Abstract
The death of trees is an ecological process that promotes regeneration, organic matter recycling, and the structure of communities. However, diverse biotic and abiotic factors can disturb this process. Dendroctonus bark beetles (Curculionidae: Scolytinae) are natural inhabitants of pine forests, some of which produce periodic outbreaks, killing thousands of trees in the process. These insects spend almost their entire life cycle under tree bark, where they reproduce and feed on phloem. Tunneling and feeding of the beetles result in the death of the tree and an alteration of the resident microbiota as well as the introduction of microbes that the beetles vector. To understand how microbial communities in subcortical tissues of pines change after they are colonized by the bark beetle Dendroctonus rhizophagus, we compare both the bacterial and fungal community structures in two colonization stages of Pinus arizonica (Arizona pine) employing Illumina MiSeq. Our findings showed significant differences in diversity and the dominance of bacterial community in the two colonization stages with Shannon (P = 0.004) and Simpson (P = 0.0006) indices, respectively, but not in species richness with Chao1 (P = 0.19). In contrast, fungal communities in both stages showed significant differences in species richness with Chao1 (P = 0.0003) and a diversity with Shannon index (P = 0.038), but not in the dominance with the Simpson index (P = 0.12). The β-diversity also showed significant changes in the structure of bacterial and fungal communities along the colonization stages, maintaining the dominant members in both cases. Our results suggest that microbial communities present in the Arizona pine at the tree early colonization stage by bark beetle change predictably over time.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The death of trees is a natural and complex ecological process that promotes the regeneration, organic matter recycling, and the community structure within forests [1]. Several biotic (e.g., insects, diseases) and abiotic (e.g., changes in temperature, levels of precipitation, fire) factors can lead to disturbances and the loss of biodiversity [1,2,3]. Dendroctonus bark beetles (Curculionidae: Scolytinae) are natural inhabitants of coniferous forests (Pinaceae) that play an important role in the process of forest recycling and regeneration [4].
The life cycle consists of adults emerging from host trees and dispersing in the forest to locate and colonize new trees. Females attract mates, and together, they build galleries where females oviposit, and then larvae develop, feeding on phloem tissue [5]. Some species use secondary chemicals as cues for host-recognition, or for synergizing aggregation pheromones employed by bark beetles to carry out mass attacks and overcoming the host tree’s defenses [6]. Populations of some species undergo periodic outbreaks during which they can kill thousands of healthy trees, causing significant disturbance in woodlands and economical losses to the forestry industry [5, 7].
Significant changes in the tree physiology occur as result of host-colonization by Dendroctonus bark beetles. Firstly, boring and feeding of bark beetles induce resinosis, and the allocation of terpenes, phenolic compounds, carbohydrates, and minerals, as well as the formation of traumatic resin ducts. In addition, severe damage occurs in vascular tissues due to the introduction of microbes that penetrate phloem, as well as cavitation caused by a water column breakage of the plant. This promotes foliar moisture declination, as photosynthesis ceases, along with needles desiccation [8,9,10,11,12].
From colonization to death, many microorganisms (bacteria, yeasts, archaea, and filamentous fungi) coexist in the subcortical habitat. These microbes come from several sources, some are endophytes or epiphytes [13,14,15], while others are vectored into the tree by bark beetle in their guts [16,17,18,19]; on their exoskeletons [17]; or, for some, in specialized structures (e.g., mycangium) [20, 21], as well as of other contamination sources, such as the frass of other insects or environmental contamination [22, 23].
Despite several studies characterizing the microbial communities in healthy tissues of coniferous trees [24,25,26,27,28], it is to our knowledge that there have been no approaches done that evaluate how these communities present in subcortical tissues change after colonization by insects. Given that bark beetles and the associated microbes are a strong selective pressure that triggers severe physiological changes in trees [29], we hypothesize that microbial communities change as they die. Arizona pine is a preferential host of Dendroctonus rhizophagus in the Sierra Madre Occidental in Mexico, which is an aggressive species that does not produce massive attacks on mature pine trees, but a pair or two—in some cases—colonizes and kills saplings < 10 cm diameter and < 3 m high from 11 pine species, including the Arizona pine [30]. Because D. rhizophagus is a species with a univoltine (i.e., from egg to adult is completed within a year) and synchronous (i.e., the same developmental stage occurs at the same time throughout its distribution range) life cycle, we characterized and compared the changes in the microbial community (bacterial, archaeal, and fungal) present in the subcortical tissue of Arizona pine saplings in two well-defined colonization stages: when paired female and male are present into tree prior to oviposition (early stage) and when their offspring is found in pupa stage (late stage).
Materials and Methods
Site Location, Collection, and Processing of Samples
The bacterial, archaeal, and fungal communities were recovered from subcortical tissues of Arizona pines saplings (approx. 10 years) attacked by D. rhizophagus in San Juanito, Bocoyna Municipality, Chihuahua State (27° 45′ 11″ N 107° 38′ 06″ W, 2288 masl). The phloem was collected near to insect gallery just prior to oviposition and while the color of the sapling foliage was green (early colonization stage; early August) (Supplementary Fig. 1A-C). It was also collected near to pupal chamber when broods of the couple reached the pupa stage, other arthropods were present into trees, foliage ranged from yellow to red, and trees were nearly completely dead (late colonization stage; early April) (Supplementary Fig. 1D-G). The phloem was carefully scraped using sterile fine forceps and scalpels, placed in sterile 50-ml tubes, and stored at 4 °C for transport to the laboratory, where they were immediately processed. Three replicates were prepared for each stage, and three pines for each replicate were selected and pooled.
The tissues of each replicate were rinsed for 30 s with 70% ethanol. A volume of 30 ml of phosphate-buffered saline (PBS) (137 mM NaCl, 2.7 mM KCl, 10 mM Na2HPO4, 2 mM KH2PO4, pH 7.4) and 0.1% Triton X-100 [17] was added to tubes. The samples were shaken overnight (180 rpm), and then the pooled microbial suspensions were centrifuged for three mins at 8000×g in sterile 2-ml screw-cap tubes to concentrate the biomass.
Metagenomic DNA Extraction, Library Preparation, and Sequencing
Metagenomic DNA was extracted using the method described by Gonzalez-Escobedo et al. [28]. DNA was quantified using a NanoDrop 2000c Spectrophotometer (Thermo Scientific, Wilmington, DE) and was observed in an agarose gel 1.0%. The extracted DNA samples were amplified for the V3-V4 region of 16S rRNA gene using the primer pairs Bakt_341F (5′-CCTACGGGNGGCWGCAG-3′) and Bakt_805R (5′-GACTACHVGGGTATCTAATCC-3′) for characterizing the bacterial community [31], and the internal transcriber spacer (ITS) region, using ITS1 (5′-GCATCGATGAAGAACGCAGC-3′) and ITS4 (5′-TCCTCCGCTTATTGATATGC-3′) primer pairs for characterizing the fungal community [32]. The 16S rRNA and ITS amplicons were sequenced using paired-end 2 × 300 bp on an Illumina MiSeq sequencer (Illumina Inc., San Diego, CA, USA) at Macrogen Inc. (Seoul, South Korea).
Bioinformatic Analysis
Illumina-paired reads were merged using FLASH v 1.2.11 [33] with default settings. Sequence files were then converted from fastq to fasta. Merged reads were processed with VSEARCH v 2.4.3 [34] and Quantitative Insights Into Microbial Ecology (QIIME) v 1.8 [35]. Bacterial/archaeal and fungal reads were each pooled, de-replicated, and checked for chimeras. Chimeras, singletons, and doubletons were discarded, and then, reads were clustered into operational taxonomic units (OTUs) at 97% similarity and an OTU table was constructed for each. Taxonomy was assigned to each OTU using the RDP naïve Bayesian classifier (bacterial/archaeal) [36] and UNITE (fungal) [37] databases and corroborated manually against the EzBioCloud (bacterial/archaeal) [38] and GenBank (bacterial/archaeal/fungal) [39]. Sequences that matched chloroplasts were manually removed, and then, for each OTU table (bacteria, archaea, and fungi), a heat map at the genus level of samples of colonization stages was constructed, with the information of the relative abundance presented as a percentage. In addition, the bacterial and fungal OTUs and assigned genera distributions across two stages were presented in Venn diagrams that were generated with the web application Venny [40]. Unfortunately, due to the extremely low number of OTUs for the archaeal community, diversity analyses were not carried out for it.
Bacterial and fungal communities were homogenized with respect to the sample with the lowest reads. To calculate α-diversity within these communities in both stages, we estimated species richness, using the observed OTU number and Chao1 [41], species diversity with Shannon [42], and the dominance with Simpson index [43] in QIIME. The diversity indices of early and late colonization stages were compared, using the Mann-Whitney U test to evaluate statistical significance between the samples (P < 0.05). To calculate the sequence coverage obtained for the 16S rRNA and ITS region datasets, Good’s coverage estimator was used [44], along with rarefaction curves that were generated with QIIME to compare relative levels of bacterial and fungal OTU diversity across all samples.
To measure the degree of differentiation of the bacterial and fungal communities in the two colonization stages, we estimated β-diversity using the Bray-Curtis dissimilarity index. We performed, using the Bray-Curtis matrix, a hierarchical cluster analysis with the unweighted paired group method with arithmetic mean (UPGMA) and a principal coordinate analysis (PCoA) to assess the dissimilarities or distances among communities during the ordination solution. The dendrogram and plots corresponding were visualized in NTSYSpc v 2.02 [45]. Significant differences in the community composition of bacterial and fungal of the two colonization stages were tested by mean of a permutational multivariate analysis of variance (PERMANOVA), after 999 random permutations in PAST v 3.11 [46].
Sequence Accession Numbers
The Illumina sequence data reported here were deposited in the NCBI Sequence Read Archive (http://www.ncbi.nlm.nih.gov/sra) under accession number SRP149017.
Results
Sequencing Results
A total of 1,111,959 (bacteria), 1716 (archaea), and 1,122,557 (fungi) high-quality reads were obtained from all samples at the two colonization stages. The OTU97% number represented in these reads was 682 for bacteria, 9 for archaea, and 634 for fungi (Table 1). Of these, 92%, 100%, and 72% were assigned to the taxonomic category of genus of these microbial groups, respectively; the remaining OTUs97% (8% bacteria and 28% fungi) were classified into taxonomic categories above the genus level (from family to phylum).
Microbial Community Composition
A total of 20 distinct phyla, 40 orders, 54 families, and 60 bacterial genera were identified from total OTUs97% in all samples of two stages. The phylum Proteobacteria comprised 93.80% of the relative abundance of all OTUs97%, while 19 phyla represented the remaining 6.20% (Supplementary Fig. 2A). At the family level, Enterobacteriaceae was the most abundant (69.10%) taxonomic group, followed by Pseudomonadaceae (12.29%), Xanthomonadaceae (6.13%), Acetobacteraceae (3.25%), Sphingomonadaceae (2.60%), and Acidobacteriaceae (1.18%), and 48 families with abundances comprising < 1.00% of total reads each (Supplementary Fig. 3A).
At the genus level, Rahnella was the dominant taxa (63.23%) in all the samples tested, followed by Pseudomonas (13.05%), Pseudoxanthomonas (5.90%), Enterobacter (4.24%), Kosakonia (1.92%), Sphingomonas (1.69%), Pantoea (1.22%), Burkholderia (1.12%) (Fig. 1a), and other low-frequency genera (< 1.0%). Substantial changes in relative abundance were shown in the most abundant bacterial genera between colonization stages (Supplementary Fig. 4A). Only two archaeal genera, Methanoculleus and Methanosaeta, belonging to the Euryarchaeota phylum, were detected in both stages (Fig. 1b).
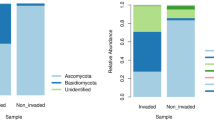
On the other hand, a total of five phyla, 33 orders, 54 families, and 83 genera were identified from total OTUs97% in all samples of the fungal of both colonization stages. Ascomycota was the most abundant phylum (92.12%), followed by Basidiomycota (7.28%), Zygomycota (0.54%), Glomeromycota (0.025%), and Chytridiomycota (0.02%) (Supplementary Fig. 2B). At the family level, Saccharomycetaceae (38.89%), Pichiaceae (18.34%), Trichocomaceae (10.10%), Helotiaceae (6.61), Ophiostomataceae (5.26%), Dothioraceae (3.82%), Mycosphaerellaceae (2.79%), Herpotrichiellaceae (2.58%), Sporobolomycetaceae (2.58%), Clavicipitaceae (1.58%), Rhizopogonaceae (1.58%), and Venturiaceae (1.10%) were the most abundant groups, followed by the 42 families present in low frequencies (< 1.00% of relative abundance) (Supplementary Fig. 3A).
At the genus level, Cyberlindnera (40.51%), Candida (19.1%), Penicillium (10.52%), Ceratocystiopsis (4.44%), Chalara (3.89%), Sydowia (3.81%), Cladosporium (2.91%), Rhodotorula (1.94%), Phaeomoniella (1.74%), Rhizopogon (1.65%), Cordyceps (1.64%), and Ophiostoma (1.04%) were the most abundant taxa (Fig. 1c), while the remaining genera were present in frequencies of < 0.1% of total reads each. Significant changes were observed in the relative abundance of fungal community (Supplementary Fig. 4B).
The Venn diagrams displayed the bacterial and fungal OTUs/genera shared and unique between both colonization stages (Fig. 2). In bacteria, 473 OTUs/50 genera were shared between two stages, and 72 OTUs/4 genera and 137 OTUs/6 genera were exclusive of early and late colonization stages, respectively (detailed information on bacterial genera available in Supplementary Table 1). On the other hand, in fungi, 91 OTUs/34 genera were shared between both stages, and 379 OTUs/33 genera and 164 OTUs/16 genera were exclusive of early and late colonization stages, respectively (detailed information on the fungal genera available in Supplementary Table 2).
Venn diagrams showing the distribution of (a) bacterial and (b) fungal OTUs97% associated with Arizona pine in early and late colonization stages by Dendroctonus rhizophagus. The number in parentheses represents the bacterial and fungal genera shared and unique, defined from taxonomic assignment of OTUs97%, between colonization stages
α- and β-diversity
Rarefaction curves of bacterial and fungal samples all tended to approach the saturation plateau (Fig. 3), while Good’s coverage values were also > 99.9% also for all samples (Table 1), indicating that the sequencing effort was enough to recover most of the bacterial and fungal diversity.
No significant differences were found in the species richness of bacterial communities between colonization stages with Chao1, because the richness was almost similar (PChao1 = 0.19). However, significant differences were observed in the diversity (PShannon = 0.004) and dominance (PSimpson = 0.0006), with Rahnella being the most abundant and dominant in the early stage, and this genus together with Pseudomonas and Pseudoxanthomonas in the late stage. On the other hand, the fungal α-diversity showed significant differences between colonization stages in the species richness (PChao1 = 0.0003) and diversity (PShannon = 0.038), given that the taxa number ranged from 244 to 281 in the early stage and 116.5 to 131.5 in the late stage. The dominance did not show differences (PSimpson = 0.12), being Cyberlindnera, Candida, and Penicillium the dominant genera in both stages.
The three first coordinates of PCoA analysis, using the Bray-Curtis dissimilarity matrix, explained 95.49 and 91.62% of the total observed variation in the analyses of bacterial (Fig. 4a) and fungal (Fig. 4c) communities, respectively. PCoA analyses showed significant differences in the β-diversity of bacterial (PERMANOVA; P = 0.02) and fungal (PERMANOVA; P = 0.0001) communities between colonization stages. In addition, the cluster analysis, also based also in the Bray-Curtis dissimilarity matrix, showed likewise that the composition of bacterial and fungal communities was different depending on the colonization stage (Fig. 4b, d).
Discussion
The present study analyzes the structure of microbial communities in subcortical tissues of Arizona pine saplings and the changes produced in them as a result of the attack and colonization carried out by the bark beetle D. rhizophagus. Our findings showed that the number of bacterial (20) and fungal (5) phyla detected in this study were almost constant across of the two colonization stages analyzed. However, the bacterial phyla number was over three times that of those detected (7) in a previous study that analyzed tissues of healthy sapling of Arizona pine [28]; whereas the fungal phyla number could not be compared because they have not been studied in healthy saplings.
Proteobacteria was the dominant bacterial phylum in the two colonization stages analyzed, which was also found to be the dominant group in decaying wood of conifers and other trees (e.g., Keteleeria evelyniana, Picea abies, Fagus sylvatica). This was done using both conventional molecular methods and next-generation sequencing (NGS) technologies [47,48,49,50]. The dominance of this phylum in wood samples under an advanced decaying state is assumed to be a reflection of their capacities to degrade available nutrients in acidic environments and with oxidative stress caused by the activity of wood-decay fungi [51, 52]. In fact, the presence of Archaea belonging to the Euryarchaeota phylum, mainly of the genera Methanoculleus and Methanosaeta, suggests a favorable environment for these members. In addition, these Archaea have also been detected in association with mycorrhizal fungi in Picea abies, Alnus glutinosa, and Pinus sylvestris [53, 54]. Other studies have reported Euryarchaeota from soil, which have the capacity to produce methane and act as decomposers of organic matter [55, 56], which may be a possible contribution to these Archaea in attacked pines by bark beetles.
The high abundance of Ascomycota members (e.g., Cyberlindnera, Candida, and Penicillium) in tissues of Arizona pines attacked by D. rhizophagus could be explained by their natural presence in healthy trees as endophytes [57, 58] and due to a number of them being pioneer species that can rapidly colonize new substrates [59], such as recently colonized pines and its death by the action of these insects. Whereas the Ascomycota was the most abundant in the two stages analyzed; no studies have reported this phylum in the tissues of trees colonized by insects. Previous studies have reported Basidiomycota as the main fungi in decaying tissues of fallen trunks, branches, stumps, littered leaves, and needles of pines and other conifers [60, 61]. However, all these studies were done in advanced stages of natural tree death, not in trees colonized by insects.
As such, it is not possible to make direct comparisons on α-diversity with previous studies, as, to our knowledge, no studies have characterized bacterial and fungal communities in tissues of pines attacked by other bark beetles. Our findings show that the diversity of bacterial and fungal genera increases as the state of decomposition of subcortical tissues in Arizona pine progresses (Table 1), favoring the presence of both bacteria (e.g., Azospirillum, Anaeromyxobacter) and fungi (e.g., Phoma, Neonectria) opportunists that produce enzymes that degrade the wood polymers, such as cellulose, lignin, and hemicellulose [62].
The presence of Rahnella as the dominant member in the early stage, and of this genus, together with Pseudomonas and Pseudoxanthomonas in the late stage, suggests that from the beginning of the attack of the bark beetle, an alteration of the endophytic microbial communities occurs in the subcortical tissue of these pine trees. In fact, it is well-known that during tree colonization, bark beetles alter the physical and mechanical defenses, along with the physiology of the tree and introduce microorganisms that might modify the microbiome of healthy pines [63,64,65]. Unfortunately, there are not studies about endophytic fungi in healthy Arizona pine tissues. Hence, it is not possible to know whether the dominance of the fungal genera (Candida, Cyberlindnera, and Penicillium) found in this study was a result from the action of this bark beetle.
An interesting aspect that should be noted is that dominant bacterial genera in the early and late stages of colonization of the subcortical tissues of Arizona pine are not the most frequent members in the endophytic bacterial community of healthy pine saplings. In contrast, they are the dominant bacteria in the D. rhizophagus gut across their different developmental stages [18, 66, 67], and they also constitute the gut core bacteriome of several species of this genus [19]. In addition, they have also been frequently recovered from other species of these bark beetles (e.g., D. ponderosae and D. simplex) [17, 63]. These aspects might be applicable to fungal dominant genera, Cyberlindnera and Candida, which have also been recovered from the gut and other parts of the body of Dendroctonus species [16, 67], despite of their frequency in subcortical tissues of healthy tree saplings being unknown.
Overall, results of β-diversity show that bacterial and fungal communities are segregated according to the colonization stage, resulting in significant differences in the community structure of these microbes (Figs. 3 and 4). This change may be due to physiological state of the tree [68, 69], or changes in different physicochemical factors, such as the temperature, pH, moisture, and organic matter content [70, 71], as well as regulatory processes (quorum sensing) and competition with other microbial groups [72], which may compromise the presence of some genera and favor the presence of others in this changing environment. However, despite this situation, the dominant bacteria and fungi maintain their prevalence, not only in the subcortical environment, but also in the gut, apparently performing specific metabolic functions for the benefit of both microorganisms and insects, as it has been well-demonstrated in other Dendroctonus species [20, 67, 73].
In summary, our findings provide evidence that microbial communities in subcortical tissues of Arizona pine are strongly influenced by the colonization stage of D. rhizophagus, resulting in a change in the structure of microbial communities as the decomposition state progresses. Several dominant members of these communities seem to be resident of trees as endophytes and are well-adapted to micro-environmental conditions in the subcortical tissues. The structure of whole microbial community varies as tree decomposition increases, apparently as a result of the shift in physicochemical conditions that can affect the microbiota composition, as well as microbial interactions. Finally, this research provides the basis for further studies to test metabolic functions in vivo of particular members of these microbial communities.
References
Franklin JF, Shugart HH, Harmon ME (1987) Tree death as an ecological process. BioScience 37:550–556
Castello JD, Leopold DJ, Smallidge PJ (1995) Pathogens, patterns, and processes in forest ecosystems. BioScience 45:16–24
Anderegg WRL, Kane JM, Anderegg LDL (2013) Consequences of widespread tree mortality triggered by drought and temperature stress. Nat Clim Chang 3:30–36
Wood SL (1982) The bark and ambrosia beetles of North and Central America (Coleoptera: Scolytidae), a taxonomic monograph. Great Basin Nat Mem 6:1–1356. https://doi.org/10.1038/NCLIMATE1635
Raffa KF, Aukema BH, Bentz BJ, Carroll AL, Hicke JA, Turner MG, Romme WH (2008) Cross-scale drivers of natural disturbances prone to anthropogenic amplification: the dynamics of bark beetle eruptions. BioScience 58:501–517. https://doi.org/10.1641/B580607
Blomquist GJ, Figueroa-Teran R, Aw M, Song M, Gorzalski A, Abbott NL, Chang E, Tittiger C (2010) Pheromone production in bark beetles. Insect Biochem Mol Biol 40:699–712. https://doi.org/10.1016/j.ibmb.2010.07.013
Fettig CJ, Klepzig KD, Billings RF, Munson AS, Nebeker TE, Negrón JF, Nowak JT (2007) The effectiveness of vegetation management practices for prevention and control of bark beetle infestations in coniferous forests of the western and southern United States. For Ecol Manag 238:24–53. https://doi.org/10.1016/J.FORECO.2006.10.011
Parmeter JR, Slaughter GW, Chen MM, Wood DL, Stubb HA (1989) Single and mixed inoculations of ponderosa pine with fungal associates of Dendroctonus spp. Phytopathology 79:768–772. https://doi.org/10.1094/Phyto-79-768
Paine TD, Raffa KF, Harrington TC (1997) Interactions among scolytid bark beetles, their associated fungi, and live host conifers. Annu Rev Entomol 42:179–206. https://doi.org/10.1146/annurev.ento.42.1.179
Lewinsohn D, Lewinsohn E, Bertagnolli CL, Patridge AD (1994) Blue-stain fungi and their transport structures on the Douglas fir beetle. Can J For Res 24:2275–2283. https://doi.org/10.1139/x94-292
Franceschi VR, Krokene P, Christiansen E, Krekling T (2005) Anatomical and chemical defenses of conifer bark against bark beetles and other pests. New Phytol 167:353–376. https://doi.org/10.1111/j.1469-8137.2005.01436.x
Keeling CI, Bohlmann J (2006) Diterpene resin acids in conifers. Phytochemistry 67:2415–2423. https://doi.org/10.1016/J.PHYTOCHEM.2006.08.019
Redford AJ, Bowers RM, Knight R, Linhart Y, Fierer N (2010) The ecology of the phyllosphere: geographic and phylogenetic variability in the distribution of bacteria on tree leaves. Environ Microbiol 12:2885–2893. https://doi.org/10.1111/j.1462-2920.2010.02258.x
Turner TR, James EK, Poole PS (2013) The plant microbiome. Genome Biol 14:209. https://doi.org/10.1186/gb-2013-14-6-209
Strid Y, Schroeder M, Lindahl B, Ihrmark K, Stenlid J (2014) Bark beetles have a decisive impact on fungal communities in Norway spruce stem sections. Fungal Ecol 7:47–58. https://doi.org/10.1016/j.funeco.2013.09.003
Rivera FN, González E, Gomez Z, López N, Hernández-Rodríguez C, Berkov A, Zúñiga G (2009) Gut-associated yeast in bark beetles of the genus Dendroctonus Erichson (Coleoptera: Curculionidae: Scolytinae). Biol J Linn Soc 98:325–342. https://doi.org/10.1111/j.1095-8312.2009.01289.x
Durand AA, Bergeron A, Constant P, Buffet JP, Déziel E, Guertin C (2015) Surveying the endomicrobiome and ectomicrobiome of bark beetles: the case of Dendroctonus simplex. Sci Rep 5:17190. https://doi.org/10.1038/srep17190
Briones-Roblero CI, Hernández-García JA, Gonzalez-Escobedo R, Soto-Robles LV, Rivera-Orduña FN, Zúñiga G (2017) Structure and dynamics of the gut bacterial microbiota of the bark beetle, Dendroctonus rhizophagus (Curculionidae: Scolytinae) across their life stages. PLoS One 12:e0175470. https://doi.org/10.1371/journal.pone.0175470
Hernández-García JA, Briones-Roblero CI, Rivera-Orduña FN, Zúñiga G (2017) Revealing the gut bacteriome of Dendroctonus bark beetles (Curculionidae: Scolytinae): diversity, core members and co-evolutionary patterns. Sci Rep 7(13864):13864. https://doi.org/10.1038/s41598-017-14031-6
Bleiker KP, Potter SE, Lauzon CR, Six DL (2009) Transport of fungal symbionts by mountain pine beetles. Can Entomol 141:503–514. https://doi.org/10.4039/n09-034
Bracewell RR, Six DL (2014) Broadscale specificity in a bark beetle-fungal symbiosis: a spatio-temporal analysis of the mycangial fungi of the western pine beetle. Microb Ecol 68:859–870. https://doi.org/10.1007/s00248-014-0449-7
Klepzig KD, Hofstetter RW (2011) From attack to emergence: interactions between the southern pine beetle, mites, microbes, and trees. In: Coulson RN, Klepzig KD (eds) Southern pine beetle II. General Technical Report SRS-140. Department of Agriculture Forest Service, Southern Research Station, Asheville, pp 141–152
Hofstetter RW, Dinkins-Bookwalter J, Davis TS, Klepzig KD (2015) Symbiotic associations of bark beetles. In: Vega FE, Hofstetter RW (eds) Bark Beetles. Elsevier, Amsterdam, pp 209–245. https://doi.org/10.1016/B978-0-12-417156-5.00006-X
Carrell AA, Frank AC (2014) Pinus flexilis and Picea engelmannii share a simple and consistent needle endophyte microbiota with a potential role in nitrogen fixation. Front Microbiol 5:333. https://doi.org/10.3389/fmicb.2014.00333
Carrell AA, Frank AC (2015) Bacterial endophyte communities in the foliage of coast redwood and giant sequoia. Front Microbiol 6:e01008. https://doi.org/10.3389/fmicb.2015.01008
Carrell AA, Carper DL, Frank AC (2016) Subalpine conifers in different geographical locations host highly similar foliar bacterial endophyte communities. FEMS Microbiol Ecol 92:fiw124. https://doi.org/10.1093/femsec/fiw124
Rúa MA, Wilson EC, Steele S, Munters AR, Hoeksema JD, Frank AC (2016) Associations between ectomycorrhizal fungi and bacterial needle endophytes in Pinus radiata: implications for biotic selection of microbial communities. Front Microbiol 7:399. https://doi.org/10.3389/fmicb.2016.00399
Gonzalez-Escobedo R, Briones-Roblero CI, Pineda-Mendoza RM, Rivera-Orduña FN, Zúñiga G (2018) Bacteriome from Pinus arizonica and P. durangensis: diversity, comparison of assemblages, and overlapping degree with the gut bacterial community of a bark beetle that kills pines. Front Microbiol 9:77. https://doi.org/10.3389/fmicb.2018.00077
Roth M, Hussain A, Cale JA, Erbilgin N (2018) Successful colonization of lodgepole pine trees by mountain pine beetle increased monoterpene production and exhausted carbohydrate reserves. J Chem Ecol 44:209–214. https://doi.org/10.1007/s10886-017-0922-0
Mendoza MG, Salinas-Moreno Y, Olivo-Martínez A, Zúñiga G (2011) Factors influencing the geographical distribution of Dendroctonus rhizophagus (Coleoptera: Curculionidae: Scolytinae) in the Sierra Madre Occidental, Mexico. Environ Entomol 40:549–559. https://doi.org/10.1603/EN10059
Herlemann DP, Labrenz M, Jürgens K, Bertilsson S, Waniek JJ, Andersson AF (2011) Transitions in bacterial communities along the 2000 km salinity gradient of the Baltic Sea. ISME J 5:1571–1579. https://doi.org/10.1038/ismej.2011.41
Bokulich NA, Mills DA (2013) Improved selection of internal transcribed spacer-specific primers enables quantitative, ultra-high-throughput profiling of fungal communities. Appl Environ Microbiol 79:2519–2526. https://doi.org/10.1128/AEM.03870-12
Magoc T, Salzberg SL (2011) FLASH: fast length adjustment of short reads to improve genome assemblies. Bioinformatics 27:2957–2963. https://doi.org/10.1093/bioinformatics/btr507
Rognes T, Flouri T, Nichols B, Quince C, Mahé F (2016) VSEARCH: a versatile open source tool for metagenomics. PeerJ 4:e2584. https://doi.org/10.7717/peerj.2584
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. https://doi.org/10.1038/nmeth.f.303
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267. https://doi.org/10.1128/AEM.00062-07
Kõljalg U, Nilsson RH, Abarenkov K, Tedersoo L, Taylor AFS, Bahram M, Bates ST, Bruns TD, Bengtsson-Palme J, Callaghan TM, Douglas B, Drenkhan T, Eberhardt U, Dueñas M, Grebenc T, Griffith GW, Hartmann M, Kirk PM, Kohout P, Larsson E, Lindahl BD, Lücking R, Martín MP, Matheny PB, Nguyen NH, Niskanen T, Oja J, Peay KG, Peintner U, Peterson M, Põldmaa K, Saag L, Saar I, Schüßler A, Scott JA, Senés C, Smith ME, Suija A, Taylor DL, Telleria MT, Weiss M, Larsson KH (2013) Towards a unified paradigm for sequence-based identification of Fungi. Mol Ecol 22:5271–5277. https://doi.org/10.1111/mec.12481
Yoon SH, Ha SM, Kwon S, Lim J, Kim Y, Seo H, Chun J (2017) Introducing EzBioCloud: a taxonomically united database of 16S rRNA and whole genome assemblies. Int J Syst Evol Microbiol 67:1613–1617. https://doi.org/10.1099/ijsem.0.001755
Altschul SF, Gish W, Miller W, Myers EW, Lipman DJ (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Oliveros JC (2007) VENNY. An interactive tool for comparing lists with Venn diagrams. http://bioinfogp.cnb.csic.es/tools/venny/index.html Accessed 26 March 2018
Chao A (1984) Non-parametric estimation of the number of classes in a population. Scand J Stat 11:265–270
Shannon CA (1948) Mathematical theory of communication. Bell Syst Tech J 27:379–423
Simpson EH (1949) Measurement of diversity. Nature 163:688–688
Good IJ (1953) The population frequencies of species and the estimation of population parameters. Biometrika 40:237–264
Rohlf FJ (1997) NTSYSpc Numerical taxonomy and multivariate analysis system version 2.0 user guide. Applied Biostatistics Inc., East Setauket
Hammer Ø, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:1–9
Zhang HB, Yang MX, Tu R (2008) Unexpectedly high bacterial diversity in decaying wood of a conifer as revealed by a molecular method. Int Biodeterior Biodegrad 62:471–474 https://doi.org/10.1016/j.ibiod.2008.06.001
Sun H, Terhonen E, Kasanen R, Asiegbu FO (2014) Diversity and community structure of primary wood-inhabiting bacteria in boreal forest. Geomicrobiol J 31:315–324. https://doi.org/10.1080/01490451.2013.827763
Hoppe B, Krger K, Kahl T, Arnstadt T, Buscot F, Bauhus J, Wubet T (2015) A pyrosequencing insight into sprawling bacterial diversity and community dynamics in decaying deadwood logs of Fagus sylvatica and Picea abies. Sci Rep 5:9456. https://doi.org/10.1038/srep09456
Rinta-Kanto J, Sinkko H, Rajala T et al (2016) Natural decay process affects the abundance and community structure of bacteria and Archaea in Picea abies logs. FEMS Microbiol Ecol 92:403–410. https://doi.org/10.1093/femsec/fiw087
de Boer W, van der Wal A (2008) Interactions between saprotrophic basidiomycetes and bacteria. In: Boddy L, Frankland JC, van West P (eds) Ecology of saprotrophic Basidiomycetes. Elsevier, Amsterdam, pp 143–153
Valášková V, de Boer W, Gunnewiek PJK, Pospíšek M, Baldrian P (2009) Phylogenetic composition and properties of bacteria coexisting with the fungus Hypholoma fasciculare in decaying wood. ISME J 3:1218–1221. https://doi.org/10.1038/ismej.2009.64
Bomberg M, Timonen S (2007) Distribution of Cren- and Euryarchaeota in scots pine mycorrhizospheres and boreal forest humus. Microb Ecol 54:406–416. https://doi.org/10.1007/s00248-007-9232-3
Bomberg M, Timonen S (2009) Effect of tree species and mycorrhizal colonization on the archaeal population of boreal forest rhizospheres. Appl Environ Microbiol 75:308–315. https://doi.org/10.1128/AEM.01739-08
Bates ST, Berg-Lyons DB, Caporosa JG, Walters WA, Knight R, Fierer N (2011) Examining the global distribution of dominant archaeal populations in soils. ISME J 5:908–917. https://doi.org/10.1038/ismej.2010.171
Angel R, Claus P, Conrad R (2012) Methanogenic archaea are globally ubiquitous in aerated soils and become active under wet anoxic conditions. ISME J 6:847–862. https://doi.org/10.1038/ismej.2011
Parfitt D, Hunt J, Dockrell D, Rogers HJ, Boddy L (2010) Do all trees carry the seeds of their own destruction? PCR reveals numerous wood decay fungi latently present in sapwood of a wide range of angiosperm trees. Fungal Ecol 3:338–346. https://doi.org/10.1016/j.funeco.2010.02.001
Rajala T, Peltoniemi M, Pennanen T, Mäkipää R (2012) Fungal community dynamics in relation to substrate quality of decaying Norway spruce (Picea abies [L.] Karst.) logs in boreal forests. FEMS Microbiol Ecol 81:494–505. https://doi.org/10.1111/j.1574-6941.2012.01376.x
Siitonen J (2001) Forest management, coarse woody debris and saproxylic organisms: Fennoscandian boreal forests as an example. Ecol Bull 49:11–41. https://doi.org/10.2307/20113262
Lundell TK, Mäkelä MR, de Vries RP, Hildén KS (2014) Chapter eleven - genomics, lifestyles and future prospects of wood-decay and litter-decomposing Basidiomycota. In: Martin FM (ed) Advances in botanical research. Elsevier Academic Press, London, pp 329–370. https://doi.org/10.1016/B978-0-12-397940-7.00011-2
Riley R, Salamov AA, Brown DW, Nagy LG, Floudas D, Held BW, Levasseur A, Lombard V, Morin E, Otillar R, Lindquist EA, Sun H, LaButti KM, Schmutz J, Jabbour D, Luo H, Baker SE, Pisabarro AG, Walton JD, Blanchette RA, Henrissat B, Martin F, Cullen D, Hibbett DS, Grigoriev IV (2014) Extensive sampling of basidiomycete genomes demonstrates inadequacy of the white-rot/brown-rot paradigm for wood decay fungi. Proc Natl Acad Sci U S A 111:9923–9928 https://doi.org/10.1073/pnas.1400592111
Johnston SR, Boddy L, Weightman AJ (2016) Bacteria in decomposing wood and their interactions with wood-decay fungi. FEMS Microbiol Ecol 92:fiw179. https://doi.org/10.1093/femsec/fiw179
Adams AS, Aylward FO, Adams SM, Erbilgin N, Aukema BH, Currie CR, Suen G, Raffa KF (2013) Mountain pine beetles colonizing historical and naïve host trees are associated with a bacterial community highly enriched in genes contributing to terpene metabolism. Appl Environ Microbiol 79:3468–3475. https://doi.org/10.1128/AEM.00068-13
Cheng CH, Xu LT, Xu DD, Lou QZ, Lu M, Sun JH (2016) Does cryptic microbiota mitigate pine resistance to an invasive beetle–fungus complex? Implications for invasion potential. Sci Rep 6:33110. https://doi.org/10.1038/srep33110
Wielkopolan B, Obrępalska-Stęplowska A (2016) Three-way interaction among plants, bacteria, and coleopteran insects. Planta 244:313–332. https://doi.org/10.1007/s00425-016-2543-1
Morales-Jiménez J, Zúñiga G, Ramírez-Saad HC, Hernández-Rodríguez C (2012) Gut-associated bacteria throughout the life cycle of the bark beetle Dendroctonus rhizophagus Thomas and Bright (Curculionidae: Scolytinae) and their cellulolytic activities. Microb Ecol 64:268–278. https://doi.org/10.1007/s00248-011-9999-0
Briones-Roblero CI, Rodríguez-Díaz R, Santiago-Cruz JA, Zúñiga G, Rivera-Orduña FN (2017) Degradation capacities of bacteria and yeasts isolated from the gut of Dendroctonus rhizophagus (Curculionidae: Scolytinae). Folia Microbiol 62:1–9. https://doi.org/10.1007/s12223-016-0469-4
Giordano L, Gonthier P, Varese GC, Miserere L, Nicolotti G (2009) Mycobiota inhabiting sapwood of healthy and declining scots pine (Pinus sylvestris L.) trees in the Alps. Fungal Divers 38:69–83
Proença DN, Francisco R, Kublik S, Schöler A, Vestergaard G, Schloter M, Morais PV (2017) The microbiome of endophytic, wood colonizing bacteria from pine trees as affected by pine wilt disease. Sci Rep 7:4205. https://doi.org/10.1038/s41598-017-04141-6
Rayner ADM, Boddy L (1988) Fungal decomposition of wood: its biology and ecology. John Willey & Sons, Chichester
Fukasawa Y, Osono T, Takeda H (2009) Microfungus communities of Japanese beech logs at different stages of decay in a cool temperate deciduous forest. Can J For Res 39:1606–1614. https://doi.org/10.1139/X09-080
Hodge A, Fitter AH (2013) Microbial mediation of plant competition and community structure. Funct Ecol 27:865–875. https://doi.org/10.1111/1365-2435.12002
Zhou F, Lou Q, Wang B, Xu L, Cheng C, Lu M, Sun J (2016) Altered carbohydrates allocation by associated bacteria-fungi interactions in a bark beetle-microbe symbiosis. Sci Rep 6:20135. https://doi.org/10.1038/srep20135
Acknowledgments
The authors would like to thank the COPAMEX for allowing the use of its facilities. This work was part of RG-E Ph.D. dissertation. He is a fellow of CONACyT (275347) and “Beca de Estímulo Institucional de Formación de Investigadores” (BEIFI-IPN). We thank the anonymous reviewers for their valuable comments on our manuscript.
Funding
This work was supported by the Secretaría de Investigación y Posgrado del Instituto Politécnico Nacional (SIP 20180686).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Rights and permissions
About this article
Cite this article
Gonzalez-Escobedo, R., Briones-Roblero, C.I., López, M.F. et al. Changes in the Microbial Community of Pinus arizonica Saplings After Being Colonized by the Bark Beetle Dendroctonus rhizophagus (Curculionidae: Scolytinae). Microb Ecol 78, 102–112 (2019). https://doi.org/10.1007/s00248-018-1274-1
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-018-1274-1