Abstract
Eukaryotic cells use G protein-coupled receptors (GPCRs) to convert external stimuli into internal signals to elicit cellular responses. However, how mutations in GPCR-coding genes affect GPCR activation and downstream signaling pathways remain poorly understood. Approaches such as deep mutational scanning show promise in investigations of GPCRs, but a high-throughput method to measure rhodopsin activation has yet to be achieved. Here, we scale up a fluorescent reporter assay in budding yeast that we engineered to study rhodopsin’s light-activated signal transduction. Using this approach, we measured the mutational effects of over 1200 individual human rhodopsin mutants, generated by low-frequency random mutagenesis of the GPCR rhodopsin (RHO) gene. Analysis of the data in the context of rhodopsin’s three-dimensional structure reveals that transmembrane helices are generally less tolerant to mutations compared to flanking helices that face the lipid bilayer, which suggest that mutational tolerance is contingent on both the local environment surrounding specific residues and the specific position of these residues in the protein structure. Comparison of functional scores from our screen to clinically identified rhodopsin disease variants found many pathogenic mutants to be loss of function. Lastly, functional scores from our assay were consistent with a complex counterion mechanism involved in ligand-binding and rhodopsin activation. Our results demonstrate that deep mutational scanning is possible for rhodopsin activation and can be an effective method for revealing properties of mutational tolerance that may be generalizable to other transmembrane proteins.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Engineered expression systems that measure the activity of protein variants on a large-scale hold significant potential for linking genotype to phenotype and are powerful tools for studying the molecular evolution of proteins. This is especially true for investigations of monogenic diseases where a single mutation can result in pathogenicity (Antonarakis and Beckmann 2006). One of the most diverse and intensely studied gene products associated with monogenic diseases are G protein-coupled receptors (GPCRs). As cell surface receptors, GPCRs function by detecting molecular cues in the cellular microenvironment to elicit cellular and organismal responses. Mutations in GPCR-coding genes have been found to affect agonist binding, interactions with downstream effectors such as G proteins, subcellular localization, and even GPCR biogenesis and maturation (Stoy and Gurevich 2015). Thus, insights into how GPCR mutations affect any of these processes can provide a better understanding of disease phenotypes and reveal molecular mechanisms that constrain GPCR evolution.
Our current understanding of how GPCRs function comes from both experimental and computational approaches, each presenting strengths and challenges. The most obvious trade-off is reliability versus scalability. At present, experimentally testing variants in live cells are considered to be more reliable than computational methods (Richards et al. 2015; Sun et al. 2016) but are comparatively low throughput. Computational approaches (Kuntz et al. 2022), on the other hand, can rapidly generate predictions concerning variant effects, but lack precision (Weile and Roth 2018). Challenged by these trade-offs, studies aimed at investigating membrane protein properties and their variant effects would benefit greatly from high-throughput experimental assays.
To better understand how missense mutations affect molecular mechanisms of protein function and contribute towards disease, multiplexed assays of variant effect (MAVE) have emerged as a high-throughput approach for accurately mapping many missense variants to function (Starita et al. 2017; Weile and Roth 2018). MAVE combines mutagenesis with high-throughput selection assays to determine the functional effect for thousands, and even millions, of variants in single experiments (Ernst et al. 2010; Fowler et al. 2010, 2014; Hietpas et al. 2013; Weile et al. 2017; Starita et al. 2017). Many MAVE studies to date have primarily focused on globular cytosolic proteins, which have consistently demonstrated that mutational tolerance is associated with solvent accessibility (Guo et al. 2004; Adkar et al. 2012; McLaughlin et al. 2012; Doolan and Colby 2015; Gray et al. 2017). In the case of membrane proteins, such as GPCRs, high-throughput functional characterization is challenging because function is dependent on proper trafficking to the membrane, engagement with an extracellular ligand or signal, and complex conformational changes that facilitate channel opening or interactions with downstream cytoplasmic proteins. Despite these challenges, there have been notable recent advances in high-throughput approaches applied to studying the functional impact of mutations in membrane proteins, including GPCRs (Elazar et al. 2016; Jones et al. 2020; Kozek et al. 2020), but this has not yet been achieved for studies of rhodopsin activation.
Rhodopsin is a GPCR expressed in the rod photoreceptors of the vertebrate eye that, when coupled to an 11-cis retinal chromophore initiated the first critical step in dim-light vision. As an integral membrane protein that requires an exogenous light-sensitive chromophore for function, it has been challenging for the development of high-throughput approaches. Here, we successfully scale up a cell-based fluorescent assay that we previously engineered to measure rhodopsin activation (Scott et al. 2019). Unlike earlier high-throughput studies of rhodopsin variants, which rely on measuring cell surface expression (Wan et al. 2019; Penn et al. 2020; Mckee et al. 2021; Roushar et al. 2022), our assay measures rhodopsin function directly by quantifying receptor-mediated activation via a fluorescent reporter in yeast. Combining our cell-based assay with fluorescence-activated cell sorting and deep sequencing, we were able to screen a library of random mutations in order to obtain functional scores for 1,205 out of 2,156 possible human rhodopsin missense variants accessible through a single-nucleotide change. When compared with RHO disease variants, the majority of variants identified in our screen were found to negatively impact function, with some found to be neutral or to exhibit functional gains. Mapping our variant effect data onto rhodopsin’s three-dimensional structure revealed that mutational tolerance appears inversely correlated with helical packing, which is a property that may be extended to other transmembrane proteins. Lastly, our analyses of variants in rhodopsin’s ligand-binding pocket support a complex counterion mechanism of rhodopsin activation.
Materials & Methods
Yeast Strain
The yeast strain used in this study, BS017, has the genotype W303 MATa, ura3Δ, far1Δ, mfa2::KanMX-pFUS1 mCherry, sst2::HygB, ste2::TRP1, gpa1::Gpa1Gαi1-LEU2 (Scott et al. 2019). These genetic modifications enable the coupling of heterologously expressed rhodopsin to the yeast-mating pathway, and rhodopsin activation to be measured via a fluorescent reporter in vivo.
Library Construction and Cloning
Human RHO libraries were generated by error-prone PCR (GeneMorph II Random Mutagenesis Kit, Agilent). The human RHO sequence (NCBI Accession: NM_000539.3) was used as the template DNA. Primers used to amplify the human RHO gene added flanking AarI restriction sites which facilitated Golden Gate Assembly of mutated PCR products into a single-copy yeast expression vector, which contained a strong constitutive promoter and the URA3 gene to facilitate auxotrophic selection (pRS316 pTDH3). To optimize our libraries for single-nucleotide variants, we tuned the starting template DNA concentration and PCR cycle number (Di Roberto et al. 2017) such that the number of mutations per gene was very low (Table S4 and S5). Using this method, we generated libraries with average 0.9 ± 0.9 mutations per gene (“750 ng-20cyc,” Library 1) and 0.5 ± 0.7 mutations per gene (“1000 ng-30cyc,” Library 2) (Table S4 and S5). We selected these libraries for their low mutation rate (Figure S1) knowing that many genes will not be mutated, as we wanted to discern effects of single mutations. Each library was then transformed into NEB 5-alpha Competent E. coli cells (New England BioLabs) and grown overnight in 250 mL LB carbenicillin (BioShop Canada) followed by plasmid extraction using PureLink HiPure Plasmid Filter Maxiprep Kit (Invitrogen). Test plating of transformed E. coli cells on selective agar plates estimated at least 5 × 105 transformants for each library.
High-Efficiency Transformation of Plasmid Libraries into Yeast Cells
To transform our variant libraries into yeast cells, we used a high-efficiency lithium acetate transformation method (Dong et al. 2010). In brief, yeast cultures were grown to log phase (OD600 ≅ 0.5) in 300 mL YPD (BioShop Canada), resuspended in 0.1 M LiAc/TE to a final volume of 4 mL, and transformed with 48 µg of one of the plasmid libraries via incubation in 30 mL 40% PEG 3350/0.1 M LiAc/TE followed by heat shock. Transformed yeast were cultured in 250 mL SC-URA (BioShop Canada) for 18 h in a shaking incubator at 30 °C. Test plating 1:10, 1:100, and 1:1000 dilutions on SC-URA agar plates were performed to estimate at least 5 × 105 transformants for each library transformation.
Light Activation of Human Rhodopsin Variants in Yeast and Fluorescence-Activated Cell Sorting (FACS)
Yeast transformed with a plasmid library of human rhodopsin variants were diluted to OD600 in 5 mL SC-URA media containing 5 µM 9-cis retinal in a LightSafe 50 mL centrifuge tube (Sigma-Aldrich). Incubation was followed by light exposure and then addition of retinal for a total of six hours, as previously described (Scott et al. 2019). To discriminate between loss of function, mildly deleterious, “wild-type,” and gain of function variants, FACS was performed on a BD Influx (BD Biosciences) where cells were sorted into 4 populations (bins) based on per cell mCherry fluorescence intensity (Figure S2). Using the fluorescence intensity of cells expressing wild-type human rhodopsin and P23H loss of function mutant as controls (Tam and Moritz 2006; Krebs et al. 2010; Chen et al. 2014; Sakami et al. 2014), bins were qualitatively drawn such that wild-type-like variants were primarily sorted into the second “Mid” bin, while loss of function variants fell into the “UltraLow” and “Low” bins (Figure S2). The “High” bin was drawn to resolve gain of function variants. Following sorting, the same light activation and sorting regime was repeated at least one additional time for each sorted subpopulation, to promote sorting refinement (Table S1 and S2). Following the final sorting round, sorted yeast were added to 100 mL SC-URA media and incubated for 24 h. Some sorted cells were also plated to individually assess responses to light following sorting, followed by Sanger sequencing of the mutant rhodopsin genes to evaluate the success of the sorting method.
Plasmid Extraction from Sorted Populations and Sample Preparations for Sequencing
The sorted yeast populations were centrifuged, resuspended in 4 mL Buffer SCE (1 M sorbitol, 0.1 M sodium citrate, 60 mM EDTA, pH 7.0). 8 mg Zymolyase (BioShop Canada) and 30 μL β-mercaptoethanol (Sigma-Aldrich) was added, and the mixture was incubated for 1 h at 37 °C to digest the yeast cell wall and form spheroplasts. Plasmid DNA was then extracted using a QIAPrep Spin Midiprep Kit (Qiagen). To increase plasmid concentrations, the extracted plasmids from each sorted population were transformed into DH5-alpha competent E. coli cells (New England BioLabs) and purified using PureLink HiPure Plasmid Filter Maxiprep Kit (Invitrogen). Serial dilutions prior to plasmid DNA extraction estimated approximately 5,000–90,000 plasmids from each of the sorted yeast populations were recovered (Table S1 and S2).
Illumina Sequencing and Variant Calling
Unsorted and sorted mutation libraries (Table S3) were prepared and sequenced at The Centre for Applied Genomics (Hospital for Sick Children, Toronto ON). In brief, adapters were added with Nextera XT DNA Library Preparation Kit (Illumina), and sequences were obtained with a HiSeq 2500 high-throughput flowcell with 125 bp paired-end reads. Raw reads were trimmed with Trimmomatic v0.36 under default settings. Surviving paired reads were assembled using BWA-mem v0.7.12 against the complete wild-type rhodopsin-containing plasmid sequence, and variants were identified using bcftools v1.6 mpileup command, implemented in a custom pipeline. From the vcf files, the alternate allele count for each alternate allele was divided by the sum of the wild-type allele count and for each specific nucleotide, then multiplied by 100 to get a percentage of how many reads were alternate (mutant) nucleotides. This was performed for each sorted population (UltraLow/Low/Mid/High) for both sorted and unsorted libraries. A heuristic cut-off was determined to be mutations that were above the 90th percentile of plasmid backbone mutations and were also > 25% enriched versus the unsorted library. Sorted sequences from both libraries were combined, and the enrichment was tallied across all populations. A functional score was assigned to each population, based on the average function relative to the wild-type-like population (Figure S2), which was determined by individually testing colonies post-sorting (UltraLow = 0.2, Low = 0.4, Mid = 1.0, High = 1.2) (Supplemental File 2). The “final adjusted activity” was determined by normalizing the functional score of each mutation based on the enrichment of the mutation in each sorted population. This helped to assign scores to sequences that appeared in multiple populations.
Results
Cell Sorting Strategy and High Coverage Deep Sequencing Functionally Characterizes 1,205 Rhodopsin Mutants
Coupling of rhodopsin to an engineered mating pathway in yeast enables measurements of rhodopsin activation using a transcriptionally activated fluorescent readout (Fig. 1A). To determine the variant effect for rhodopsin mutants, we expressed two human RHO gene libraries in our engineered yeast strain and applied FACS-Seq (Fig. 1B and Materials & Methods). Sorted plasmid libraries were then investigated by deep sequencing using Illumina sequencing methods. Samples had > 99% of 125-bp paired-end reads mapped to the human rhodopsin plasmid sequence, with approximately 3 × 107 mapped reads per sorted sample (Table S3), which represented an average coverage of 5.5 × 105 reads at each position in the plasmid (Figure S3). This high sequence coverage of both the rhodopsin gene and the plasmid allowed us to filter out mutations that are likely to have resulted from sequencing and assembly artifacts, and to better identify mutations that have been enriched in sorted populations. Analysis of the reads post-sorting confirmed that mutations were enriched in the rhodopsin gene over the plasmid backbone (Figure S4). Previous studies using error-prone PCR to conduct deep mutational scanning have deemed significance based either on enrichment alone (Romero et al. 2015), or on an estimated error rate in the plasmid antibiotic selection marker (Shin et al. 2014). Here, we opted for a more conservative cut-off (see Materials & Methods). After combining the results of all sorted populations, we identified a total of 1,205 missense mutations.
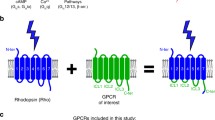
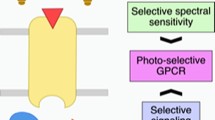
Deep mutational scan of human rhodopsin in yeast. A An engineered yeast-mating pathway for measuring rhodopsin activation in response to light. In the presence of 9-cis retinal chromophore (L) and activation by light, rhodopsin undergoes a conformational change, promoting a Gpa1-Gα protein chimera to undergo GDP to GTP exchange. Gβ and Gγ are then released, initiating MAP kinase signal transduction, pFUS1 transcriptional initiation, and mCherry expression. B Human rhodopsin library construction workflow. Random mutagenesis by error-prone PCR was used to introduce mutations into the human rhodopsin gene. Mutated genes are ligated into a linearized yeast expression vector to generate circular plasmids and transformed into E. coli competent cells. A small sample of the transformed E. coli are plated, which allows for individual colonies to be picked at random. These colonies have their plasmids extracted and sequenced to estimate mutational diversity in each library (Table S5 and S6). The full plasmid library is transformed into yeast, and subjected to 9-cis retinal ligand, followed by light activation, and FACS. Cells were sorted into four gates based on mCherry fluorescence. Finally, plasmids were extracted from each sorted bin and deep sequenced
Sequence-Function Mapping of Rhodopsin Activation
To analyze our collection of rhodopsin mutants, we generated a sequence-function map for all variants identified in our screen. We observed amino acid replacements at 345 out of 348 sites across the protein and 1,205 of 2,424 possible missense single-nucleotide polymorphisms (SNPs) from human rhodopsin (Fig. 2A). To validate the functional scores, we picked 22 missense variants, introduced these mutations into a wild-type rhodopsin plasmid using standard site-directed mutagenesis, and tested each of their responses to light (Figure S5) using a previously reported method (Scott et al. 2019). The functional scores for these 22 mutants range from complete loss of function to superactive. In addition, these 22 mutants have been previously studied and/or reported in clinical variant databases, such as gnomAD (Karczewski et al. 2020) and ClinVar (Landrum et al. 2018) (Supplemental File 3). When the normalized responses for these 22 mutants were compared to the function scores determined by our deep mutational scan, the results from the two methods were found to be correlated (R2 = 0.6272 in Fig. 2B) which indicated the functional scores assigned to sorted populations could help predict the function of individual mutations.
Human rhodopsin functional scores. A A sequence-function map showing the scores associated with each single amino acid substitution generated in the random-mutagenesis library across sites in rhodopsin. Colors are scaled from red for low scores associated with loss of function mutations, to green for wild-type-like, to blue representing activation above wild-type levels. Amino acid states reachable with one SNP, but not characterized in the screen, are colored gray. Amino acid states for more than one SNP from wild-type human rhodopsin are blacked out. A filled black ‘X’ indicates the wild-type residue in human rhodopsin. B Scatter plot of human rhodopsin activation for 22 mutants determined by the deep mutational scan (Library Scores) and flow cytometry results for each mutant tested individually (Normalized mCherry Fluorescence) with error bars showing standard deviation. C Distribution of mutational effect for the SNPs from (A). Black bars represent nonsense mutations
To evaluate how the variant effect for all variants identified in the library are distributed, we categorized each variant based on their functional scores (Fig. 2C). The resulting distribution is bimodal but heavily left skewed, revealing that most variants are deleterious while a small proportion exhibits functional gains. As expected, almost all nonsense mutants are highly deleterious, scoring below 0.4 (Fig. 2C). The eight nonsense variants that scored above 0.4 all introduced a stop codon at or after residue C322, near the end of the RHO gene, which would result in rhodopsin variants with truncated C-termini (Fig. 2C, Supplemental File 3).
Buried Transmembrane Helices Appear More Functionally Conserved than Flanking Helices
Using our data, we sought to understand whether the position where amino acid replacements occur within the protein structure affects mutational tolerance. Due to the stochastic nature of the error-prone PCR method, we focused analyses on general protein domains in rhodopsin. We observed that rhodopsin’s N-terminal tail, helices 2, 3, 4, and 7, and extracellular loop (ECLoop) 2 are the most intolerant to mutations. Helices 1, 5, 6, and intracellular loops 1, 2, and 3 are moderately tolerant, while ECLoop 3 and the C-terminal tail are the most tolerant (Fig. 3A). When we overlaid this information onto the rhodopsin crystal structure, we observed that internal helices 2, 3, and 7 buried towards the center of the protein are mutationally constrained while external helices 1, 5, and 6 that flank the protein core are much more variable (Fig. 3B). Indeed, a direct comparison of the functional scores for internal versus external helices support our observations (Fig. 3C). We note that helix 4 is the only exception, which is positioned as a flanking helix but is mutationally intolerant.
Heterogeneity of mutational tolerance across rhodopsin domains is dependent on position. A Violin plots show the distribution of functional scores for rhodopsin mutants in their respective domains. Black dots in the violins denote the median of functional scores for sites in that domain. Median scores for each domain are also displayed with a gradient of colors reflecting the median functional score ranging from red (lower) to blue (higher). B Domains in the rhodopsin 1U19 crystal structure colored by median functional scores. C Internal helices in rhodopsin’s structure (helices 2, 3, and 7) compared to external “flanking” helices (helices 1, 4, 5, and 6). D Violin plot demonstrating that functional scores in rhodopsin are associated with surface accessibility (relative exposure of sites)
We reasoned that the internal helices might be more functionally conserved compared to external helices because helices closer to the protein center experience dense helical packing. Thus, we calculated the relative surface exposure of each position in rhodopsin using the areaSES command in UCSF Chimera (Pettersen et al. 2004) normalized to the relative exposure value of each amino acid Gly-X-Gly tripeptides (Bendell et al. 2014) and found an association between mutational tolerance and surface exposure (Fig. 3D). These results are functionally consistent because internal helices help to maintain dense side-chain interactions, critical hydrogen bond networks, and the retinal-binding pocket (Janz and Farrens 2004). However, it was exceptionally interesting that helices 5 and 6 are tolerant to mutations (Figs. 3A and 3B) because these two helices are known to undergo the greatest conformational changes when rhodopsin transitions from the dark (inactive) to the metarhodopsin II (active) state (Figure S6). Further, we found the C-terminal tail, a region known to interact with downstream G proteins (Kang et al. 2018; Tsai et al. 2019), to be mutationally tolerant (Figs. 2A and B).
Clinical Variants Cluster
We also explored whether there is any relationship between variant effect, pathogenicity, and where in the protein amino acid replacements occurred. We compared our dataset to a list of clinical rhodopsin variants comprised of data from ClinVar (Landrum et al. 2018), OMIM (Hamosh et al. 2005), and LOVD (Fokkema et al. 2011) (Supplemental File 3). Of the 173 mutants in the collection, we functionally characterized 96 in our screen. Compared with the total distribution of missense mutations in our library, the clinical mutations fell disproportionally into the lower functional classes (Fig. 4A). When categorized according to structural domain, we observed that clinical variants are unevenly distributed across the protein sequence, disproportionately represented within ECLoop 2 and the C-terminal tail. ECLoop 2 and the C-terminal tail contain very high numbers of clinical variants, followed by the N-terminus, helix 1, and helix 5 (Fig. 4B). On the other hand, clinical variants seem to occur less within ICLoops 1 and 3, and ECLoop 3 (Fig. 4B). These results suggest that indeed some clustering of clinical variants exist but not necessarily within the mutationally intolerant domains (Fig. 3).
Clinical variants are disproportionally loss of function. A Proportion of missense variants colored and categorized based on functional score (top). Proportion of clinically identified variants characterized in this study, colored, and categorized based on score (bottom). Number of variants within each category is shown. B Number of clinical variants organized by domain. Stacked bars represent the number of clinical variants characterized by deep mutational scanning in this study (colored) and clinical variants not functionally characterized (gray)
Complex Counterion Mechanism of Rhodopsin Activation Involves Multiple Sites
Previous studies have proposed a complex counterion mechanism where E113 and E181 act as counterions to the Schiff-base link between the retinal chromophore and K296 at different stages of rhodopsin activation (Yan et al. 2003). Our data for variant effect at these three sites (Fig. 5A) in human rhodopsin were consistent with this complex counterion mechanism. We observed mutation-specific functional variability at K296 where K296N, T, Q, and E maintain wild-type-like activity while K296M and R promotes functional loss (Fig. 4B). Of these six K296 mutants, K296E and K296M have been previously studied in detail and were shown to be constitutively active (Robinson et al. 1992; Yang et al. 1997). E113 is known to serve as the primary counterion to the protonated Schiff base located at the covalent bond between K296 and retinal (Sakmar et al. 1989; Kim et al. 2004), while E181 may additionally serve as another counterion involved in a complex counterion shift mechanism (Fig. 5B) (Yan et al. 2003). In our study, we found that E113 is intolerant to mutations except when mutated to Ala (Fig. 5B). However, at site E181, we observed only a mild reduction in function when mutated to Asp while E181K completely abolishes activity (Fig. 5B). For all three sites, synonymous mutations maintained wild-type-like activation levels. Thus, the role of the E113 counterion appears to be highly critical for receptor activation, while E181 is involved but is more tolerant to mutations.
Functional characterization of single amino acid substitutions in human rhodopsin supports complex counterion mechanism. A Structure of rhodopsin focused on the retinal-binding site. 11-cis-retinal is shown in black and is connected to K296 (red) via a Schiff-base linkage. E113 and E181 are highlighted in magenta (PDB: 1U19). B Stick structure showing the complex counterion mechanism, with functional effects of mutations found in our screen color coded as in Fig. 2
Discussion
In this study, we were able to successfully scale up our cell-based fluorescent assay of rhodopsin activation, and to use it to initiate investigations of residue-specific impact of mutations across the visual pigment rhodopsin. Here, we discuss our interpretation of the data and the implications of our observations for other transmembrane proteins.
Mutational Robustness Across Transmembrane Helices Depends on Position
Measuring the functional effect of many missense variants across sites for a protein can provide insight into the relationship between mutability across protein domains. Our results suggest that functional determinants of missense mutations are contingent on both composition and context, the identity of the replaced amino acid and its location within a protein’s three-dimensional structure. Factors such as hydrophobicity, available volume in the core, and steric interactions have long been known as determinants of protein structure (Bowie et al., 1990; Lim & Sauer 1989, 1991). The general rule that the core of globular proteins is more intolerant to mutations compared to surface residues is supported by deep mutational scanning studies of CcdB bacterial toxin and human DNA repair enzyme 3-methyladenine DNA glycosylase (Guo et al. 2004; Adkar et al. 2012; McLaughlin et al. 2012; Doolan and Colby 2015; Gray et al. 2017). However, for integral membrane proteins it remains unclear whether the same rules apply because transmembrane proteins are constrained differently by the lipid bilayer where interactions with hydrophobic side chains promote the formation of stable structures. This has resulted in the idea that transmembrane proteins can be viewed as “inside-out” (Engelman and Zaccai 1980), with hydrophobic residues facing the lipid bilayer and hydrophilic residues facing inwards. In addition, helical packing density and the number of helices per monomer are highly diverse among transmembrane proteins (Eilers et al. 2000; Gimpelev et al. 2004; Lin et al. 2019). With the exception of some recent studies (Glazer et al. 2020; Penn et al. 2020; Jones et al. 2020; Mckee et al. 2021; Roushar et al. 2022), the mutational tolerance of transmembrane proteins remains largely uncharacterized.
In the present study, our results reveal heterogeneity in mutational tolerance among domains in rhodopsin. Internal helices located closer to rhodopsin’s center appear more intolerant to random amino acid change compared to external helices. This result suggests that regions of high helical density may be less tolerant to the introduction of bulky side chains (Schlinkmann et al. 2012; Heydenreich et al. 2015; Athanasiou et al. 2018), which raises the possibility that the physical constraint attributed to dense side-chain packing learned from soluble proteins may be applicable to transmembrane proteins. However, while globular proteins often have tightly packed cores (Chothia 1975; Richards 1977), many transmembrane proteins function as transporters where sufficient volume is required to form a path for molecules to cross the lipid bilayer (Duncan et al. 2019). Near rhodopsin’s center is the retinal-binding pocket where packing arrangements organize to accommodate access, binding, and release of the retinal ligand. Consistent with previous studies (Nakayama and Khorana 1991), data from our screen shows that many amino acid replacements of proximal residues that point into the retinal-binding pocket reduce receptor activation (Figure S7). We also found that these inward facing residues are neither exclusively hydrophobic nor mutationally restricted to hydrophobic residues (Figure S7). These findings suggest that the positional preference for hydrophobic residues examined for soluble proteins (Lim and Sauer 1989) may be quite different for transmembrane proteins, supporting the idea that transmembrane proteins may indeed be thought of as “inside-out” (Engelman and Zaccai 1980).
An exception to our general observations is that we found helix 4 to be mutationally intolerant. Atomic force microscopy topographs (Liang et al. 2003), disulfide cross-linking (Knepp et al. 2012), and synthetic peptide treatment (Jastrzebska et al. 2015) of rhodopsin in native photoreceptor membranes suggest that helices 1, 4, 5, and 8 serve as interaction interfaces for rhodopsin dimerization, which leads us to speculate that these interactions may serve as an explanation for the observed mutational intolerance of helix 4. Similar results concerning helix 4 can also be found from a deep mutational scan of the β2-adrenergic receptor (Jones et al. 2020), another Class A GPCR.
Of special interest is the high degree of mutational tolerance observed for the C-terminal tail and helix 8 for both rhodopsin in the present study and the β2-adrenergic receptor (Jones et al. 2020). This observation is surprising because the GPCR C-terminal tail is known to form intimate contacts with downstream effector proteins such as G proteins (Kang et al. 2018; Tsai et al. 2019) and helix 8 has been suggested to endow some GPCRs with mechanosensation (Erdogmus et al. 2019), an alternative function that has been suggested for opsins (Leung and Montell 2017). One possible reason for this may come from a limitation in our system, which heterologously expresses human rhodopsin in yeast cells instead of a more physiologically relevant cell type. Indeed, earlier studies have shown that rhodopsin is post-translationally modified differently when expressed in yeast compared to mammalian cells (Mollaaghababa et al. 1996). However, considering that mutational tolerance for the C-terminus tail and helix 8 was observed for both the β2-adrenergic receptor (Jones et al. 2020) and rhodopsin despite differences in the approach, mutational completeness, receptor, and model systems, it is also possible that the C-terminal tail of GPCRs are intrinsically tolerant to mutations. Testing more conditions, such as dark versus light or ligand versus no ligand treatments, may reveal reasons behind these observations. Towards this point, differences in membrane expression have previously been observed for rhodopsin variants under retinal versus no retinal conditions (Penn et al. 2020). Furthermore, no deep mutational scanning studies of GPCRs have been conducted in their native cell types, which may serve as an unexplored factor that can yield further insight into the determinants of transmembrane protein mutability.
Functional Variability in the Retinal-Binding Pocket is Consistent with a Complex Counterion Model
Our data is also consistent with a well-studied region of rhodopsin, three sites (K296, E113, E181) involved in a complex mechanism of rhodopsin activation. In rhodopsin, K296 binds retinal through a Schiff-base covalent bond (Hargrave et al. 1983) stabilized by a counterion at site E113 (Sakmar et al. 1989; Zhukovsky and Oprian 1989). The proposed mechanism involves transferring a proton from E181 to E113, switching the counterion from E113 to E181 when rhodopsin transitions from the dark state to the Meta I photointermediate (Yan et al. 2003b). Consistent with previous studies (Keen et al. 1991; Cohen et al. 1992; Robinson et al. 1992; Yang et al. 1997), we observed mutational variability at K296, which suggests that the covalent bond is not required for receptor activation. This contrasts with E113, a highly conserved residue that, even when mutated to E113D, which differs by only a single CH2 in side-chain length, strongly perturbs activation. For E181, mutating to Asp showed only a mild reduction in activation, but only minor changes to side-chain length at E181 are tolerated. E113 is known to be conserved in vertebrate rhodopsins while almost all and invertebrate rhodopsins have either Glu or Asp at position 181 (Terakita et al. 2000, 2004).
Assays of Molecular Function are Important for Predicting Pathogenic Variant Phenotypes
In this study, we used an assay that measured changes in rhodopsin molecular function, which differs from previous high-throughput studies of rhodopsin that measured receptor expression at the plasma membrane (Wan et al. 2019; Penn et al. 2020). Although easier to design experimentally, quantifying plasma membrane expression precludes the accurate classification of variants that are properly expressed at the cell surface but fail to activate. For instance, some variants, such as M39R and N55K (Ramon et al. 2014), alter signaling but escape degradation and are properly trafficked to the plasma membrane. The data presented here will serve as a useful resource when combined with alternative approaches to studying rhodopsin variant effect to better inform and predict disease phenotypes for clinical interpretation.
References
Adkar BV, Tripathi A, Sahoo A et al (2012) Protein model discrimination using mutational sensitivity derived from deep sequencing. Structure 20:371–381. https://doi.org/10.1016/j.str.2011.11.021
Antonarakis SE, Beckmann JS (2006) Focus on monogenic disorders. Nat Rev Genet 7:277–282. https://doi.org/10.1038/nrg1826
Athanasiou D, Aguila M, Bellingham J et al (2018) The molecular and cellular basis of rhodopsin retinitis pigmentosa reveals potential strategies for therapy. Prog Retin Eye Res 62:1–23. https://doi.org/10.1016/j.preteyeres.2017.10.002
Bendell CJ, Liu S, Aumentado-Armstrong T et al (2014) Transient protein-protein interface prediction: datasets, features, algorithms, and the RAD-T predictor. BMC Bioinform 15:1–12. https://doi.org/10.1186/1471-2105-15-82
Bowie JU, Reidhaar-Olson JF, Lim WA, Sauer RT (1990) Deciphering the message in protein sequences: tolerance to amino acid substitutions. Science 247:1306–1310. https://doi.org/10.1126/science.2315699
Chen Y, Jastrzebska B, Cao P et al (2014) Inherent instability of the retinitis pigmentosa P23H mutant opsin. J Biol Chem 289:9288–9303. https://doi.org/10.1074/jbc.M114.551713
Chothia C (1975) Structural invariants in protein folding. Nature 254:304–308. https://doi.org/10.1038/254304a0
Cohen GB, Oprian DD, Robinson PR (1992) Mechanism of activation and inactivation of Opsin: role of Glu113 and Lys296. Biochemistry 31:12592–12601. https://doi.org/10.1021/bi00165a008
Dong S, Rogan SC, Roth BL (2010) Directed molecular evolution of DREADDs: a generic approach to creating next-generation RASSLs. Nat Protoc 5:561–573. https://doi.org/10.1038/nprot.2009.239
Doolan KM, Colby DW (2015) Conformation-dependent epitopes recognized by prion protein antibodies probed using mutational scanning and deep sequencing. J Mol Biol 427:328–340. https://doi.org/10.1016/j.jmb.2014.10.024
Duncan AL, Song W, Sansom MSP (2019) Lipid-dependent regulation of ion channels and G protein-coupled receptors: insights from structures and simulations. Annu Rev Pharmacol Toxicol 60:31–50. https://doi.org/10.1146/annurev-pharmtox-010919
Eilers M, Shekar SC, Shieh T et al (2000) Internal packing of helical membrane proteins. Proc Natl Acad Sci 97:5796–5801. https://doi.org/10.1073/pnas.97.11.5796
Elazar A, Weinstein J, Biran I et al (2016) Mutational scanning reveals the determinants of protein insertion and association energetics in the plasma membrane. Elife 5:1–19. https://doi.org/10.7554/elife.12125
Engelman DM, Zaccai G (1980) Bacteriorhodopsin is an inside-out protein. Proc Natl Acad Sci USA 77:5894–5898. https://doi.org/10.1073/pnas.77.10.5894
Erdogmus S, Storch U, Danner L, et al (2019) Helix 8 is the essential structural motif of mechanosensitive GPCRs. Nat Commun. https://doi.org/10.1038/s41467-019-13722-0
Ernst A, Gfeller D, Kan Z et al (2010) Coevolution of PDZ domain-ligand interactions analyzed by high-throughput phage display and deep sequencing. Mol Biosyst 6:1782–1790. https://doi.org/10.1039/c0mb00061b
Fokkema IFAC, Taschner PEM, Schaafsma GCP et al (2011) LOVD vol 2.0: The next generation in gene variant databases. Hum Mutat 32:557–563. https://doi.org/10.1002/humu.21438
Fowler DM, Araya CL, Fleishman SJ et al (2010) High-resolution mapping of protein sequence-function relationships. Nat Methods 7:741–746. https://doi.org/10.1038/nmeth.1492
Fowler DM, Stephany JJ, Fields S (2014) Measuring the activity of protein variants on a large scale using deep mutational scanning. Nat Protoc 9:2267–2284. https://doi.org/10.1038/nprot.2014.153
Gimpelev M, Forrest LR, Murray D, Honig B (2004) Helical packing patterns in membrane and soluble proteins. Biophys J 87:4075–4086. https://doi.org/10.1529/biophysj.104.049288
Glazer AM, Kroncke BM, Matreyek KA, et al (2020) Deep mutational scan of an SCN5A voltage sensor. Circ Genom Precis Med 20–29. https://doi.org/10.1161/CIRCGEN.119.002786
Gray VE, Hause RJ, Fowler DM (2017) Analysis of large-scale mutagenesis data to assess the impact of single amino acid substitutions. Genetics 207:53–61. https://doi.org/10.1534/genetics.117.300064
Guo HH, Choe J, Loeb LA (2004) Protein tolerance to random amino acid change. Proc Natl Acad Sci 101:9205–9210. https://doi.org/10.1073/pnas.0403255101
Hamosh A, Scott AF, Amberger JS et al (2005) Online mendelian inheritance in man (OMIM), a knowledgebase of human genes and genetic disorders. Nucleic Acids Res 33:514–517. https://doi.org/10.1093/nar/gki033
Hargrave PA, McDowell JH, Curtis DR et al (1983) The structure of bovine rhodopsin. Biophys Struct Mech 9:235–244. https://doi.org/10.1007/BF00535659
Heydenreich FM, Vuckovic Z, Matkovic M, Veprintsev DB (2015) Stabilization of G protein-coupled receptors by point mutations. Front Pharmacol 6:1–15. https://doi.org/10.3389/fphar.2015.00082
Hietpas RT, Bank C, Jensen JD, Bolon DNA (2013) Shifting fitness landscapes in response to altered environments. Evolution (n y) 67:3512–3522. https://doi.org/10.1111/evo.12207
Janz JM, Farrens DL (2004) Role of the retinal hydrogen bond network in rhodopsin Schiff base stability and hydrolysis. J Biol Chem 279:55886–55894. https://doi.org/10.1074/jbc.M408766200
Jastrzebska B, Chen Y, Orban T et al (2015) Disruption of rhodopsin dimerization with synthetic peptides targeting an interaction interface. J Biol Chem 290:25728–25744. https://doi.org/10.1074/jbc.M115.662684
Jones EM, Lubock NB, Venkatakrishnan A et al (2020) Structural and functional characterization of G protein–coupled receptors with deep mutational scanning. Elife 9:1–28. https://doi.org/10.7554/eLife.54895
Kang Y, Kuybeda O, De Waal PW et al (2018) Cryo-EM structure of human rhodopsin bound to an inhibitory G protein. Nature 558:553–558. https://doi.org/10.1038/s41586-018-0215-y
Karczewski KJ, Francioli LC, Tiao G et al (2020) The mutational constraint spectrum quantified from variation in 141,456 humans. Nature 581:434–443. https://doi.org/10.1038/s41586-020-2308-7
Keen TJ, Inglehearn CF, Lester DH et al (1991) Autosomal dominant retinitis pigmentosa: four new mutations in rhodopsin, one of them in the retinal attachment site. Genomics 11:199–205. https://doi.org/10.1016/0888-7543(91)90119-Y
Kim JM, Altenbach C, Kono M et al (2004) Structural origins of constitutive activation in rhodopsin: role of the K296/E113 salt bridge. Proc Natl Acad Sci U S A 101:12508–12513. https://doi.org/10.1073/pnas.0404519101
Knepp AM, Periole X, Marrink SJ et al (2012) Rhodopsin forms a dimer with cytoplasmic helix 8 contacts in native membranes. Biochemistry 51:1819–1821. https://doi.org/10.1021/bi3001598
Kozek KA, Glazer AM, Ng CA et al (2020) High-throughput discovery of trafficking-deficient variants in the cardiac potassium channel KV11.1. Heart Rhythm 17:2180–2189. https://doi.org/10.1016/j.hrthm.2020.05.041
Krebs MP, Holden DC, Joshi P et al (2010) Molecular mechanisms of rhodopsin retinitis pigmentosa and the efficacy of pharmacological rescue. J Mol Biol 395:1063–1078. https://doi.org/10.1016/j.jmb.2009.11.015
Kuntz CP, Woods H, McKee AG et al (2022) Towards generalizable predictions for G protein-coupled receptor variant expression. Biophys J 121:2712–2720. https://doi.org/10.1016/J.BPJ.2022.06.018
Landrum MJ, Lee JM, Benson M et al (2018) ClinVar: Improving access to variant interpretations and supporting evidence. Nucleic Acids Res 46:D1062–D1067. https://doi.org/10.1093/nar/gkx1153
Leung NY, Montell C (2017) Unconventional Roles of Opsins. Annu Rev Cell Dev Biol 33:241–264. https://doi.org/10.1146/annurev-cellbio-100616-060432
Liang Y, Fotiadis D, Filipek S et al (2003) Organization of the G protein-coupled receptors rhodopsin and opsin in native membranes. J Biol Chem 278:21655–21662. https://doi.org/10.1074/jbc.M302536200
Lim WA, Sauer RT (1989) Alternative packing arrangements in the hydrophobic core of λrepresser. Nature 339:31–36. https://doi.org/10.1038/339031a0
Lim WA, Sauer RT (1991) The role of internal packing interactions in determining the structure and stability of a protein. J Mol Biol 219:359–376. https://doi.org/10.1016/0022-2836(91)90570-V
Lin Y-C, Guo YR, Miyagi A et al (2019) Force-induced conformational changes in PIEZO1. Nature 573:230–234. https://doi.org/10.1038/s41586-019-1499-2
Mckee AG, Kuntz CP, Ortega JT et al (2021) Systematic profiling of temperature- and retinal-sensitive rhodopsin variants by deep mutational scanning. J Biol Chem 297:101359. https://doi.org/10.1016/j.jbc.2021.101359
McLaughlin RN, Poelwijk FJ, Raman A et al (2012) The spatial architecture of protein function and adaptation. Nature 491:138–142. https://doi.org/10.1038/nature11500
Mollaaghababa R, Davidson FF, Kaiser C, Khorana HG (1996) Structure and function in rhodopsin: expression of functional mammalian opsin in Saccharomyces cerevisiae. Proc Natl Acad Sci U S A 93:11482–11486. https://doi.org/10.1073/pnas.93.21.11482
Nakayama TA, Khorana HG (1991) Mapping of the amino acids in membrane-embedded helices that interact with the retinal chromophore in bovine rhodopsin. J Biol Chem 266:4269–4275. https://doi.org/10.1016/S0021-9258(20)64317-4
Penn WD, McKee AG, Kuntz CP et al (2020) Probing biophysical sequence constraints within the transmembrane domains of rhodopsin by deep mutational scanning. Sci Adv 6:1–10. https://doi.org/10.1126/sciadv.aay7505
Pettersen EF, Goddard TD, Huang CC et al (2004) UCSF Chimera - a visualization system for exploratory research and analysis. J Comput Chem 25:1605–1612. https://doi.org/10.1002/jcc.20084
Ramon E, Cordomi A, Aguilà M et al (2014) Differential light-induced responses in sectorial inherited retinal degeneration. J Biol Chem 289:35918–35928. https://doi.org/10.1074/jbc.M114.609958
Richards FM (1977) Areas, volumes, packing and protein structure. Annu Rev Biophys Bioeng 6:151–176
Richards S, Aziz N, Bale S et al (2015) Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the american college of medical genetics and genomics and the association for molecular pathology. Genet Med 17:405–424. https://doi.org/10.1038/gim.2015.30
Di Roberto RB, Scott BM, Peisajovich SG (2017) Directed Evolution Methods to Rewire Signaling Networks. 1596: 321–337
Robinson PR, Cohen GB, Zhukovsky EA, Oprian DD (1992) Constitutively active mutants of rhodopsin. Neuron 9:719–725. https://doi.org/10.1016/0896-6273(92)90034-B
Romero PA, Tran TM, Abate AR (2015) Dissecting enzyme function with microfluidic-based deep mutational scanning. Proc Natl Acad Sci U S A 112:7159–7164. https://doi.org/10.1073/pnas.1422285112
Roushar FJ, Mckee AG, Kuntz CP, et al (2022) Molecular basis for variations in the sensitivity of pathogenic rhodopsin variants to 9-cis-retinal. https://doi.org/10.1016/j.jbc.2022.102266
Sakami S, Kolesnikov AV, Kefalov VJ, Palczewski K (2014) P23H opsin knock-in mice reveal a novel step in retinal rod disc morphogenesis. Hum Mol Genet 23:1723–1741. https://doi.org/10.1093/hmg/ddt561
Sakmar TP, Franke RR, Khorana HG (1989) Glutamic acid-113 serves as the retinylidene Schiff base counterion in bovine rhodopsin. Proc Natl Acad Sci U S A 86:8309–8313. https://doi.org/10.1073/pnas.86.21.8309
Schlinkmann KM, Honegger A, Tureci E et al (2012) Critical features for biosynthesis, stability, and functionality of a G protein-coupled receptor uncovered by all-versus-all mutations. Proc Natl Acad Sci 109:9810–9815. https://doi.org/10.1073/pnas.1202107109
Scott BM, Chen SK, Bhattacharyya N et al (2019) Coupling of human rhodopsin to a yeast signaling pathway enables characterization of mutations associated with retinal disease. Genetics 211:597–615. https://doi.org/10.1534/genetics.118.301733
Shin HS, Cho Y, Choe DH et al (2014) Exploring the functional residues in a flavin-binding fluorescent protein using deep mutational scanning. PLOS ONE 9:1–10. https://doi.org/10.1371/journal.pone.0097817
Starita LM, Ahituv N, Dunham MJ et al (2017) Variant interpretation: functional assays to the rescue. Am J Hum Genet 101:315–325. https://doi.org/10.1016/j.ajhg.2017.07.014
Stoy H, Gurevich VV (2015) How genetic errors in GPCRs affect their function: possible therapeutic strategies. Genes Dis 2:108–132. https://doi.org/10.1016/j.gendis.2015.02.005
Sun S, Yang F, Tan G et al (2016) An extended set of yeast-based functional assays accurately identifies human disease mutations. Genome Res 26:670–680. https://doi.org/10.1101/gr.192526.115
Tam BM, Moritz OL (2006) Characterization of rhodopsin P23H-induced retinal degeneration in a Xenopus laevis model of retinitis pigmentosa. Invest Ophthalmol vis Sci 47:3234–3241. https://doi.org/10.1167/iovs.06-0213
Terakita A, Yamashita T, Shichida Y (2000) Highly conserved glutamic acid in the extracellular IV-V loop in rhodopsins acts as the counterion in retinochrome, a member of the rhodopsin family. Proc Natl Acad Sci 97:14263–14267. https://doi.org/10.1073/pnas.260349597
Terakita A, Koyanagi M, Tsukamoto H et al (2004) Counterion displacement in the molecular evolution of the rhodopsin family. Nat Struct Mol Biol 11:284–289. https://doi.org/10.1038/nsmb731
Tsai CJ, Marino J, Adaixo R et al (2019) Cryo-EM structure of the rhodopsin-Gαi- βγ complex reveals binding of the rhodopsin C-terminal tail to the gβ subunit. Elife 8:1–19. https://doi.org/10.7554/eLife.46041.001
Wan A, Place E, Pierce EA, Comander J (2019) Characterizing variants of unknown significance in rhodopsin: a functional genomics approach. Hum Mutat 40:1127–1144. https://doi.org/10.1002/humu.23762
Weile J, Roth FP (2018) Multiplexed assays of variant effects contribute to a growing genotype–phenotype atlas. Hum Genet 137:665–678. https://doi.org/10.1007/s00439-018-1916-x
Weile J, Sun S, Cote AG et al (2017) A framework for exhaustively mapping functional missense variants. Mol Syst Biol 13:957. https://doi.org/10.15252/msb.20177908
Yan ECY, Kazmi MA, Ganim Z et al (2003) Retinal counterion switch in the photoactivation of the G protein-coupled receptor rhodopsin. Proc Natl Acad Sci 100:9262–9267. https://doi.org/10.1073/pnas.1531970100
Yang T, Snider BB, Oprian DD (1997) Synthesis and characterization of a novel retinylamine analog inhibitor of constitutively active rhodopsin mutants found in patients with autosomal dominant retinitis pigmentosa. Proc Natl Acad Sci 94:13559–13564. https://doi.org/10.1073/pnas.94.25.13559
Zhukovsky E, Oprian D (1989) Effect of carboxylic acid side chains on the absorption maximum of visual pigments. Science 246:928–930. https://doi.org/10.1126/science.2573154
Acknowledgements
This work was by Ontario Graduate Scholarships (B.M.S., S.K.C.), a Vision Science Research Program Fellowship (S.K.C.), and an NSERC Discovery Grant (B.S.W.C.).
Author information
Authors and Affiliations
Contributions
Conceptualization: BMS, SKC, SGP, and BSWC; Methodology and investigation: BMS, SKC, and JL; Bioinformatics analyses: BMS and RKS; Visualization: AVN; Clinical variant data acquisition and curation: EH; Writing: SKC and BSWC, with input from BMS; Supervision: BSWC and SGP; All authors have read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing Interests
All authors declare no competing interests.
Additional information
Handling editor: David Liberles.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Scott, B.M., Chen, S.K., Van Nynatten, A. et al. Scaling up Functional Analyses of the G Protein-Coupled Receptor Rhodopsin. J Mol Evol 92, 61–71 (2024). https://doi.org/10.1007/s00239-024-10154-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-024-10154-3