Abstract
The information content and responsiveness of microbial biofilm community structure, as an integrative indicator of water quality, was assessed against short-term changes in oxygen and nutrient loading in an open-water estuarine setting. Biofilms were grown for 7-day periods on artificial substrates in the Pensacola Bay estuary, Florida, in the vicinity of a wastewater treatment plant (WWTP) outfall and a nearby reference site. Substrates were deployed floating at the surface and near the benthos in 5.4 m of water. Three sampling events covered a 1-month period coincident with declining seasonal WWTP flow and increasing dissolved oxygen (DO) levels in the bottom waters. Biomass accumulation in benthic biofilms appeared to be controlled by oxygen rather than nutrients. The overriding effect of DO was also seen in DNA fingerprints of community structure by terminal restriction fragment length polymorphism (T-RFLP) of amplified 16S rRNA genes. Ribotype diversity in benthic biofilms at both sites dramatically increased during the transition from hypoxic to normoxic. Terminal restriction fragment length polymorphism patterns showed pronounced differences between benthic and surface biofilm communities from the same site in terms of signal type, strength, and diversity, but minor differences between sites. Sequencing of 16S rRNA gene clone libraries from benthic biofilms at the WWTP site suggested that low DO levels favored sulfate-reducing prokaryotes (SRP), which decreased with rising oxygen levels and increasing overall diversity. A 91-bp ribotype in the CfoI-restricted 16S rRNA gene T-RFLP profiles, indicative of SRP, tracked the decrease in relative SRP abundance over time.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Microbial biofilms are complex communities that can be comprised of bacteria, protozoa, microalgae, and micrometazoa existing in an extracellular polysaccharide matrix. Their development is affected by biotic and abiotic characteristics of the milieu and is therefore integrative of environmental conditions [44]. Many microorganisms in estuarine environments are particle-attached or associated with surfaces [9, 10]. The tremendous surface area available within intertidal marshes, sea grasses, and oyster reefs in addition to shell, sand, and mud benthos makes the contribution of biofilms to estuarine ecosystem function significant.
The use of the biomass and community structure of the eukaryotic microalgal part of periphytic biofilms has been formalized as a standard method in freshwaters [13, 46, 49] although the prokaryotic part of this indicator system has been largely ignored. Due to the tremendous metabolic diversity of prokaryotes, the information content of this component of biofilm community structure is potentially valuable to assess ecosystem condition. Standardization of procedures and validation of responses of prokaryotes would allow the use of biofilms as indicators in not only aerobic photic zones but also in light-limited and low dissolved-oxygen (DO) zones common to most coastal estuaries.
To assess this potential, we hypothesized that estuarine biofilm prokaryotic communities would reflect differences in water-quality parameters over short spatial and temporal scales. The potential of bacterial communities responding to changing environmental conditions has been documented in a limited number of in situ and field studies. Different successions of bacterial populations in microbial mats and a mudflat beach were observed as a response to experimental petroleum exposure [1, 3, 39]. Microbial mats have been reported to reveal noticeable seasonal, site-, and depth-specific differences [29]. Previous work by our group [31] has documented significant differences in habitat-specific microbial diversity of 7-day-old biofilms for two very closely located but distinct benthic habitats: an oyster reef and a muddy sand bottom. In addition to this habitat fidelity, Moss et al. [28] found temporal stability of biofilm prokaryotic community structure within an undisturbed habitat during summer months (often considered an index period for determining ecosystem status). In this report, we studied the response of developing biofilm prokaryotic communities to changing environmental conditions over three consecutive sampling periods.
We took advantage of a “natural experiment” involving a site at a wastewater treatment plant (WWTP) outfall whose effluent volume decreased over time, and a nearby reference site intended to isolate any effect from the added nutrients and organics from the WWTP. Sampling encompassed a 1-month summer period during which DO concentrations in bottom waters at both sites increased from severe hypoxia (0–3 mg DO l−1) to normoxic (3–7 mg DO l−1) conditions.
DO availability appeared to dominate the response of overall bacterial diversity and relative sulfate-reducing prokaryote (SRP) abundance, masking differences due to the WWTP effluent. Although we are far from fully understanding what controls the structure and function of biofilm communities, this study adds to our knowledge of how the prokaryotic community composition of nascent microbial biofilms responds to changes in ambient water conditions.
Materials and Methods
Study Area and Sample Collection
Nascent biofilms were collected at two locations (site 1 and site 2) in Santa Rosa Sound, Florida. Site 1 (30°33′69″N; 87°13′24″W) was an established monitoring station for the outfall of a WWTP. Site 2 (30°34′42″N; 87°11′17″W) was situated 2 km away with equivalent depth (5.4 m) and similar bathymetric features but was separated by a short point of land. Temperature, salinity, and DO were measured with hand-held meters (YSI Inc., Yellow Springs, Ohio, USA or Hach-Hydrolab Corporation, Loveland, CO, USA) in addition to conductivity, temperature, and DO recording sondes (Hach-Hydrolab) for continuous data recording 15 cm over the benthos. Water temperatures ranged between 28.1 and 30.8°C. The oxygen data sonde at site 2 did not pass QA/QC, and the DO recorded by hand held meters was reported. Samples for nutrient analysis of the water column were analyzed by US EPA standard methods (300.00, 300.01, and 300.03) on a Bran–Luebbe autoanalyzer. The data on the volume and nutrient content of the WWTP effluent were provided by the Emerald Coast Utilities Authority.
Polyvinylchloride frames were constructed to hold acrylic plates, which served as artificial substrates for biofilm growth [44]. Surface samplers measured 57 × 15 cm overall, holding 12.7 × 9.5 cm plates 3 cm apart. Benthic samplers measured 28 × 28 cm overall, holding an acrylic rack for 19.5 × 9.5 cm plates 3 cm apart. The difference in size was to account for an anticipated greater biomass on the surface plates. Concrete was molded onto the frames to serve as an anchor and to collect biofilms 10–15 cm above the bottom. An additional set of plates was floated at the surface just under the air–water interface tethered to a benthic unit. Sampling was performed over a 1-month time course with three 7-day growth periods: 13–20 August, 22–29 August, and 5–12 September.
Biofilm Biomass Determination by Elemental Analysis
Biofilm material was scraped from plates, collected on pre-ashed glass fiber filters (Whatman GF/F; nominal pore size 0.7 μm), and frozen at −80°C until analysis. Samples were dried under vacuum and combusted for analysis of carbon and nitrogen content in an elemental analyzer (CE Elantech Flash EA 1200). Carbon and nitrogen content of the biomass was expressed as micromole per square centimeter of plate area.
Genomic DNA Extraction
Biofilm material was collected on membrane filters (Millipore Duropore; pore size 0.2 μm) and frozen at −80°C until analysis. Material on filters was washed off with 2% NaCl and harvested by centrifugation at 7,000 rpm for 10 min in a microcentrifuge. Genomic DNA was extracted by bead beating (15 s, speed 4.5 m s−1) using the FastDNA Spin Kit according to the protocol of the manufacturer (Qbiogene, Montreal, Canada). DNA was extracted from at least three independent plates and pooled before PCR amplification to incorporate any among-slide variability.
Amplification of 16S rRNA Genes
Bacterial small subunit rRNA genes were amplified from purified genomic DNA using the bacteria-specific forward primer 27F (5′-AGAGTTTGATC(A/C)TGGCTCAG-3′, corresponding to Escherichia coli positions 8 to 25; 22) and the universal reverse primer 1492R (5′-GG(C/T)TACCTTGTTACGACTT-3′, corresponding to E. coli positions 1510 to 1492; 22). Two versions of the forward primer 27F were used with different modifications at the 5′ end: For terminal restriction fragment length (T-RFLP) analysis, 6-carboxyfluorescein (6-FAM) was added to allow the detection of terminal fragments; for cloning, the 4-bp sequence CACC was added to allow directional insertion into the vector. PCR amplification was performed in 50 μl reaction mixtures containing 5 ng of sample DNA, 1.25 U of ExTaq polymerase (Takara Mirus Bio, Madison, WI, USA), 1× ExTaq buffer (supplied; with 2 mM MgCl final concentration), 0.2 mM of each deoxynucleoside triphosphate, 2.5 μg bovine serum albumin (Roche Diagnostics, Indianapolis, IN, USA), 2% dimethyl sulfoxide (Acros Organics, Morris Plains, New Jersey, USA), and each primer at 0.2 μM. Identical reaction mixtures without target DNA were used as negative controls and did not yield products. The PCR conditions used were 1.5 min of initial denaturation (94°C) followed by 30 cycles of 94°C for 1 min (denaturation), 45°C for 1 min (annealing), and 72°C for 90 s (elongation) in a TC-312 thermal cycler (Techne Inc., NJ, USA) with a final extension step at 72°C for 7 min. Product lengths were in the expected range of 1,500 to 1,600 bp.
Gel Extraction
All PCR products were separated on 0.8% agarose gels, excised, and purified using the QIAquick Gel Extraction Kit (QIAGEN, Valencia, CA, USA) by following the manufacturer’s instructions.
T-RFLP Analysis
Gel-purified 16S rRNA gene PCR products were enzymatically digested in separate reactions for 4 h at 37°C using 10 U each of CfoI, MspI, or RsaI (Roche Diagnostics). The digested DNA was precipitated with 0.1 volumes of 3 M sodium acetate (pH 5.2) and 2.5 volumes of 95% ethanol at −20°C (overnight) followed by centrifugation at 4°C at 16,000×g for 30 min. The DNA pellet was washed with 1 ml of ice-cold 70% ethanol followed by centrifugation for 5 min at 4°C. After removing all ethanol, the DNA pellet was dried and resuspended in 10 μl water. A 150- to 250-ng DNA aliquot was mixed with 5 μl deionized formamide and 0.5 μl internal lane standard (Mapmarker 1000, Bioventures Inc., Murfreesboro, TN, USA). Samples were denatured at 95°C for 5 min and immediately chilled on ice. The fluorescent labels of the terminal fragments were detected on an ABI Prism 310 Genetic Analyzer (Applied Biosystems) in GeneScan mode (15 kV, 5-s injection, 60°C for 40 min). Terminal restriction fragments (T-RFs), which were <50 bp, >1000 bp, or with a peak height less than 50 fluorescence units, were excluded from the analysis. Fragment sizes were estimated by using the Local Southern Method in GeneScan 3.1 software (Applied Biosystems).
Statistical Analysis of T-RFLP Data
Terminal restriction fragment length polymorphism patterns were analyzed using the DAX software (Van Mierlo Software Consultancy, Eindhoven, The Netherlands). Only peaks over a threshold of 150 DAX units were considered. Peaks from different sites were considered identical if their fragment lengths were within a relative tolerance of 1%. This “binned” dataset was subjected to principal component analysis using PC-ORD version 4 (standard parameters; 25).
Cloning and Sequence Analysis
The 4-bp sequence CACC at the 5′ end of each 16S rRNA amplicon (introduced by the forward primer) allowed directional cloning into pENTR/TOPO vectors (Invitrogen Corp., Carlsbad, CA, USA) by providing the recognition site for topoisomerase I. PCR products were cloned in accordance with the manufacturer’s instructions, and ligation mixtures were used to transform competent E. coli TOP10 (supplied with the cloning kit). Recombinants were selected by using LB agarose plates containing 50 μg kanamycin ml−1. E. coli transformants carrying plasmids with inserts were grown overnight in LB broth with 50 μg kanamycin ml−1. Plasmid DNA was prepared from the clones with a QIAprep Spin Miniprep Kit (QIAGEN, Valencia, CA, USA) according to the manufacturer’s instructions. The insert orientation was verified by PCR amplification (with 1 μl of a 100-fold diluted plasmid preparation as template and PCR conditions as described above for 16S rRNA gene amplification) using primers 27F and M13R (supplied with the cloning kit). Only clones with the desired insert orientation (for which a PCR product was obtained) were sequenced. Sequencing was carried out at the Genomics Technology Support Facility (Michigan State University, Lansing, MI, USA) by using the M13F-20 primer (5′-GTAAAACGACGGCCAG-3′). Sequences with a range of about 600–800 bases were obtained for most clones. The sequenced clone libraries comprised 90 (13–20 August), 100 (22–29 August), and 98 (5–12 September) sequences for benthic biofilms from site 1 and 100 sequences for the benthic biofilm (13–20 August) from site 2. Unaligned sequences were entered into the Blast search program (Blastcl3) of the National Center for Biotechnology Information to obtain closest matches for identification of clones to their lowest possible taxonomic identity.
Results
Growth of Biofilms and Water-Quality Parameters
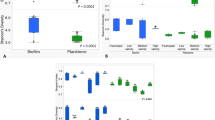
Biofilms from site 1 were developed under higher nutrient and higher organic loads than those from site 2 during the first sampling interval (Fig. 1A, B). The effluent volume from the Pensacola Beach WWTP decreased over the time course studied (Fig. 1A). Dissolved nitrate concentrations were elevated for the benthic water compared to the surface water for the August 20th and 29th incubation periods at both sites but decreased over time (Fig. 1B). Benthic concentrations of nitrate decreased over time in accordance with the decrease in effluent volume. Phosphate levels were higher at both benthic sites relative to surface waters during the first interval but were variable over the remaining 1-month period (data not shown). Salinity values were higher at the benthos for both sites between the 13th and 20th of August (Fig. 2A). This stratification of the water column contributed to sustained low concentrations of DO in the bottom water during the first incubation with minimum values of 0.11 mg/l (site 1) and 0.38 mg/l (site 2). Stratification has also been known to limit benthos-derived nutrients from reaching surface waters. With the subsequent incubations, the stratification broke down, and the variance in DO values at the benthos increased over time (Fig. 2B). Point measurements during the day at site 2 also showed increasing DO concentrations with the subsequent incubations. The dissolved oxygen at the surface was fairly constant ranging between 5.8 and 8 mg/l during the study period.
Effluent volume from Pensacola Beach WWTP over a 3-month period (A) and water nitrate levels over the sampling period at sites 1 and 2 for benthos and surface (B). The 7-day biofilm growth periods are indicated by gray bars. Flow is the amount of wastewater discharged from the WWTP in millions of gallons per day (MGD). The nitrate concentrations were measured in micrograms per liter (μg/L).
Salinity (A) and dissolved oxygen (B) time courses for sample sites 1 and 2 [surface (filled squares) and benthos (filled diamonds)]. The salinity values are given in parts per thousand (ppt), whereas the dissolved oxygen concentrations in milligrams per liter (mg l−1). The 7-day biofilm growth periods are indicated by shaded bars.
Elemental analysis documented relative trends in biofilm biomass (as carbon and nitrogen) at both sites over the study period (Table 1). For the August 20th and 29th incubation periods, biomass was similar between sites for both benthic and surface biofilms, respectively. Surface biofilms had higher biomass than benthic biofilms for the incubation ending on the 29th of August. The biofilms that were recovered on the 12th of September had greater biomass than the first incubation periods for all but the surface sample at site 2, and surface biomass was approximately double than that of benthic biomass. Increased biomass in the benthic samples corresponded to increased oxygen availability. Increased biomass in the surface biofilms is likely a response to destratification of the water column and alleviation of nutrient limitation under high light levels, which is typical for stratified, late summer estuarine water columns.
Biofilm Community Comparisons
Based on 16S rRNA gene T-RFLP fingerprints, low diversity was found in benthic biofilms for the 13–20 August sampling period during the lowest DO recordings (Figs. 2B and 3A), with the WWTP site having twice the number of CfoI-generated peaks as the reference site. Two CfoI-generated T-RFs with 91 and 564 bp were dominant for both site 1 (out of 19 peaks) and site 2 (out of eight peaks). 16S rRNA gene diversities were substantially higher and more similar in benthic biofilms with increasing oxygen concentrations between 22–29 August (45 peaks for site 1, 52 peaks for site 2) and 5–12 September (57 peaks for site 1, 61 peaks for site 2). Patterns obtained with MspI and RsaI showed the identical tendencies in terms of diversity (data not shown). Surface biofilms from the first sampling event were also more dissimilar between sites than with subsequent samples, with a higher number of peaks at site 1 than site 2.
Summary of T-RFLP profiles from PCR-amplified 16S rRNA genes from 7-day-old biofilms at the sample sites 1 and 2 (benthos or surface). Fingerprints are shown for a time scale comprising three growth periods. PCR products were restricted with CfoI. The fragment lengths of prominent peaks are indicated. Fluorescence intensity is shown on the y-axis, whereas fragment length (in bp) is shown on the x-axis.
Principal component analysis of T-RFLP patterns (incorporating the data from all three restriction digestions) supported the visually obvious difference between benthic and surface biofilms (Fig. 4). For benthic biofilms grown under maximum effluent flow (20 August), T-RFLP patterns separated along the axis of the second component from other benthic samples and were identical for both sites (Fig. 4) relative to the other benthic samples, which act as outgroups for these low-DO samples.
Principal component (PC) analysis of T-RFLPs generated from amplified 16S rRNA marker genes from 7-day-old biofilm communities. Biofilms were grown either near the benthos [site 1 (filled squares), site 2 (empty squares)] or near the surface [site 1 (filled triangles), site 2 (empty triangles)]. Dates indicate the corresponding harvest times of the biofilms. The percentage variance explained by PC axes is indicated.
Phylogenetic Community Composition of Benthic Biofilms from Site 1
Sequencing of 16S rRNA gene clones from benthic biofilms grown at site 1 allowed an affiliation of individual community members to phylogenetic groups (Fig. 5). The majority of clones showed highest sequence similarity to the α, γ, and δ subgroups of Proteobacteria or the Cytophaga–Flavobacterium–Bacteroides (CFB) group. Clones that could not be affiliated to these most abundant phylogenetic groups were classified as “Others” including members of the phyla Verrucomicrobia, Pseudomonas, and Holophaga–Acidobacterium.
Relative abundances of phylogenetic groups from benthic biofilms at location 1. The phylogenetic affiliation was derived from sequencing of cloned 16S rRNA genes. Biofilm community members were classified in the α, γ, or δ subdivision of Proteobacteria, the Cytophaga–Flexibacter–Bacteroides (CFB) cluster, and other unknown phylogenetic groups.
The most dramatic change over time occurred in the δ-Proteobacteria. Members of this group in the biofilm grown on 13–20 August accounted for 36% of all clones compared to 16 and 17% for 22–29 August and 5–12 September, respectively. The next largest shifts were an increase in the γ- and α-Proteobacteria as the δ-Proteobacteria decreased (Fig. 5).
The majority of the two dominant phylogenetic groups from site 1, the γ and δ subclasses of Proteobacteria, showed homology either to the functional groups of SRP or the sulfide-oxidizing prokaryotes (SOP) based on their 16S rRNA gene sequences. In the biofilm from the 13–20 August sampling period, potential SRP and SOP represented 36 and 37% of all clones, respectively (Fig. 6). Potential SOP decreased over time from 36 to 27% of all clones. However, the major change observed was within the SRP with the relative abundance of potential SRP changing from 36 to 10% and 14% over time.
Sequences from benthic biofilms harvested at site 1, which were shared between different sampling periods or found more than once in a single clone library, are shown in Table 2. The clone library retrieved from the 13–20 August biofilm showed considerable redundancy correlating with the highly uneven T-RFLP profile: The majority of the γ- and δ-Proteobacteria showed closest similarity to the endosymbiont of Ridgeia piscesae and the uncultured delta proteobacterium clone KorMud-V8C77, making up 21.1 and 24.5% of all sequences, respectively. All other community members, which are not given in Table 2, were found only once in the particular sampling period.
Relative Peak Area of Characteristic T-RF of 91 bp
Nearly without exception, all 16S rRNA gene sequences showing homology to SRP have a CfoI restriction site between positions 94 and 97 starting from the 5′-end of the 27F primer. The resulting T-RF in the actual fingerprint showed up at 91 bp. The sharp drop in the relative abundance of potential SRP over the time course correlated with a similarly dramatic relative decrease of this 91 bp T-RF in the corresponding benthic community fingerprint from site 1. Figure 7 shows the peak areas of this T-RF as a relative percentage of the total peak areas of all peaks in the corresponding fingerprints from all sites.
Relative peak areas of the 91-bp terminal restriction fragments from CfoI-generated 16S rRNA gene T-RFLP fingerprints from biofilms grown at the indicated sites and over the given time periods. The peak areas were calculated as a percentage of the total peak area in the corresponding T-RFLP pattern.
Comparison of Benthic Biofilms from Sites 1 and 2 for 22–29 August
The benthic biofilm from site 2 grown on 22–29 August was further analyzed by cloning and sequencing of amplified 16S rRNA genes to study the effect of the WWTP effluent on the community structure. The substantially higher number 16S rRNA gene sequences from this biofilm could be assigned to the members of the CFB group, when compared to the benthic biofilm from site 1 during the same sampling period (Table 3), at the cost of the γ- and δ-Proteobacteria. Within the δ- and γ-Proteobacteria, 10 and 6 sequences showed similarities to potential SRP and 33 and 28 to potential SOP for sites 1 and 2, respectively. The libraries from site 1 and site 2 with 100 clones each shared only two identical sequences (see Table 2) despite overall similarity in T-RFLP profiles (Fig. 3).
Discussion
This study shows a response of prokaryotic community composition of microbial biofilms to increasing oxygenation and reduction in WWTP effluent volume. This temporal responsiveness of biofilm communities to changing environmental parameters adds a new dimension to a previous finding that 7-day-old estuarine biofilms can exhibit substantial community composition differences over a short spatial scale, dependent on habitat differences where they are incubated [31]. The prokaryotic component of biofilms seems to integrate ambient water conditions early in their development.
The outfall site of a WWTP (site 1) was chosen for two reasons: (1) The fact that effluent volume normally decreases after a summer peak, providing changes in the surrounding water where biofilms were grown; and (2) the potential to study the effect of elevated nutrient and organic loading compared to a reference site with similar hydrologic features. The nitrate concentration was half of what was measured at site 1 (Fig. 1B). The measured nitrate levels suggested that the impact of the outfall was most acute at the benthos and not in the surface water although the T-RFLP fingerprints show a distinct difference between surface sites with maximal WWTP flow that is lost with lower flow and destratification of the water column. Although the dissolved organic matter and the total organic load at the two sites were not determined, elevated dissolved organic carbon (DOC) was likely at the outfall site as well. The existence of moderate site-specific differences in the community composition is exemplified for the benthic biofilms grown on 22–29 August (Table 3).
Sulfate-reducing Prokaryotes
The two overlying gradients at the benthos over the time course, decreasing nutrient and increasing DO concentrations, had dramatic effects on the development of the biofilm communities. Although absolute certainty is elusive in field experiments, the severe hypoxia during 13–20 August in combination with nutrients and DOC from elevated WWTP effluent volume would have created conditions very favorable for SRP, which represented 36% of the total benthic biofilm community at site 1. In the biofilm from the 13–20 August sampling period, more than 24% of all clones showed 98% similarity to both the uncultured delta proteobacterium clone KorMud-V8C77 (first identified from an intertidal mudflat-sediment vegetated by Suaeda japonica) and the Desulfuromonadales bacterium clone SIMO-2442 (first identified in a salt marsh). Dissimilatory sulfate-reducing bacteria are common in estuarine and coastal marine sediments [21, 35, 47] as well as in WWTP systems [23, 40]. They use sulfate as an electron acceptor in the anaerobic oxidation of organic substrates such as lactate, acetate, and propionate, which are mainly the products of fermentative organisms [2, 48]. Although sulfate reduction is principally an anaerobic process, SRP are oxygen tolerant and are readily recovered from oxic habitats [11, 26, 27]. Their activity can dominate anaerobic terminal oxidation of organic matter in high-sulfate environments (>10 mM), degrading up to 50% of all organic matter in coastal marine sediments [21]. Sulfate-reducing prokaryotes in biofilms and sediments have been quantified using fluorescent in situ hybridization (FISH) in other studies showing that SRP can constitute a substantial part of bacterial communities in organic-rich, low-DO environments. For example, around 3–5% of total cells stained by 4′,6-diamidino-2-phenylindole were found to be SRP in a well-established aerated sludge [42]. Okabe et al. [33] report SRP abundances around 10% in their mature wastewater biofilms as determined by FISH analysis explaining their higher numbers as a result of lower DO concentrations when compared to the other wastewater systems mentioned above. Purdy et al. [34, 36] report that SRP account for up to 11 and 24% of the total bacterial communities in the anaerobic sediments of the Colne estuary (United Kingdom) and the River Tama estuary (Japan), respectively. Both estuaries were impacted by treated municipal sewage effluent and other cultural enrichments.
The primers 27F and 1492R used for amplification of 16S rRNA genes are commonly used for phylogenetic community analyses and have not been reported to have a particular bias for SRP although they do integrate all PCR-inherent biases [4, 24].
We found that the change in relative proportions of potential SRP as determined from the clone libraries from site 1 was reflected in the relative peak areas of the CfoI-generated 16S rRNA gene T-RF at 91 bp (Figs. 6 and 7). As this peak was nearly exclusively assigned to potential SRP through in silico digestion of obtained 16S rRNA gene sequences, it appears to indicate their abundance quite well. The same correlation was presented by Urakawa et al. [45] studying microbial communities from Japanese coastal sediments.
Sulfur-oxidizing Bacteria
Sulfate-reducing prokaryotes produce, as a consequence of their metabolic activity, large amounts of sulfide ions and thus create an opportunity for SOP. These organisms represented another dominant group of early colonizers in the 7-day-old biofilms as suggested by the 16S rRNA gene sequence data. This close association of sulfate-reducing bacteria and sulfur-oxidizing bacteria was not surprising considering the fact that both phylogenetically diverse groups of microorganisms are adapted to the conditions found in the bottom waters (e.g., rich in organic matter, neutral pH, and low DO). Whereas the diversity and ecophysiology of SRP have been intensively investigated [32, 33, 40], comparatively little is known about the diversity and ecological importance of SOP involved in the sulfur cycle. Sulfur-oxidizing populations involved in the sulfur cycle are primarily responsible for the oxidation of sulfide and/or elemental sulfur to sulfate in the presence of oxygen [18]. Members of this phylogenetically diverse group have been found in a wide variety of freshwater and marine habitats, such as sediments and microbial mats [6, 7, 15], hydrothermal vent systems [43], ground water and caves [5, 41], and also in wastewater biofilms [19, 22] where reduced sulfur is available. These organisms are also found as symbionts in a wide variety of marine invertebrate taxa [12, 16, 38].
The overwhelming majority of clones in the benthic biofilms from site 1 classified as S-oxidizing γ-Proteobacteria showed close homology to lithoautotrophic bacterial endosymbionts. Free-living sulfide-oxidizing autotrophs have been identified in a variety of sulfide-rich habitats, such as shallow-water marine sediments [15], sewage outfall areas [14, 20], pulp mill effluent sites [37], anoxic basins [14], sea-grass beds [17], and hydrothermal vents [8, 30]. In the biofilm from the 13–20 August sampling period, 21% of all clones show closest similarity to the S-oxidizing symbiont of the hydrothermal-vent tubeworm R. piscesae. In silico restriction of the obtained 16S rRNA gene sequence using the CfoI recognition codon revealed that this clone produces a theoretical T-RF at 564 bp, which can be clearly seen in the corresponding community profile of the 13–20 August biofilm (Fig. 3A).
Concluding Remarks
In summary, we report shifts in biofilm community structure of 7-day-old biofilms associated with changes in DO and local enrichment from a WWTP effluent in an estuarine environment. Among the environmental factors measured, low DO emerged as the dominant factor in this study influencing microbial population dynamics, with differences due to WWTP effluent as a secondary effect. The observations support the hypothesis that nascent microbial biofilm communities cannot only change at short spatial scales but also at short temporal scales. The results raise questions as to whether the microbial pool serving for recruitment of new biofilm assemblages is highly dynamic or if the development process itself is selective of colonizers and responds to changing conditions. Isolating the effect of anthropogenic stresses from other environmental parameters requires further, higher resolution work.
References
Abed, RMM, Safi, NMD, Köster, J, De Beer, D, El-Nahhal, Y, Rullkötter, J, Garcia-Pichel, F (2002) Microbial diversity of a heavily polluted microbial mat and its community changes following degradation of petroleum compounds. Appl Environ Microbiol 68: 1674–1683
Balba, MT, Nedwell, DB (1982) Microbial metabolism of acetate, propionate and butyrate in anoxic sediments from the Colne Point saltmarsh, Essex, UK. J Gen Microbiol 128: 1415–1422
Bordenave, S, Fourçans, A, Blanchard S, Goñi, MS, Caumette, P, Duran, R (2004) Structure and functional analyses of bacterial communities changes in microbial mats following petroleum exposure. Ophelia 58: 195–204
Brakenhoff, RH, Schoenmakers, JGG, Lubsen, NH (1991) Chimeric cDNA clones: a novel PCR artifact. Nucleic Acids Res 19: 1949
Brigmon, RL, Martin, HW, Morris, TL, Britton, G, Zam, SG (1994) Biogeochemical ecology of Thiothrix spp. in underwater limestone caves. Geomicrobiol J 12: 141–159
Brinkhoff, T, Muyzer, G, Wirsen, CO, Kuever, J (1999) Thiomicrospira kuenenii sp. nov. and Thiomicrospira frisia sp. nov., two mesophilic obligately chemolithoautotrophic sulfur-oxidizing bacteria isolated from an intertidal mud flat. Int J Syst Bacteriol 49: 385–392
Brinkhoff, T, Muyzer, G, Wirsen, CO, Kuever, J (1999) Thiomicrospira chilensis sp. nov., a mesophilic obligately chemolithoautotrophic sulfur-oxidizing bacterium isolated from a Thioploca mat. Int J Syst Bacteriol 49: 875–879
Cavanaugh, CM (1983) Symbiotic chemoautotrophic bacteria in marine invertebrates from sulphide-rich habitats. Nature 302: 58–61
Crump, BC, Baross, JA (1996) Particle-attached bacteria and heterotrophic plankton in the Columbia River estuary. Mar Ecol Prog Ser 138: 265–273
Crump, BC, Simenstad, CA, Baross, JA (1998) Particle-attached bacteria dominate the Columbia River estuary. Aquat Microb Ecol 14: 7–18
Cypionka, H, Widdel, F, Pfennig, N (1985) Survival of sulfate-reducing bacteria after oxygen stress growth in sulfate-free oxygen-sulfide gradients. FEMS Microbiol Ecol 31: 39–45
Distel, DL, Lane, DJ, Olsen, GJ, Giovanni, SJ, Pace, B, Pace, NR, Stahl, DA, Felbeck, H (1988) Sulfur-oxidizing bacterial endosymbionts: analysis of phylogeny and specificity by 16S rRNA sequences. J Bacteriol 170: 2506–2510
Eaton, AD, Clesceri, LS, Greenberg, AE (1995) Standard Methods for the Examination of Water and Wastewater, 19th edn. American Public Health Association; American Water Works Association; Water Environment Federation, Washington, D.C.
Felbeck, H, Childress, JJ, Somero, GN (1981) Calvin–Benson cycle and sulphide oxidation enzymes in animals from sulphide-rich habitats. Nature 293: 291–293
Fenchel, T (1969) The ecology of marine microbenthos IV. Structure and function of the benthic ecosystem, its chemical and physical factors and the microfauna communities with special reference to the ciliated Protozoa. Ophelia 6: 1–182
Fenchel, T, Perry, T, Thane, A (1977) Anaerobiosis and symbiosis with bacteria in free-living ciliates. J Protozool 24: 154–163
Fisher, MR, Hand, SC (1984) Chemoautotrophic symbionts in the bivalve Lucina floridana from seagrass beds. Biol Bull 167: 445–459
Friedrich, CG, Rother, D, Bardischewsky, F, Quentmeier, A, Fischer, J (2001) Oxidation of reduced inorganic sulfur compounds by bacteria: emergence of a common mechanism? Appl Environ Microbiol 67: 2873–2882
Ito, T, Sugita, K, Okabe, S (2004) Isolation, characterization, and in situ detection of a novel chemolithoautotrophic sulfur-oxidizing bacterium in wastewater biofilms growing under microaerophilic conditions. Appl Environ Microbiol 70: 3122–3129
Johnson, MA, Diouris, M, Le Pennec, M (1994) Endosymbiotic bacterial contribution in the carbon nutrition of Loripes lucinalis (Mollusca: Bivalvia). Symbiosis 17: 1–13
Jørgensen, BB, Bak, F (1991) Pathways and microbiology of thiosulfate transformations and sulfate reduction in a marine sediment (Kattegat, Denmark). Appl Environ Microbiol 57: 847–856
Lane, DJ (1991) 16S/23S rRNA sequencing. In: Stackebrandt, E, Goodfellow, M (Eds.) Nucleic Acid Techniques in Bacterial Systematics, John Wiley & Sons, Chichester, United Kingdom, pp 115–175
Lens, PN, De Poorter, M-P, Cronenberg, CC, Verstraete, WH (1995) Sulfate reducing and methane producing bacteria in aerobic wastewater treatment systems. Water Res 29: 871–880
Liesack, W, Weyland, H, Stackebrandt, E (1991) Potential risks of gene amplification by PCR as determined by 16S rDNA analysis of a mixed-culture of strict barophilic bacteria. Microb Ecol 21: 191–198
McCune, B, Mefford, MJ (1999) PC-ORD: Multivariate Analysis of Ecological Data. MjM Software Design, Gleneden Beach, Oregon, USA
Minz, D, Fishbain, S, Green, SJ, Muyzer, G, Cohen, Y, Rittmann, BE, Stahl, DA (1999) Unexpected population distribution in a microbial mat community: sulfate-reducing bacteria localized to the highly oxic chemocline in contrast to a eukaryotic preference for anoxia. Appl Environ Microbiol 65: 4659–4665
Minz, D, Flax, JL, Green, SJ, Muyzer, G, Cohen, Y, Wagner, M, Rittmann, BE, Stahl, DA (1999) Diversity of sulfate-reducing bacteria in oxic and anoxic regions of a microbial mat characterized by comparative analysis of dissimilatory sulfite reductase genes. Appl Environ Microbiol 65: 4666–4671
Moss, JA, Nocker, A, Lepo, JE, Snyder, RA (2006) Stability and change in estuarine biofilm bacterial community diversity. Appl Environ Microbiol 72: 5679–5688
Navarrete, A, Urmenta, J, Cantu, JM, Vegas, E, White, DC, Guerrero, R (2004) Signature lipid biomarkers of microbial mats of the Ebro Delta (Spain), Camargue and Étang (France): an assessment of biomass and activity. Ophelia 58: 175–188
Nelson, DC, Fisher, CR (1995) Chemoautotrophic and methanotrophic endosymbiotic bacteria at deep-sea vents and seeps. In: Karl, DM (Ed.) The Microbiology of Deep-Sea Hydrothermal Vents, CRC Press, Boca Raton, FL, pp 125–167
Nocker, A, Lepo, JE, Snyder, RA (2004) Influence of an oyster reef on development of the microbial heterotrophic community of an estuarine biofilm. Appl Environ Microbiol 70: 6834–6845
Okabe, S, Itoh, T, Satoh, H (2003) Sulfate-reducing bacterial community structure and their contribution to carbon mineralization in a wastewater biofilm growing under microaerophilic conditions. Appl Microbiol Biotechnol 63: 322–334
Okabe, S, Itoh, T, Satoh, H, Watanabe, Y (1999) Analyses of spatial distributions of sulfate-reducing bacteria and their activity in aerobic wastewater biofilms. Appl Environ Microbiol 65: 5107–5116
Purdy, KJ, Munson, MA, Embley, TM, Nedwell, DB (2003) Use of 16S rRNA-targeted oligonucleotide probes to investigate function and phylogeny of sulfate-reducing bacteria and methanogenic archaea in a UK estuary. FEMS Microbiol Ecol 44: 361–371
Purdy, KJ, Embley, TM, Nedwell, DB (2002) The distribution and activity of sulphate reducing bacteria in estuarine and coastal marine sediments. Antonie Van Leeuwenhoek 8: 181–187
Purdy, KJ, Nedwell, DB, Embley, TM, Takii, S (2001) Using 16S rRNA-targeted oligonucleotide probes to investigate the distribution of sulphate-reducing bacteria in a Japanese estuary. FEMS Microbiol Ecol 36: 165–168
Reid, R (1980) Aspects of the biology of a gutless species of Solemya (Bivalvia: Protobranchia). Can J Zool 58: 386–393
Rinke, C, Schmitz-Esser, S, Stoecker, K, Nussbaumer, AD, Molnár, DA, Vanura, Wagner, KM, Horn, M, Ott, JA, Brigh, M (2006) “Candidatus Thiobios zoothamnicoli,” an ectosymbiotic bacterium covering the giant marine ciliate Zoothamnium niveum. Appl Environ Microbiol 72: 2014–2021
Röling, WFM, Milner, MG, Martin Jones, D, Fratepietro, F, Swannell, RPJ, Daniel, F, Head, IM (2004) Bacterial community dynamics and hydrocarbon degradation during a field-scale evaluation of bioremediation on a mudflat beach contaminated with buried oil. Appl Environ Microbiol 70: 2603–2613
Santegoeds CM, Ferdelman, TG, Muyzer, G, de Beer, D (1998) Structural and functional dynamics of sulfate-reducing populations in bacterial biofilms. Appl Environ Microbiol 64: 3731–3739
Sarbu, SM, Kinkle, BK, Vlasceanu, L, Kane, TC, Popa, R (1994) Microbiological characterization of a sulfide-rich groundwater ecosystem. Geomicrobiol J 12: 175–182
Schramm, A, Larsen, LH, Revsbech, NP, Ramsing, NB, Amann, R, Schleifer, K-H (1996) Structure and function of a nitrifying biofilm as determined by in situ hybridization and the use of microelectrodes. Appl Environ Microbiol 62: 4641–4647
Sievert, SM, Heidorn, T, Kuever, J (2000) Halothiobacillus kellyi sp. nov., a mesophilic, obligately chemolithoautotrophic, sulfur-oxidizing bacterium isolated from a shallow-water hydrothermal vent in the Aegean sea, and emended description of the genus Halothiobacillus. Int J Syst Evol Microbiol 50: 1229–1237
Snyder, RA, Lewis, MA, Nocker, A, Lepo, JE (2004) Microbial biofilms as integrative sensors of environmental quality. In: Bortone, SA (Ed.) Estuarine Indicators, CRC Press, Boca Raton, FL, pp 111–126
Urakawa, H, Yoshida, T, Nishimura, M, Ohwada, K (2001) Characterization of microbial community in surface marine sediments using terminal restriction fragment length polymorphism (T-RFLP) analysis and quinone profiling. Mar Ecol Prog Ser 220: 47–57
U.S. Environmental Protection Agency (1997) Revision to Rapid Bioassessment Protocols for Use in Streams and Rivers: Periphyton, Benthic Macroinvertebrates, and Fish. EPA 841-D-97-002. United States Environmental Protection Agency, Washington, D.C.
Widdel, F, Bak, F (1992) Gram-negative mesophilic sulfate-reducing bacteria. In: Balows, A, Trüper, HG, Dworkin, M, Harder, W, Schleifer, K-H (Eds.) The Procaryotes: A Handbook on the Biology of Bacteria: Ecophysiology, Identification, Application, Springer-Verlag, NY
Winfrey, MR, Ward, DM (1983) Substrates for sulfate reduction and methane production in intertidal sediments. Appl Environ Microbiol 45: 193–199
Wu, J (1999) A generic index of diatom assemblages as bioindicator of pollution in the Keelung River of Taiwan. Hydrobiologia 397: 79–87
Acknowledgments
We thank Jeff Alison, Laura Pennington, Natasha Rondon, and Matt Wagner for their field assistance and Melissa Hagy for the data management.
This research was supported by a grant from the U.S. Environmental Protection Agency (US EPA)’s Science to Achieve Results (STAR) Estuarine and Great Lakes (EaGLe) Coastal Initiative through funding to the CEER-GOM Project, US EPA Agreement EPA/R-8294580.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Nocker, A., Lepo, J.E., Martin, L.L. et al. Response of Estuarine Biofilm Microbial Community Development to Changes in Dissolved Oxygen and Nutrient Concentrations. Microb Ecol 54, 532–542 (2007). https://doi.org/10.1007/s00248-007-9236-z
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00248-007-9236-z