Abstract
We report the first investigation of a deep subpermafrost microbial ecosystem, a terrestrial analog for the Martian subsurface. Our multidisciplinary team analyzed fracture water collected at 890 and 1,130 m depths beneath a 540-m-thick permafrost layer at the Lupin Au mine (Nunavut, Canada). 14C, 3H, and noble gas isotope analyses suggest that the Na–Ca–Cl, suboxic, fracture water represents a mixture of geologically ancient brine, ~25-kyr-old, meteoric water and a minor modern talik-water component. Microbial planktonic concentrations were ~103 cells mL−1. Analysis of the 16S rRNA gene from extracted DNA and enrichment cultures revealed 42 unique operational taxonomic units in 11 genera with Desulfosporosinus, Halothiobacillus, and Pseudomonas representing the most prominent phylotypes and failed to detect Archaea. The abundance of terminally branched and midchain-branched saturated fatty acids (5 to 15 mol%) was consistent with the abundance of Gram-positive bacteria in the clone libraries. Geochemical data, the ubiquinone (UQ) abundance (3 to 11 mol%), and the presence of both aerobic and anaerobic bacteria indicated that the environment was suboxic, not anoxic. Stable sulfur isotope analyses of the fracture water detected the presence of microbial sulfate reduction, and analyses of the vein-filling pyrite indicated that it was in isotopic equilibrium with the dissolved sulfide. Free energy calculations revealed that sulfate reduction and sulfide oxidation via denitrification and not methanogenesis were the most thermodynamically viable consistent with the principal metabolisms inferred from the 16S rRNA community composition and with CH4 isotopic compositions. The sulfate-reducing bacteria most likely colonized the subsurface during the Pleistocene or earlier, whereas aerobic bacteria may have entered the fracture water networks either during deglaciation prior to permafrost formation 9,000 years ago or from the nearby talik through the hydrologic gradient created during mine dewatering. Although the absence of methanogens from this subsurface ecosystem is somewhat surprising, it may be attributable to an energy bottleneck that restricts their migration from surface permafrost deposits where they are frequently reported. These results have implications for the biological origin of CH4 on Mars.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Interest in the possibility of subsurface life on Mars has been fueled recently by the detection of Martian gullies suggesting modern transient groundwater expulsion [1, 2] and Martian atmospheric CH4, which is believed to originate from active, nonvolcanic, subsurface, potentially biological emissions [3–6]. Because of the lethal UV irradiation at the Martian surface [7, 8] and the fact that water at or near the Martian surface is locked in permafrost, most investigations have focused on readily accessible terrestrial permafrost environments and the microorganisms inhabiting them [9–15]. Based upon heat flow models for Mars, Clifford and Parker [16] hypothesized that water, if present, would be liquid at kilometers depth beneath the Martian permafrost, and Boston et al. [17] speculated that chemoautolithotrophic communities could exist in this realm. Yet little is known about the microbial community structure of analogous subpermafrost environments on Earth. The Canadian subpermafrost environment serves as an excellent analog of the Martian deep subsurface because of its low temperatures, overlying permafrost that is hundreds of meters thick, and very low organic content and saline water [18]. Our only glimpse of this environment comes from investigations of perennial saline springs on Axel Heiberg Island in the Canadian Arctic that have yielded bacterial communities dominated by aerobic sulfur-metabolizing bacteria, such as Thiomicrospira arctica and Thiomicrospira psychrophilia [19, 20], but until this study, no direct sampling of subpermafrost environments has been reported.
We collected and analyzed fracture water from the Lupin Au mine, which is located in the Canadian Shield roughly 80 km south of the Arctic Circle (65°48′ N, 111°15′ W) and sits within a continuous permafrost zone that extends from the base of a 1-m-thick active zone to a depth of roughly 540 m [21]. The mine is located within the Contwoyto Formation, which is a low porosity and low hydraulic conductivity, Archaean metasedimentary formation comprised of metagreywacke and slate interbedded with a Au-bearing, amphibolite-grade, banded-iron formation [22]. The Lupin Au mine also lies adjacent to Contwoyto Lake, which is deep enough to prevent ice freezing to the bottom during the winter and as a result maintains an unfrozen hydraulic conduit or “talik” through the permafrost (Fig. 1). A combination of culture-dependent (direct plating and enrichment) and culture-independent (DNA and membrane lipid analyses) approaches were used to characterize the microbial community structure and abundance in boreholes tapping the water-bearing fractures, while geochemical and isotopic data were utilized to constrain the principal metabolic processes of these microbial communities and the origin of the fracture water.
Cross-section showing the spatial relationship between the mine and the assumed talik below Contwoyto Lake (the mine workings are delineated down to about the 1,400-m level by the dashed contours). Additionally, the projections of the research boreholes at 570, 890, and 1,130 m levels extending closest to the talik are shown. The dashed vertical line indicates the approximate location of the shoreline. The large arrow indicates the approximate path of talik water into the mine as a result of dewatering. The cross-sectional view is to the NE (302°; adapted from Ruskeeniemi et al. [24])
Materials and Methods
Site Description
The Lupin Au mine was a largely dry mine with water emanating predominately from just two long horizontal exploration boreholes: one located at the 1,130-m-deep level, 1130-192, and one located at the 890-m-deep level, 890-188, both of which discharged water at a rate of 6 to 8 L min−1 [21]. Other slowly leaking boreholes existed at the 1,130 level, but with the exception of 1130-64 and 1130-175, all had been plugged after being drilled in 1992–1993. These boreholes extend for 400 to 500+ m to the east behind the rock face (Fig. 1), well beyond the 2- to 3-m-deep tunnel damage zone [23]. Borehole camera surveys revealed the presence of open fractures with submillimeter to millimeter apertures and gas typically emanating from smaller fracture sets. The open flowing boreholes were sealed with margot-type compression plugs set at 3 m behind the rock face and fitted with valves and pressure gauges in December 2002. The other boreholes were rehabilitated with the same type of plugs in February 2003. Observations on the increase in water pressure after sealing and the variation in salinity over time indicated that the intersected fractures were not well connected hydrologically [24] between boreholes even though the boreholes did intersect a north–south trending fracture network and that mixing between two or three formerly independent fractures was most likely occurring within some of the boreholes. Borehole 1130-267, drilled at the end of a newly excavated exploration tunnel in 2003 to a distance of several hundred meters eastward, was sealed with the margot-type device soon after completion. Borehole 1130-273, which was completed the same year and was drilled northward toward the Contwoyto Lake, was treated in the same way. The high hydraulic head measured from borehole 1130-273 immediately after sealing recorded only a minor decline in the groundwater table. These two boreholes were the least exposed to mine air. Isolation of single fractures within a borehole was not feasible in this study.
Sampling Methods
During May 2004, March 2005, and October 2005, water samples were collected from boreholes 890–188, 1130–197, 1130–192, 1130–267, and 1130–273. Samples were also collected from 1130–64, a very slowly dripping open ceiling borehole with the lowest salinity of all the borehole water, and from 1130–175, an open, flowing, and bubbling borehole located in the floor (referred to by the miners as the Fountain of Youth) and partially filled with debris that yielded very saline water with the highest recorded temperature (13°C) and from very slowing dripping seeps at the 390-, 550-, and 570-m levels. The pH, temperature, and conductivity were measured with a Hanna Instruments portable combination tester with a high range conductivity probe (HI 98130). The Eh was measured with a Hanna Instruments portable, Pt electrode, and ORP tester (HI 98201), which is calibrated against a Ag/AgCl internal redox couple. The O2, Fe, and H2S were measured with Chemetrics CHEMets colorimetric kits. Samples for geochemical analyses were collected and preserved following the procedures outlined in Onstott et al. [25]. Water was collected for cell counts in sterile 20-mL headspace vials sealed with a crimp septum and flushed with ultrapure sterile N2. To maintain anoxic conditions, the vials were filled using two needles: one connected by sterile tubing to the borehole valve and another to allow N2 to escape while water flowed in the vial. Two milliliters of 0.2-μm-filtered formalin was added as fixative. Dissolved gas (H2, N2, He, CO, and C1–4) samples were collected following the procedure of Ward et al. [26]. Water samples for aerobic enrichments were collected in sterile 10-mL serum vials. Water samples for anaerobic enrichments were directly injected into 30-mL pressure tubes preloaded with 1 mL of media and 2 bars of 80% N2 and 20% CO2 headspace (more details on media are presented below).
Using autoclaved tubing and brass fittings connected directly to the borehole valves, water was filtered with two Cole Parmer in-line, capsule, 0.3-μm filters (EW-02909-60, EW-02908-90) at a rate of 150 to 300 mL min−1 for 3 to 6 days. The total volume of water filtered varied from 300 to 800 L. The in-line capsule filters were shipped to Yellowknife, NWT, Canada; here, they were frozen on dry ice and then shipped to Princeton University and the University of Tennessee where they were stored at −80°C. DNA was extracted from the EW-02909-60 filters at Princeton University. Lipids were extracted from the EW-02908-90 filters at the University of Tennessee.
Cell Enumeration
Cells were enumerated at the University of Rhode Island using epifluorescence microscopy according to the method of Noble and Fuhrman [27]. Briefly, 15 mL of water were filtered onto a 0.02-μm aluminum oxide filter, stained with SYBR GREEN, and counted under UV excitation. At least 250 fields of view were counted for each sample. The cell concentration was calculated from the area of the filter, the number and area of the fields of view counted, and the volume of water filtered. The standard deviation was calculated assuming a Poisson distribution [28].
Enrichment Culturing
Borehole water was diluted in low nutrient marine broth or 0.1× tryptic soy broth (TSB), spread on plates, and incubated at 0°C, 4°C, or 20°C aerobically for 2 months at Michigan State University. Subsequently, colonies were screened for unique morphologies. Several different colonies of each morphotype were streaked on new plates for isolation and further characterization. DNA extractions were performed with Qiagen DNEasy Blood & Tissue Kit. The complete 16S rRNA gene was sequenced at the Michigan State University Research Technology Support Facility with primers 8f, 357f, 926f, 1114f, 519r, 907r, 1392r, and 1541r [29]. Salt tolerance was tested in 10 mL of 0.1× TSB with NaCl added to a concentration of 0.05%, 5%, 7.5%, 10%, 12.5%, and 15% (w/v) at 22°C inoculated with 60 μL of cultures and incubated for 60 days with shaking at 200 rpm. Growth temperature range was tested on 0.1× tryptic soy agar or marine agar with incubation at 0°C, 4°C, 10°C, 22°C, 30°C, 37°C, and 42°C for up to 2 months.
Anaerobic media for Fe(III) reducers, sulfate reducers (SRB), fermenters, acetogens, and methanogens were prepared using previously published recipes [30, 31]. The media for fermenting bacteria utilized a dilute 0.3-g L−1 combination of yeast extract, glucose, tryptone, and peptone as C and N sources. Hydrous ferric oxide was the electron acceptor for the Fe(III)-reducing media and lactate, acetate, and pyruvate were utilized for the electron donors with a minor amount of yeast extract added for additional nutrients. The SRB media used the same combination of electron donors as the Fe(III)-reducing media. One type of methanogenic media used acetate and methanol for the principal carbon sources along with a minor amount of yeast extract. For acetogens and autotrophic methanogens, H2 was the electron donor and CO2 the electron acceptor (80:20% H2/CO2 headspace) and a minor amount of yeast extract was provided as an additional nutrient source. The anaerobic media were dispensed and sealed in pressure tubes under an 80% N2 and 20% CO2 headspace (with the exception of the acetogen and autotrophic methanogen media) and autoclaved at 121°C for 35 min. Ten milliliters of H2 was added as an additional electron donor to half of the fermenting, Fe(III)-reducing, and SRB media. One milliliter of a sterile 25% NaCl solution was injected into all media (10 mL) in the field after we had measured the electrical conductivity of the borehole water and confirmed its salinity. In the field 0.3 mL of sterile, 2.5% cysteine-sulfide solution was injected into the methanogenic, acetogenic, or fermenting bacteria media (10 mL) when necessary to remove any residual dissolved O2 either before or after inoculation of the media. For the SRB media, 0.2 mL of sterile 2.5% cysteine-HCl solution was used for this purpose. In all cases, these reductants produced colorless media by the second day of incubation. The inoculated anaerobic media were returned to Princeton University and the University of Tennessee in coolers with blue ice.
At Princeton University and the University of Tennessee, the anaerobic enrichments were incubated at 4°C for several months and positive enrichments were subjected to 16S rRNA analyses. The procedure used at Princeton University for one of the Clostridia enrichments is the same as that described below. At the University of Tennessee, the DNA was extracted from the enrichments with the Mo Bio UltraClean DNA extraction kit, eluted into 10 mM Tris–ethylenediaminetetraacetic acid and stored at −20°C. Polymerase chain reaction (PCR) was performed on these extracts using 341f and 519r followed by denaturing gradient gel electrophoresis (DGGE) and sequencing.
16S rRNA Gene Analyses
Filter housings were cracked open with a hammer wrapped in baked Al foil and filters were removed with heat-sterilized tweezers beneath a high-efficiency particulate air (HEPA) filtered laminar flow hood. DNA extractions were performed with Mo Bio UltraClean soil DNA extraction kit. DNA was eluted into 10 mM Tris-HCl and stored at −20°C (working) or −80°C (storage).
PCR was performed on all extracts using either Takara Ex Taq or Roche Diagnostics Taq. The recipe consisted of 1/10 volume PCR buffer, 200 μM per deoxyribonucleotide triphosphate, 400 nM primer, and 1–2 μL template. Cycling consisted of denaturation for 2 min at 95°C, annealing 45 s at 51°C, extension 2 min at 72°C for 30–35 cycles, and final extension for 5 min at 72°C. The following primers were used: a21f [32], b27f [33], and u1492r [34] on a Stratagene Robocycler 96 or a Perkin-Elmer GeneAmp 9600. Amplicon was verified and assessed for size with 2% agarose gel electrophoresis for 30 min at 100 V followed by UV visualization with ethidium bromide. Amplicon not exposed to UV or EtBr was purified by Qiagen PCR Purification Kit. The 16S rRNA amplicons obtained from environmental DNAs and enrichments were cloned with an Invitrogen TOPO-TA Cloning kit. Clones were verified with M13 PCR using the above conditions and visualized with 2% gel electrophoresis as above. Clones examined with restriction fragment length polymorphism were treated with RsaI at 37°C for 2 to 3 h and denatured at 65°C for 5 min followed by examination and also visualization by 2% agarose gel electrophoresis as above. Clones selected to be sequenced were extracted with Qiagen Spin Miniprep 96 kits and sent to Florida State University, the University of Wisconsin Madison, or GeneWiz, Inc. (South Plainsfield, NJ, USA) for sequencing. PCR of just the reagents failed to yield any detectable DNA product. PCR of 105 cells of Gram-positive Desulfotomaculum putei yielded readily visible, but weak, bands on the electrophoretic agarose gel confirming the sensitivity of the procedure.
Multiple displacement amplification (MDA) of DNA extracts, D. putei, and negative controls was performed with a Qiagen Repli-G kit. All items to be placed in the UV, HEPA filtered laminar flow through hood, and lab coats to be worn during the procedure were autoclaved immediately before use. Coat sleeves were fixed to latex gloves with laboratory tape to limit skin cell contamination. The hood was cleaned with Ambion DNA Zap followed by two washes with 70% ethanol in Millipore filtered D.I. water and irradiated with UV for 45 min prior to use. MDA reactions were mixed using the 5-μL template DNA protocol and allowed to react at 30°C for 16 h in a Perkin-Elmer 9600 followed by a 5-min denaturation at 65°C. MDA products were queried with 16S rRNA PCR as listed above for DNA extracts and with a universal primer pair, e341f (5′-CCT ACG GGI GGC UGC A-3′) and e926r (5′CCG ICI ATT IIT TTI AGT TT-3′) [35]. MDA products of negative controls failed to produce archaeal 16S rRNA PCR products even though they yielded ~1 μg of DNA and failed to produce bacterial 16S rRNA PCR products with the exception of that from the human skin bacterium Propionibacterium acnes. MDA product from ~104 cells of DNA from D. putei yielded readily visible bands on an electrophoretic gel. A nested procedure that has met with limited success in other studies [36] was not employed in this study.
Possible chimeras and their breakpoints were identified with Bellerophon [37] and Ribosomal Database Project (RDP) ChimeraCheck [38]. The closest matches available in GenBank were identified by using BLASTN [39]. In addition, phylogenetic identity and sequence similarity was determined using the RDP’s Seqmatch which calculates a sequence similarity as the percent sequence identity over all pairwise comparable positions [38]. Operational taxonomic units (OTUs) were distinguished at the ≥99% identity level between environmental and enrichment sequences. Trees were constructed using the RDP's Tree Builder [38]. Briefly, sequences were first aligned with RDP's aligner and a distance matrix was generated with the Jukes-Cantor corrected distance model (using only alignment model positions and ignoring alignment inserts). A tree was then created using Weighbor.
Lipid Analyses
Filtered fracture water cartridges for membrane lipid analyses were stored at −80°C until analyzed at the University of Tennessee. The filter membrane was removed from the cartridge by placing the cartridge in a sterile whirl-Pak bag and breaking its nylon exterior. The filter was transferred to a muffled glass centrifuge tube using a solvent-rinsed forceps. The total lipids were extracted using methanol/chloroform/water buffer (2:1:0.8, v/v/v) and subsequently fractionated on a silicic acid column with only the polar lipids then transesterified into phospholipid fatty acid (PLFA) methyl esters [40]. The PLFA methyl esters were separated, quantified, and identified by gas chromatography (GC)–mass spectrometry [40]. Archaeal lipids were not analyzed.
Quinones were determined directly from the neutral lipid fraction collected during fractionation of the total lipids [40]. Quinones were analyzed using high-performance liquid chromatography coupled to atmospheric pressure photo-ionization or atmospheric pressure chemical ionization tandem mass spectrometry [41, 42].
Geochemical Analyses
Cations were analyzed by an Optima 4300 DV inductively coupled plasma with atomic emission spectrometer (Perkin-Elmer, Wellesley, MA, USA), and inorganic anions, acetate, formate, and propionate were measured by a DX-320 ion chromatograph (Dionex, Sunnyvale, CA, USA) at Princeton University. Total organic carbon (TOC) samples were acidified using sulfuric acid. The TOC samples were split and filtered with 0.45 μm PTFE filters to obtain the DOC sample. The TOC and DOC samples were combusted at 680°C and the CO2 analyzed by a NDIR detector (Tekmar Achrmann Apollo 9000) at Shaw Environmental.
One-liter amber bottles fixed with 1 mL of saturated HgCl2 were used to collect water samples for organic analyses. A 125-mL volume from these bottles was extracted with 75 mL of a 2:1 chloroform/methanol mixture and the organic phase was collected. The aqueous phase was extracted two more times with 10 mL aliquots of chloroform. The chloroform phase was combined with the original extract and then concentrated down to 1 mL in a Zymark TurboVac II under a stream of N2 at 40°C. The organic phase (2.5 µL split less injection) was analyzed by GC/flame ionization detector at Shaw Environmental using a Varian 3800 GC equipped with a laminar cup splitter inlet liner and a Stabilwax column (30 m, 0.32 mm ID, 1.00 μm df). The injector temperature was set at 260°C and the detector at 290°C. The column temperature (165°C) was held for 4.0 min and then ramped to 250°C at 12°C min−1 and held for 9.0 min using a constant column flow rate of 3.0 mL min−1.
The dissolved inorganic carbon (DIC) sample was filtered, acidified with phosphoric acid, sparged with carbon free air, and the resulting CO2 is analyzed by the NDIR detector (LI-6252, LiCOR Biosciences, Lincoln, NE, USA). Ammonia analyses were performed at ACTLABS utilizing an ion-specific electrode (Metrohm 789 Robotic System with PHOENIX gas-sensing electrode) with a detection limit of 1.2 μM.
Dissolved gas (H2, N2, He, CO, and C1–4) samples were measured with either a reduced gas analyzer (CO and H2; Trace Analytical, Sparks, MD, USA) [25] or a gas chromatograph equipped with a flame ionization detector (C1–4) and a thermal conductivity detector (N2, H2, and He) [26]. The concentrations of dissolved gases were derived from the gas volume abundance, the ratio of water to gas flow rates, and Henry’s law constants following the procedure of Andrews and Wilson [43].
Geochemical Modeling
To determine if the inferences drawn about the metabolic niches of these microorganisms are valid, the Gibbs free energy (ΔG) and free energy fluence (FEF) [44], for the relevant reactions within the system, were calculated for fracture water samples 890-188 and 1130-192 which span the range of temperature, pH, dissolved gas concentrations, and salinity. In the case of hematite and magnetite reduction, the electron donor diffusivity was multiplied by the 0.5% porosity of the rock strata [21] when calculating the FEF. The ΔG and FEF for HS− oxidation by O2 were calculated assuming the 10−7 M detection limit for O2 concentration and hence are maximum estimates.
Mineral–water–microbial reactions were modeled with the React module of the Geochemist’s Workbench (GWB) Standard Version 7.0 software package (University of Illinois). Rates for radiolytic reactions were calculated by combining pore water dosage rates estimated using the chemical and physical properties of the Contwoyto Formation and the formula discussed in Lin et al. [45, 46] with radiolytic reactions and experimentally determined rates for H, O, and Cl species published by Bjergbakke et al. [47, 48], for C species published by Cai et al. [49], and for N species reported by Raymond [50] and simultaneously solving these equations for a water chemistry corresponding to that of the Lupin fracture water using Kineticus version 2 (Copyright 1995–2002 James C. Ianni). The radiolytic H2, H2O2, O2, HO −2 , H+, formate, and NO −3 production rates were then utilized in GWB react models. In the GWB react code, the oxidizing species, H2O2, O2, and HO −2 , were reacted with pyrrhotite to produce hematite and SO 2−4 .
Sulfur Isotope Analyses
Unfiltered borehole water was drawn directly into 60 mL polycarbonate luer-lock syringes that previously were washed and preloaded with 10 mL of 1.5 M CdCl2 solution in order to fix HS− and H2S as CdS. Three syringes were collected at each borehole, providing a total of 150 mL of groundwater for sulfur isotopic analyses. Samples were refrigerated during storage and shipped to Indiana University in insulated containers with ice packs.
CdS precipitate was separated from reacted groundwater by vacuum filtration using pre-annealed PALL Tissuquartz membrane filters. The resulting filtered groundwater was saved for recovery of dissolved SO 2−4 . Recovered solids were dried at 60°C in an oven overnight.
Dissolved SO 2−4 was recovered from groundwater samples by precipitation of BaSO4. Concentrated HNO3 was added in drops to reduce water pH below 5 for all filtered samples. After acidification, 0.4 M BaCl2 was added in sufficient volume to give five times excess Ba concentration compared to the estimated concentration of SO 2−4 in the sample. Samples were heated to 80°C without stirring for 5 to 6 h in order to precipitate BaSO4. Resulting precipitates were separated from supernatant by centrifugation with several rinses of D.I. water and then transferred by pipette to a small glass vial and dried at 60°C for 12 h.
Sequential extraction of sulfur species from rock and minerals samples were done following the methods described in Bruchert and Pratt [51] and Lefticariu et al. [52]. S0 (ES), acid volatile sulfur (AVS), and chrome-reducible sulfur (CRS) were purified in the form of AgS for isotopic measurement.
Sulfur isotopic compositions were determined with a Finnigan MAT 252 mass spectrometer using continuous flow from a CE Instruments EA 110 elemental analyzer inlet. Sulfur isotopic compositions are expressed in standard delta (δ) notation given by δ34S = [(R sample/R standard − 1) × 1,000], where R = 34S/32S. Sulfur isotopic values are expressed as per mil difference from Vienna Canyon Diablo troilite (VCDT). Accuracy and reproducibility were assessed using replicate samples and using three standards to normalize measurements (NBS 127 BaSO4 at 20 ± 0.5‰, Indiana EMR chalcopyrite at 1.1 ± 0.5‰, and Indiana standard ERE AgS at −3.9 ± 0.5‰) run repeatedly during each cycle of mass spectrometer operation. Isotopic compositions are considered to be accurate to ±0.1‰. Duplicate groundwater samples collected over a period of several hours on the same day varied by ±0.6‰ are inferred to reflect actual variability of sulfur isotopic compositions in water flowing from a single borehole.
Results
Geochemistry Analyses
The borehole water temperature, Cl− concentration, and pH ranged from 7.5°C to 11.4°C, 0.05 to 0.7 M, and 7.9 to 9.2, respectively (Table 1) with the coldest, least saline, and most alkaline water located at the 890 level and the warmest, most saline, and least alkaline water located at the 1,130 level. The borehole water redox state as determined by Pt electrode was indicative of suboxic, rather than anoxic, conditions (Table 1). The dominant potential electron donors were CH4, CO, and HS− at 10−5 M, whereas other detectable electron donors (H2, acetate, formate, and Fe2+) were all at submicromolar concentrations. The dominant potential electron acceptors were HCO −3 , NO −3 , and SO 2−4 at 10−4 to 10−5 M, whereas AsO 3−4 and Fe3+ were at submicromolar concentrations and the dissolved O2 and NO −2 were below the detection limit of 1.5 × 10−7 and 2.1 × 10−7 M, respectively. Concentrations of other dissolved gases, e.g., He, CO, CH4, and N2, and hydrocarbons increased with increasing salinity and temperature (Table 1). The N2 concentrations ranged from two to six times higher than that predicted from equilibration with the atmosphere at 0°C (~800 μM). The DOC, total dissolved N (NH3/NH +4 + NO −3 ) and PO 3−4 ranged from 1.5 to 3.6 × 10−4, from 5.5 to 6.7 × 10−5, and from 1.3 to 3.5 × 10−5 M, respectively, and the corresponding average C/N/P proportions were 13:2:1. Although formate, butanoate, pentanoate, and hexanoate were the principal organic acids detected in some of the samples (Table 1), the combined concentration of the organic acids comprised less than 10% of the measured DOC indicating the presence of substantial amounts of unclassified organic species, most likely nonvolatile hydrocarbons.
S Isotopic Analyses
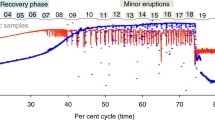
Saline seeps from the 570, 550, and 390 levels within the permafrost zone yielded less than 1‰ difference in the isotopic composition of SO 2−4 and HS− despite ranging from about 1‰ to 9‰ in the isotopic composition of SO 2−4 (Fig. 2). Borehole 890-188 yielded SO 2−4 with an isotopic composition of 11‰ but no detectable HS−. Borehole 1130-197 contained SO 2−4 with an isotopic composition of ~8‰ and no detectable HS−, whereas borehole 1130-195 contained SO 2−4 and HS− both with an isotopic composition of ~8‰. Boreholes 1130-217 and 1130-198 contained SO 2−4 with an isotopic composition ranging from 11‰ to 15‰ and HS− that was 2.5‰ to 3‰ lighter than the corresponding SO 2−4 . Boreholes 1130-260 and 1130-267 contained SO 2−4 with an isotopic composition ranging from 13‰ to 16‰ and HS− that was 5‰ to 10‰ lighter. Sample 1130-64 was sampled three times during a period of 24 h with SO 2−4 that varied from 15‰ to 16‰ and HS− that ranged from 1‰ to 2‰ lighter than SO 2−4 .
Top S isotope analyses of dissolved sulfate and sulfide for borehole water. Samples are arranged by depth in the mine with the highest levels at the left and the deepest levels at the right of the figure. Adjacent boreholes inferred to intersect the same fracture zone are indicated by a pair bracket below the sampling identification. Bottom S isotope analyses of ES, AVS, and CRS for various lithologies and vein-filling minerals from the Contwoyto Formation. Samples are arranged by rock type with amphibolites and garnetiferous phyllites on the left, meta-lutites in the middle, and vein-filling minerals or vein-rich phyllite on the right
Sulfur isotopic compositions of rock and mineral samples from Lupin Au mine revealed sulfur isotopic compositions of S0 and AVS (predominantly pyrrhotite) that were similar to each other and ranged from 2‰ to 6‰ (Fig. 2). In contrast, the isotopic composition of CRS (predominantly pyrite) ranges from −1‰ to 14‰ with the isotopically heaviest pyrite occurring in the vein-filling minerals and vein-rich phyllite. The sulfur isotopic composition of CRS is lighter than the isotopic composition of S0 or AVS for most rock samples, whereas for the vein-filling minerals and vein-rich phyllite, which contain no detectable S0, the CRS is isotopically heavier than the AVS. It is notable that the only three samples from the Contwoyto Formation with isotopic compositions greater than 4‰ for CRS are the two vein-filling minerals and the vein-rich phyllite. The unusually heavy isotopic composition of the vein-filling quartz sample suggests precipitation from fluids with a sulfide isotopic composition similar to the fracture water sampled at boreholes 1130-64 and 1130-198. The sulfur isotopic composition of the CRS from the vein-filling calcite sample and the vein-rich phyllite overlaps the 6‰ to 9.5‰ values of sulfide from the other subpermafrost fracture water samples.
Free Energy of Possible Metabolic Activities
The ΔG and FEF values for 25 microbial redox reactions are presented for the coldest, least saline, and most alkaline borehole, 890-188, and the warmer, most saline, and least alkaline borehole, 1130-192 (Table 2). HS− oxidation coupled to NO −3 reduction yielded the greatest FEF value for both boreholes with oxidation of HS− coupled to NO −3 reduction only as far as NO −2 being only slightly less favorable. The FEF values indicate that the second most favorable metabolism is denitrification coupled to oxidation of either H2 or acetate. If we assume that the dissolved O2 concentrations are much lower than our detection limit (see Table 1), then S0 or SO 2−4 reduction is the third most favorable metabolism with little difference in the ΔG and FEF values for the two water chemistries. The ΔG values for autotrophic and aceticlastic methanogenesis were slightly less than the minimal value proposed for ATP synthesis [53], whereas the ΔG values for methanogenesis and acetogenesis with CO as an electron donor were highly favorable. Thermodynamically, the least favorable metabolism is Fe3+ reduction not only because of the small and sometimes positive ΔG values but also because the low rock porosity would limit accessibility to solid phase ferric oxides. Furthermore, hematite has not been reported as a mineral phase in the Contwoyto Formation. The Contwoyto Formation does contain banded iron formation, but the Fe3+ oxides have been replaced by Fe2+-bearing hornblende, grunerite, and sulfides [22].
Microbial Community Phylogeny
Phylogenetic analyses of cultured and total-community genomic DNA provided 126 16S rRNA gene sequences that comprised 42 OTUs, which included the α-, β-, γ-, and δ-Proteobacteria, Firmicutes, Actinobacteria, Bacteroidetes/Chlorobi, and Spirochaetes (Tables 3, 4 and 5; Fig. 3a–c). The 18 short 16S rRNA sequences for the anaerobic enrichments did not enable us to clearly define their OTUs; however, seven could be correlated with OTUs identified in either the aerobic enrichments or the environmental clone library. Thirty-four of the 16S rRNA gene sequences were obtained after PCR from MDA of DNA extracts from environmental samples. MDA has been proposed as a method of amplifying DNA from samples from difficult microbial environments [54], since MDA procedures will produce microgram quantities of DNA from even single cells [55]. It has also been demonstrated that the proportion of quality genomic DNA in the final product rises in direct proportion to the quantity of starting template provided, so in low biomass systems, contamination is a primary consideration [56]. MDA of three DNA extracts, two of which were unsuccessful with 16S rRNA PCR, produced between 1 and 10 μg of DNA. Only the extract that had initially worked in 16S rRNA PCR from borehole 1130-197 produced 100% bacterial 16S rRNA clones appropriate to the system (Table 3). The 16S rRNA PCR on the MDA product from borehole 890-188 DNA extract yielded ~33% useful bacterial sequences (Table 3) and ~66% nonspecific vector product or 16S rRNA genes from human skin bacterium P. acnes. The 16S rRNA PCR on the MDA product from borehole 1130-192 DNA extract yielded only nonspecific vector product and 16S rRNA genes from the human skin bacterium P. acnes, similar to results from the negative controls. Archaeal genes were not detected in either the total-community genomic DNA extracts or enrichment cultures. The anaerobic enrichments designed specifically for methanogens failed to yield any positive signs of growth.
Phylogenetic trees of 16S ribosomal RNA gene sequences from a bulk community DNA from 1,130 m level boreholes, b cultured isolates from 1,130 m level boreholes, and c bulk community DNA and cultured isolates from 890 m level borehole. Lupin clones and isolates are designated by names containing the following information: (depth)–(borehole)–(number of clones represented)–(clone or isolate identifier). Sample names that include MDA were based upon PCR from MDA product. Reference sequences from the RDP database are also shown. Bootstrap values are shown as percent of 100
The closest relatives to many of the Lupin OTUs included microorganisms potentially involved in the metabolism of sulfur, nitrogen, and metals. SRB constituted the most common clade of phylotypes found in the total-community genomic DNA extracts (32 clones out of the 62 environmental clones from the 1,130 level and five out of the ten environmental clones from the 880 level) and the anaerobic enrichments (five out of 15 sequences from the 1,130 level; Table 5) originating from boreholes 1130-197, 1130-267, and 1130-273. The clade was comprised of five OTUs, all of which were 97–98% identical to Desulfosporosinus species (see Fig. 3a). Members of Desulfosporosinus are obligately anaerobic, chemoheterotrophic SO 2−4 reducers that belong to the Firmicutes phylum. Desulfosporosinus species are known to reduce SO 2−4 , S0, AsO 3−4 , NO −3 , and Fe3+, to grow autotrophically with H2, CO2, and SO 2−4 [57], to grow homoacetogenically, and to grow fermentatively with acetate as the end product [58]. To date, Desulfosporosinus species have not been found to disproportionate S2O 2−3 [59]. A mesophilic species belonging to the genus Desulfosporosinus has previously been isolated from permafrost [60], has been found living anaerobically within a biofilm in a flooded, pH ~7.2–8.6, carbonate-hosted Zn/Pb mine in Wisconsin [61] and reducing SO 2−4 and U6+ in uranium mine sediments [62]. Three additional OTUs, each represented by a single clone from the environmental DNA, were found in the water from borehole 1130-197 that belong to the genus Desulfuromonas, a well-known chemoheterotrophic S0 and Fe3+ reducer. Desulfuromonas has been found previously in the subsurface at Cerro Negro, New Mexico [63] and in Arctic and Antarctic sediments [64, 65].
The second most abundant clade (12 of 72 clones from environmental DNA) is comprised of two OTUs found in the water from boreholes 1130-273 and 1130-197. This clade exhibited 95–97% identity to species of Halothiobacillus, an obligately aerobic, halotolerant, or halophilic chemolithotrophic γ-Proteobacteria [66]. Halothiobacillus species are known to oxidize HS−, S0, S4O 2−6 , and S2O 2−3 all the way to SO 2−4 , although S0, SO 2−3 , or polythionates may transiently accumulate from incomplete oxidation.
A variety of Pseudomonas species were found in the aerobic enrichments from borehole water from both the 1,130 and 890 levels (Table 4) in the clone libraries of the environmental DNA from the boreholes of the 1,130 level (Table 3) and in one anaerobic enrichment on Fe3+ reducing media of water from 1130-192 (Table 5).
Within the anaerobic enrichments incubated at 4°C, the second most abundant OTUs belonged to the Veillonellaceae family (Table 5). These OTUs, plus the Fusibacter and Clostridium OTUs, all fall within the order Clostridiales. The remaining taxonomic groups in the anaerobic 4°C enrichments are the Paludibacter and Rikenellaceae OTUs which, like two environmental clones, fall within the Bacteroidetes/Chlorobi superphylum.
Characterization of aerobic enrichments revealed that 83% of the isolates tested were adapted to low-temperature conditions and were capable of sustained growth at 0°C. Although species from the 4°C anaerobic enrichments have not been isolated and their growth temperature ranges examined in detail, their growth at 4°C indicates that they are at least psychrotolerant. In addition, aerobic microorganisms isolated from the high-salinity samples (boreholes 1130-175 and 1130-192) were tolerant to 7.5% NaCl.
Of the 42 OTUs identified in this study, eight isolates (Table 4) were confirmed as being adapted to low-temperature conditions and five additional OTUs exhibited >98% identity with species or environmental sequences from polar environments. One aerobic isolate and uncultured clone were both 98–99% identical to a marine Pseudomonas sp. D5044 [DQ480134] and had >97% identity to Pseudomonas isolates from Arctic and Antarctic environments including seawater [67], lake water [68], and oil-contaminated Arctic and Antarctic sea ice (DQ521388, DQ533960) [69]. Two aerobic isolates from the 890-188 borehole water and one anaerobic enrichment from the low-salinity 1130-64 borehole water yielded Brevundimonas species which have been discovered in soil samples in Antarctica [70]. Six uncharacterized aerobic isolates were related to Rhodococcus species (Table 4), which have been isolated from Antarctic soils and cold Himalayan desert soil [70, 71]. Two aerobic isolates within the genus Devosia, one of which was shown to be cold-adapted, were >98% identical to microorganisms isolated from glaciers, Arctic sea ice, and Antarctic lakes [72–74]. One aerobic isolate was 99% identical to the cold-tolerant, facultatively anaerobic, marine Shewanella baltica that is capable of reducing NO −3 , Fe3+, and sulfur compounds [75, 76]. One uncultured OTU was 97% identical to 16S rRNA gene sequences reported from refrigerated South African Au mine drilling water [77]. One uncultured OTU was >98% identical to an Acinetobacter 16S rRNA gene sequence [AJ551148] from a deep sea environment near Antarctica. The Microbacterium isolates (Table 3) were 99% identical to glacial ice bacteria [78].
Besides their relatedness to psychrotolerant or psychrophilic species from polar environments, a number of Lupin OTUs have closest relatives that were isolated from hydrocarbon-bearing environments or are associated with species that degrade hydrocarbons, such as the Pseudomonas species isolated from oil-contaminated sea ice mentioned above. Environmental clone 1130-197-1-MDA-bur1 (Fig. 3a; uncultured β-Proteobacteria in Table 3) is 97% identical to Pusillimonas noertemannii [79], which is a petroleum hydrocarbon-degrading β-Proteobacterium. Environmental clone 890-3-MDA-sph1 belonged to the genus Sphingomonas, which contain hydrocarbon degrading species [80] and have recently been discovered as contaminants from drilling fluids on deep ice cores at Lake Vostok, Antarctica [81]. The most abundant aerobic isolate belonged to Rhodococcus species (Table 4), which are known for their ability to degrade petroleum; benzene, toluene, ethylbenzene, and o-, m-, and p-xylenes; and aromatic hydrocarbons at low temperatures [82–84]. Two aerobic isolates were 97% identical to Hydrogenophaga species isolated from activated sludge [85], which can degrade polychlorinated biphenyls [86] and have been found as members of an aerobic benzene-degrading consortia, along with Pseudomonas and Rhodococcus species, in O2-depleted groundwater [87]. One environmental clone fell within the phylum Spirochaetes of which one member, Spirochaeta smaragdinae, is a strict anaerobe residing in an oil reservoir [88] and other members have been found in oil fields in the North Sea [89].
Microbial Lipid and Quinone Composition and Cellular Concentration
PLFA concentrations ranged from 0.25 to 2.4 pmol L−1 equivalent to 6 × 103 to 6 × 104 bacterial cells L−1 using the conversion factor of 2.5 × 104 cells pmol−1 PLFA [90]. These concentrations overlap the 1–12 pmol L−1 values reported for the fracture water samples from the South Africa Au mines [91]. Borehole 1130-192, from which the most saline fracture water emanated (Table 1), yielded the lowest PLFA concentration and was dominated by normal saturates (Table 6), whereas borehole 890-188, from which the least saline fracture water emanated, had similar relative abundances of both normal saturates and monounsaturates. The fracture water from borehole 1130-197 was dominated by monounsaturates, which are typically associated with Proteobacteria [92] but lacked the midchain-branched saturates and midchain-branched monounsaturates detected in 890-188. Boreholes 890-188 and 1130-197 are at identical locations relative to the geological units and intersected the same fracture zone, despite the differences in their fracture water salinity. Another saline borehole, 1130-191, yielded mostly mid-chain-branched and terminally branched saturates (data not shown) indicative of Gram-positive bacteria (Firmicutes) [93]. Cyclopropyl to monounsaturate fatty acid ratios ranged from 0 to 0.2 (Table 6) consistent with a lack of potential nutritional stress. The trans/cis ratio of 15.6 in fracture water 1130-197 (Table 6), however, indicated environmental conditions toxic enough to trigger the cells to change isomeric configuration in order to increase membrane ordering [94, 95].
Analyses of respiratory quinones indicated that menaquinone (MK) 7 was the most prominent followed by menaquinone 6 (Table 7). Menaquinones 6 and 7 comprised 80% to 93% of the quinone profiles, whereas ubiquinone 6, the principal ubiquinone identified in these samples, represented 1% to 11% of quinone profiles (Table 7). Ubiquinone 6 is associated with aerobic respiration and may indicate fungi, yeast, or Gram-negative facultative anaerobes [96]. The UQ/MK ratio (Table 6) ranged from 0.03 for borehole 1130-197, which yielded the lowest measured Eh value, to 0.13 for borehole 1130-192, which yielded a greater Eh value (Table 1).
The cell concentrations values were quite low ranging from 360 to 3,000 cells mL−1 for the >0.02-μm fraction (Table 1). Because no Archaea were detected in either the 16S rRNA clone libraries or the enrichments of the fracture water, the PLFA and quinone analyses should both reflect the total microbial community structure and cell concentration. The slightly higher estimates of cell concentrations derived from direct cell counts may mean that the cell PLFA−1 ratio is significantly greater than the 25,000 value based upon shallow aquifer samples [90] or it may reflect the fact that the 0.02-μm pore-size filter used for cell counts captured more cells than the 0.3-µm filters used for the PLFA, quinone, and DNA extracts. In Greenland ice core samples, cells that are capable of passing though 0.2-µm filters and of being cultivated predominate [97]. Similarly, the Lupin subpermafrost water may also contain large numbers of ultrasmall cells. Nonetheless, detecting the same dominating genera of Desulfosporosinus and Pseudomonas in the clone libraries from both (1) the filtered uncultured samples and (2) the unfiltered cultured samples suggests that the principal microbial community members have been identified.
Discussion
The geochemical, 16S rRNA, PLFA and quinone analyses all suggest that the fracture water is predominantly suboxic. Enrichments and total DNA results detected both aerobic and anaerobic bacteria and failed to detect any methanogens. PLFA analyses indicated the presence of Gram-positive bacteria, some anaerobic Gram-negative bacteria, and metal or sulfate-reducing bacteria, and possibly other anaerobic prokaryotes as well [98]. The dominant quinones identified are found in both aerobic and anaerobic α-Proteobacteria, Actinobacteria, Flavobacterium, Desulfovibrio, Desulfotomaculum, Rhodococcus, and Bacillus, among other phylogenetic groups [96, 99]. In addition, ubiquinones indicative of Gram-negative bacteria and eukaryotes that utilize O2 as an electron acceptor were found [96, 99] consistent with the positive aerobic enrichments. The midpoint potential for menaquinone/menaquinol is −74 mV [100], whereas the midpoint potential for ubiquinone/ubiquinol is +110 mV [99, 101]. The relative abundance of ubiquinones to total quinones (Table 7) ranged from 3% for borehole 1130-197 to 7% for borehole 890-188 to 11% for borehole 1130-192, a trend which is consistent with their measured Pe values of 0.3 (15 mV), 0.6 (32 mV), and 0.8 (42 mV), respectively (Table 1). The suboxic nature of the Lupin fracture water ecology suggests that the subpermafrost fracture network is not completely isolated from the overlying oxygenic biosphere and that a pathway between the two realms exists.
Hydrobiogeochemical Model
Geochemical and isotopic analyses of the Lupin borehole water indicate that the subpermafrost fracture water is most likely a mixture [102]. Here, we assume that it represents a mixture of a small fraction of highly saline brine (similar to brines reported from other parts of the Canadian shield [18]) and a much larger fraction of paleometeoric water because this model is compatible with the isotopic data [102, 103]. Utilizing the in situ generation model of Lippmann et al. [104] and the 2-ppm U concentration, porosity, and density of the Contwoyto Formation [24], the 1.5-mM He observed in the most saline fracture water (1130-192 in Table 1) would require ~40 Gyr of accumulation at in situ temperatures or ~90 Myr of accumulation for temperatures >100°C, the temperature at which He will readily diffuse out of most U-bearing minerals on million-year time scales. This time scale precludes marine water invasion during the Holocene as a source of the saline water. The exclusion of recent marine surface water is also consistent with the geological evidence indicating that an ice-dammed lake occupied the basin of Contwoyto Lake and the valley of the Burnside River until about 9,000 years BP [105]. It is reasonable to assume that during and shortly after deglaciation, the upper part of the bedrock was saturated with dilute and cold melt water. The high hydraulic pressures experienced during the most recent subglacial conditions may have forced melt water to considerable depths along fracture zones, such as the one assumed to be located beneath Contwoyto Lake and intersected by the boreholes approximately 500 m further west (Fig. 1). The DIC in the Lupin fracture water, which would primarily originate with the paleometeoric component, yielded 14C ages of 23.0 to 26.8 ka (3.50 to 5.58 pMC; Table 1), consistent with such a glacial recharge component [103]. Assuming that the 550-m-thick permafrost is impermeable, this recharge episode must have occurred before the formation of the permafrost.
Although the fracture networks appear to be discontinuous [24], the occasional appearance of <1 tritium unit of 3H during the 7-year monitoring period in boreholes 890-188 and 1130-197 suggests that some limited migration of modern meteoric water is occurring in these boreholes. The most likely source for the modern component is a talik beneath Contwoyto Lake, with mixing induced by dewatering of the fracture network as the mine expanded its subsurface foot print (Fig. 1).
Utilizing the major element composition of brine characterized at the Miramar Con mine in Yellowknife [106] and Contwoyto Lake water [21] as end members, a mixing model was constructed to approximate the affects of freshwater talik or paleometeoric melt water interactions with the saline water-bearing fractures (Fig. 1). The dissolved concentrations of gases, organic acids, Ca2+, and Na+ in the brine end member were adjusted to match those observed for the Lupin fracture water at the observed salinity. The dissolved gases of the Contwoyto Lake were assumed to represent atmospheric equilibration at 1°C. By assuming a temperature of 11.5°C for the brine end member, the mixing model yielded temperatures and pH values that closely match those observed for Lupin borehole water at the observed salinity (Fig. 4). According to reaction simulations, the observed decrease in pH with increasing temperature and major cation concentrations in the Lupin borehole water (Table 1) is consistent with equilibration with albite, monohydrocalcite, chalcedony, and chlorite. The He concentration derived for the brine end member by matching the He concentrations of the Lupin fracture water was 5.5 mM, which is equivalent to a ~300-Myr accumulation age at temperatures greater than ~100°C.
ΔG for microbial redox reactions, pH, and T as a function of chlorinity generated from GWB simulated mixing/reaction model. The dashed horizontal line at −20 kJ mol−1 represents the minimum energy required for ATP synthesis [53]
Two significant conclusions follow from this mixing/reaction model. The first is that the dissolved O2 from cold fresh Contwoyto Lake water should yield O2 concentrations of hundreds of micromolars for water with the salinity values observed in the Lupin borehole water. This result combined with the occasional 3H detected in some boreholes and the small, but detectable, ubiquinone lipid signatures suggests that aerobic metabolism must be occurring intermittently along the flow path from the talik into the fractures. Furthermore, because the predicted concentrations of dissolved O2 greatly exceed the observed concentrations of HS− and SO 2−4 , heterotrophic aerobic activity, not just the chemolithotrophic activity of Halothiobacillus, must be occurring. This is consistent with the observed aerobic enrichments. The low the SO 2−4 concentrations in the saline fracture water exceed that of Contwoyto Lake, 26 μM [24], the only potential modern fluid source. The SO 2−4 concentrations also do not exhibit any obvious correlation with salinity and their O isotope values are more consistent with predominantly anaerobic oxidation [102]. The SO 2−4 , therefore, must originate from within the Contwoyto Formation. One potential SO 2−4 source is oxidation of sulfides by radiolytically produced H2O2, O2, and HO −2 from H2O at an estimated rate of ~10−4 M Myr−1 [52, 107] within the Contwoyto Formation. The similarity in the δ34S values of SO 2−4 and those of vein-filling pyrite is consistent with this hypothesis. The rate of radiolytic electron acceptor production is closely matched by radiolytic production of H2 and formate [49] at 8 × 10−4 and 3 × 10−4 M Myr−1, respectively. Steady-state reaction simulations indicate that microbial SO 2−4 reduction at rates of 10−4 M Myr−1 reproduce the observed SO 2−4 , H2, and formate concentrations while yielding ΔG values of 66 and 69 kJ (mol of SO 2−4 )−1 for the reduction of SO 2−4 by H2 and formate, respectively. Using the maintenance energy rate model of Price and Sowers [108], which is based upon respiration rates inferred from geochemical models, a 10−4-M Myr−1 respiration rate could sustain 106 cells mL−1 of SRB, a value which is much greater than the observed planktonic biodensities.
The second conclusion comes from analysis of the ΔG for the reactions in Table 2 as a function of mixing (Fig. 4). Because of the elevated pH resulting from mineral water interaction at lower temperature and salinity, the ΔG for the methanogenic and Fe3+-reducing reactions become progressively unfavorable at lower salinity or as the talik is approached. In the case of the methanogenic reactions, the ΔG values are insufficient to provide the −20 kJ mol−1 required for ATP synthesis [53]. This behavior is distinct from the ΔG values for SO 2−4 reduction, NO −3 reduction, and aerobic oxidation of HS−, which exhibit little variation with salinity, temperature, and pH.
Even for the more saline fracture water, ΔG of SO 2−4 reduction in this system is high enough that SRBs can out-compete methanogens for acetate, formate, and H2. Indeed, the ΔGs for some methanogenic reactions, while negative, were among the least favorable of all the reactions examined (Table 2), perhaps explaining why methanogenic Archaea were not detected in the DNA extracts. In addition, isotopic analyses of the CH4 are consistent with a thermogenic, rather than a methanogenic, origin [103]. S2O 2−3 disproportionation is also not competitive with SO 2−4 reduction, consistent with the absence of microorganisms bearing these metabolisms in the 16S rRNA gene clone libraries. The ΔG for the acetogenic reaction is less than the minimum ΔG value required for ATP synthesis [53] again consistent with the paucity of homoacetogens in the clone libraries of the enrichments and the DNA extracts.
Inferred Metabolisms
The range of physiological properties observed from the enrichments and inferred from the 16S rRNA gene phylogeny is compatible with the above hydrogeochemical model. Like the perennial springs of Axel Heiberg Island [19] and the subglacial brine of Taylor Glacier, Antarctica [109, 110], the Lupin OTUs were dominated by 16S rRNA gene sequences related to microorganisms capable of both reduction and oxidation of sulfur species, suggesting a possibly complete subsurface sulfur cycle.
The anaerobic enrichments revealed a psychrotolerant Desulfosporosinus species capable of reducing SO 2−4 and the presence of HS− that is 10‰ to 11‰ isotopically lighter than coexisting SO 2−4 in fracture water from borehole 1130-267 is consistent with in situ SRB activity (Table 1; Fig. 2). Although Desulfosporosinus was prevalent in water from boreholes 1130-197 and 890-188, their δ34S of the SO 2−4 was not as high as that of 1130-267 (Fig. 2). This lack of significant S isotopic enrichment in these boreholes may reflect the fact that their SO 2−4 concentrations were <100 μM, which is less than the 200-μM SO 2−4 concentrations above which significant isotopic enrichment is typically observed [111], whereas the SO 2−4 concentration for 1130-267 was 160 μM.
Following reduction, sulfur compounds (HS−, S0, S4O 2−6 , and S2O 2−3 ) could be oxidized by Halothiobacillus species all the way to SO 2−4 , but if this is occurring, then it would have to be doing so without significantly increasing the O isotopic signature of the SO 2−4 [102]. The oxidation of HS− by reduction of NO −3 to NO −2 appears to be a thermodynamically viable alternative to O2 reduction (Table 7), but anaerobic HS− oxidation has not been reported for the genera Halothiobacillus. Thioalkalivibrio denitrificans (also a γ-Proteobacterium isolated from an alkaline lake) is capable of oxidizing S0 while reducing NO −2 and N2O to N2 [112] and perhaps the Halothiobacillus in the Lupin subpermafrost water has adopted a similar metabolic pathway. If true, then what is the source of the NO −3 in the Lupin fracture water? Water from the overlying permafrost zone yielded 7 to 40 mM NO −3 concentrations suggestive of residue from explosive compounds as has been found to be the case in other underground excavations [21, 113, 114]. Because the boreholes sampled in this study occur well beneath the permafrost horizon and extend well beyond the damage zone associated with tunnel excavation, explosive residues seem a less likely source for this NO −3 . NO −3 residue does occur in the drilling water at lesser concentrations and may represent a source. The most recently drilled boreholes, 1130-267 and 1130-273, were drilled 2 years prior to sampling and yielded 5.2 and 3.1 μM NO −3 , respectively, whereas 1130-192, which has been flowing since it was first drilled over a decade ago, yielded 63 μM NO −3 . In addition, borehole 1130-192 did not yield any detectable 3H, indicating that it is not contaminated by drilling water infiltrating the fracture system from other parts of the mine. In fact, all the sampled boreholes were flowing freely or through leaking plugs prior to instrumentation with flow rates of 2–8 L min−1, which would allow the effective removal of drilling contamination. Therefore, the low NO −3 concentrations observed in the fracture water do not appear to be caused by mining contamination. The absence of 16S rDNA genes associated with nitrifiers seems to rule out microbial oxidation of NH +4 by trace O2 as well. Unfortunately, the low NO −3 concentrations precluded the use of stable isotope analyses to address its origin in the subpermafrost saline water [102] or to what extent the potentially heterotrophic denitrifying bacteria identified in the total DNA are active.
Implications for Martian Subsurface Ecosystems
This “energy bottleneck” for methanogens as determined from the hydrobiogeochemical model has interesting implications for the origin of CH4 that has recently been reported in the Martian atmosphere [3, 4]. If the Martian subsurface brine chemistry bears any resemblance to that at Lupin, then psychrotolerant sulfur-metabolizing microorganisms sustained by radiolytic reactions, not methanogens, may dominate beneath the permafrost. The abundance of potential psychrotolerant microorganisms in both enrichments and 16S rRNA gene clone libraries suggests that the subsurface environment, which has stored thermogenic gases for millennia, was colonized when the surface environment entered the glacial period of the Pleistocene. Similar colonization of the Martian subsurface would have occurred during the late Noachian as the cryosphere began to encompass the planet. Microorganisms that managed to colonize the fracture zones at depth would be selected from salt-tolerant aerobic and anaerobic microorganisms residing in talik water by high alkalinity, increasing salinity, and diminishing O2. This environment, however, produces an energy bottleneck near the surface that filters out methanogens and perhaps Fe3+ reducers, but permits aerobic, denitrifying, S0 reducing, and SO 2−4 reducing metabolisms through. This model would also explain the lower levels of electron donors relative to that of electron acceptors as the aerobic microorganisms would strip the water of electron donors as until O2 was consumed. The Halothiobacillus, unless it has adopted an anaerobic pathway, could only survive long term by O2 leakage into the subpermafrost fracture systems. Methanogens probably do exist in the active layer at Lupin Au mine as they do in many permafrost sites in the Arctic. If the same were true of Mars during the late Noachian, then they may have adapted to the more stringent surface environment of Mars today and thus explain the seasonal fluctuation of the Martian atmospheric CH4 [5, 6].
References
Malin MC, Edgett KS (2000) Evidence for recent groundwater seepage and surface runoff on Mars. Science 288:2330–2335
Malin MC, Edgett KS, Posiolova LV, McColley SM, Noe Dobrea EZ (2006) Present-day impact cratering rate and contemporary gully activity on Mars. Science 314:1573–1577
Krasnopolsky VA, Maillard JP, Owen TC (2004) Detection of methane in the Martian atmosphere: evidence for life? Icarus 172:537–547
Onstott TC, McGown D, Kessler J, Sherwood Lollar B, Lehmann KK, Clifford SM (2006) Martian CH4: sources, flux and detection. Astrobiology 6:377–395
Mumma MJ, Villanueva GL, Novak RE, Hewagama T, Bonev BP, DiSanti MA, Mandell AM, Smith MD (2009) Strong release of methane on Mars in northern summer 2003. Science 323:1024–1045
Geminale A, Formisano V, Giuranna M (2009) Methane in Martian atmosphere: average spatial, diurnal, and seasonal behaviour. Planet Space Sci 56:1194–1203
Schuerger AC, Mancinelli RL, Kern RG, Rothschild LJ, McKay CP (2003) Survival of endospores of Bacillus subtilis on spacecraft surfaces under simulated Martian environments: implications for the forward contamination of Mars. Icarus 165:253–276
Schuerger AC, Richards JT, Newcombe DA, Venkateswaran K (2006) Rapid inactivation of seven Bacillus spp. under simulated Mars UV irradiation. Icarus 181:52–62
Bakermans C, Tsapin AI, Souza-Egipsy V, Gilichinsky DA, Nealson KH (2003) Reproduction and metabolism at −10°C of bacteria isolated from Siberian permafrost. Environ Microbiol 5:321–326
Bakermans C, Ayala-del-Rio L, Ponder M, Vishnivetskaya TA, Gilichinsky DA, Thomashow MF, Tiedje JM (2006) Psychrobacter cryohalolentis sp. nov. and Psychrobacter arcticus sp. nov., isolated from Siberian permafrost. Int J Syst Evol Microbiol 56:125–1291
Gilichinsky D, Rivkina E, Bakermans C, Shcherbakova V, Petrovskaya L, Ozerskaya S, Ivanushkina N, Kochkina G, Laurinavichuis K, Pecheritsina S, Fattakhova R, Tiedje JM (2005) Biodiversity of cryopegs in permafrost. FEMS Microbiol Ecol 53:117–128
Gilichinsky DA, Wilson GS, Friedmann EI, Mckay CP, Sletten RS, Rivkina EM, Vishnivetskaya TA, Erokhina LG, Ivanushkina NE, Kochkina GA, Shcherbakova VA, Soina VS, Sprina EV, Vorobyova EA, Fyodorov-Davydov DG, Hallet B, Ozerskaya SM, Sorokovikov VA, Laurinavichyus KS, Shatilovich AV, Chanton JP, Ostroumov VE, Tiedje JM (2007) Microbial populations in Antarctic permafrost: biodiversity, state, age, and implication for astrobiology. Astrobiology 7:275–311
Morozova D, Möhlmann D, Wagner D (2007) Survival of methanogenic Archaea from Siberian permafrost under simulated martian thermal conditions. Orig Life Evol Biosph 37:189–200
Rivkina E, Laurinavichius K, McGrath J, Tiedje J, Shcherbakova V, Gilichinsky D (2004) Microbial life in permafrost. Adv Space Res 33:1215–1221
Smith JJ, Tow LA, Stafford W, Cary C, Cowan DA (2006) Bacterial diversity in three different Antarctic cold desert mineral soils. Microb Ecol 51:413–421
Clifford SM, Parker TJ (2001) The evolution of the Martian hydrosphere: implications for the fate of a primordial ocean and the current state of the northern plains. Icarus 154:40–79
Boston PJ, Ivanov MV, McKay CP (1992) On the possibility of chemosynthetic ecosystems. Icarus 95:300–308
Frape SK, Fritz P, McNutt RH (1984) Water–rock interaction and chemistry of groundwaters from the Canadian Shield. Geochim Cosmochim Acta 48:1617–1627
Perreault NN, Andersen DT, Pollard WH, Greer CW, Whyte LG (2007) Characterization of the prokaryotic diversity in cold saline perennial springs of the Canadian high arctic. Appl Environ Microbiol 73:1532–1543
Perreault N, Greer C, Andersen D, Tille S, Lacrampe-Couloume G, Sherwood Lollar B, Whyte L (2008) Heterotrophic and autotrophic microbial populations in cold perennial springs of the high arctic. Appl Environ Microbiol 74:6898–6907
Ruskeeniemi T, Paananen M, Ahonen L, Kaija J, Kuivamaki A, Frape S, Moren L, Degnan P (2002) Permafrost at Lupin: report of phase I. Geological Survey of Finland Nuclear Waste Disposal Research Report YST-112
Bullis HR, Hureau RA, Penner BD (1994) Distribution of gold and sulfides at Lupin, Northwest Territories. Econ Geol 89:1217–1227
Rubel AP, Sonntag C, Lippmann J, Pearson FJ, Gautschi A (2002) Solute transport in formations of very low permeability: profiles of stable isotope and dissovled noble gas contents of pore water in the Opalinus Clay, Mont Terri, Switzerland. Geochim Cosmochim Acta 66:1311–1321
Ruskeeniemi T, Ahonen L, Paananen M, Frape S, Stotler R, Hobbs M, Kaija J, Degnan P, Blomqvist R, Jensen M, Lehto K, Moren L, Puigdomenech I, Snellman M (2004) Permafrost at Lupin: report of phase II. Geological Survey of Finland Nuclear Waste Disposal Research Report YST-119
Onstott TC, Lin L-H, Davidson M, Mislowack B, Borcsik M, Hall J, Slater GF, Ward J, Sherwood Lollar B, Lippmann-Pipke J, Boice E, Pratt LM, Pfiffner SM, Moser DP, Gihring T, Kieft TL, Phelps TJ, van Heerden E, Litthaur D, DeFlaun M, Rothmel R (2006) The origin and age of biogeochemical trends in deep fracture water of the Witwatersrand Basin, South Africa. Geomicrobiol J 23:369–414
Ward JA, Slater GF, Moser DP, Lin L-H, Lacrampe-Couloume G, Bonin AP, Davidson M, Hall JA, Mislowack B, Bellamy RES, Onstott TC, Sherwood Lollar B (2004) Microbial hydrocarbon gases in the Witwatersrand Basin, South Africa: implications for the deep biosphere. Geochim Cosmochim Acta 68:3239–3250
Noble RT, Fuhrman JA (1998) Use of SYBR Green I for rapid epifluorescence counts of marine viruses and bacteria. Aquat Microb Ecol 14:113–118
Kirchman D, Sigda J, Kapuscinski R, Mitchell R (1982) Statistical analysis of the direct count method for enumerating bacteria. Appl Environ Microbiol 44:376–382
Lane DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, Chichester, pp 115–175
Onstott TC, Moser DP, Fredrickson JK, Brockman FJ, Pfiffner SM, Phelps TJ, White DC, Peacock A, Balkwill D, Hoover R, Krumholz LR, Borscik M, Kieft TL, Wilson RB (2003) Indigenous versus contaminant microbes in ultradeep mines. Environ Microbiol 5:1168–1191
Pfiffner SM, Onstott TC, Ruskeeniemi T, MT T, Bakermans C, McGown D, Chan E, Johnson A, Phelps TJ, Puil ML, Difurio SA, Pratt LM, Stotler R, Frape S, Telling J, Sherwood-Lollar B, Neill I, Zerbin B (2008) Challenges for coring deep permafrost on Earth and Mars. Astrobiology 8:623–638
DeLong EF (1992) Archaea in coastal marine environments. Proc Natl Acad Sci U S A 89:5685–5689
Giovannoni SJ (1991) The polymerase chain reaction. In: Stackebrandt E, Goodfellow M (eds) Modern microbiological methods: nucleic acids techniques in bacterial systematics. Wiley, New York, pp 177–203
Lane DJ (1985) 16S/23S sequencing. In: Stackenbrandt E, Goodfellow M (eds) Modern microbiological methods: nucleic acid techniques in bacterial systematics. Wiley, New York, pp 115–175
Forney LJ, Zhou X, Brown CJ (2004) Molecular microbial ecology: land of the one-eyed king. Curr Opin Microbiol 7:210–220
Takai K, Moser DP, DeFlaun MF, Onstott TC, Fredrickson JK (2001) Archaeal diversity in waters from deep South African gold mines. Appl Environ Microbiol 67:5750–5760
Huber T, Faulkner G, Hugenholtz P (2004) Bellerophon; a program to detect chimeric sequences in multiple sequence alignments. Bioinformatics 20:2317–2319
Cole J, Chai B, Marsh T, Farris R, Wang Q, Kulam S, Chandra S, McGarrell D, Schmidt T, Gairity G, Tiedje J (2003) The Ribosomal Database Project (RDP-II): previewing a new autoaligner that allows regular updates and the new prokaryotic taxonomy. Nucleic Acids Res 31:442–443
Altschul SF, Thomas LM, Schäffer AA, Zhang JH, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402
White DC, Ringelberg DB (1998) Signature lipid biomarker analysis. In: Burlage RS, Atlas R, Stahl DA, Geesey G, Sayler G (eds) Techniques in microbial ecology. Oxford University Press, New York
Lytle CA, Gan YDM, Salone K, White DC (2004) Sensitive characterization of microbial ubiquinones from biofilms by electrospray/mas spectrometry. Environ Microbiol 3:265–272
Geyer R, Peacock AD, White DC, Lytle CA, Van Verkel GJ (2004) Atmospheric pressure chemical ionization and atmospheric pressure photoionization for simultaneous mass spectrometric analysis of microbial respiratory ubiquinones and menaquinones. J Mass Spectrom 39:922–929
Andrews JN, Wilson GB (1987) The composition of dissolved gases in deep groundwaters and groundwater degassing. Geol Assoc Can Spec Pap 33:245–252
Onstott TC (2004) Impact of CO2 injections on deep subsurface microbial ecosystems and potential ramifications for the surface biosphere. In: Thomas DC, Benson SM (eds) The CO2 capture and storage project, vol II. Lawrence Berkeley National Laboratory, Berkeley, pp 1207–1239
Lin L-H, Hall JA, Lippmann J, Ward JA, Sherwood-Lollar B, Onstott TC (2005b) Radiolytic H2 in the continental crust: nuclear power for deep subsurface microbial communities. Geochem Geophys Geosys 6:Q07003. doi:10.1029/2004GC000907
Lin L-H, Slater G, Sherwood Lollar B, Lacrampe-Couloume G, Onstott TC (2005a) The yield and isotopic composition of radiolytic H2, a potential energy source for the deep subsurface biosphere. Geochim Cosmochim Acta 69:893–903
Bjergbakke E, Draganic ZD, Sehested K, Draganic IG (1989a) Radiolytic products in waters. 1. Computer-simulation of some radiolytic processes in the laboratory. Radiochim Acta 48:65–71
Bjergbakke E, Draganic ZD, Sehested K, Draganic IG (1989b) Radiolytic products in waters. 2. Computer-simulation of some radiolytic processes in nature. Radiochim Acta 48:73–77
Cai Z, Li X, Katsumura Y, Urabe O (2001) Radiolysis of bicarbonate and carbonate aqueous solutions: product analysis and simulation of radiolytic processes. Nucl Technol 136:231–240
Raymond R (2006) Radiolytic destruction of ammonia as a subsurface source of nitrate, B.A. dissertation. Princeton University, Princeton, N.J., USA. p 50
Bruchert V, Pratt LM (1996) Contemporaneous early diagenetic formation of organic and inorganic sulfur in estuarine sediments from St. Andrews Bay, Florida. Geochim Cosmochim Acta 60:2325–2332
Lefticariu L, Pratt LM, Ripley EM (2006) Mineralogic and sulfur isotopic effects accompanying oxidation of pyrite in millimolar solutions of hydrogen peroxide at temperatures from 4 to 150°C. Geochim Cosmochim Acta 70:4889–4905
Shink B (1997) Energetics of syntrophic cooperation in methanogenic degradation. Microbiol Mol Biol Rev 61:262–280
Abulencia CB, Wyborski DL, Garcia JA, Podar M, Chen W, Chang SH, Chang HW, Watson D, Brodie EL, Hazen TC, Keller M (2006) Environmental whole genome amplification to access microbial populations in contaminated sediments. Appl Environ Microbiol 72:3291–3301
Spits C, Le Caignec C, De Rycke M, Van Haute L, Van Steirteghem A, Liebars I, Sermon K (2006) Optimization and evaluation of single-cell whole genome multiple displacement amplification. Hum Mutat 27:496–503
Bergen AW, Qi Y, Haque KA, Welch RA, Chanock SJ (2005) Effects of DNA mass on multiple displacement whole genome amplification and genotyping performance. BMC Biotechnol 5:24
Klemps R, Cypionka H, Widdel F, Pfennig N (1985) Growth with hydrogen, and further physiological characteristics of Desulfotomaculum species. Arch Microbiol 143:203–208
Stackebrandt E, Schumann P, Schüler E, Hippe H (2003) Reclassification of Desulfotomaculum auripigmentum as Desulfosporosinus auripigmeni corrig., comb. nov. Int J Syst Evol Microbiol 53:1439–1443
Jackson BE, McInerney MJ (2000) Thiosulfate Disproportionation by Desulfotomaculum thermobenzoicum. Appl Environ Microbiol 66:3650–3653
Vatsurina A, Badrutdinova D, Schumann P, Spring S, Vainshtein M (2008) Desulfosporosinus hippei sp. nov., a mesophilic sulfate-reducing bacterium isolated from permafrost. Int J Syst Evol Microbiol 58:1228–1232
Labrenz M, Banfeld JF (2004) Sulfate-reducing bacteria-dominated biofilms that precipitate ZnS in a subsurface circumneutral-pH mine drainage system. Microb Ecol 47:205–217
Suzuki Y, Kelly SD, Kemner KM, Banfield JF (2002) Microbial populations stimulated for hexavalent uranium reduction in uranium mine sediment. Appl Environ Microbiol 69:1337–1346
Kovacik WP, Takai K, Mormile MR, McKinley JP, Brockman FJ, Fredrickson JK, Holben WE (2006) Molecular analysis of deep subsurface Cretaceous rock indicates abundant Fe(III)- and S0-reducing bacteria in a sulfate-rich environment. Environ Microbiol 8:141–155
Purdy KJ, Nedwell DB, Embley TM (2003) Analysis of the sulfate-reducing bacterial and methanogenic archaeal populations in contrasting Antarctic sediments. Appl Environ Microbiol 69:3181–3191
Vandieken V, Mussmann M, Niemann H, Jorgensen BB (2006) Desulfuromonas svalbardensis sp. nov. and Desulfuromusa ferrireducens sp. nov., psychrophilic, Fe(III)-reducing bacteria isolated from Arctic sediments, Svalbard. Int J Syst Evol Microbiol 56:1133–1139
Sievert SM, Heidorn T, Kuever J (2000) Halothiobacillus kellyi sp. nov., a mesophilic, obligately chemolithoautotrophic, sulfur oxidizinig bacterium isolated from a shallow-water hydrothermal vent in the Aegean Sea, and emended description of the genus Halothiobacillus. Int J Syst Evol Microbiol 50:1229–1237
Mergaert J, Verhelst A, Cnockaert MC, Tan TL, Swings J (2001) Characterization of facultative oligotrophic bacteria from polar seas by analysis of their fatty acids and 16S rDNA sequences. Syst Appl Microbiol 24:98–107
Gilbert JA, Hill PJ, Dodd CE, Laybourn-Parry J (2004) Demonstration of antifreeze protein activity in Antarctic lake bacteria. Microbiol Mol Biol Rev 150:171–180
Gerdes B, Brinkmeyer R, Dieckmann G, Helmke E (2005) Influence of crude oil on changes of bacterial communities in Arctic sea-ice. FEMS Microbiol Ecol 53:129–139
Shivaji S, Reddy GSN, Aduri RP, Kutty R, Ravenschlag K (2004) Bacterial diversity of a soil sample from Schirmacher Oasis, Antarctica. Cell Mol Biol 50:525–536
Mayilraj S, Krishnamurthi S, Saha P, Saini HS (2006) Rhodococcus kroppenstedtii sp nov., a novel actinobacterium isolated from a cold desert of the Himalayas, India. Int J Syst Evol Microbiol 56:979–982
Brinkmeyer R, Knittel K, Jurgens J, Weyland H, Amann R, Helmke E (2003) Diversity and structure of bacterial communities in Arctic versus Antarctic pack ice. Appl Environ Microbiol 69:6610–6619
Foght J, Aislabie J, Turner S, Brown CE, Ryburn J, Saul DJ, Lawson W (2004) Culturable bacteria in subglacial sediments and ice from two Southern Hemisphere glaciers. Microb Ecol 47:329–340
Van Trappen S, Mergaert J, Van Eygen S, Dawyndt P, Cnockaert MC, Swings J (2002) Diversity of 746 heterotrophic bacteria isolated from microbial mats from ten Antarctic lakes. Syst Appl Microbiol 25:603–610
Ziemke F, Höfle MG, Lalucat J, Roselló-Mora R (1998) Reclassification of Shewanella putrefaciens Owen’s genomic group II as Shewanella baltica sp. nov. Int J Syst Bacteriol 48:179–186
Vogel B, Venkateswaran K, Satomi M, Gram L (2005) Identification of Shewanella baltica as the most important H2S-producing species during iced storage of Danish marine fish. Appl Environ Microbiol 71:6689–6697
Gihring TM, Moser DP, Lin L-H, Davidson M, Onstott TC, Morgan L, Milleson M, Kieft TL, Trimarco E, Balkwill DL, Dollhopf ME (2006) The distribution of microbial taxa in the subsurface water of the Kalahari Shield, South Africa. Geomicrobiol J 23:415–430
Behrendt U, Ulrich A, Schumann P (2001) Description of Microbacterium foliorum sp. nov. and Microbacterium phyllosphaerae sp. nov., isolated from the phyllosphere of grasses and the surface litter after mulching the sward, and reclassification of Aureobacterium resistens (Funke et al. 1998) as Microbacterium resistens comb. nov. Int J Syst Evol Microbiol 51:1267–1276
Stolz A, Burger S, Kuhm A, Kampfer P, Busse HJ (2005) Pusillimonas noertemannii gen. nov., sp. nov., a new member of the family Alcaligenaceae that degrades substituted salicylates. Int J Syst Evol Microbiol 55:1077–1081
Popp N, Schlomann M, Mau M (2006) Bacterial diversity in the active stage of a bioremediation system for mineral oil hydrocarbon contaminated soils. Microbiol Mol Biol Rev 152:3291–3304
Alekhina IA, Marie D, Petit JR, Lukin VV, Zubkov VM, Bulat SA (2007) Molecular analysis of bacterial diversity in kerosene-based drilling fluid from the deep ice borehole at Vostok, East Antarctica. FEMS Microbiol Ecol 59:289–299
Margesin R, Fonteyne PA, Redl B (2005) Low-temperature biodegradation of high amounts of phenol by Rhodococcus spp. and basidiomycetous yeasts. Res Microbiol 156:68–75
Belousova NI, Shkidchenko AN (2004) Low-temperature microbial degradation of oil products differing in the extent of condensation. Appl Biochem Microbiol 40:262–265
Margesin R, Labbe D, Schinner F, Greer CW, Whyte LG (2003) Characterization of hydrocarbon-degrading microbial populations in contaminated and pristine alpine soils. Appl Environ Microbiol 69:3085–3092
Kampfer P, Schulze R, Jackel U, Malik KA, Amann R, Spring S (2005) Hydrogenophaga defluvii sp. nov. and Hydrogenophaga atypica sp. nov., isolated from activated sludge. Int J Syst Evol Microbiol 55:341–344
Lambo AJ, Patel TR (2007) Biodegradation of polychlorinated biphenyls in Aroclor 1232 and production of metabolites from 2, 4, 4′-trichlorobiphenyl at low temperature by psychrotolerant Hydrogenophaga sp strain IA3-A. J Appl Microbiol 102:1318–1329
Fahy A, McGenity TJ, Timmis KN, Ball AS (2006) Heterogeneous aerobic benzene-degrading communities in oxygen-depleted groundwaters. FEMS Microbiol Ecol 58:260–270
Magot M, Fardeau M-L, Arnauld O, Lanau C, Ollivier B, Thomas P, Patel BK (1997) Spirochaeta smaragdinae sp. nov., a new mesophilic strictly anaerobic spirochete from an oil field. FEMS Microbiol Lett 155:185–191
Dahle H, Garshol F, Madsen M, Birkeland N-K (2008) Microbial community structure analysis of produced water from a high-temperature North Sea oil-field. Antonie Van Leeuwenhoek 93:37–49
Balkwill D, Leach F, Wilson J, McNabb J, White D (1988) Equivalence of microbial biomass measures based on membrane lipid and cell wall components, adenosine triphosphate, and direct counts in subsurface sediments. Microb Ecol 16:73–84
Pfiffner SM, Cantu JM, Smithgall A, Peacock A, White DC, Moser DP, Onstott TC, van Heerden E (2006) Phospholipid fatty acid profiles and biodensity estimates for water, rock and air samples recovered from Witwatersrand Basin mines. Geomicrobiol J 23:431–442
Kaneda T (1991) Iso and anteiso fatty acids in bacteria: biosynthesis, function, and taxonomic significance. Microbiol Rev 55:288–302
Kohring LL, Ringelberg DB, Devereux R, Stahl M, Mittleman M, White DC (1994) Comparison of phylogenetic relationships based on phospholipids fatty acid profiles and ribosomal RNA sequence similarities among dissimilatory sulfate-reducing bacteria. FEMS Microbiol Lett 119:303–308
Guckert JB, Hood MA, White DC (1986) Phospholipid, ester-linked fatty acid profile changes during nutrient depletion of Vibrio cholerae: increases in the trans/cis and proportions of cyclopropyl fatty acids. Appl Environ Microbiol 52:794–801
Sikkema J, Bont JAMD, Poolman B (1995) Mechanisms of membrane toxicity of hydrocarbons. Microbiol Rev 59:201–222
Collins MD, Jones D (1981) Distrubition of isoprenoid quinone structyral types in bacteria and their taxonomic implications. Microbiol Rev 45:316–354
Miteva VI, Brenchley JE (2005) Detection and isolation of ultrasmall microorganisms from a 120, 000-year-old Greenland glacier ice core. Appl Environ Microbiol 71:7806–7818
Tunlid A, White DC (1992) Biochemical analysis of biomass, community structure, nutritional status and metabolic activity of the microbial community in soil. In: Bollag J-M, Stotzky G, editors. Soil Biochemistry Vol. 7. New York: Marcel Dekker Inc., P. 229–262
Hiraishi A, Ueda Y, Ishihara J (1998) Quinone profiling of bacterial communities in natural and synthetic sewage activated sludge for enhanced phosphate removal. Appl Environ Microbiol 64:992–995
Roy C, Lancaster D (2001) Structure and function of succinate:quinone oxidoreductases and the role of quinol:fumarate reductases in fumarate respiration. In: Zannoni D (ed) Respiration in Archaea and bacteria: diversity of prokaryotic electron transport carriers. Kluwer Academic, Dordrecht, pp 57–85
Schultz BE, Chan SI (1998) Thermodynamics of electron transfer in Escherichia coli cytochrome bo3. Proc Natl Acad Sci U S A 95:11643–11648
Stotler RL, Frape SK, Ruskeeniemi T, Ahonen A, Onstott TC, Hobbs MY (2009b) The hydrogeochemistry of groundwaters in and below the base of thick permafrost at Lupin, Nunavut, Canada. J Hydrol (373:80–95)
Stotler, RL (2008) “Evolution of Canadian Shield Groundwaters and Gases: Influence of Deep Permafrost”, PhD Dissertation, Department of Earth and Environmental Sciences, University of Waterloo, Waterloo, Ontario, Canada, 265p
Lippmann J, Stute M, Torgersen T, Moser DP, Hall J, Lin L, Borcsik M, Bellamy RES, Onstott TC (2003) Dating ultra-deep mine waters with noble gases and 36Cl, Witwatersrand Basin, South Africa. Geochim Cosmochim Acta 67:4597–4619
Dyke AS, Prest VK (1987) Late Wisconsian and Holocene history of the Laurentide ice sheet. Geogr Phys Quat 41:237–263
Frape SK, Blyth A, Blomqvist R, McNutt RH, Gascoyne M (2005) Deep fluids in the continents: II. Crystalline rocks. In: Drever JI (ed) Surface and ground water, weathering, and soils, vol. 5. Elsevier, Oxford, pp 542–580
Lin LH, Wang PL, Rumble D, Lippmann-Pipke J, Boice E, Pratt LM, Sherwood Lollar B, Brodie EL, Hazen TC, Andersen GL, DeSantis TZ, Moser DP, Kershaw D, Onstott TC (2006) Long-term sustainability of a high-energy, low-diversity crustal biome. Science 314:479–482
Price PB, Sowers T (2004) Temperature dependence of metabolic rates for microbial growth, maintenance, and survival. Proc Natl Acad Sci U S A 101:4631–4646
Mikucki JA, Priscu JC (2007) Bacterial diversity associated with Blood Falls, a subglacial outflow from the Taylor Glacier, Antarctica. Appl Environ Microbiol 73:4029–4039
Mikucki JA, Pearson A, Johnston DT, Turchyn AV, Farquhar J, Schrag DP, Anbar AD, Priscu JC, Lee PA (2009) A contemporary microbially maintained subglacial ferrous “ocean”. Science 324:397–400
Habicht KS, Gade M, Thamdrup B, Berg P, Canfield DE (2002) Calibration of sulfate levels in the Archean ocean. Science 298:2372–2374
Sorokin D, Kuenen J, Jetten M (2001) Denitrification at extremely high pH values by the alkaliphilic, obligately chemolithoautotrophic, sulfur-oxidizing bacterium Thioalkalivibrio denitrificans strain ALJD. Arch Microbiol 175:94–101
Gascoyne M, Thomas DA (1997) Impact of blasting on groundwater composition in a fracture in Canada's Underground Research Laboratory. J Geophys Res Solid Earth 102:573–584
Stroes-Gascoyne S, Gascoyne M (1998) The introduction of microbial nutrients into a nuclear waste disposal vault during excavation and operation. Environ Sci Technol 32:317–326
Acknowledgments
This work was supported by the NASA Astrobiology Institute through award NNA04CC03A to the IPTAI Team codirected by LMP and TCO, through award NCC21275 to Steve D’Hondt of the University of Rhode Island, and through award NCC21274 to Mike Thomashow of Michigan State University. Funding was also provided by the NASA Astrobiology Institute, NAS NRC (Postdoctoral Associateship 0385260) to C.B. We are indebted to Echo Bay Mines Ltd., Kinross Inc. and the staff of Lupin Au Mine for providing logistical support. The research was carried out in cooperation with a project group (Geological Survey of Finland, the University of Waterloo), which is studying groundwater chemistry under deep permafrost conditions and is funded by Posiva Oy, Finland, Svensk kärnbränslehantering (SKB), Sweden, and the Nuclear Waste Management Organization/Ontario Power Generation, Canada.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Onstott, T.C., McGown, D.J., Bakermans, C. et al. Microbial Communities in Subpermafrost Saline Fracture Water at the Lupin Au Mine, Nunavut, Canada. Microb Ecol 58, 786–807 (2009). https://doi.org/10.1007/s00248-009-9553-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-009-9553-5