Abstract
Phosphorus (P) is a key biological element with important and unique biogeochemical cycling in natural ecosystems. Anthropogenic phosphorus inputs have been shown to greatly affect natural ecosystems, and this has been shown to be especially true of freshwater systems. While the importance of microbial communities in the P cycle is widely accepted, the role, composition, and relationship to P of these communities in freshwater systems still hold many secrets. Here, we investigated combined bacterial and archaeal communities utilizing 16S ribosomal RNA (rRNA) gene sequencing and computationally predicted functional metagenomes (PFMs) in 25 streams representing a strong P gradient. We discovered that 16S rRNA community structure and PFMs demonstrate a degree of decoupling between structure and function in the system. While we found that total phosphorus (TP) was correlated to the structure and functional capability of bacterial and archaeal communities in the system, turbidity had a stronger, but largely independent, correlation. At TP levels of approximately 55 μg/L, we see sharp differences in the abundance of numerous ecologically important taxa related to vegetation, agriculture, sediment, and other ecosystem inhabitants.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Microbial communities are extremely important in environmental and biogeochemical processes in nature with diversity being a key component [1,2,3]. It is well established that in addition to being P consumers, microbial communities affect the amount of available P for other ecosystem inhabitants through accessing P that is unavailable to other organisms and making it available [2,3,4,5]. Recent culture-based work looking at microbial communities and stoichiometric relationships between nutrients in freshwaters has shown that community assemblies shift dependent on phosphorus availability as well [6, 7]. This demonstrates a complex relationship where microbial community assemblages affect environmental P and environmental P affects microbial community assemblages in return [8, 9].
Shifts in environmental P may have impacts on community structure, diversity, and functional capability as well [10]. The results of previous studies exploring the relationship between species composition and function using a wide range of methods have been inconsistent. If we consider a strong correlation between community structure and function to be “coupled,” and a disconnect between the two to be decoupled, several studies have found community structure and function coupling [11, 12], others show decoupling [10, 13, 14], and many show a mixture of results [15, 16]. A mixture of coupling and decoupling or varying degrees of coupling seems the most likely response as some studies finding coupling such as in Garcia-Palacios et al. [11] hint at decoupled aspects and studies finding decoupling do not present the decouple as a complete disconnect. A large degree of uncertainty exists in lotic systems about the resiliency of microbial communities to environmental stress [17]. A decoupling of structure and function could indicate potential differences in community resiliency to environmental conditions. Functional redundancy of aquatic microbial communities may also be a good indicator of community resiliency to environmental stress [18, 19].
Environment-driven microbial community structuring in aquatic systems has been studied extensively since the 1970s. Until recently, this study has largely been conducted using microscopy and low-resolution community fingerprinting methods such as DGGE and TRFLP while relatively little work has been performed using high-throughput approaches such as next generation sequencing (NGS) technologies to better capture complex microbial communities [20]. Studies of microbial diversity have been a keystone of aquatic ecosystem research, but lotic systems have been underrepresented when compared to marine and lentic systems [21]. River and stream ecosystems are sensitive to anthropogenic P inputs often resulting in eutrophication under increased P levels [22,23,24]. Decreasing the gap in understanding between the P in lotic systems and microbial community structure has the potential to act as the foundation for bringing the mechanisms of this relationship to light.
The purpose of this study was to examine microbial community structure, function, and diversity across a total phosphorus (TP) gradient in freshwater streams. We hypothesized that (1) in an environment displaying a strong TP gradient, TP would be a strong correlate of community structure and (2) the overall functional capability of the sampled communities would be strongly coupled to their structural assembly in their relationship to TP. We tested these hypotheses by using next generation sequencing to look at microbial community structure and predicted functional capability in a system of streams representing a strong TP gradient.
Methods
Sampling
The study area was a collection of streams and rivers along the Oklahoma-Arkansas border, an area with known P enrichment problems [25,26,27]. We selected sampling sites with TP levels ranging from 18 to 163 μg/L. Sampling was performed in October of 2014. At each of the 25 sampling sites (Fig. S1), 50 mL of water was collected approximately 10 cm below the water surface and vacuum filtered with approximately 10 kPa of pressure through two filter sizes. A stacked filtration through a 1-μm PALL-type A/E glass fiber filter was conducted and then a PALL SUPOR 200 0.2-μm filter for bacterial and archaeal collection. Only the 0.2-μm filters were used in this study. Filters were stored at 4 °C less than 7 days for transport and then transferred to −80 °C until processed for DNA extraction.
Environmental Data
Total phosphorus (TP), total N (TN), sestonic chlorophyll-a, total suspended solids (TSS), turbidity, pH, specific conductance, dissolved oxygen, and temperature were measured at each site at the same time when bacterial and archaeal samples were collected. Molar concentrations of carbon (C), P, and N were calculated from dissolved organic carbon (DOC), TP, and TN and used to calculate ratios of C/P and C/N.
Unfiltered water samples were used for TP and TN. TP was digested with persulfate in an autoclave at 121 °C for an hour then analyzed on a Lachat 8500 series 2 using the ascorbic acid-molybdate method [28]. TN was digested with persulfate and sodium hydroxide (NaOH) in an autoclave at 121 °C for an hour. Samples were cooled and boric acid solution was added, then the sample was analyzed on a Lachat 8500 series 2 using the sulfanilamide cadmium reduction method [28]. For sestonic chlorophyll-a, 1 L of water was filtered onto a 0.45-μm filter then frozen. A 90% ethanol extraction was performed in a heated (78 °C) water bath for 10 min [29]. Total suspended solids (TSS), turbidity, pH, specific conductance, dissolved oxygen, and temperature were measured at each site at the same time when bacterial and archaeal samples were collected using a YSI EXO2 multiparameter data sonde (Yellow Springs, OH).
TP was measured at each site in August 2014 and October 2014 because of the likelihood that antecedent P conditions would drive bacterial and archaeal community composition as estimated by the October 2014 sampling. The October 2014 sampling event was a low water event resulting in abnormally high point data TP levels at some sites; thus, an average TP spanning August and October sampling events was used for subsequent analyses. This average was more representative of typical stream TP levels while still capturing the increase in TP leading into sampling.
Catchments were delineated in ArcGIS version 10 by utilizing the geographic coordinates of each stream reach to define a catchment outlet. We estimated the contributing area of each outlet on the basis of flow accumulation values derived from a 30-m digital elevation model (DEM) from the USGS National Elevation Dataset. Land use in each catchment was estimated from the National Land Cover Data (NLCD) raster, which was based on the 2011 Landsat Thematic Mapper.
Library Preparation
DNA was extracted from the 0.2-μm filters using a Mo Bio PowerWater® DNA extraction kit with the manufacturer’s protocol. Quantification of extractions was performed using a Qubit 3.0 fluorometric system, and samples showing no DNA extraction were discarded from the study at this time. DNA yields were less than 1 ng/μL causing us some initial concern about coverage but resulted in sequence depth of ~70,000 up to ~250,000 sequences per site and good sampling coverage per the rarefaction curves generated downstream (Fig. S2.A).
An initial PCR amplification of the 16S ribosomal RNA (rRNA) gene V4–V5 region was conducted using 515F forward and 926R reverse primers [30] modified to include adapters for future indexing. PCR was done using 2X Platinum™ Green Master Mix from Invitrogen. PCR specifications were 1 cycle for HotStart step at 94 °C for 2:00 min, then 30 cycles of 94 °C denaturation step for 0:45 min, 50 °C annealing step for 1:00 min, and 72 °C elongation step for 1:30 min. Successful PCR was identified through electrophoresis gel. PCR cleanup was conducted using an Agencourt AMPure XP kit and with the standard protocol from Beckman Coulter Life Sciences. Final PCR product quantification was conducted using Qubit 3.0.
A second round of PCR amplification was run in order to add unique indices to each sample as well as Illumina sequencing adapters. PCR was again done using 2X Platinum™ Green Master Mix kit from Invitrogen. PCR specifications for the second round of PCR were 1 cycle for HotStart step at 94 °C for 2:00 min, then 8 cycles of 94 °C denaturation step for 0:45 min, 59 °C annealing step for 1:00 min, and 72 °C elongation step for 1:30 min. Cleanup and quantification were performed in the same manner as the first round of PCR. Samples were then pooled so that 10 ng of DNA from each sample was present in the final library.
Sequencing
Sequencing was performed on an Illumina MiSeq system using a MiSeq Reagent Kit v3 2X300 with paired-end reads. Libraries were spiked with 20% PhiX Control.
Sequence Processing
Initial sequence processing including filtering and demultiplexing was conducted through the 16S metagenomic pipeline in Illumina BaseSpace. Paired-end read fastq files for each sample were extracted for downstream processing. Additional sequence processing was done using Quantitative Insights Into Microbial Ecology (QIIME) [31]. Paired-end reads were aligned first using the join-fastq algorithm from ea-utils [32]. Resulting sequences were then filtered at a Phred score of 20. Chimeric sequences were identified and removed using the USEARCH algorithm [33]. OTU picking was performed open reference using uclust against the Greengenes 13_8 database with a 0.97 similarity cutoff followed by de novo OTU picking [34]. Alpha diversity metrics were computed by QIIME as part of this pipeline using a rarefaction depth of 70,500 allowing use of all sites included in this study.
Construction of Predicted Functional Metagenomes
Predicted functional metagenomes (PFMs) were constructed from 16S rRNA gene sequence data using Phylogenetic Investigation of Communities by Reconstruction of Unobserved States (PICRUSt) [35]. OTU tables were normalized by count, and the PFMs were generated using the Kyoto Encyclopedia of Genes and Genomes (KEGG) database [36, 37]. PFMs were then grouped at KEGG level 3 representing the most diverse and specific functional groupings available.
Statistical Analysis Software
All analyses were performed in the R software package v.3.2.3 [38] with appropriate packages and scripts. OTU table BIOM files from QIIME were either exported to tab delimited format directly from QIIME or were imported for use in R using the phyloseq package version 1.14.0 in R [39].
Ordinations, Models, and Comparisons
Non-metric multidimensional scaling (NMDS) ordinations were constructed using the vegan package version 2.4–0 [40] to describe community dissimilarity in unconstrained space. OTU tables were log10(x + 1) transformed, and distances were computed as both Bray-Curtis and UniFrac. NMDS for Bray-Curtis distance 16S rRNA data and PFMs were performed on three axes, and 16S rRNA UniFrac distance was also performed on three axes. Redundancy analysis (RDA) models were also built in the vegan package in order to describe the community structure ordinations in environmentally constrained space. NMDS and RDA plots were created using the ggplot2 package in R [41]. Environmental gradients were built on NMDS ordinations using ordisurf from the vegan package. ordisurf uses general additive model (GAM) model building to overlay environmental variables in the ordination space [42]. RDA model selection was performed by starting with an initial model including all collected variables and manually removing collinear variables in an effort to maximize adjusted R 2 and minimize the magnitude of difference between R 2 and adjusted R 2.
16S rRNA and PFM data was compared with a Mantel test using the mantel function in the vegan package with the Pearson method and 1000 permutations in order to test for similarity between the data sets [43]. PROcrustean randomization TEST of community environment concordance (PROTEST), a potentially more sensitive detection method than a Mantel test, was also used to compare NMDS ordinations as well as RDAs using the protest function in the vegan package [44]. PROTEST uses scaling and rotations to maximize alignment in ordinations as a multivariate measure of concordance in species abundance and environmental datasets along with permutation-based significance testing [44]. All PROTEST analyses were performed with 1000 permutations.
Environmental Groupings and Analysis
Because preliminary environmental fitting against NMDS ordinations as well as RDA models suggested that turbidity and TP were both important in the system; sites were divided into turbidity and TP groups of Low, Med, and High for each variable independently. Group cutoffs were determined at apparent breakpoints in the distribution of our collected turbidity and TP data. Low for TP was set for sites below 40 μg/L and High as above 70 μg/L. Low for turbidity was set for sites below 1 Nephelometric Turbidity Units (NTU) and High as above 2.0 NTU. The designation of groupings for each site can be viewed in Table S1. Multivariate ANOVA-type analyses including PERMANOVA and ANOSIM were then performed with 1000 permutations using the vegan package to test for significant relationships between and among groupings.
Network Construction and Analysis
In order to visualize site relatedness and clustering, networks were constructed from phyloseq imported data using the network package version 1.13.0 and igraph package version 1.0.1 in R and Bray-Curtis distance [45,46,47]. The Bray-Curtis cutoff used for each network was the smallest value that allowed for the inclusion of all 25 sites. For 16S rRNA data, this value was 0.72, and for PFM data it was 0.028. Network clustering for each network was performed using a spin-glass model and simulated annealing via the cluster_spinglass function in the igraph package. Clustering from networks was compared using the normalized mutual information method (NMI) via the compare function in igraph package. Heatmaps were built using the heatmap.2 function in the gplots package version 3.0.1 [48]. Heatmap dendrograms were built using Bray-Curtis distance and uclust WPGMA clustering. Taxa that do not represent more than 1.5% of relative abundance in any site were removed post clustering for visual clarity.
Total Phosphorus Range of Greatest Impact
Threshold analysis for TP and turbidity was performed using the TITAN 2.1 package in R [49]. Threshold Indicator Taxa Analysis (TITAN) identifies environmental variable values maximizing taxa frequency and abundance using bootstrapping to identify reliable indicator taxa and the sum of indicator taxa value Z-scores to identify the environmental values representing the peak of increase or decline of the taxa [50]. The number of bootstraps performed in our TITAN 2.1 analyses was 500. For genus-level analysis, referenced genera observed more than three times were used, and for OTU-level analysis, OTUs observed more than 15 times were used. The same analysis was performed for turbidity as well.
Functional Redundancy Testing
Functional redundancy can be represented by the relationship between functional and species diversity with less correlation representing more redundancy [19, 51]. In order to test for functional redundancy in the system, Shannon diversity scores were computed for all sites in both the 16S rRNA and PFM data sets. Shannon diversity scores for each data set were plotted against each other, and potential correlations were explored using a GAM through the mgcv package version 1.8-11 in R [52]. GAM models were also used to model Shannon diversity correlation with turbidity for both 16S rRNA and PFM data.
Results
Environmental Relationship to Community Structure and Function
Vector fitting of environmental variables onto the NMDS ordinations identified turbidity and TP as the strongest and most significant environmental factors. The NMDS ordinations overlaid with turbidity and TP help to show trends of placement in ordination space with environment gradients for both 16S rRNA data and PFM data (Fig. 1). Contours for both turbidity and TP fit well with the distribution of sites in ordination space albeit in somewhat orthogonal directions. The turbidity gradient represents an almost linear fit to the ordination space distribution for the PFM data whereas more smoothing was involved in the 16S rRNA data. 16S rRNA UniFrac and Bray-Curtis ordinations were very similar with a PROTEST correlation statistic of 0.88 (p = 0.001).
NMDS ordinations. a 16S rRNA community with Bray-Curtis distance. Yellow to red gradient represents turbidity environmental fit, and gray lines represent TP (stress 0.067). b Predicted functional metagenome with Bray-Curtis distance. Yellow to red lines represent turbidity environmental fit, and gray lines represent TP (stress 0.030)
The RDA model for 16S rRNA data included turbidity, TN, pasture land coverage, and C/P ratios (Fig. 2a). TP was collinear with TN so it was removed from the model. The model had an adjusted R 2 value of 0.107, and an ANOVA-like permutation test for the constraining variables was significant with a p value of 0.001. The RDA model for the PFM data included turbidity, TP, developed land coverage, C/N ratios, and C/P ratios (Fig. 2b). The model-adjusted R 2 was 0.394, and an ANOVA-like permutation test for the constraining variables was significant with a p value of 0.001.
PERMANOVA and ANOSIM on the 16S rRNA data for TP grouping were both significant. Only PERMANOVA was significant for TP grouping on the PFM data where ANOSIM was not (Table 1). PERMANOVA and ANOSIM on the 16S rRNA data for turbidity groupings were also both significant. The analyses were also significant for the PFMs and turbidity groupings (Table 1).
Structure and Function Relationship
The OTU table and PFM, and dissimilarity matrices of each, were prepared for the structural and predicted functional capacity aspects of the communities. A Mantel test of 16S rRNA and PFM data returns no significant correlation between the dissimilarity matrices of the two data sets. The PROTEST correlation statistic for the RDA models was 0.57 (p = 0.01), and PROcrustes error plotting shows varying magnitudes and directions of change in ordination space (Fig. S3).
Linear and GAM modeling proved unproductive for diversity relationship to TP. A GAM model for 16S rRNA diversity shows a significant negative association between turbidity and Shannon diversity index values (R 2 adj. = 0.322, p = 0.008) (Fig. S4.A). A GAM model of PFM diversity against turbidity shows a negative association with turbidity as well (R 2 adj. = 0.593, p < 0.001) (Fig. S4.B).
16S rRNA and PFM networks by site relatedness were also visually quite different with very different clustering. In addition to being visually different, network clustering differed statistically with an NMI score of 0.430. Networks for 16s rRNA and PFMs appeared to both cluster well visually with turbidity groupings rather than TP groupings (Fig. S5). Many nodes differ in linked partners, and clusters differed in composition, number of connections to other clusters, and connection sites.
Total Phosphorus Range of Greatest Impact
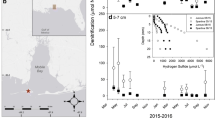
TITAN 2.1 identified five separate indicator genera decreasing in abundance (decreasers) and eight indicator genera increasing in abundance (increasers) in response to TP levels. OTU-based analysis identified a large number of OTUs in both the increaser and decreaser categories. Increasers and decreasers intersect at just below 55 μg/L of TP (Fig. 3). The 95% confidence intervals for the analysis identify a range from 52 to 58 μg/L TP for decreasers and 74.5 to 79.6 μg/L for increasers. In terms of sheer number of taxa, decreasers were much more prevalent than increasers. A TITAN analysis performed on turbidity indicated a strong threshold value right around 2.0 NTU mostly of decreasers, but interestingly, Sediminibacterium saw a significant increase (Fig. S6).
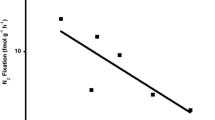
TITAN 2.1 analysis. a Plot of pure and reliable indicator taxa along the TP gradient. Black symbols correspond to genera that declined with increasing TP (z−), whereas open symbols correspond to those that increased (z+). Symbols are sized in proportion to the magnitude of the response (z-score). Horizontal lines represent 5th and 95th quantiles of values of TP resulting in the largest change in genus z-scores among 1000 bootstrap replicates. b Plot of sum z-scores for genus-level taxa. Steep slopes indicate major change points in abundance. c Plot of pure and reliable OTUs changing abundance around the threshold region. d Plot of sum z-scores for OTU-level taxa
Diversity and Community Taxonomy
The heatmap in Fig. 4 illustrates abundance trends across sites at the order level. Several unique genera displayed strong differences in relative abundance between groupings based on TP. Relative abundance of Acinetobacter from the order Pseudomonales varied with TP grouping increasing from very little (0.89%) in high TP sites to relatively much more in low TP sites (9.84%) (Fig. S2.B). Bacteria of the family Cytophagaceae were also much more abundant in high TP sites (Fig. S2.B). Abundance of bacteria of the genus Fluvicola from the order Flavobacteriales also increased with TP showing greater abundance in higher groupings.
Discussion
Turbidity
Turbidity was an important factor in our analyses despite utilizing only the 0.2- to 1-μm fraction which likely removed many larger particle-associated bacteria. We would expect this to limit a large amount of particle-associated bacteria, but the turbidity factor still had a strong relationship to both the structure and the functional capacity of the studied communities.
The primary ecological effect of turbidity in stream systems is to limit light availability [53]. Interactions between light availability and phosphorus on algal growth have previously been demonstrated as important for algal growth and community composition [54]. Turbidity also introduces sediment to the system and, along with it, particle-associated organisms. We did see soil-associated taxa such as Pseudomonas, Cytophaga, Micrococcus, Bacillus, and Agrobacterium present in our communities despite the focus on the 0.2- to 1-μm fraction. It is important to note, however, that the highest turbidity Nephelometric Turbidity Unit (NTU) values from the study sites were still relatively low (0.5–3.5 NTU) when compared to large rivers, lakes, or estuaries that are considered turbid, where turbidity typically exceeds >10–20 NTU [55]. Sampling for this study also occurred under base flow conditions representing the likely lowest turbidity conditions.
These factors make it difficult to separate out precisely what factors turbidity may be representing. Turbidity showed no correlation to total chlorophyll, and although it did correlate with total suspended solids and was collinear with several terrestrial type factors such as catchment area and flow, that is not the whole picture. Turbidity can be a poor measurement of sedimentation [56], and preliminary data exploration shows that sediment was likely not as prevalent in this sampling event as it is during other times of the year with similar turbidity measures. Total suspended solids only represented a small proportion of the turbidity measure in our data and was lower (\( \overline{x} \) = 2.0 mg/L, s = 1.75) than 10 historical measurements on all sites over the course of 2 years (\( \overline{x} \) = 3.7 mg/L, s = 6.87) within the same system as well.
The water column for the most turbid sites possessed an uncharacteristic and unidentified gray tint to the naked eye that was not readily apparent during other months of the year. This may be indicative of a biological factor such as the bacterial colonization of mineral particles or bacterial aggregates increasing turbidity scores. Therefore, turbidity here is probably a complex mixture of terrestrial and biological factors but further work is needed in order to explain this phenomenon.
Structure and Function Relationship
In addition to visual differences, the results of multiple analyses including the Mantel test, PROcrustes, and PROTEST analyses together indicate that structure and predicted functional capacity likely have features that are both coupled and decoupled in this system. Rather than seeing complete coupling or decoupling, we instead see indications of a degree of some decoupling. This appears to support the inconsistent results seen in previous studies and may mean a different result in terms of the degree of coupling when focusing on different specific ecosystem functional traits as opposed to the entire functional metagenome [15, 16].
Diversity relationship to turbidity appears to be one strongly coupled facet in this system. This indicates a potentially large environmental impact on taxa diversity with ongoing pollution and eutrophication of streams and rivers [57] which according to our results could lead to reduced functional diversity as well because the two appear to share a coupled diversity response. This is in contrast to the diversity relationship to TP which resulted in no productive linear or GAM models.
Functional Redundancy
We considered one possible explanation for any decoupling that we saw between structure and function to be functional redundancy. The GAM model for redundancy indicates a strong relationship between functional capability and species diversity in this system (Fig. S7). There does not appear to be much functional redundancy which could imply that environmental changes affecting diversity could have major implications for these communities.
The similarity of 16S rRNA UniFrac and Bray-Curtis ordinations seems to indicate that these communities have a strong evolutionary structural component and may maintain and develop unique communities over longer periods of time than would be expected in this type of fast moving lotic system. A recent study in catchment bacterioplankton found evidence that spatial factors are more important than temporal factors in determining community composition in lotic river systems supporting the development of unique community structures based on location and environmental parameters [58].
Nutrient Stoichiometric Relationships
The appearance of C/P and C/N ratios in our models is supported in previous research including research into changes in biomass incorporation based on C/P ratios [4]. The purpose of our study was to explore microbial communities across a gradient of TP, but the effects of C/P ratios should continue to be explored. Our results indicate only a relationship but not the direction of the relationship. Interestingly, the directions of TP and C/P ratios in the models were almost antithetical which is also supported in the literature [59]. The same was true for TN and C/N ratios. DOC by itself was not a strong correlate in any of our analyses although possibly due to collinearity.
Total Phosphorus Range of Greatest Impact
The steep decline of taxa near the ~55-μg/L TP value identified by TITAN analysis fell in the middle of our “middle” grouping for TP. This value would suggest that for microbial communities, any lotic freshwater site with a TP level of greater than 55 μg/L might be considered as a high TP site. This number is higher than threshold values found for other taxa in other systems. Previous work on benthic periphyton has indicated a slightly lower threshold for benthic periphyton taxa using regression-tree analysis in a wide-scale study in Canada [60], and an application of TITAN on benthic periphyton in Texas revealed an assemblage level threshold of approximately 20 μg/L TP [61]. Previous study of algal biomass has also indicated a lower threshold of 25 μg/L depending on light availability; however, this refers only to soluble reactive phosphorus and not to total phosphorus [54].
Diversity and Community Taxonomy
QIIME diversity analysis and TITAN threshold analysis both identified taxa that could be of interest to P dynamics in this system. The QIIME diversity analysis identified Acinetobacter which has been known to accumulate polyphosphates and polyhydroxyalkanoates in low nutrient environments [62] allowing it to thrive in environments with relatively low concentrations of orthophosphates (e.g., [63]). The lower TP sites may give Acinetobacter a competitive advantage over taxa that are more dependent on readily available P.
Within the threshold range of TP identified by TITAN analysis, we see a decrease in genus Cytophaga, a microbe associated with chitin consumption and with large organic matter decomposition [64] as well as Agrobacterium, Nesterenkonia, and Dysgonomonas. We see a significant increase in Clavibacter, a genus containing pythopathic pathogens affecting agriculture [65], and in Cloacibacterium, a genus originally isolated from wastewater in the central USA [66].
The heatmap in Fig. 4 illustrates how some sites are dominated by a few taxa where other sites are distributed much more evenly. The review performed by Zeglin [20] indicates a significant differentiation of many of these taxa’s parental phyla across stream “compartments” with relatively consistent taxonomic abundance within compartments. However, our results would seem to indicate quite a bit of variation in those phyla within the same compartment (water column) among these different sites.
Conclusions
Although TP may not be the strongest factor involved in the structure of microbial communities in this system during this October 2014 sampling, it remains undoubtedly an important factor in this stream and river system representing a strong P gradient. Testing via three separate methods including MANOVA-like tests, ordination, and RDA modeling, all identify TP as a factor in community and functional capability assembly.
Turbidity displays a strong relationship to our system despite the potential limiting of large particle-associated microbes by using the 0.2- to 1-μm fraction. It is easy to see how a strong or complex factor such as turbidity, missing factors, or data resolution limitations may mask other important factors in these systems.
We found that TP is important to both community structure and the PFM of the combined bacterial and archaeal community in this lotic freshwater system but in a decoupled fashion which is contrary to what we expected. These results indicate that looking expressly at community structure without function may show an incomplete ecosystem picture.
Our results showing a lack of functional redundancy in this system and the potential impacts of diversity change illustrate why further study of lotic freshwater microbial communities in the context of P is important to our understanding of the microbial ecology and biogeochemical nutrient cycling in these systems. It is important that we begin to look at the mechanisms for P biogeochemical cycling in lotic freshwater microbial communities beyond the characterization and modeling of structure, function, and diversity in order to better understand how they structure in relation to environmental P uptake and mobilization as members of a larger ecosystem. While these mechanisms are not addressed as part of this initial study, we have demonstrated that PFMs along with metagenome exploration may offer excellent tools to begin this investigation.
References
Schlesinger WH, Bernhardt ES (2013) Front matter. In: Biogeochem, Third Ed. Academic Press, Boston, pp i–ii
Van Der Heijden MG, Bardgett RD, Van Straalen NM (2008) The unseen majority: soil microbes as drivers of plant diversity and productivity in terrestrial ecosystems. Ecol Lett 11:296–310
Cotner JB, Biddanda BA (2002) Small players, large role: microbial influence on biogeochemical processes in pelagic aquatic ecosystems. Ecosystems 5:105–121. doi:10.1007/s10021-001-0059-3
Scott T, Cotner J, LaPara T (2012) Variable stoichiometry and homeostatic regulation of bacterial biomass elemental composition. Front Microbiol 3. doi:10.3389/fmicb.2012.00042
Richardson AE, Simpson RJ (2011) Soil microorganisms mediating phosphorus availability update on microbial phosphorus. Plant Physiol 156:989–996
Godwin CM, Cotner JB (2014) Carbon: phosphorus homeostasis of aquatic bacterial assemblages is mediated by shifts in assemblage composition. Aquat Microb Ecol 73:245–258
Godwin CM, Whitaker EA, Cotner JB (2017) Growth rate and resource imbalance interactively control biomass stoichiometry and elemental quotas of aquatic bacteria. Ecology 98:820–829
Tapia-Torres Y, Rodríguez-Torres MD, Elser JJ et al (2016) How to live with phosphorus scarcity in soil and sediment: lessons from bacteria. Appl Environ Microbiol 82:4652–4662. doi:10.1128/AEM.00160-16
Su J-Q, Ding L-J, Xue K et al (2015) Long-term balanced fertilization increases the soil microbial functional diversity in a phosphorus-limited paddy soil. Mol Ecol 24:136–150. doi:10.1111/mec.13010
Louca S, Parfrey LW, Doebeli M (2016) Decoupling function and taxonomy in the global ocean microbiome. Science 353:1272–1277
García-Palacios P, Vandegehuchte ML, Shaw EA et al (2015) Are there links between responses of soil microbes and ecosystem functioning to elevated CO2, N deposition and warming? A global perspective. Glob Chang Biol 21:1590–1600
Logue JB, Stedmon CA, Kellerman AM et al (2016) Experimental insights into the importance of aquatic bacterial community composition to the degradation of dissolved organic matter. ISME J 10:533–545
Pagaling E, Strathdee F, Spears BM et al (2014) Community history affects the predictability of microbial ecosystem development ISME J 8:19–30. doi:10.1038/ismej.2013.150
Sunagawa S, Coelho LP, Chaffron S et al (2015) Structure and function of the global ocean microbiome. Science 348:1261359. doi:10.1126/science.1261359
Tilman D, Knops J, Wedin D et al (1997) The influence of functional diversity and composition on ecosystem processes. Science 277:1300–1302
Xu Z, Malmer D, Langille MG et al (2014) Which is more important for classifying microbial communities: who’s there or what they can do. ISME J 8:2357–2359
Shade A, Peter H, Allison SD et al (2012) Fundamentals of microbial community resistance and resilience. Front Microbiol 3:417
Allison SD, Martiny JB (2008) Resistance, resilience, and redundancy in microbial communities. Proc Natl Acad Sci 105:11512–11519
Micheli F, Halpern BS (2005) Low functional redundancy in coastal marine assemblages. Ecol Lett 8:391–400
Zeglin LH (2015) Stream microbial diversity in response to environmental changes: review and synthesis of existing research. Front Microbiol 6:454
Zinger L, Gobet A, Pommier T (2012) Two decades of describing the unseen majority of aquatic microbial diversity. Mol Ecol 21:1878–1896. doi:10.1111/j.1365-294X.2011.05362.x
Anderson DM, Glibert PM, Burkholder JM (2002) Harmful algal blooms and eutrophication: nutrient sources, composition, and consequences. Estuaries 25:704–726. doi:10.1007/BF02804901
Correll DL (1998) The role of phosphorus in the eutrophication of receiving waters: a review. J Environ Qual 27:261. doi:10.2134/jeq1998.00472425002700020004x
Sharpley AN, Chapra SC, Wedepohl R et al (1994) Managing agricultural phosphorus for protection of surface waters: issues and options. J Environ Qual 23:437–451
Green WR, Haggard BE (2001) Phosphorus and nitrogen concentrations and loads at Illinois River south of Siloam Springs, Arkansas, 1997–1999
Haggard BE, Soerens TS (2006) Sediment phosphorus release at a small impoundment on the Illinois River, Arkansas and Oklahoma, USA. Ecol Eng 28:280–287. doi:10.1016/j.ecoleng.2006.07.013
Haggard BE (2010) Phosphorus concentrations, loads, and sources within the Illinois River drainage area, northwest Arkansas, 1997–2008. J Environ Qual 39:2113–2120
American Public Health Association (1998) Standard methods for the examination of water and wastewater. APHA-AWWA-WEF, Washington, D.C
Biggs BJF, Kilroy C Stream periphyton monitoring manual. Prepared for the New Zealand Ministry for the Environment
Caporaso JG, Lauber CL, Walters WA et al (2012) Ultra-high-throughput microbial community analysis on the Illumina HiSeq and MiSeq platforms. ISME J 6:1621–1624
Caporaso JG, Kuczynski J, Stombaugh J et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Aronesty E (2013) Comparison of sequencing utility programs. Open Bioinform J 7
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
DeSantis TZ, Hugenholtz P, Larsen N et al (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Langille MG, Zaneveld J, Caporaso JG et al (2013) Predictive functional profiling of microbial communities using 16S rRNA marker gene sequences. Nat Biotechnol 31:814–821
Kanehisa M, Goto S (2000) KEGG: kyoto encyclopedia of genes and genomes. Nucleic Acids Res 28:27–30
Kanehisa M, Sato Y, Kawashima M, et al (2015) KEGG as a reference resource for gene and protein annotation. Nucleic Acids Res gkv1070
R Core Team (2015) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
McMurdie PJ, Holmes S (2013) Phyloseq: an R package for reproducible interactive analysis and graphics of microbiome census data. PLoS One 8:e61217
Oksanen J, Blanchet FG, Kindt R, et al (2016) Vegan: community ecology package. R package version 2.4–1
Wickham H (2006) ggplot: an implementation of the grammar of graphics. R Package Version 210
Marra G, Wood SN (2011) Practical variable selection for generalized additive models. Comput Stat Data Anal 55:2372–2387
Smouse PE, Long JC, Sokal RR (1986) Multiple regression and correlation extensions of the Mantel test of matrix correspondence. Syst Zool 35:627–632
Jackson DA (1995) PROTEST: a PROcrustean randomization TEST of community environment concordance. Ecoscience 2:297–303
Butts CT (2008) Network: a package for managing relational data in R. J Stat Softw 24:1–36
Butts CT, Hunter D, Handcock M, et al (2015) Network: Classes for Relational Data
Csardi G, Nepusz T (2006) The igraph software package for complex network research. Int J Complex Syst 1695:1–9
Warnes GR, Bolker B, Bonebakker L, et al (2016) gplots: Various R programming tools for plotting data. R Package Version 301 2
Baker ME, King RS, Kahle D (2015) TITAN2: Threshold Indicator Taxa Analysis (Version 2.1). https://rdrr.io/cran/TITAN2/. Accessed 20 June 2016
Baker ME, King RS (2010) A new method for detecting and interpreting biodiversity and ecological community thresholds. Methods Ecol Evol 1:25–37
Miki T, Yokokawa T, Matsui K (2014) Biodiversity and multifunctionality in a microbial community: a novel theoretical approach to quantify functional redundancy. Proc R Soc B 281:20132498. doi:10.1098/rspb.2013.2498
Wood SN (2001) mgcv: GAMs and generalized ridge regression for R. R News 1:20–25
Lloyd DS, Koenings JP, Laperriere JD (1987) Effects of turbidity in fresh waters of Alaska. N Am J Fish Manag 7:18–33
Hill WR, Fanta SE, Roberts BJ (2009) Quantifying phosphorus and light effects in stream algae. Limnol Oceanogr 54:368–380
Chapman DV, Unesco and World Health Organization and United Nations Environment Programme (eds) (1996) Water quality assessments: a guide to the use of biota, sediments, and water in environmental monitoring. E & FN Spon, London
Davies-Colley RJ, Smith DG (2001) Turbidity suspended sediment, and water clarity: a review
Torsvik V, Øvreås L, Thingstad TF (2002) Prokaryotic diversity—magnitude, dynamics, and controlling factors. Science 296:1064–1066. doi:10.1126/science.1071698
Read DS, Gweon HS, Bowes MJ et al (2015) Catchment-scale biogeography of riverine bacterioplankton. ISME J 9:516–526
Cardinale BJ, Hillebrand H, Harpole WS et al (2009) Separating the influence of resource “availability” from resource “imbalance” on productivity–diversity relationships. Ecol Lett 12:475–487. doi:10.1111/j.1461-0248.2009.01317.x
Chambers PA, McGoldrick DJ, Brua RB et al (2012) Development of environmental thresholds for nitrogen and phosphorus in streams. J Environ Qual 41:7–20
Taylor JM, King RS, Pease AA, Winemiller KO (2014) Nonlinear response of stream ecosystem structure to low-level phosphorus enrichment. Freshw Biol 59:969–984
Singh V, Stapleton RD (2002) Biotransformations bioremediation technology for health and environmental protection. Elsevier Science Ltd., Amsterdam. http://0-www.sciencedirect.com.fama.us.es/science/publication?issn=00796352&volume=36
Saralov AI, Mol’kov DV, Bannikova OM, Solomennyĭ AP, Chikin SM (2001) Intracellular accumulation of monomer precursors of polyphosphates and polyhydroxyalkanoates from Acinetobacter calcoaceticus and Escherichia coli. Mikrobiologiia 70:737–744
Cottrell MT, Kirchman DL (2000) Natural assemblages of marine proteobacteria and members of the Cytophaga-Flavobacter cluster consuming low-and high-molecular-weight dissolved organic matter. Appl Environ Microbiol 66:1692–1697
Gartemann K-H, Kirchner O, Engemann J et al (2003) Clavibacter michiganensis subsp. michiganensis: first steps in the understanding of virulence of a gram-positive phytopathogenic bacterium. J Biotechnol 106:179–191
Allen TD, Lawson PA, Collins MD et al (2006) Cloacibacterium normanense gen. nov., sp. nov., a novel bacterium in the family Flavobacteriaceae isolated from municipal wastewater. Int J Syst Evol Microbiol 56:1311–1316. doi:10.1099/ijs.0.64218-0
Acknowledgements
The authors thank Morgan Bettcher, Stephen Cook, Stephen Elser, Katherine Hooker, Lauren Housley, and Caleb Robbins for their help in collecting field samples. We also thank Owen Lind and J. Thad Scott for assistance with internal review. We acknowledge the research support by Baylor University Office of Research and Baylor University Center for Reservoir and Aquatic Systems Research (CRASR).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflict of interest.
Statement of Data Availability
Sequence data that support the findings of this study have been deposited in GenBank with the BioProject accession code PRJNA350288. The environmental data that support the findings of this study are available from the corresponding author upon reasonable request.
Rights and permissions
About this article
Cite this article
LeBrun, E.S., King, R.S., Back, J.A. et al. Microbial Community Structure and Function Decoupling Across a Phosphorus Gradient in Streams. Microb Ecol 75, 64–73 (2018). https://doi.org/10.1007/s00248-017-1039-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-017-1039-2