Abstract
Pathogens currently threaten the existence of many amphibian species. In efforts to combat global declines, researchers have characterized the amphibian cutaneous microbiome as a resource for disease management. Characterization of microbial communities has become useful in studying the links between organismal health and the host microbiome. Hellbender salamanders (Cryptobranchus alleganiensis) provide an ideal system to explore the cutaneous microbiome as this species requires extensive conservation management across its range. In addition, the Ozark hellbender subspecies (Cryptobranchus alleganiensis bishopi) exhibits chronic wounds hypothesized to be caused by bacterial infections, whereas the eastern hellbender (Cryptobranchus alleganiensis alleganiensis) does not. We assessed the cutaneous bacterial microbiome of both subspecies at two locations in the state of Missouri, USA. Through 16S rRNA gene-based amplicon sequencing, we detected more than 1000 distinct operational taxonomic units (OTUs) in the cutaneous and environmental bacterial microbiome. Phylogenetic and abundance-based dissimilarity matrices identified differences in the bacterial communities between the two subspecies, but only the abundance-based dissimilarity matrix identified differences between wounds and healthy skin on Ozark hellbenders. The higher abundance of OTUs on Ozark wounds suggests that commensal bacteria present on the skin and environment may be opportunistically colonizing the wounds. This brief exploration of the hellbender cutaneous bacterial microbiome provides foundational support for future studies seeking to understand the hellbender cutaneous bacterial microbiome and the role of the bacterial microbiota on chronic wounds of Ozark hellbenders.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Amphibian declines due to disease (such as chytridiomycosis and Ranavirus infections) have gained a great deal of attention within the past 20 years, and recent analyses have attempted to characterize the microbiome on the skin of amphibians as a way to link microbiome composition and disease susceptibility [1–3]. Next-generation DNA sequencing technologies have greatly increased the efficiency of microbial ecology studies. These technologies have opened up the possibility to characterize the microbiome of a diverse array of habitat types. Recent studies exploring bacterial communities, their functions, and their influence on their habitat have revolutionized our understanding of the microbial world in general. For example, microbiome characterization as a way to study disease in wildlife now includes studies characterizing the fecal [4], skin [5], respiratory system [6], and even whole colony microbiota [7]. Because disease has become a major factor of worldwide amphibian decline [8], expanding the use of this technology to amphibian taxa is crucial for future conservation efforts and decisions.
Hellbenders (Cryptobranchus alleganiensis) are fully aquatic salamanders that have a range spanning the eastern USA. These salamanders are restricted to lotic habitats that are fast flowing, cool, and have abundant rock cover [9]. In the past 20 years, this species has been facing population declines throughout the range associated with habitat loss, water quality disruptions, harvesting, and disease [10–13]. The eastern hellbender subspecies (Cryptobranchus alleganiensis alleganiensis) is broadly distributed throughout the Appalachian Mountain region and also the Midwest including Kentucky, Ohio, Indiana, Illinois, and Missouri. Eastern hellbenders are now classified as threatened or endangered in many of these states. The Ozark hellbender subspecies (Cryptobranchus alleganiensis bishopi) is endemic to southern Missouri and northern Arkansas and has also suffered population declines and local extirpations. Because of their limited distribution and population sizes, Ozark hellbenders are now federally listed under the Endangered Species Act [14, 15]. The state of Missouri is the only state within the range of hellbenders where both subspecies are present, though not sympatric.
Ozark hellbenders currently experience disease caused by unknown pathogen(s), as manifested by chronic cutaneous wounds throughout the body [16–18]. While ranaviruses and amphibian chytrid fungus have not been associated with these wounds, a bacterial pathogen has been suspected [17, 19]. Nickerson et al. [17] cultured and identified bacteria and fungi from six Ozark hellbender wounds showing signs of repressed tissue regeneration. The authors were able to identify representatives of a total of four bacterial and two fungal phyla from which some bacterial isolates were categorized as opportunistic pathogens. However, certain colonies were not identifiable and some inoculates exhibited no growth, as many bacteria cannot grow in culture mediums [20, 21]. Thus, they likely identified only a small subset of the total microbial diversity within the wounds of Ozark hellbenders.
Unlike Ozark hellbenders, chronic skin lesions have not been observed on eastern hellbenders in Missouri (C. a. alleganiensis) (J. Briggler, unpublished data). This key difference provides an ideal opportunity to explore the cutaneous microbiome between the two subspecies. While fungi and viruses could also be found on the skin of hellbenders, we chose to concentrate on the exploration of the cutaneous bacterial communities based on suspected bacterial involvement in hellbenders [17] and amphibians in general [1, 22–25]. The threatened status of hellbenders, and the presence of chronic wounds in only one subspecies, creates an opportunity to comparatively explore the cutaneous bacterial communities using a culture-independent approach (eg, next-generation sequencing). Next-generation sequencing technologies and statistical tools are more sensitive in the detection and quantification of rare species in microbial communities [26]. Assessing the bacterial communities between the intact skin and wounds of Ozark hellbenders could provide relevant information regarding the colonization of wounds by opportunistic bacteria. In addition, understanding the structure of the microbial communities between the two hellbender subspecies could inform future conservation decisions.
In the present study, we characterize the bacterial microbiomes of the two hellbender subspecies and between healthy skin and wounds of Ozark hellbenders using next-generation sequencing. When comparing Ozark and eastern hellbenders, we predicted that subspecies’ identity should be the primary driver of the bacterial microbiome composition on the skin of these salamanders. In addition, microbiome communities will differ based on Ozark hellbender swab location (ie, wound or healthy skin).
Materials and Methods
Sample Collection
Hellbenders were sampled from November 5 to 6, 2013. Samples for the eastern subspecies were collected from the Niangua River in central Missouri and the Ozark subspecies from the North Fork of the White River in southern Missouri. All individuals were captured by hand and rinsed with 1 L of sterile water. Rinsing the animal prior to swabbing was intended to ensure that the sample primarily included skin-associated microorganisms as opposed to those associated with the water column [27]. Following rinsing, the hellbenders were swabbed with sterile cotton-tipped swabs (Medline Industries Inc., Mundelein, Illinois) for 30 s in a rotating motion while applying minimum pressure on the skin. From each hellbender, two healthy skin samples were collected: one from the dorsum and one from the plantar surface of one foot free of wounds. Sample swabs from all wounds present on the hellbender’s body were also collected. Immediately after collection, the sample swabs were stored in 1.5-mL microcentrifuge tubes, placed on dry ice, and moved to a −80 °C freezer within 48 h. All hellbenders were returned to their location of capture within the river following sampling. River water samples were obtained from each locality by collecting 4 L of water from each river 1 to 10 m upstream from where hellbender sampling occurred. Water samples were stored on dry ice for up to 12 h until filtration occurred in an aseptic environment using a Whatman glass microfiber filter (GE Healthcare, Chicago, IL). The filters were stored in 15-mL centrifuge tubes and placed in a −80 °C freezer until DNA isolation occurred.
DNA Extractions, Amplification, and Sequencing
The two river samples were extracted using a PowerWater DNA Isolation Kit (MoBio Laboratories Inc., Carlsbad, CA) following their protocol for organisms that are difficult to lyse. Extraction of the swab samples was performed using a PowerSoil DNA Isolation Kit (MoBio Laboratories Inc., Carlsbad, CA) with a modified protocol (found in Supplementary Material, p. 1). The 16S rRNA gene V2 region was amplified for bacteria by using the primer set 27F/338R as in Fierer et al. [28]. Library preparation was performed in two sequential PCRs per sample, one in which a combination of the amplifying primers and partial Illumina adaptors were attached (performed in duplicate for each sample) and a second one in which dual-index barcodes were attached to each end with the remaining portion of the sequencing adaptors (Table S2). The initial PCRs consisted of 5 μL of template DNA, 5 μL of 5xMyTaq Reaction Buffer (Bioline, Tauton, MA, USA), 0.5 μL of MyTaq DNA Polymerase (Bioline, Tauton, MA, USA), 1 μL of 10 mM forward and reverse primers, and 13.5 μL of PCR water (MoBio Laboratories, Carlsbad, CA, USA) for a total of 25 μL per reaction. PCR conditions for these reactions were 95 °C for 2 min, 30 cycles of 94 °C for 45 s, 50 °C for 60 s, and 72 °C for 90 s, followed by 72 °C for 10 min. Following the initial PCR, the duplicate PCR products were pooled and cleaned using an UltraClean PCR Clean-Up kit (MoBio Laboratories, Carlsbad, CA, USA) following manufacturer’s protocol. The second PCR consisted of 5 μL of clean PCR product from the initial PCR, 5 μL of 5xMyTaq Reaction Buffer, 0.5 μL of MyTaq DNA Polymerase, 1 μL of 10 mM forward and reverse barcoded primers (specific per sample), and 13.5 μL of PCR Water for a total of 25 μL. PCR conditions consisted of 95 °C for 2 min, 10 cycles of 94 °C for 45 s, 65 °C for 60 s, and 72 °C for 90 s, followed by 72 °C for 10 min. PCR products were quantified using a Qubit Fluorometer (Invitrogen, NY) and pooled in equimolar amounts to be cleaned using the UltraClean PCR Clean-Up kit. The cleaned sample pool was sent to the Purdue Genomics Core Facility and sequenced on one lane of the Illumina HiSeq 2500 (Illumina Inc., San Diego, CA, USA) as 150 bp paired-end reads in rapid mode.
Sequence and Statistical Analysis
Resulting reads were processed using Trimmomatic [29] to remove Illumina adapter sequences and low quality bases (below threshold quality of phred-20) from both ends of the reads. The reads were further processed using a custom Python program (Supplementary Material, p. 13) that removed any reads fewer than 150 bp in length, to standardize read length, and renamed the reads with a name compatible with our chosen pipeline. The forward read file was processed using the Quantitative Insights Into Microbial Ecology version 1.8.0 (QIIME) pipeline [30]. Reads were grouped into operational taxonomic units (OTUs) at the standard 97 % identity using UCLUST [31]. To avoid including any OTUs generated by sequencer errors, such as base miscalls or chimeras, OTUs that were represented by a frequency fewer than 0.005 % of the total reads or were absent from more than one sample were removed [32]. The most abundant read per OTU was chosen as the representative sequence for that OTU in later analyses. Representative sequences were aligned to the pre-aligned Greengenes 13_5 release database [33] using PyNAST [34] at a minimum percent identity match of 75 %. Taxonomy was assigned to each OTU representative sequence using the UCLUST consensus taxonomy assigner and the Greengenes 13_5 OTU database at 80 % confidence [35]. The aligned representative OTU sequences were then used to generate a phylogenetic tree through FastTree to be used in later phylogenetic based distance matrix calculations [36]. Trimmomatic filtered reads and metadata were deposited in the European Bioinformatics Institute (primary accession no. PRJEB9832). OTUs were assigned and quality filtered from all environmental and animal samples. Following OTU assignment, the generated OTU table was rarified to 40,030 sequences per sample based on the sample with the lowest read count. To evaluate the diversity differences between the two river samples, OTU relative abundances were compared between the two river samples by computing a Pearson correlation coefficient.
Alpha diversity metrics (Chao 1, phylogenetic diversity index, Shannon-Wiener index, and observed species) for each eastern dorsum, eastern foot, Ozark dorsum, and Ozark wound sample were calculated within the QIIME pipeline. All four metrics were included to account for richness and evenness differences in alpha diversity between samples. Chao 1 is a richness estimator based on the number of rare classes found in samples [37]. The phylogenetic diversity index is a richness measure that uses the minimum total length of the phylogenetic branches represented by the bacterial species in a sample [38]. Finally, the Shannon-Wiener index is a quantitative measure that accounts for the frequency that each bacterial species is observed within a sample (community diversity) [39]. Calculated alpha diversity metrics were uploaded to R for subsequent analysis.
Feet Versus Dorsum Community Comparisons
To test for alpha diversity differences between sample types in eastern (dorsum versus feet) and Ozark hellbenders (dorsum versus wounds), a linear mixed model was applied through the package lme4 [40]. Sample type was included as a fixed variable and individual identity as a random variable. Beta diversity (using the unweighted UniFrac, weighted UniFrac, and Bray-Curtis dissimilarity metrics) was calculated independently for eastern and Ozark hellbender samples to produce between-sample dissimilarity matrices within QIIME. The UniFrac metrics use the phylogenetic distances between observed OTUs in each community when calculating distance between communities based on presence/absence (unweighted) or abundance (weighted) [41]. Alternatively, the Bray-Curtis metric incorporates OTU abundance only in the calculation of dissimilarity between samples [42]. Unweighted UniFrac and Bray-Curtis dissimilarity matrices were used to perform principal coordinate analysis (PCoA) in QIIME. To test for community differences between sample types for Ozark and eastern hellbenders, the dissimilarity matrices were uploaded to R. The dissimilarity matrices were used to perform a two-way Adonis test using both individual identity and sample types as grouping variables under the package vegan [43]. Additionally, a two-way PERMANOVA was calculated for wound versus dorsum Ozark hellbender comparisons with individual identity as a second variable in Past 3.10 [44].
To derive the OTUs that differentiate between Ozark hellbender dorsum and wounds, the core OTUs present in 95 % of samples in each sample type were first identified. This strict cutoff was chosen as it has been applied in other studies with low sample sizes, ensuring that only common OTUs represented across sample types are included [45]. Venn diagrams were created to display the number of core OTUs shared between sample types using the package VennDiagram in R. The linear discriminant analysis effect size (LEfSe) algorithm described in Segata et al. [46] was used to test for significant differences in core OTU relative abundance between dorsum and wounds. The LEfSe algorithm identifies the OTUs whose abundance statistically differs between the subspecies through a nonparametric factorial Kruskal-Wallis sum rank test (α < 0.05). Subsequently, generated effect sizes are calculated through a linear discriminant analysis (LDA) for those divergent OTUs. The effect sizes represent the magnitude of the association of each relevant core OTU to the category assigned. Because taxonomy data was not inputted into LEfSe, the resulting LDA scores reflect core OTU differences only. Core OTU taxonomy data was retrieved from Greengenes assignments performed in QIIME with an additional search of sequences in the Ribosomal Database Project (RDP) to confirm taxonomy or resolve unassigned sequences [47].
Subspecies Community Comparisons
Subspecies comparisons of alpha diversity metrics were performed using a Student’s t test since each sample derived from a unique individual. Beta diversity (using the unweighted UniFrac, weighted UniFrac, and Bray-Curtis dissimilarity metrics) was calculated for eastern hellbender dorsum and Ozark hellbender dorsum samples to produce between-sample dissimilarity matrices within QIIME. Unweighted UniFrac and Bray-Curtis dissimilarity matrices were used to perform the PCoA’s. To test for community differences between subspecies, the dissimilarity matrices were uploaded to R and used to calculate one-way ANOSIM and Adonis tests with subspecies identity as a grouping variable. Core OTUs present in 95 % of all Ozark and eastern dorsum samples were identified independently for each subspecies. The lists were used to produce Venn diagrams of shared core OTUs between both subspecies, inputted into LEfSe to test for differences in core OTU relative abundance between subspecies, and used to identify taxonomy as described above.
Results
We sampled five unique eastern and six unique Ozark hellbenders providing us with ten eastern healthy skin swabs, 11 Ozark healthy skin swabs, and nine Ozark wound swabs (Table S1). We removed one Ozark hellbender foot sample (9F) from the analysis due to a low number of sequences after the filtering process (n = 304). From each river, we collected one water sample. After assessing for quality, the filtered sequence set analyzed contained a total of 15,590,201 filtered reads of ∼150 bp length that were used in the remaining analyses. Our OTU list comparisons for the two river water samples showed high similarity (Pearson correlation value of r = 0.94 with a p < 0.001).
Feet Versus Dorsum Community Comparisons
Alpha metrics (Table S3) between eastern hellbender dorsum and feet samples were comparable (Table 1). The two-way Adonis test performed on the dissimilarity matrices produced for the feet and dorsum of eastern hellbenders showed no significant community differences for sample type (unweighted UniFrac: R = 0.08, p = 0.321; weighted UniFrac: R = 0.08, p = 0.182; Bray-Curtis: R = 0.08, p = 0.159) but significant differences between individuals (unweighted UniFrac: R = 0.65, p = 0.023; weighted UniFrac: R = 0.74, p = 0.008; Bray-Curtis: R = 0.75, p = 0.002). PCoAs generated from beta diversity metrics display no evident pattern between eastern foot and dorsum samples (Fig. S2). Finding no differences between sample types in healthy eastern hellbenders allowed us to continue comparisons between the healthy dorsum of eastern and Ozark hellbenders and between the Ozark dorsum and wound samples.
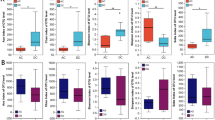
Alpha metric comparisons between Ozark hellbender dorsum and wound swabs revealed that wound sample richness outnumbered dorsum richness (Table 1 and Fig. 1). However, Shannon-Wiener and phylogenetic diversity comparisons between Ozark dorsum and wound swabs were comparable (Table 1). Community comparisons between Ozark hellbender wound and dorsum swabs using the two-way Adonis tests resulted in differences between sample type and individuals for both unweighted UniFrac (sample type R = 0.10, p = 0.019; individual R = 0.47, p = 0.004) and Bray-Curtis distance matrices (sample type R = 0.17, p = 0.001; individual R = 0.53, p = 0.007). The two-way Adonis test on the weighted UniFrac distance matrix between Ozark wound and dorsum swabs returned no differences between sample type but differences between individuals (sample type R = 0.06, p = 0.337; individual R = 0.50, p = 0.034). The two-way PERMANOVA comparisons revealed significant differences between Ozark sample type and individuals for the Bray-Curtis distance matrix (type: pseudo-F = 0.84, p = 0.023; individual: pseudo-F = 0.53, p = 0.050; interaction: pseudo-F = −0.32, p = 0.962) and non-significant differences for the unweighted UniFrac (type: pseudo-F = 0.81, p = 0.061; individual: pseudo-F = 0.76, p = 0.062; interaction: pseudo-F = 0.03, p = 0.278) or weighted UniFrac distance matrices (type: pseudo-F = 0.18, p = 0.663; individual: pseudo-F = 0.30, p = 0.448; interaction: pseudo-F = −0.36, p = 0.961). These results indicate that community dissimilarity between Ozark dorsum and wound swabs is due to differences in the abundance of shared OTUs observed within each sample type. Our PCoA plots comparing Ozark hellbender dorsum skin and wound swab communities support our statistical results by displaying no distinct grouping by sample type for coordinates calculated from the unweighted UniFrac distance matrix (Fig. 2b), but distinct grouping from the Bray-Curtis distance matrix (Fig. 3b).
Subspecies Community Comparisons
Alpha metrics (Table S3) between hellbender subspecies dorsum samples were comparable (Table 1). We noted community differences between the sampled eastern and Ozark hellbender dorsum communities for all metrics testing with both Adonis (unweighted UniFrac: R = 0.25, p = 0.024; weighted UniFrac: R = 0.26, p = 0.019; Bray-Curtis = 0.26, p = 0.011) and ANOSIM tests (unweighted UniFrac: R = 0.34, p = 0.020; weighted UniFrac: R = 0.39, p = 0.006; Bray-Curtis = 0.48, p = 0.008). Our PCoA plots comparing both hellbender subspecies dorsum swabs show distinct grouping by subspecies identity for coordinates derived from unweighted UniFrac (Fig. 2a) and Bray-Curtis distance matrices (Fig. 3a). These results suggest that the cutaneous microbiome of eastern and Ozark hellbenders differs in both composition and abundance of OTUs.
Core Bacterial Microbiome Results
Our core microbiome cutoff identified 70 core OTUs from eastern hellbender dorsum skin swabs, 112 core OTUs from Ozark hellbender dorsum skin swabs, and 321 core OTUs from Ozark hellbender wound swabs. There was an overlap in the core OTUs identified between Ozark and eastern hellbenders dorsum and Ozark hellbender dorsum and wound samples (Fig. 4). Phylum-level richness of the core bacterial microbiome of eastern and Ozark hellbenders was similar, while the wound core bacterial microbiome contained higher richness with a larger percentage of unclassified bacteria (Table S4).
The LEfSe algorithm identified 117 distinct core OTUs between Ozark and eastern hellbenders (Table S5). Thirty-seven OTUs were associated with eastern hellbenders, whereas 80 were associated with Ozark hellbenders. OTUs identified as Cetobacterium somerae, Leptospira sp., Bacteroides sp., Alistipes sp., and an unclassified species in the family Enterobacteriaceae were assigned with high LDA effect sizes to eastern hellbenders (>4.0). OTUs identified as Bacillariophyta sp., an unclassified species in the family Sphingomonadaceae, Dechloromonas sp., an unclassified species in the class Betaproteobacteria, Massilia sp., and Novosphingobium sp. received the highest associations to the Ozark hellbenders (LDA>3.2). The LEfSe algorithm identified 29 core OTUs to be discriminant biomarkers between wounds and healthy dorsum swabs collected from Ozark hellbenders, with 26 wound-associated and 3 dorsum-associated bacteria (Table 2). Four OTUs with unknown taxonomy (denovo206854, denovo37779, denovo40873, and denovo84139) were associated with wounds and exhibited high relative abundance across wound swabs. For OTUs whose taxonomy is known Acidovorax sp., Flavobacterium sp., Aquabacterium sp., Chryseobacterium sp., and Acinetobacter sp. also displayed high effect size and high relative abundance within wounds compared to intact skin.
Discussion
Microbiome studies across diverse amphibian taxa provide evidence for an association between hosts and their microbial symbionts [5, 49]. Our study is unique among other amphibian cutaneous microbiota projects because it builds on a previous culture-based approach characterizing the wounds of Ozark hellbenders in the North Fork of the White River, MO [17]. Compared to the work performed by Nickerson et al. [17], the number of bacterial OTUs described in this culture-independent study is significantly greater. We also included samples from eastern hellbenders and intact skin from the Ozark hellbenders from which we obtained wound samples. This allowed us to compare the bacteria of wounds to healthy skin sites and of affected to healthy populations. From our samples, we were able to identify unclassified, culture-specific, and unculturable bacteria [48], which could not be identified using the methods of Nickerson et al. [17].
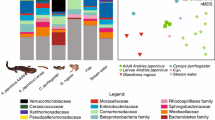
The two hellbender subspecies sampled in our study differed in their skin-associated bacterial communities (Figs. 2a and 3a). Although the differences observed could be due to other factors (e.g., difference in age, sex, exposure to different sediment bacteria), the results of this study are consistent with previous bacterial microbiome evaluations on the skin of amphibians [5, 49–51]. The cutaneous bacterial communities of amphibians have been strongly associated with host species identity; thus, indicating that skin bacterial communities are specific to each amphibian species [5, 49]. In hellbenders, phylogenetic assessments across the species range offer strong support for paraphyly for the two currently known subspecies [52, 53]. Through this perspective, differences in the bacterial microbiome between the subspecies could be driven by the same speciation forces already acting between them [54]. However, these observations are based on individuals from two proximate rivers within the large range of the species and could be due to environmental differences between the two streams. While the communities that we identified from the river water were similar, they do not reflect all environmental sources of bacteria (i.e., substrate). Thus, characterizing the skin and environmental bacterial microbiome throughout the entire range of hellbenders will allow to test if symbiotic differences follow patterns similar to those of genetic differentiation between host populations [55].
While Ozark and eastern hellbenders shared a subset of their core OTUs, the Ozarks displayed a higher OTU richness than the easterns (Fig. 4). These core OTUs are specific to the populations sampled, and it is unknown whether they reflect the core bacterial microbiota of each subspecies across the range. Therefore, investigating the bacterial communities of hellbenders across the range of both subspecies could also reduce the list of core OTUs. Identifying the subset that is common between all localities where these salamanders are found could provide insight to current conservation approaches. Captive rearing and translocations have become common conservation tools for hellbenders across the range [56, 57], and these methods have been associated with disruption of the natural microbiome of amphibians [58]. Given that bacterial symbionts protect the amphibian host against pathogens [25], maintaining diversity of the core microbiome may increase the survivorship of reintroduced individuals. Thus, it is important to minimize the disruption of the microbiome when moving individuals from source to supplemented populations. The composition of our hellbender skin bacterial microbiome is comparable to that of other North American amphibians and fish (Fig. S1) [49, 51, 59], suggesting a close association between bacterial symbionts and these taxa across evolutionary time. Similarities between hellbenders compared to other aquatic vertebrate taxa and the existence of an extensive captive rearing effort in this species, suggest that future investigations in wild and captive hellbenders could provide knowledge applicable to conservation of other amphibians.
Wound samples contained higher OTU richness than dorsum samples within Ozark hellbenders (Table 1 and Fig. 1), while community structure differed between the two sample types based on abundance only (Figs. 2b and 3b). The differences in OTU richness and community structure observed between these sample types can be attributed to the pathophysiology of the chronic wounds. Adult hellbenders tend to acquire injuries during the mating season, when territorial bouts occur between conspecifics or as a result of predator attacks [16, 60]. While eastern hellbenders are able to heal completely as evidenced by a lack of chronic wounds, Ozark hellbenders are unable to clear their injuries retaining an open port of colonization [18]. We cannot determine whether the OTUs that we identified in wounds are the cause of wound retardation in this subspecies, but we can note that these open areas provide a novel habitat that commensal and environmental bacteria can exploit. As a result, the exposed tissue could provide a habitat through which a richer community is maintained compared to that of the skin.
Apart from differences in OTU richness between Ozark hellbender wounds and dorsum skin, we were also able to identify OTUs that differed in their abundance between both sample types (Table 2). These wound-assigned OTUs were identified to the genus Acidovirax within the family Comamonadaceae, the genus Flavobacterium within the family Flavobacteriaceae, and the genus Acinetobacter within the family Moraxellaceae. These families have been previously described as common members of the amphibian skin microbiome [5, 24], indicating that the OTUs identified on the wounds could derive from the healthy skin. Acidovorax sp. has been recovered from the skin of other amphibians [61–63], including identification on Italian stream frogs (Rana italica) infected with amphibian chytrid fungus (Batrachochytrium dendrobatidis) [1]. The genus Flavobacterium has been recovered from several amphibian species and has been suspected to possess protective roles in the microbiome of the amphibian host [1, 61, 63, 64]; however, the same genus has been associated with dermatosepticemia in amphibians [65–67]. Acinetobacter sp. has been recovered from skin lesions and inner organs of the hellbender’s sister species, Andrias davidianus, in individuals that perished from Ranavirus infections [68]. While our methodology makes it difficult to declare these wound-associated species as pathogens or protective symbionts, it is likely that some of these OTUs contribute to the chronic wound status. Therefore, future bacterial evaluations on hellbenders and other amphibians should consider these OTUs (Table 2) as they could correlate with host health status.
We found differences in the bacterial community between the two subspecies of hellbenders that coincide with the divergence of these two groups. Our findings serve as a starting point towards future microbiome assessments in this species, which could inform the conservation of hellbenders and other amphibians. The negative effects of disease on amphibians plus the concomitant linkage between the cutaneous bacterial microbiome and host health drive the need to utilize these taxa as a model for microbiome assessment. The current focus of amphibian bacterial microbiota studies resides mostly within the realm of chytridiomycosis resistance research [22, 69]. Susceptibility to chytridiomycosis has been found to be mediated in part by members of the skin bacterial communities, and current conservation approaches to mediate disease include probiotic applications [70]. However, we have shown that high-throughput techniques can be employed to assess the bacterial microbiota between groups of amphibians which are not related to devastating diseases.
References
Federici E, Rossi R, Fidati L, Paracucchi R, Scargetta S, Montalbani E, Franzetti A, La Porta G, Fagotti A, Simonceli F, Cenci G, Di Rosa I (2015) Characterization of the skin microbiota in Italian stream frogs (Rana italica) infected and uninfected by a cutaneous parasitic disease. Microbes Environ 30:262–269
Loudon AH, Holland JA, Umile TP, Burzynski EA, Minbiole KPC, Harris RN (2014) Interactions between amphibians’ symbiotic bacteria cause the production of emergent anti-fungal metabolites. Front Microbiol 5:441
Vredenburg VT, Briggs CJ, Harris RN (2011) Host-pathogen dynamics of amphibian chytridiomycosis: the role of the skin microbiome in health and disease. Fungal diseases: an emerging challenge to human, animal, and plant health. National Academic Press, Washington D.C., pp 342–355
Barker CJ, Gillett A, Polkinghorne A, Timms P (2013) Investigation of the koala (Phascolarctos cinereus) hindgut microbiome via 16S pyrosequencing. Vet Microbiol 167:554–564
Kueneman JG, Parfrey LW, Woodhams DC, Archer HM, Knight R, McKenzie VJ (2014) The amphibian skin-associated microbiome across species, space and life history stages. Mol Ecol 23:1238–1250
Shabbir MZ, Park J, Muhammad K, Rabbani M, Rana MY, Harvill ET (2014) Culture independent analysis of respiratory microbiome of houbara bustard (Chlamydotis undulata) revealed organisms of public health significance. Int J Agric Biol 16:222–226
Ishak HD, Plowes R, Sen R, Kellner K, Meyer E, Estrada DA, Dowd SE, Mueller UG (2011) Bacterial diversity in Solenopsis invicta and Solenopsis geminata ant colonies characterized by 16S amplicon 454 pyrosequencing. Microb Ecol 61:821–831
Daszak P, Cunningham AA, Hyatt AD (2003) Infectious disease and amphibian population declines. Divers Distrib 9:141–150
Nickerson MA, Mays CE (1973) The hellbenders: North American ‘giant salamanders’. Milwaukee Public Museum, Milwaukee
Foster RL, McMillan AM, Roblee KJ (2009) Population status of hellbender salamanders (Cryptobranchus alleganiensis) in the Allegheny River drainage of New York State. J Herpetol 43:579–588
Hecht-Kardasz KA, Nickerson MA, Freake M, Colclough P (2012) Population structure of the hellbender (Cryptobranchus alleganiensis) in a Great Smoky Mountains stream. Bull Fla Museum Nat Hist 51:227
Briggler J, Utrup J, Davidson C, Humphries J, Groves J, Johnson T, Ettling J, Wanner M, Traylor-Holzer K, Reed D, Lindgren V, Byers O (2007) Hellbender population and habitat viability assessment: final report. IUCN/SSC Conservation Breeding Specialist Group, Apple Valley
Burgmeier NG, Unger SD, Sutton TM, Williams RN (2011) Population status of the eastern hellbender (Cryptobranchus alleganiensis alleganiensis) in Indiana. J Herpetol 45:195–201
Federal Register (2011) Endangered and threatened wildlife and plants; endangered status for the Ozark hellbender salamander. Fed Commun Comm 76:61956–61978
Wheeler BA, Prosen E, Mathis A, Wilkinson RF (2003) Population declines of a long-lived salamander: a 20+-year study of hellbenders, Cryptobranchus alleganiensis. Biol Conserv 109:151–156
Hiler WR, Wheeler BA, Trauth SE (2005) Abnormalities in the Ozark hellbender (Cryptobranchus alleganiensis bishopi) in Arkansas: a comparison between two rivers with a historical perspective. J Arkansas Acad Sci 59:88–94
Nickerson CA, Ott CM, Castro SL, Garcia VM, Molina TC, Briggler JT, Pitt AL, Tavano JJ, Byram JK, Barrila J et al (2011) Evaluation of microorganisms cultured from injured and repressed tissue regeneration sites in endangered giant aquatic Ozark hellbender salamanders. PLoS One 6, e28906
Wheeler BA, McCallum ML, Trauth SE (2002) Abnormalities in the Ozark hellbender, Cryptobranchus alleganiensis bishopi. J Arkansas Acad Sci 56:250–252
Irwin K (2008) Ozark hellbender long-term monitoring SWG project. Arkansas Game and Fish Commission, Benton
Hill GT, Mitkowski NA, Aldrich-Wolfe L, Emele LR, Jurkonie DD, Ficke A, Maldonado-Ramirez S, Lynch ST, Nelson EB (2000) Methods for assessing the composition and diversity of soil microbial communities. Appl Soil Ecol 15:25–36
Pace NR (1997) A molecular view of microbial diversity and the biosphere. Science 276:734–740
Jani AJ, Briggs CJ (2014) The pathogen Batrachochytrium dendrobatidis disturbs the frog skin microbiome during a natural epidemic and experimental infection. Proc Natl Acad Sci U S A 111:E5049–E5058
Fitzpatrick BM, Allison AL (2014) Similarity and differentiation between bacteria associated with skin of salamanders (Plethodon jordani) and free-living assemblages. FEMS Microbiol Ecol 88:482–494
Loudon AH, Woodhams DC, Parfrey LW, Archer H, Knight R, McKenzie V, Harris RN (2014) Microbial community dynamics and effect of environmental microbial reservoirs on red-backed salamanders (Plethodon cinereus). ISME J 8:830–840
Woodhams DC, Brandt H, Baumgartner S, Kielgast J, Küpfer E, Tobler U, Davis LR, Schmidt BR, Bel C, Hodel S, Knight R, McKenzie V (2014) Interacting symbionts and immunity in the amphibian skin mucosome predict disease risk and probiotic effectiveness. PLoS One 9, e96375
Caporaso JG, Lauber CL, Walters WA, Berg-Lyons D, Lozupone CA, Turnbaugh PJ, Fierer N, Knight R (2011) Global patterns of 16S rRNA diversity at a depth of millions of sequences per sample. Proc Natl Acad Sci U S A 108:4516–4522
Culp CE, Iii JOF, Belden LK (2007) Identification of the natubal bacterial microflora on the skin of eastern newts, bullfrog tadpoles and redback salamanders. Herpetologica 63:66–71
Fierer N, Hamady M, Lauber CL, Knight R (2008) The influence of sex, handedness, and washing on the diversity of hand surface bacteria. Proc Natl Acad Sci U S A 105:17994–17999
Bolger D, Lohse M, Usadel B (2014) Trimmomatic: a flexible trimmer for illumina sequence data. Bioinformatics 30:2114–2120
Caporaso JG, Kuczynski J, Stombaugh J, Bittinger K, Bushman FD, Costello EK, Fierer N, Pena AG, Goodrich JK, Gordon JI et al (2010) QIIME allows analysis of high-throughput community sequencing data. Nat Methods 7:335–336
Edgar RC (2010) Search and clustering orders of magnitude faster than BLAST. Bioinformatics 26:2460–2461
Bokulich NA, Subramanian S, Faith JJ, Gevers D, Gordon JI, Knight R, Mills DA, Caporaso JG (2013) Quality-filtering vastly improves diversity estimates from Illumina amplicon sequencing. Nat Methods 10:57–59
DeSantis TZ, Hugenholtz P, Larsen N, Rojas M, Brodie EL, Keller K, Huber T, Dalevi D, Hu P, Andersen GL (2006) Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB. Appl Environ Microbiol 72:5069–5072
Caporaso JG, Bittinger K, Bushman FD, DeSantis TZ, Andersen GL, Knight R (2010) PyNAST: a flexible tool for aligning sequences to a template alignment. Bioinformatics 26:266–267
Wang Q, Garrity GM, Tiedje JM, Cole JR (2007) Naïve Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy. Appl Environ Microbiol 73:5261–5267
Price MN, Dehal PS, Arkin AP (2010) FastTree 2—approximately maximum-likelihood trees for large alignments. PLoS One 5, e9490
Chao A (1984) Nonparametric estimation of the number of classes in a population. Scand J Statist 11:265–270
Faith DP, Baker AM (2006) Phylogenetic diversity (PD) and biodiversity conservation: some bioinformatics challenges. Evol Bioinform Online 2:121–128
Shannon CE, Weaver W (1949) The mathematical theory of communication. University of Illinois Press, Champaign
Bates D, Maechler M, Bolker BM, Walker SC (2015) Fitting linear mixed-effects models using lme4. J Stat Softw 67:1–48
Lozupone C, Lladser ME, Knights D, Stombaugh J, Knight R (2011) UniFrac: an effective distance metric for microbial community comparison. ISME J 5:169–172
Rees GN, Baldwin DS, Watson GO, Perryman S, Nielsen DL (2004) Ordination and significance testing of microbial community composition derived from terminal restriction fragment length polymorphisms: application of multivariate statistics. Anton Leeuw Int J G 86:339–347
Okasen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O’Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoencs E, Wagner H (2016) Vegan: community ecology package. R package version 2.4-0
Hammer O, Harper DAT, Ryan PD (2001) PAST: paleontological statistics software package for education and data analysis. Palaeontol Electron 4:9
Longo AV, Savage AE, Hewson I, Zamudio KR (2015) Seasonal and ontogenetic variation of skin microbial communities and relationships to natural disease dynamics in declining amphibians. Roy Soc Open Sci 2:140377
Segata N, Izard J, Waldron L, Gevers D, Miropolsky L, Garrett WS, Huttenhower C (2011) Metagenomic biomarker discovery and explanation. Genome Biol 12:R60
Cole JR, Wang Q, Fish JA, Chai B, McGarrell DM, Sun Y, Brown CT, Porras-Alfaro A, Kuske CR, Tiedje JM (2014) Ribosomal Database Project: data and tools for high throughput rRNA analysis. Nucleic Acids Res 42:D633–D642
Atlas RM (2010) Handbook of microbiological media. CRC Press, Taylor and Francis Group, Boca Raton
McKenzie VJ, Bowers RM, Fierer N, Knight R, Lauber CL (2012) Co-habiting amphibian species harbor unique skin bacterial communities in wild populations. ISME J 6:588–596
Bataille A, Lee-Cruz L, Tripathi B, Kim H, Waldman B (2016) Microbiome variation across amphibian skin regions: implications for chytridiomycosis mitigation efforts. Microb Ecol 71:221–232
Walke JB, Becker MH, Loftus SC, House LL, Cormier G, Jensen RV, Belden LK (2014) Amphibian skin may select for rare environmental microbes. ISME J 8:2207–2217
Crowhurst RS, Faries KM, Collantes J, Briggler JT, Koppelman JB, Eggert LS (2011) Genetic relationships of hellbenders in the Ozark highlands of Missouri and conservation implications for the Ozark subspecies (Cryptobranchus alleganiensis bishopi). Conserv Genet 12:637–646
Feist SM, Briggler JT, Koppelman JB, Eggert LS (2014) Within-river gene flow in the hellbender (Cryptobranchus alleganiensis) and implications for restorative release. Conserv Genet 15:953–966
Zilber-Rosenberg I, Rosenberg E (2008) Role of microorganisms in the evolution of animals and plants: the hologenome theory of evolution. FEMS Microbiol Rev 32:723–735
Unger SD, Rhodes OE, Sutton TM, Williams RN (2013) Population genetics of the Eastern Hellbender (Cryptobranchus alleganiensis alleganiensis) across multiple spatial scales. PLoS One 8, e74180
Olson ZH, Burgmeier NG, Zollner PA, Williams RN (2013) Survival estimates for adult Eastern Hellbenders and their utility for conservation. J Herpetol 47:71–74
Bodinof CM, Briggler JT, Junge RE, Mong T, Beringer J, Wanner MD, Schuette CD, Ettling J, Millspaugh JJ (2012) Survival and body condition of captive-reared juvenile Ozark hellbenders (Cryptobranchus alleganiensis bishopi) following translocation to the wild. Copeia 2012:150–159
Becker MH, Richards-Zawacki CL, Gratwicke B, Belden LK (2014) The effect of captivity on the cutaneous bacterial community of the critically endangered Panamanian golden frog (Atelopus zeteki). Biol Conserv 176:199–206
Merrifield DL, Rodiles A (2015) The fish microbiome and its interactions with mucosal tissues. In: Mucosal health in aquaculture. Academic, Oxford, UK, pp 273–295
Pfingsten RA (1989) The status and distribution of the hellbender, Cryptobranchus alleganiensis, in Ohio. Ohio J Sci 89:3
Lauer A, Simon MA, Banning JL, Lam BA, Harris RN (2008) Diversity of cutaneous bacteria with antifungal activity isolated from female four-toed salamanders. ISME J 2:145–157
Michaels CJ, Antwis RE, Preziosi RF (2014) Impact of plant cover on fitness and behavioural traits of captive red-eyed tree frogs (Agalychnis callidryas). PLoS One 9(4):295207
Roth T, Foley J, Worth J, Piovia-Scott J, Pope K, Lawler S (2013) Bacterial flora on Cascades frogs in the Klamath mountains of California. Comp Immunol Microb 36:591–598
Lam BA, Walke JB, Vredenburg VT, Harris RN (2010) Proportion of individuals with anti-Batrachochytrium dendrobatidis skin bacteria is associated with population persistence in the frog Rana muscosa. Biol Conserv 143:529–531
Olson ME, Gard S, Brown M, Hampton R, Morck DW (1992) Flavobacterium indologenes infection in leopard frogs. JAVMA J Am Vet Med A 201:1766–1770
Taylor SK, Williams ES, Thorne ET, Mills KW, Withers DI, Pier AC (1999) Causes of mortality of the Wyoming toad. J Wildl Dis 35:49–57
Densmore CL, Green DE (2007) Diseases of amphibians. ILAR J 48:235–254
Geng Y, Wang KY, Zhou ZY, Li CW, Wang J, He M, Yin ZQ, Lai WM (2011) First report of a ranavirus associated with morbidity and mortality in farmed Chinese giant salamanders (Andrias davidianus). J Comp Pathol 145:95–102
Becker MH, Harris RN (2010) Cutaneous bacteria of the redback salamander prevent morbidity associated with a lethal disease. PLoS One 5, e10957
Bletz MC, Loudon AH, Becker MH, Bell SC, Woodhams DC, Minbiole KPC, Harris RN (2013) Mitigating amphibian chytridiomycosis with bioaugmentation: characteristics of effective probiotics and strategies for their selection and use. Ecol Lett 16:807–820
Acknowledgments
We would like to thank members of the Williams lab for assistance in revising this document. Special thanks also go to Jyothi Thimmapuram from the Purdue Bioinformatics Core, Phillip San Miguel and Paul Parker from the Purdue Genomics Core for assistance in project design, Bart Kraus for assistance in field collection, and Ardith Wang for assistance in sequencing preparation. Special thanks to the Missouri Department of Conservation for their interest and support of this project.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Funding
Funding for this study was provided by Purdue University.
Conflict of Interest
The authors declare that they have no conflict of interest.
Ethical Approval
All applicable international, national, and/or institutional guidelines for the care and use of animals were followed.
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
ESM 1
(DOCX 187 kb)
Rights and permissions
About this article
Cite this article
Hernández-Gómez, O., Kimble, S.J.A., Briggler, J.T. et al. Characterization of the Cutaneous Bacterial Communities of Two Giant Salamander Subspecies. Microb Ecol 73, 445–454 (2017). https://doi.org/10.1007/s00248-016-0859-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-016-0859-9