Abstract
Mineral salt bricks are often used in cow raising as compensation for mineral losses to improve milk yield, growth, and metabolic activity. Generally, effects of minerals are partially thought to result from improvement of microbial metabolism, but their influence on the rumen microbiota has rarely been documented to date. In this study, we investigated the response of microbiota to mineral salt in heifer and adult cows and evaluated ruminal fermentation and enteric methane emissions of cows fed mineral salts. Twelve lactating Holstein cows and twelve heifers fed a total mixed ration (TMR) diet were randomly allocated into two groups, respectively: a treatment group comprising half of the adults and heifers that were fed mineral salt and a control group containing the other half fed a diet with no mineral salt supplement. Enteric methane emissions were reduced by 9.6% (P < 0.05) in adults ingesting a mineral salt diet, while concentrations of ruminal ammonia, butyrate, and propionate were increased to a significant extent (P < 0.05). Enteric methane emissions were also reduced in heifers ingesting a mineral salt diet, but not to a significant extent (P > 0.05). Moreover, the concentrations of ammonia and volatile fatty acids (VFAs) were not significantly altered in heifers (P > 0.05). Based on these results, we performed high-throughput sequencing to explore the bacterial and archaeal communities of the rumen samples. Succiniclasticum and Prevotella, two propionate-producing bacteria, were predominant in samples of both adults and heifers. At the phylotype level, mineral salt intake led to a significant shift from Succiniclasticum to Prevotella and Prevotellaceae populations in adults. In contrast, reduced abundance of Succiniclasticum and Prevotella phylotypes was observed, with no marked shift in propionate-producing bacteria in heifers. Methanogenic archaea were not significantly abundant between groups, either in adult cows or heifers. The shift of Succiniclasticum to Prevotella and Prevotellaceae in adults suggests a response of microbiota to mineral salt that contributes to higher propionate production, which competes for hydrogen utilized by methanogens. Our data collectively indicate that a mineral salt diet can alter interactions of bacterial taxa that result in enteric methane reduction, and this effect is also influenced in an age-dependent manner.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Enteric fermentation in ruminants generates methane, regarded as a byproduct of the digestive process, with 2–12% loss of ingested gross energy (GE) [1, 2] and significantly contributes to global warming [3, 4]. Several studies are underway to develop nutritional or additive-inhibiting strategies based on the principle of anti-methanogenesis to reduce methane emission in ruminants, including the use of electron receptors, such as fumarate, nitrates, sulfates, and nitroethane [5, 6]; chemical inhibitors, such as cyclodextrin [7]; ionophores, such as monensin [8]; plant bioactive compounds, such as tannin and saponin [9, 10]; and biotic agents [11, 12]. However, the effects of these additives are inconsistent, with some exerting toxicity in host animals at sufficiently high doses to reduce enteric methane emissions, in addition to concerns about toxic residuals in animal products sold to the public. Consequently, these techniques have yet to achieve widespread applicability. A strategy of nutritional regulation is believed to present an effective solution.
Mineral salt bricks are used to supply necessary mineral nutrients to animals as a feed supplement and generally aid in improving growth, fattening, milk yield, metabolism, and even disease prevention [13–15]. The mineral residual is partially taken up by the systemic circulation, and the remainder is absorbed by microbes, potentially resulting in changes in microbiota in the rumen. Methane is a byproduct released by methanogens during metabolic activity, a process known as methanogenesis [16], which also involves other microbial participants within the network of metabolic substrates [17]. So, it is very interesting to question if these related microbe responds to minerals and what do they have an effect on ruminal fermentation and enteric methane production. In a previous study by our group, ingestion of diets mixed with mineral salt brick powder led to reduced enteric methane emission from lactating cows (unpublished). Accordingly, we speculate that the effects of mineral salt on microbial communities may be related to methane mitigation. However, to our knowledge, no reports to date have focused on the response of the microbial community in the rumen to mineral salt intake, and the shifts in microbial composition, interactions, and abundance require further investigation.
In the present study, we applied high-throughput sequencing to analyze the effects of mineral salt intake on the diversity and composition of bacterial and archaeal communities in the rumen. Age was additionally included as a factor in a simultaneous investigation on heifers and lactating cows. Our main objective was to establish the responses of microbiota to mineral salt, based on key taxa classification and interaction patterns, with a view to explaining the mechanisms underlying the effects of mineral salt on ruminal fermentation and enteric methane mitigation.
Materials and Methods
Experimental Design
Twelve healthy lactating Holstein cows (3 to 4 years old, 682.3 ± 12.7 kg body weight) and twelve heifers (10 months old, 304.9 ± 11.8 kg body weight) were divided into two groups, with those in the treatment group fed a fresh diet containing mineral salts for 1 month. This study was performed in Shandong Province (118° 41′ N, 37° 04′ E) from September 2014 to October 2014, when the ambient temperature ranged from 25 to 30 °C inside the barn. The experimental period continued for 40 days, with the initial 10 days as the adaptive phase to comfort these animals. All the lactating cows chosen for experiment have produced two or three offspring and were 4 or 5 months postpartum. A commercial mineral brick (Tithebarn, Cheshire, UK) was ground into powder to produce mineral salt. The dose of minerals supplied was calculated according to the recommendation of the producer with 2 g minerals contained in 1 kg fresh diet. The mineral salt brick mainly contained Mg (5000 mg/kg), Co (75 mg/kg), Cu (1000 mg/kg), Fe (1500 mg/kg), Mn (500 mg/kg), Se (30 mg/kg), Zn (1000 mg/kg), I (150 mg/kg), and Na (38%). All animals were fed a TMR diet based on corn silage and alfalfa, had free access to food and water, and had no metabolic disorders or antibiotic treatments. During the whole experimental period, we canceled the premix supply, and the requirement for minerals just met with the diet. The ingredients and compositions of the diet are shown in supplementary Table S1.
Feed Sample, Enteric Methane, and Ruminal Fluid Analysis
The feed samples were dried in an oven at 105 °C for 24 h prior to grinding. The drying samples were ground through a 1-mm sieve before further analysis. Dry matter (DM), ash, neutral detergent fiber (NDF), acid detergent fiber (ADF), ether extract (EE), gross energy (GE), and total phosphorus (TP) were analyzed following the national standard methods in China. Total nitrogen (TN) and total carbon (TC) contents were analyzed by element analyzer (Elementar, Heraeus, Germany).
Enteric methane emissions were measured using the sulfur hexafluoride tracer technique according to the protocol of Lassey [18]. In the adaptive phase, permeation tubes filled with 1.5 g of sulfur hexafluoride were placed into the rumen for equilibrium. The release rate of each permeation tube was determined by weighing the tubes over time (about 4 mg/day). The sampling apparatus consisted of a U-shaped stainless steel vessel and a filter (50 μm) connected by a capillary tube. The U-shaped canister was fixed around the neck, while the filter was placed near to the cattle’s mouth and nostrils. The filter was used to keep the capillary line from plugging. Before use, the U-shaped vessel was first vacuumed, and the gas sampling time was regulated by the valve and the length of capillary tube. After a 24-h collection, the pressure inside was tested. If normal, pure N2 (99.9999%) was flushed into the vessel with a final pressure of 0.15 MPa. Thereafter, the gas samples were transferred to gas bags. In this study, eructated gas was collected in five consecutive days. A gas chromatograph fitted with an electron capture detector and flame ionization detector (Shimadzu, Kyoto, Japan) was used for determining sulfur hexafluoride and methane, respectively, according to the method reported by Boadi et al. [19]. The column and injector temperatures were 100 and 200 °C, respectively. Nitrogen was used as the carrier gas with a flow rate of 200 mL/s, and air and hydrogen flow rates set at 50 and 60 mL/s, respectively. The final methane output was calculated with the following equation: Q = R × C 1/C 2, where Q represents the methane emission volume per day, R the sulfur hexafluoride release rate, C 1 the methane concentration in samples, and C 2 the sulfur hexafluoride concentration in samples.
Ruminal fluid was aspirated using a flexible plastic tube and the first 100 mL discarded to avoid contamination of saliva. The remaining fluid was strained through a four-layer cheesecloth and transferred to a sterile centrifuge tube. Following immediate measurement of pH with a portable pH meter (IQ, CO, USA), the fluid was transferred into 2-mL tubes and immediately frozen by dipping of liquid nitrogen. Thereafter, the ruminal fluid samples were stored in −80 °C. Volatile fatty acid (VFA) concentrations were analyzed using a gas chromatograph (GC-14B, Shimadzu, Japan) according to the method of Hoskin et al. [20]. Ammonia-N levels were determined with a colorimetric technique using a spectrophotometer (Hash DR 6000, CO, USA) following the method of Chaney and Marbach [21].
DNA Extraction, MiSeq Sequencing and Bioinformatics Analysis
Total microbial DNA was extracted using a commercially available kit (Qiagen, Dusseldorf, Germany) according to the manufacturer’s instructions, and the concentrations detected at A260/A280 nanometer with a spectrophotometer (Bio-Rad, CA, USA). Amplification and sequencing of the hypervariable region of the 16S ribosomal RNA (rRNA) gene was performed using validated, region-specific primers optimized for the Illumina MiSeq platform [22, 23]. For bacterial 16S rRNA gene amplification, the primers used were 338F (5′–ACTCCTACGGGAGGCAGCA–3′) and 806R (5′–GGACTACHVGGGTWTCTAAT–3′). For archaeal 16S rRNA gene amplification, the primers Arch524F (5′–TGYCAGCCGCCGCGGTAA–3′) and Arch958R (5′–YCCGGCGTTGAVTCCAATT–3′) were employed. PCR reactions were performed in a triplicate 20-μL mixture containing 4 μL of 5×FastPfu Buffer, 2 μL of 2.5 mM dNTPs, 0.8 μL of each primer (5 μM), 0.4 μL of FastPfu Polymerase, and 10 ng of template DNA. Amplicons were extracted from 2% agarose gels, purified using the AxyPrep DNA Gel Extraction Kit (Axygen Biosciences, CA, USA) according to the manufacturer’s instructions, and quantified using QuantiFluor™-ST (Promega, WI, USA). Purified amplicons were pooled and paired-end sequenced (2 × 250) on an Illumina MiSeq platform according to standard protocols.
Raw FASTQ files were demultiplexed and quality-filtered using QIIME (version 1.17) using the following criteria: (i) 250-bp reads were truncated at any site receiving an average quality score of <20 over a 10-bp sliding window, discarding the truncated reads shorter than 50 bp; (ii) exact barcode matching, two-nucleotide mismatches in primer matching, and reads containing ambiguous characters were removed; and (iii) only sequences that showed >10 bp overlaps were assembled according to the overlap sequence. Reads that could not be assembled were discarded. Operational taxonomic units (OTUs) were clustered with 97% similarity cutoff using UPARSE (version 7.1 http://drive5.com/uparse/) and chimeric sequences identified and removed using UCHIME. The phylogenetic affiliation of each 16S rRNA gene sequence was analyzed via RDP classifier (http://rdp.cme.msu.edu/) against the SILVA (SSU119) 16S rRNA database using a confidence threshold of 70% [24].
Statistical Analysis
Statistical analyses were performed using analysis of variance (ANOVA) in SPSS 20.0 (IBM Statistics, NY, USA). Mean values were compared using the Bonferroni test for multiple comparisons. Differences between mean values were considered significant at P < 0.05. Bioinformatic analyses were performed using the R package. Non-metric multidimensional scaling (NMDS) was conducted to show the separation of samples, and the significance between the groups was tested using a vegan package based on the Bray-Curtis distance matrix. The correlation matrix was performed using the gpairs package, Venn diagram was plotted using the gplot package, and network analysis was performed using the igraph package. Linear discriminant analysis effect size (LEfSe) was performed using the LEfSe tool (http://huttenhower.sph.harvard.edu/galaxy/root?tool_id=lefse_upload) [25]. Redundancy analysis (RDA) was conducted with the vegan package to determine the relationships of microbial taxa, sample separation, and ruminal fermentation.
Accession Number
The sequencing data [BioProject ID: PRJNA348656] were deposited in GenBank database with sample accession numbers from SAMN05913198 to SAMN05913245.
Results
Animal Performance, Enteric Methane Emissions, and Ruminal Fermentation
The effects of mineral salt intake on performance, enteric methane emission, and fermentation are shown in Table 1. We observed no significant differences in DM intake and body weight between the treatment and control groups of heifers (P > 0.05) and adults (P > 0.05). The body weight gain of heifers was not significantly different between groups (P > 0.05).
Emissions of enteric methane (g/day) were reduced for both heifers and adults fed a diet containing mineral salt, although these differences did not appear significant between groups. The differences between the adult groups were moderately significant (P = 0.10), compared with those between the heifer groups (P = 0.61). Similar trends were observed for calculated methane emission (g/kg DM) and methane energy loss. Based on DM intake, higher methane emission and energy loss were observed in heifers, indicating distinct metabolic efficiency of the diet due to immature rumen development in younger animals.
Investigation of fermentative parameters showed that ammonia-N and VFA concentrations in adult cows were increased to a higher extent than those in the heifer group after mineral salt intake, in particular, a marked increase in ammonia-N and propionate levels (P < 0.05). Although the concentrations of acetate and butyrate were also increased, the differences in these parameters between groups were not significant (P > 0.05). The acetate to propionate (A:P) ratio in adults fed a mineral salt diet was markedly decreased (P < 0.05). Compared with adults, heifers displayed relatively constant ammonia-N and VFA concentrations after mineral salt intake. The pH was not obviously influenced by mineral salt in both the heifer and adult groups.
Sequencing Depth and Estimation of Alpha Diversity
We investigated both bacterial and archaeal communities using the high-throughput Illumina sequencing technique. Valid reads contained more than 18,000 sequences for each sample. For bacteria, a total of 779,669 high-quality reads remained, with an average of 32,486 reads per sample, ranging from 18,000 to 36,000. These sequences were assigned to 1704 OTUs based on 97% similarity. For archaea, 707,021 high-quality reads remained, with an average of 29,459 reads per sample, ranging from 20,000 to 37,400. These sequences were assigned to 381 OTUs based on 97% similarity. To eliminate uneven distribution of sequencing data, we adopted random subsampling for downstream diversity analysis. The subsampling threshold was set by the lowest number of sequences obtained in the samples, with cutoff values of 20,254 and 18,084 for bacteria and archaea, respectively.
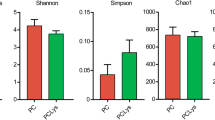
Subsampling sequences yielded sufficient resolution of bacterial and archaeal communities, as evident from rarefaction curve analysis (Fig. S1). Bacterial and archaeal community diversities were measured using both the Shannon-Wiener and Simpson indices. Bacterial and archaeal indices were not significantly different for both heifers and adults fed a mineral salt diet, compared to their counterparts (P > 0.05). The OTU numbers and estimated ACE and Chao1 values indicated an inverse trend for heifers and adults fed a mineral salt diet. Specifically, these values were increased for bacteria but decreased for archaea in adults, and the opposite trend was observed in heifers (Table 2).
Microbiota Distribution and Core OTUs Influenced by Mineral Salt
Overall, 25 phyla of bacterial taxa were detected in the samples, among which Bacteroidetes and Firmicutes were the most dominant, accounting for 92% of the total reads (Fig. S2a). Proteobacteria, Tenericutes, and Spirochaetae constituted 5% of the total reads (each showing 1.5–2% abundance). Prevotella_1, Succiniclasticum, Rikenellaceae_RC9_gut, and Christensenellaceae_R-7 were more predominantly detected between groups using ANOVA (Table 3; Fig. S2b), which listed the genus taxa accounting for at least 85% abundance in each sample. For archaea, only three phyla were detected, with Euryarchaeota and an unclassified group accounting for more than 98% abundance (Fig. S2c), but no significant differences observed between the groups at the genus level (Table 3). Methanobrevibacter contributed to about 93% total abundance of archaea (Fig. S2d).
Non-metric multidimensional scaling analysis was applied to determine the effects of bacterial and archaeal compositions on sample separation, based on the Bray-Curtis similarity metric with 97% sequence identity at the phylotype level (Fig. 1). The ordination plot showed a significant effect of mineral salt on divergence of bacterial composition in samples from adult (P < 0.05) group rather than from heifer (P > 0.05) group using analysis of similarities (ANOISM). In terms of archaeal composition, samples were not separated according to their own groups (P > 0.05), indicating no effects of mineral salt on archaeal composition and shift. The effects of mineral salt on sample separation revealed key roles of bacteria in reducing enteric methane from adult fed a mineral salt diet, suggestive of differential bacterial activities or interactions related to methane mitigation depending on age.
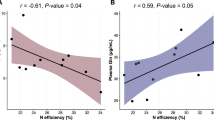
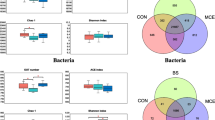
To identify specific OTUs that were differentially distributed in hosts subjected to different dietary regimes, we used LEfSe, a robust tool that distinguishes not only statistical significance but also biological relevance. Differential OTUs with abundance >0.1% were selected for further analysis. For adults, a total of 30 OTUs were significantly different between the two groups, with 18 being more abundant in the treatment group and 13 in the control group. Two predominant phylotypes of Succiniclasticum (OTU449 and OTU1336) were significantly decreased, while numerous phylotypes of Prevotella (OTU57, 68, 155, 335, 512, 809, 839, 950, 997, 1045, 1386, and 1612) and Prevotellaceae (OTU88 and 307) were increased in the treatment group (Fig. 2a, c). For heifers, 12 OTUs were differentially represented between the two groups to a significant extent, with eight being more abundant in the treatment group and four in the control group. The predominant phylotypes of Succiniclasticum (OTU449) and Prevotella (OTU1365) were decreased in the treatment group, and other phylotypes occurred in relatively low abundance of <0.5% (Fig. 2b, d). Specific archaea were not differentially distributed among hosts under different dietary regimes, indicating relatively stable composition regardless of mineral salt intake (Fig. S3). We also carried out the correlation analysis of differential abundances for Methanobrevibacter and bacteria (e.g., Succiniclasticum and Prevotella). For adult, Methanobrevibacter abundance was weakly correlated with other bacterial phylotypes (P > 0.05) and most of Succiniclasticum and Prevotella phylotypes were significantly negatively correlated in abundance (P < 0.05) (Fig. 3a). For heifer, the similar weak correlations of Methanobrevibacter were also observed with other bacterial phylotypes, but Succiniclasticum was not significantly correlated with other bacterial phylotypes in abundance (P > 0.05) (Fig. 3b). Furthermore, the bi-plot of redundancy analysis revealed a positive relationship between propionate production and Prevotella_1 abundance, which was dominant in the adult treatment group. Succiniclasticum and Succinivibrionaceae_unclassified showed a negative relationship with propionate production and were dominant only in samples of heifer groups (Fig. 4). Our results further confirmed the differential response to mineral salt in adults or heifers and the inhibitory effects of propionate-producing bacteria on methanogenesis.
Network Analysis of Microbiota
For both adult and heifer, intake of mineral salts induced effects on the rumen microbiota and altered patterns of microbial interactions, as revealed by network analysis. Succiniclasticum phylotypes (OTU449 and 1336) were key hubs shared by adults and heifers of the control groups. However, different patterns were observed in response to mineral salts, with a specific increase in abundance of other Prevotella phylotypes in adults, analogous to LEfSe data, that outcompeted Succiniclasticum (OTU449 and 1336) (Fig. 5a, b). In heifers ingesting mineral salts, network analysis disclosed no compensatory effect of Prevotella, but some dominant Bacteroidetes and unclassified Bacteroidales were increased (Fig. 5c, d; Table 3). A further search of unclassified Bacteroidales against nucleotide databases using BLASTN revealed ∼85% sequence identity with Barnesiella involved in succinate production.
Relationships of bacterial communities. The networks are based on the adjacency matrix of microbial abundance and correlation analysis at the phylotype level for adult control (a), adult treatment (b), heifer control (c), and heifer treatment (d). The nodes represent the bacterial taxa dominant in the samples, with the type and average abundance grouped by color and size. The edges represent the correlation between bacterial taxa, with red as positive and green as negative direction, respectively. The width of lines represents the strength of correlation
Discussion
Ruminants instinctively consume mineral salts in nature. Mineral salts are generally added to the diet as a supplementary nutrient or supplied in the form of bricks for consumption to improve animal performance [26, 27]. Despite their extensive use, the effects of mineral salts on the rumen fermentation and microbiota have not been thoroughly investigated. However, a previous study by our group revealed that mineral salt supplements are effective in methane reduction, suggesting that the microbial activity response to mineral salt is associated with methane production (unpublished).
In the present study, adult cows and heifers showed differential responses to mineral salt intake, mainly in terms of methane emission and fermentation. Adults fed a mineral salt diet displayed reduced methane emission and increased VFA and ammonia concentrations in the rumen while heifers did not show obvious changes. The rumen bacteria integrate ammonia-N into amino acids used for microbial protein synthesis, and their digestion supplies amino acids to the host animals [28]. The increased concentrations of ammonia-N indicate that mineral salts enhance the deamination activity of the rumen microbes in the degradation of food protein in adults. VFAs are extremely important for ruminants, contributing more than 70% of the energy supply. The rumen microbial fermentation produces propionate, butyrate, and acetate as the main VFAs, which are utilized for milk and meat production and supply a major proportion of the daily energy requirement of ruminants [29]. Reduced A:P ratios in the rumen are associated with increased feed efficiency and reduced methane production [30–32]. Therefore, increased propionate concentration and reduced A:P ratio suggest a shift of the metabolic pathways of some microbes related to carbohydrate degradation in adults. A previous report on bovine rumen showed that greater propionate production is accompanied by increased abundance of Prevotella sp., at the expense of simultaneous acetate and butyrate production [33]. Similar results were obtained in our study, whereby predominant Prevotella exceeded 48% of the total bacterial content in the rumen samples of adults fed a mineral salt diet (Table 3). The above findings suggest that mineral salt reduces enteric methane emissions in adults by facilitating propionate production in the rumen, which could compete with methanogens for hydrogen and contribute to the attenuation of methanogenesis [34].
16S rRNA gene sequencing of archaea revealed Methanobrevibacter as the most abundant OTU in the rumen in the present study, consistent with previous reports [35–39], but its abundance was not significantly influenced by mineral salt in either adults or heifers. This finding was not consistent with reduced enteric methane emission in adults, indicating that bacterial communities other than Methanobrevibacter are affected by mineral salt to influence methane production. The importance of bacterial community in enteric methane production was reported that different ruminotypes are linked with the low-methane emission trait in sheep [40]. Intergroup comparisons disclosed that differential OTUs are influenced by mineral salt, including phylotypes affiliated with Prevotella, Succiniclasticum, Ruminococcaceae, Butyrivibrio, Eubacterium, Saccharofermentans, Lachnospiraceae, Prevotellaceae, Fibrobacter, Bacteroidetes, unclassified Bacteroidales, Brocadiaceae, Rikenellaceae, and Erysipelotrichaceae (Fig. 2). Among the bacterial taxa, Prevotella and Succiniclasticum were the most abundant genera, belonging to the two dominant phyla of Bacteroidetes and Firmicutes, respectively. After an adaptive period to a dietary regime supplemented with mineral salt, the Succiniclasticum content was reduced significantly in both adult and heifer cows and succeeded by Prevotella in adults. Prevotella is involved in propionate production, which is further used for energy by the host [41, 42]. Therefore, the increased abundance of Prevotella phylotypes may account for the enhanced propionate concentration in the rumen of adults fed a mineral salt diet, and potentially underlie inhibition of methanogenesis, as reported previously [43, 44]. Network analysis further supported the importance of Prevotella in animals fed a mineral salt diet, showing increased abundance of Prevotella outcompeting other genera, as reflected by the negative correlations and independent occurrence (Fig. 5). Recently, Denman et al. [45] reported an enteric methane-inhibiting mechanism of bromochloromethane and confirmed its ability to shift rumen fermentation to propionate production, mediated by an increase in the populations of Prevotella and Selenomonas. It was also reported that the rumen OTUs assigned to Prevotella were promoted under chloroform treatment, which reduced enteric methane production [46].
Succiniclasticum is affiliated to the order Selenomonadales and specializes in fermenting succinate and converting it to propionate [47]. In the present study, Succiniclasticum abundance was decreased following mineral salt intake, and a simultaneous increase in Prevotella compensated for the loss of Succiniclasticum and propionate in adult cows. However, this compensatory effect was not observed in heifers. LEfSe analysis did not detect a similar boost in Prevotella phylotypes or other propionate-producing bacteria, making it difficult to be explained at the phylotype level. Some Bacteroidetes and unclassified genera of Bacteroidales were increased in heifers fed a mineral salt diet (Table 3). These taxa were related to Barnesiella, with 85% sequence similarity, based on a BLASTN search of nucleotide databases. Barnesiella produces succinate and acetate as end products of glucose metabolism [48]. The cumulative abundance of these Barnesiella-like taxa was found to be associated with the loss of Succiniclasticum abundance. Accordingly, we propose that these potential propionate-producing bacteria compensate for propionate production more or less.
Additionally, based on the same dietary regime, comparison between adults and heifers revealed differential microbial patterns in response to mineral salt, especially for phylotypes of Succiniclasticum and Prevotella. This phenomenon led to the speculation that bacterial distribution patterns and resource competition among organisms are regulated by age-dependent effects coupled with mineral-salt-induced pathways. As for the reason why high-abundant Succiniclasticum is coupled with more methane production in adult rather than in heifer, necessary evidence on metagenomics and metabonomics are still needed.
Conclusion
In summary, the effects of a mineral salt diet on the rumen microbiome were investigated using high-throughput sequencing to explore the mechanisms underlying enteric methane reduction. The response of microbiota in the rumen of adult or heifer cows to mineral salt differed mainly in terms of predominant propionate-producing bacteria. Mineral salt intake triggered a shift from predominant Succiniclasticum to Prevotella and Prevotellaceae in adults, but a similar effect was not observed in heifers, which simply maintained propionate homeostasis. Our data suggest that a mineral-salt-rich diet reduces enteric emissions mainly through enhancing the propionate-producing pathway. Moreover, this inhibitory effect is dependent on animal age. Future study will include metagenomics and metabonomics to relate biological function to the specific taxa found here, providing further insight to the response of key microbes to mineral salts.
References
Johnson K, Johnson D (1995) Methane emissions from cattle. J Anim Sci 73:2483–2492
Hook SE, Wright ADG, McBride BW (2010) Methanogens: methane producers of the rumen and mitigation strategies. Archaea 20:1–11. doi:10.1155/2010/945785
Hristov AN, Ott T, Tricarico J, Rotz A, Waghorn G, Adesogan A, Dijkstra J, Montes F, Oh J, Kebreab E, Oosting SJ, Gerber PJ, Henderson B, Makkar HP, Firkins JL (2013) Special topics-mitigation of methane and nitrous oxide emissions from animal operations: III. A review of animal management mitigation options. J Anim Sci 91:5095–5113. doi:10.2527/jas.2013-6585
Moss AR, Jouany JP, Newbold J (2000) Methane production by ruminants: its contribution to global warming. Ann de Zootech 49:231–253
Gutierrez-Banuelos H, Anderson RC, Carstens GE, Slay LJ, Ramlachan N, Horrocks SM, Callaway TR, Edrington TS, Nisbet DJ (2007) Zoonotic bacterial populations, gut fermentation characteristics and methane production in feedlot steers during oral nitroethane treatment and after the feeding of an experimental chlorate product. Anaerobe 13:21–31. doi:10.1016/j.anaerobe.2006.11.002
Brown EG, Anderson RC, Carstens GE, Gutierrez-Baňuelos H, McReynolds JL, Slay LJ, Callaway TR, Nisbet DJ (2011) Effects of oral nitroethane administration on enteric methane emissions and ruminal fermentation in cattle. Anim Feed Sci Technol 167:275–281. doi:10.1016/j.anifeedsci.2011.04.017
Lila ZA, Mohammed N, Tatsuoka N, Kanda S, Kurokawa Y, Itabashi H (2004) Effect of cyclodextrin diallyl maleate on methane production, ruminal fermentation and microbes in vitro and in vivo. Anim Sci J 75:15–22. doi:10.1111/j.1740-0929.2004.00149.x
Grainger C, Williams R, Eckard RJ, Hannah MC (2010) A high dose of monensin does not reduce methane emissions of dairy cows offered pasture supplemented with grain. J Dairy Sci 93:5300–5308. doi:10.3168/jds.2010-3154
Goel G, Makkar HPS (2012) Methane mitigation from ruminants using tannins and saponins. Trop Anim Health Prod 44:729–739. doi:10.1007/s11250-011-9966-2
Wang JK, Ye JA, Liu JX (2013) Effects of tea saponins on rumen microbiota, rumen fermentation, methane production and growth performance—a review. Trop Anim Health Prod 44:697–706. doi:10.1007/s11250-011-9960-8
Zhou M, Chung YH, Beauchemin KA, Holtshausen L, Oba M, McAllister TA, Guan LL (2011) Relationship between rumen methanogens and methane production in dairy cows fed diets supplemented with a feed enzyme additive. J Appl Microbiol 111:1148–1158. doi:10.1111/j.1365-2672.2011.05126.x
Kim SH, Mamuad LL, Kim DW, Kim SK, Lee SS (2016) Fumarate reductase-producing enterococci reduce methane production in rumen fermentation in vitro. J Microbiol Biotechnol 26:558–566. doi:10.4014/jmb.1512.12008
Morris JG (1980) Assessment of sodium requirements of grazing beef cattle: a review. J Anim Sci 50:145–152
Oconnor M, Hawke MF, Waller JE, Rotherham J, Coulter SP (2000) Salt supplementation of dairy cows. Pro New Zealand Grass Associ 62:49–53
Kincaid RL, Socha MT, Acan PAS (2004) Inorganic versus complexed trace mineral supplements on performance of dairy cows. Prod Anim Sci 20:66–73
Thauer RK, Kaster AK, Seedorf H, Bucke W, Hedderich R (2008) Methanogenic archaea: ecologically relevant differences in energy conservation. Nature 6:579–591. doi:10.1038/nrmicro1931
Ng F, Kittelmann S, Patchett ML, Attwood GT, Janssen PH, Rakonjac J, Gagic D (2015) An adhesin from hydrogen-utilizing rumen methanogen Methanobrevibacter ruminantium M1 binds a broad range of hydrogen-producing microorganisms. Environ Microbiol. doi:10.1111/1462-2920.13155
Lassey KR (2008) Livestock methane emission and its perspective in the global methane cycle. Aus J Exp Agric 48:114–118. doi:10.1071/EA07220
Boadi DA, Wittenberg KM, Kennedy AD (2002) Validation of the sulphur hexafluoride (SF6) tracer gas technique for measurement of methane and carbon dioxide production by cattle. Can J Anim Sci 82:125–131. doi:10.4141/A01-054
Hoskin SO, Stafford KJ, Barry TN (1995) Digestion, rumen fermentation and chewing behaviour of red deer fed fresh chicory and perennial ryegrass. J Agri Sci 124:289–295. doi:10.1017/S0021859600072956
Chaney AL, Marbach EP (1962) Modified reagents for determination of urea and ammonia. Clin Chem 8:130–137
Teske A, Sørensen KB (2008) Uncultured archaea in deep marine subsurface sediments: have we caught them all? ISME J 2:3–18. doi:10.1038/ismej.2007.90
Nossa CW, Oberdorf WE, Yang L, Aas JA, Paster BJ, Desantis TZ (2010) Design of 16S rRNA gene primers for 454 pyrosequencing of the human foregut microbiome. World J Gastroenterol 16:4135–4144. doi:10.3748/wjg.v16.i33.4135
Amato KR, Yeoman CJ, Kent A, Righini N, Carbonero F, Estrada A, Gaskins HR, Stumpf RM, Yildirim S, Torralba M, Gillis M, Wilson BA, Nelson KE, White BA, Leigh SR (2013) Habitat degradation impacts black howler monkey (Alouatta pigra) gastrointestinal microbiomes. ISME J 7:1344–1353. doi:10.1038/ismej.2013.16
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2010) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60. doi:10.1186/gb-2011-12-6-r60
DeFrain JM, Socha MT, Tomlinson DJ, Kluth D (2009) Effect of complexed trace minerals on the performance of lactating dairy cows on a commercial dairy. Prod Anim Sci 25:709–715
Hackbart KS, Ferreira RM, Dietsche AA, Socha MT, Shaver RD, Wiltbank MC, Fricke PM (2010) Effect of dietary organic zinc, manganese, copper, and cobalt supplementation on milk production, follicular growth, embryo quality, and tissue mineral concentrations in dairy cows. J Anim Sci 88:3856–3870. doi:10.2527/jas.2010-3055
Parmar NR, Solanki JV, Patel AB, Shah TM, Patel AK, Parnerkar S, Kumar JI, Joshi CG (2014) Metagenome of Mehsani buffalo rumen microbiota: an assessment of variation in feed-dependent phylogenetic and functional classification. J Mol Microbiol Biotechnol 24:249–261. doi:10.1159/000365054
Bergman EN (1990) Energy contributions of volatile fatty acids from the gastrointestinal tract in various species. Physiol Rev 70:567–590
Russell JB (1998) The importance of pH in the regulation of ruminal acetate to propionate ratio and methane production in vitro. J Dairy Sci 81:3222–3230. doi:10.3168/jds.S0022-0302(98)75886-2
Morgavi DP, Martin C, Boudra H (2013) Fungal secondary metabolites from Monascus spp. reduce rumen methane production in vitro and in vivo. J Anim Sci 91:848–860. doi:10.2527/jas.2012-5665
McCabe MS, Cormican P, Keogh K, O’Connor A, O’Hara E, Palladino RA, Kenny DA, Waters SM (2015) Illumina MiSeq phylogenetic amplicon sequencing shows a large reduction of an uncharacterised succinivibrionaceae and an increase of the Methanobrevibacter gottschalkii clade in feed restricted cattle. PLoS One 10:e0133234. doi:10.1371/journal.pone.0133234
Lettat A, Hassanat F, Benchaar C (2013) Corn silage in dairy cow diets to reduce ruminal methanogenesis: effects on the rumen metabolically active microbial communities. J Dairy Sci 96:5237–5248. doi:10.3168/jds.2012-6481
Morgavi DP, Forano E, Martin C, Newbold CJ (2010) Microbial ecosystem and methanogenesis in ruminants. Animal 4:1024–1036. doi:10.1017/S1751731110000546
Wright ADG, Ma X, Obispo NE (2008) Methanobrevibacter phylotypes are the dominant methanogens in sheep from Venezuela. Microbiol Ecol 56:390–394. doi:10.1007/s00248-007-9351-x
Zhou M, Sanabria EH, Guan L (2010) Characterization of rumen methanogenic community variation under different diets and host feed efficiencies using PCR-DGGE analysis. Appl Environ Microbiol 76:3776–3786. doi:10.1128/AEM.00010-10
Janssen PH, Kirs M (2008) Structure of the archaeal community of the rumen. Appl Environ Microbiol 74:3619–3625. doi:10.1128/AEM.02812-07
Seedorf H, Kittelmann S, Henderson G, Janssen PH (2014) Rim-DB: a taxonomic framework for community structure analysis of methanogenic archaea from the rumen and other intestinal environments. Peer J 2, e494. doi:10.7717/peerj.494
Henderson G, Cox F, Ganesh S, Jonker A, Young W, Collaborators GRC, Janssen PH (2015) Rumen microbial community composition varies with diet and host, but a core microbiome is found across a wide geographical range. Sci Rep 5:14567. doi:10.1038/srep14567
Kittelmann S, Pinares-Patinño CS, Seedorf H, Kirk MR, Ganesh S, McEwan JC, Janssen PH (2014) Two different bacterial community types are linked with the low-methane emission trait in sheep. PLoS One 9:e103171. doi:10.1371/journal.pone.0103171
Strobel HJ (1992) Vitamin B12-dependent propionate production by the ruminal bacterium Prevotella ruminicola. Appl Environ Microbiol 58:2331–2333
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Peña AG, Goodrich JK, Gordon JI, Huttley GA, Kelley ST, Knights D, Koenig JE, Ley RE, Lozupone CA, McDonald D, Muegge BD, Pirrung M, Reeder J, Sevinsky JR, Turnbaugh PJ, Walters WA, Widmann J, Yatsunenko T, Zaneveld J, Knight R (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336. doi:10.1038/nmeth.f.303
O’Neill BF, Deighton MH, O’Loughlin BM, Mulligan FJ, Boland TM, O’Donovan M, Lewis E (2011) Effects of a perennial ryegrass diet or total mixed ration diet offered to spring-calving Holstein-Friesian dairy cows on methane emissions, dry matter intake, and milk production. J Dairy Sci 94:1941–1951. doi:10.3168/jds.2010-3361
McCann JC, Wickersham TA, Loor JJ (2014) High-throughput methods redefine the rumen microbiome and its relationship with nutrition and metabolism. Bioinf Biol Insights 8:109–125. doi:10.4137/BBI.S15389
Denman SE, Martinez FG, Shinkai T, Mitsumori M, McSweeney CS (2015) Metagenomic analysis of the rumen microbial community following inhibition of methane formation by a halogenated methane analog. Front Microbiol 6:1087. doi:10.3389/fmicb.2015.01087
Martinez-fernandez G, Denman SE, Yang C, Cheung J, Mitsumori M, Mcsweeney CS (2016) Methane inhibition alters the microbial community, hydrogen flow and fermentation response in the rumen of cattle. Front Microbiol 7:1122. doi:10.3389/fmicb.2016.01122
Van Gylswyk NO (1995) Succiniclasticum ruminis gen. nov. sp. nov., a rumen bacterium converting succinate to propionate as sole energy yielding mechanism. Int J Syst Bacteriol 45:297–300. doi:10.1099/00207713-45-2-297
Morotomi M, Nagai F, Sakon H, Tanaka R (2008) Dialister succinatiphilus sp. nov. and Barnesiella intestinihominis sp. nov., isolated from human faeces. Int J Syst Evol Microbiol 58:2716–2720. doi:10.1099/ijs.0.2008/000810-0
Acknowledgments
This work is finacially supported by National Water Pollution Control and Treatment Science and Technology Major Project in China (2015ZX07103-007-022). We are particularly grateful to Dr. Guo Guang’s support with the data interpretation and design discussion.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
This study was performed on an experimental farm and granted permission by farm administrators. The experimental protocol was approved by the Animal Care and Use Committee of the Chinese Academy of Agricultural Sciences, and care of experimental animals was provided in accordance with Chinese standards.
Conflict of Interest
The authors declare that they have no competing interests.
Additional information
C Liu and XH Li are co-first authors.
Author contributions Conceived and designed the experiments: C Liu. Performed the experiments: C Liu, XH Li, YX Chen, ZH Cheng, QH Duan, QH Meng, XP Tao and B Shang. Analyzed the data: C Liu, XH Li, YX Chen. Contributed reagents/materials/analysis tools: XH Li and HM Dong. Wrote the paper: C Liu and XH Li. All authors agree to be accountable for all aspects of the work.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Fig. S1
Rarefaction analysis of the different samples. Rarefaction curves of OTUs clustered at 97% sequence identity across different environmental samples for bacteria and archaea. (DOCX 333 kb)
Fig. S2
The heatmap diagram showing the dominant bacterial taxa at the phylum (a) and genus (b) levels and the dominant archaeal taxa at the phylum (c) and genus (d) levels in different groups. AC: adult control; AT: adult treatment; HC: heifer control; HT: heifer treatment. (DOCX 382 kb)
Fig. S3
Differential archaeal OTUs identified by LEfSe between the treatment and control groups. Histogram shows OTUs that are more differential and abundant in the treatment (blue) or control (orange) group for adult (a and c) and heifer (b and d). (DOCX 117 kb)
Table S1
Ingredients and chemical composition of the diet (based on dry material). (DOCX 17.6 kb)
Rights and permissions
About this article
Cite this article
Liu, C., Li, X.H., Chen, Y.X. et al. Age-Related Response of Rumen Microbiota to Mineral Salt and Effects of Their Interactions on Enteric Methane Emissions in Cattle. Microb Ecol 73, 590–601 (2017). https://doi.org/10.1007/s00248-016-0888-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-016-0888-4