-solubilizing and phosphate-mineralizing abilities can co-exist in the same bacterial strain, which makes more likely the hypothesis that microorganisms expressing phosphatase activity are also mineralizing P [60]. We suggest that opportunistic microbial members use their enzymatic capabilities not only under P scarcity but also under P excess in order to convert inorganic P into organic forms.
Conclusion
These results provide an initial insight into the impact of mineral P on the rhizosphere microbiome of potato. This study suggests that excessive P amendment results in microbial conversion of P to forms that are not available to the plant translating this into less biomass gain and P uptake on modern potato cultivars. However, the confirmation of these hypotheses warrants experimental testing.
References
Chaparro JM, Sheflin AM, Manter DK, Vivanco JM (2012) Manipulating the soil microbiome to increase soil health and plant fertility. Biol Fertil Soils 48(5):489–499
Van Der Heijden MG, Scheublin TR (2007) Functional traits in mycorrhizal ecology: their use for predicting the impact of arbuscular mycorrhizal fungal communities on plant growth and ecosystem functioning. New Phytol 174(2):244–250
Junaidi J, Kallenbach CM, Byrne PF, Fonte SJ (2018) Root traits and root biomass allocation impact how wheat genotypes respond to organic amendments and earthworms. PLoS One 13(7):e0200646
Sessitsch A, Gyamfi S, Tscherko D, Gerzabek MH, Kandeler E (2005) Activity of microorganisms in the rhizosphere of herbicide treated and untreated transgenic glufosinate-tolerant and wildtype oilseed rape grown in containment. Plant Soil 266(1–2):105–116
Garbeva P, Van Elsas JD, Van Veen JA (2008) Rhizosphere microbial community and its response to plant species and soil history. Plant Soil 302(1–2):19–32
Lo CC (2010) Effect of pesticides on soil microbial community. J Environ Sci Health B 45(5):348–359
Syers, J. K., Johnston, A. E., & Curtin, D. (2008). Efficiency of soil and fertilizer phosphorus use. FAO Fertilizer and Plant Nutrition Bulletin, 18(108)
Havlin JL, Beaton JD, Tisdale SL, Nelson WL (2005) Micronutrients. Soil Fertility and Fertilizers. Pearson Education Inc., and Dorling Kindersley Publishing Inc. India
Wolf J, De Wit CT, Janssen BH, Lathwell DJ (1987) Modeling long-term crop response to fertilizer phosphorus. I. the model 1. Agron J 79(3):445–451
Tilman D, Fargione J, Wolff B, D'antonio C, Dobson A, Howarth R et al (2001) Forecasting agriculturally driven global environmental change. Science 292(5515):281–284
Zhu J, Li M, Whelan M (2018) Phosphorus activators contribute to legacy phosphorus availability in agricultural soils: a review. Sci Total Environ 612:522–537
Sattari SZ, Bouwman AF, Giller KE, van Ittersum MK (2012) Residual soil phosphorus as the missing piece in the global phosphorus crisis puzzle. Proc Natl Acad Sci 109(16):6348–6353
Wang Q, Ma M, Jiang X, Guan D, Wei D, Zhao B et al (2019) Impact of 36 years of nitrogen fertilization on microbial community composition and soil carbon cycling-related enzyme activities in rhizospheres and bulk soils in northeast China. Appl Soil Ecol 136:148–157
Castrillo G, Teixeira PJPL, Paredes SH, Law TF, de Lorenzo L, Feltcher ME et al (2017) Root microbiota drive direct integration of phosphate stress and immunity. Nature 543(7646):513–518
Hardigan MA, Laimbeer FPE, Newton L, Crisovan E, Hamilton JP, Vaillancourt B et al (2017) Genome diversity of tuber-bearing Solanum uncovers complex evolutionary history and targets of domestication in the cultivated potato. Proc Natl Acad Sci 114(46):E9999–E10008
Hawkes JG (1990) The potato: evolution, biodiversity and genetic resources. Belhaven Press, London
Friedman M (2006) Potato glycoalkaloids and metabolites: roles in the plant and in the diet. J Agric Food Chem 54(23):8655–8681
Wissuwa M, Mazzola M, Picard C (2009) Novel approaches in plant breeding for rhizosphere-related traits. Plant Soil 321(1–2):409
Perez-Jaramillo JE, Mendes R, Raaijmakers JM (2016) Impact of plant domestication on rhizosphere microbiome assembly and functions. Plant Mol Biol 90(6):635–644
Bhattacharyya PN, Jha DK (2012) Plant growth-promoting rhizobacteria (PGPR): emergence in agriculture. World J Microbiol Biotechnol 28(4):1327–1350
Pantigoso HA, Manter DK, Vivanco JM (2018) Phosphorus addition shifts the microbial community in the rhizosphere of blueberry (Vaccinium corymbosum L.). Rhizosphere 7:1–7
Williams A, Manoharan L, Rosenstock NP, Olsson PA, Hedlund K (2017) Long-term agricultural fertilization alters arbuscular mycorrhizal fungal community composition and barley (Hordeum vulgare) mycorrhizal carbon and phosphorus exchange. The New Phytologist 213(2):874–885
Engelhard M, Hurek T, Reinhold-Hurek B (2000) Preferential occurrence of diazotrophic endophytes, Azoarcus spp., in wild rice species and land races of Oryza sativa in comparison with modern races. Environ Microbiol 2(2):131–141
Van Deynze A, Zamora P, Delaux PM, Heitmann C, Jayaraman D, Rajasekar S et al (2018) Nitrogen fixation in a landrace of maize is supported by a mucilage-associated diazotrophic microbiota. PLoS Biol 16(8):e2006352
İnceoğlu Ö, Al-Soud WA, Salles JF, Semenov AV, van Elsas JD (2011) Comparative analysis of bacterial communities in a potato field as determined by pyrosequencing. PLoS One 6(8):e23321
Weinert N, Piceno Y, Ding GC, Meincke R, Heuer H, Berg G, Schloter M, Andersen G, Smalla K (2011) PhyloChip hybridization uncovered an enormous bacterial diversity in the rhizosphere of different potato cultivars: many common and few cultivar-dependent taxa. FEMS Microbiol Ecol 75(3):497–506
Pérez-Jaramillo JE, Carrión VJ, Bosse M, Ferrão LF, de Hollander M, Garcia AA et al (2017) Linking rhizosphere microbiome composition of wild and domesticated Phaseolus vulgaris to genotypic and root phenotypic traits. ISME J 11(10):2244–2257
Essah SYC, Davis JG (1996) Fertilizing potatoes in Colorado, 0.541. Colorado State University. https://extension.colostate.edu/docs/pubs/crops/00541.pdf. Accessed 26 August 2019
Lynch JM (1984) The rhizosphere form and function. Appl Soil Ecol 1193
Lane DJ, Pace B, Olsen GJ, Stahl DA, Sogin ML, Pace NR (1985) Rapid determination of 16S ribosomal RNA sequences for phylogenetic analyses. Proc Natl Acad Sci 82(20):6955–6959
Marchesi JR, Sato T, Weightman AJ, Martin TA, Fry JC, Hiom SJ, Wade WG (1998) Design and evaluation of useful bacterium-specific PCR primers that amplify genes coding for bacterial 16S rRNA. Appl Environ Microbiol 64(2):795–799
Gilbert JA, Jansson JK, Knight R (2014) The earth microbiome project: successes and aspirations. BMC Biol 12(1):69
Callahan BJ, McMurdie PJ, Rosen MJ, Han AW, Johnson AJA, Holmes SP (2016) DADA2: high-resolution sample inference from Illumina amplicon data. Nat Methods 13(7):581
Manter DK, Korsa M, Tebbe C, Delgado JA (2016) myPhyloDB: a local web server for the storage and analysis of metagenomic data. Database 2016
Martin M (2011) Cutadapt removes adapter sequences from high-throughput sequencing reads. EMBnet J 17(1):10–12
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584
Langille MG, Zaneveld J, Caporaso JG, McDonald D, Knights D, Reyes JA et al (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31(9):814
Robinson MD, McCarthy DJ, Smyth GK (2010) edgeR: a bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26(1):139–140
Rosen CJ, Kelling KA, Stark JC, Porter GA (2014) Optimizing phosphorus fertilizer management in potato production. Am J Potato Res 91(2):145–160
Schweitzer JA, Bailey JK, Fischer DG, LeRoy CJ, Lonsdorf EV, Whitham TG, Hart SC (2008) Plant–soil–microorganism interactions: heritable relationship between plant genotype and associated soil microorganisms. Ecology 89(3):773–781
Bodenhausen N, Somerville V, Desirò A, Walser JC, Borghi L, van der Heijden MG, Schlaeppi K (2019) Petunia-and Arabidopsis-specific root microbiota responses to phosphate supplementation. Phytobiomes J, PBIOMES-12
Peiffer JA, Spor A, Koren O, Jin Z, Tringe SG, Dangl JL et al (2013) Diversity and heritability of the maize rhizosphere microbiome under field conditions. Proc Natl Acad Sci 110(16):6548–6553
Kaminsky LM, Thompson GL, Trexler RV, Bell TH, Kao-Kniffin J (2018) Medicago sativa has reduced biomass and nodulation when grown with soil microbiomes conditioned to high phosphorus inputs. Phytobiomes (ja)
Hetrick BAD, Wilson GWT, Cox TS (1993) Mycorrhizal dependence of modern wheat cultivars and ancestors: a synthesis. Can J Bot 71(3):512–518
Mutch LA, Young JPW (2004) Diversity and specificity of Rhizobium leguminosarum biovar viciae on wild and cultivated legumes. Mol Ecol 13(8):2435–2444
Kiers ET, Denison RF (2008) Sanctions, cooperation, and the stability of plant-rhizosphere mutualisms. Annu Rev Ecol Evol Syst 39:215–236
Sangabriel-Conde W, Negrete-Yankelevich S, Maldonado-Mendoza IE, Trejo-Aguilar D (2014) Native maize landraces from Los Tuxtlas, Mexico show varying mycorrhizal dependency for P uptake. Biol Fertil Soils 50(2):405–414
Emmett BD, Buckley DH, Smith ME, Drinkwater LE (2018) Eighty years of maize breeding alters plant nitrogen acquisition but not rhizosphere bacterial community composition. Plant Soil. https://doi.org/10.1007/s11104-018-3744-0
Fanin N, Hättenschwiler S, Schimann H, Fromin N (2015) Interactive effects of C, N and P fertilization on soil microbial community structure and function in an Amazonian rain forest. Funct Ecol 29(1):140–150
Trabelsi D, Cherni A, Zineb AB, Dhane SF, Mhamdi R (2017) Fertilization of Phaseolus vulgaris with the Tunisian rock phosphate affects richness and structure of rhizosphere bacterial communities. Appl Soil Ecol 114:1–8
Spiers GA, McGill WB (1979) Effects of phosphorus addition and energy supply on acid phosphatase production and activity in soils. Soil Biol Biochem 11(1):3–8
Fierer N, Lauber CL, Ramirez KS, Zaneveld J, Bradford MA, Knight R (2012) Comparative metagenomic, phylogenetic and physiological analyses of soil microbial communities across nitrogen gradients. ISME J 6(5):1007
Liu XJA, van Groenigen KJ, Dijkstra P, Hungate BA (2017) Increased plant uptake of native soil nitrogen following fertilizer addition–not a priming effect? Appl Soil Ecol 114:105–110
Bardgett RD, Streeter TC, Bol R (2003) Soil microbes compete effectively with plants for organic-nitrogen inputs to temperate grasslands. Ecology 84(5):1277–1287
Fan K, Delgado-Baquerizo M, Guo X, Wang D, Wu Y, Zhu M, Yu W, Yao H, Zhu YG, Chu H (2019) Suppressed N fixation and diazotrophs after four decades of fertilization. Microbiome 7:143. https://doi.org/10.1186/s40168-019-0757-8
Preece C, Peñuelas J (2019) A return to the wild: root exudates and food security. Trends Plant Sci
Liu J, Cade-Menun BJ, Yang J, Hu Y, Liu CW, Tremblay J et al (2018) Long-term land use affects phosphorus speciation and the composition of phosphorus cycling genes in agricultural soils. Front Microbiol 9:1643
Schachtman DP, Reid RJ, Ayling SM (1998) Phosphorus uptake by plants: from soil to cell. Plant Physiol 116(2):447–453
Havlin JL, Beaton JD, Tisdale SL, Nelson WL (1999) Soil fertility and fertilizers6th edn. Prentice Hall, Upper Saddle River
Guang-Can TAO, Shu-Jun TIAN, Miao-Ying CAI, Guang-Hui XIE (2008) Phosphate-solubilizing and-mineralizing abilities of bacteria isolated from soils. Pedosphere 18(4):515–523
Acknowledgements
Special thanks to Dr. Steve Fonte and Dr. Ioannis Minas for reviewing the manuscript and for their valuable input.
Funding
This research was financially supported by the Colorado State University Agricultural Experimental Station.
Author information
Authors and Affiliations
Corresponding authors
Electronic supplementary material
ESM 1
(DOCX 315 kb)
Rights and permissions
About this article
Cite this article
Pantigoso, H.A., Manter, D.K. & Vivanco, J.M. Differential Effects of Phosphorus Fertilization on Plant Uptake and Rhizosphere Microbiome of Cultivated and Non-cultivated Potatoes. Microb Ecol 80, 169–180 (2020). https://doi.org/10.1007/s00248-020-01486-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-020-01486-w