Abstract
Hypoxia-inducible factor 1 (HIF-1) is a crucial transcriptional factor that can restore oxygen balance in the body by regulating multiple vital activities. Two HIF-1α copies were retained in cyprinid fish after experiencing a teleost-specific genome duplication. How the “divergent collaboration” of HIF-1αA and HIF-1αB proceeds in regulating mitophagy and apoptosis under hypoxic stress in cells of cyprinid fish remains unclear. In this study, zebrafish HIF-1αA/B expression plasmids were constructed and transfected into the epithelioma papulosum cyprini cells and were subjected to hypoxic stress. HIF-1αA induced apoptosis through promoting ROS generation and mitochondrial depolarization when cells were subjected to oxygen deficiency. Conversely, HIF-1αB was primarily responsible for mitophagy induction, prompting ATP production to mitigate apoptosis. HIF-1αA did not induce mitophagy in the mitochondria and lysosomes co-localization assay but it was involved in the regulation of different mitophagy pathways. Over-expression of HIF-1αA increased the expression of bnip3, fundc1, Beclin1, and foxo3, suggesting it has a dual role in mitochondrial autophagy and cell death. Each duplicated copy also experienced functional divergence and target shifting in the regulation of complexes in the mitochondrial electron transport chain (ETC). Our findings shed light on the post-subfunctionalization function of HIF-1αA and HIF-1αB in zebrafish to fine-tune regulation of mitophagy and apoptosis following hypoxia exposure.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Hypoxia-inducible factor 1 (HIF-1) is a crucial important transcriptional factor that can restore oxygen balance in the body by regulating the production of red blood cells, blood vessels, glycolysis, and apoptosis (Schofield and Ratcliffe 2004; Semenza 2014). Being a heterodimer composed of a specific α subunit and a ubiquitous β subunit, the transcriptional activity of HIF-1 predominantly depends on the stability of the α-subunit, which is functionally inhibited and degraded by prolyl hydroxylase enzymes under normoxia (Schofield and Ratcliffe 2004; Webb et al. 2009). Invertebrates possess only one HIF-α, while most jawed vertebrates possess the other two functional HIF-α isoforms, HIF-2α and HIF-3α, after having undergone two rounds of whole genome duplication (WGD) during evolution (Jiang et al. 2001; Loenarz et al. 2011; Schofield and Ratcliffe 2004). Having experienced a specific third round of WGD early in their evolution, teleosts generated two copies of each HIF-α isoform (HIF-αA/B). This was followed by the loss of one copy of each paralogous pair in most teleosts, except for cyprinids. Therefore, two HIF-1αs, HIF-1αA and HIF-1αB were retained in cyprinid fish (Rytkonen et al. 2011). (Rytkonen et al. 2013) The relaxation of negative selection on the oxygen sensing properties of the cyprinid hif-1αA coding sequence, coupled with the loss of developmental regulatory motifs in B paralogs, suggests that cyprinid paralogs may have undergone subfunctionalization, potentially serving distinct functions both in development and in responding to hypoxia (Rytkonen et al. 2013), including regulation of cellular functions like apoptosis and mitophagy.
Mitochondria play a central role in energy supply, reactive oxygen species (ROS) generation, and apoptosis and are thus a pivotal part of the cellular response to low oxygen. ROS is highly active and has irreversible effects on intracellular proteins, lipids, and DNA, causing cell damage that leads to more ROS production. Excessive ROS can inhibit the activity of PHD, an enzyme that targets and triggers degradation of HIF-1α, which leads to HIF-1α stabilization and activation, promoting HIF-mediated gene expression (Majmundar et al. 2010; Nathan and Cunningham-Bussel 2013; Sangbin et al. 2016; Zepeda et al. 2013). Meanwhile, HIF-1 can regulate the generation of ROS by unbalancing the electron flow in the Electron Transport Chain (Han et al. 2017; Kung-Chun Chiu et al. 2019). Accumulated ROS may not be quickly eliminated under hypoxia, which would exacerbate signals for autophagy (Ruth and Zvulun 2011).
Mitophagy, a cellular process that selectively removes damaged or superfluous mitochondria, is an essential process for mitochondrial quality control and dynamics (Lemasters 2005). Hypoxia is one of the key environmental stressors that can lead to mitophagy. Mitochondrial protein NIX (also known as BNIP3L) is a selective autophagy receptor that can be strongly induced by hypoxia (Zhang and Ney 2009). NIX promotes the loss of mitochondrial membrane potential (MMP) and enables the entry of mitochondria into autophagosomes for clearance (Sandoval et al. 2008). This process is also firmly established in hypoxia-induced cell death alongside BNIP3, which plays a critical role in mitochondrial depolarization and autophagy in numerous cell lines (Daido et al. 2004; Hamacher-Brady et al. 2007; Kanzawa et al. 2005). The transcriptional and translational mechanism of both NIX and BNIP3 are regulated by hypoxia and HIF-1α, and their expression is thus associated with tumor cell death (Sowter et al. 2001). The PINK1-Parkin pathway also mediates mitophagy to eliminate those mitochondria which have lost membrane potential in response to acute stress (Vincow et al. 2013). Parkin is a ubiquitin enzyme that labels damaged mitochondria by ubiquitinating outer mitochondrial membrane proteins (Tanaka et al. 2010; Yoshii et al. 2011), which are targeted by autophagy proteins ATG8 and LC3 to form autophagosomes and then subsequently degraded by lysosomes (Sylvie et al. 2017). The activation of the PINK1-Parkin pathway is hypoxia independent, but it is strongly affected by hypoxia and ROS (Liu et al. 2021; Wu et al. 2021; Xu et al. 2021; Zeb et al. 2021). FUNDC1, an outer mitochondrial membrane protein, is another mitochondrial receptor that mediates hypoxia-induced mitophagy. After being dephosphorylated, FUNDC1 induces the formation of autophagic vesicles by interacting with LC3 in response to hypoxia (Chen et al. 2014; Huifang et al. 2015; Liu et al. 2012b).
Although it has been demonstrated in mammals that HIF-1α is involved in ROS generation and hypoxia-induced mitophagy, how the specific “divergent collaboration” of HIF-1αA and HIF-1αB operates in regulating mitophagy induced by hypoxic stress in cyprinids remains unknown. To address this, we transfected zebrafish HIF-1αA and HIF-1αB into the EPC cells and investigated how they function in mitophagy regulation under induced hypoxic stress.
Material and Methods
Cell Culture and Viability Determination
Epithelioma papulosum cyprini (EPC) cells (obtained from the Center of Aquatic Cell Research and Preservation, Huazhong Agricultural University) were cultured in M199 medium (Gibco, Suzhou, China) with 10% fetal bovine serum (Hyclone, Logan, USA) and 1% Penicillin–Streptomycin (Invitrogen, Carlsbad, USA) at 28 °C. The EPC cells were passaged after reaching a minimum of 70% confluence to fresh ∼ 5 mL medium in 25 cm2 cell culture flasks (Sigma-Aldrich) with 3 replicates by a 1:2 ratio. Cell viability was detected using the CCK-8 cell counting kit (Beyotime, Shanghai, China). Cells were inoculated into 96-well plates and incubated under 20% O2, 10% O2, 3% O2, 1% O2, and 0.3% O2, respectively (InvivO2 300, Baker Ruskinn, Britain). Cell viability was measured every 6 h according to the manufacturer's instruction. The absorbance at 450 nm was detected by a multifunctional microplate reader SpectraMax M5 (Molecular Devices, Sunnyvale, CA, USA).
Plasmid Construction and Transfection
The indicated Myc-tagged Hif-1α constructs, pTK-Renilla luciferase reporter, HRE (hypoxia-responsive elements)-luciferase reporter, and pCMV-Myc vector were obtained from the Institute of Hydrobiology, Chinese Academy of Sciences. The full length of cDNA sequence of zebrafish HIF-1αA (NM_001308559.1) and HIF-1αB (NM_001310042.1) were subcloned into the pCMV-Myc vector (Clontech. Palo Alto, CA), and all recombinant plasmids were verified by DNA sequencing. We adopted the dual luciferase reporter assay system for transfection efficiency confirmation. The transfected HIF-1αA/B binds to the HRE, subsequently promoting the expression of firefly luciferase gene. Simultaneously, the constitutively expressing Renilla luciferase gene was used as an internal control for normalization. Eighteen hours before transfection, EPCs were seeded in 12-well plates with M199 medium (10% FBS medium, no antibiotics), then co-transfected with the indicated Myc-tagged HIF-1α constructs (HIF-1αA, HIF-1αB or empty vector), HRE-luciferase reporter, and pTK-Renilla luciferase reporter by using Lipofectamine 2000 transfection reagent (Invitrogen, USA). After transfection, the culture medium was removed and replaced with serum-free medium for 4 h, then replaced with 10% FBS medium, and incubated under 10% O2 for 24 h. Luciferase activity was detected by using a Dual Luciferase Reporter Assay System (Promega, Madison, WI, USA). The fraction of positively transfected cells was determined by counting. The calculation of the firefly-to-Renilla luciferase activity ratio for each sample serves as a reflection of the transfection efficiency of our target genes.
Measurement of Intracellular ROS
The reactive oxygen species (ROS) detection kit (Beyotime, Shanghai, China) was used to detect mitochondrial ROS generation. Cells were collected in a hypoxia operating system after 24 h incubation, washed twice with serum-free medium, added with diluted DCFH-DA working solution, and incubated for 20 min in an incubator at 28 °C in the dark. The cells were washed with serum-free medium for three times and then analyzed using a multifunctional microplate reader SpectraMax M5 (Molecular Devices, Sunnyvale, CA, USA). The ROS generation was also analyzed by Guava easyCyte 8HT flow cytometry (Merck Millipore, Billerica, MA, USA) at the same time.
Mitochondrial Membrane Potential Assay
The MMP was detected using a JC-1 kit (Beyotime, Shanghai, China). The transfected cells were incubated under 10% O2 for 24 h and were then collected and incubated with 500 μl JC-1 staining solution for 30 min in the dark at 28 °C and monitored under a Leica TCS SP confocal microscope. The wavelength of excitation/emission light was set to 490/530 nm for JC-1 monomer detection and was set to 525/590 nm for JC-1 polymer detection.
Apoptosis Detection
The Annexin V-FITC/PI apoptosis kit (Yeasen, Shanghai, China) was used for apoptosis detection. Cells were incubated under 10% O2 for 24 h after transfection and then approximately 5 × 105 cells were collected. After resuspension, cells were incubated with 5 μL Annexin V-FITC and 5 μL PI Staining Solution for 10 min at room temperature in the dark and analyzed by Guava easyCyte 8HT flow cytometry (Merck Millipore, Billerica, MA, USA).
Mitophagy Detection by Mitochondria and Lysosomes Co-localization Assay
EPC cells were seeded at a density 1 × 105 on the 6 micro-well glass bottom plate (n = 3) after transfection and incubated under 10% O2 for 24 h. Cells were stained with 50 nm Lyso-Tracker Red (Beyotime, Shanghai, China) at 28 °C for 2 h. This was followed by the addition of 100 nm Mito-Tracker Green (Beyotime) and an incubation at 28 °C for 30 min. Cells were then fixed with 4% paraformaldehyde. After fixation, cells were treated with DAPI (Beyotime, Shanghai, China) for 3–5 min in room temperature, then washed with PBS for 2–3 times, each time 3–5 min. After treatment, cells were monitored under a Leica TCS SP confocal microscope.
Measurement of Intracellular ATP Content
The generation of ATP in cells was measured using an enhanced ATP assay kit (Beyotime, Jiangsu, China) according to the manufacturer’s instructions. ATP is measured using a luciferase-luciferin reaction assay. Cells were treated with 10% O2 for 24 h after transfection, then split by the lysis reagent, centrifugated at 12,000 g for 5 min to obtain supernatant. The protein concentration of samples was detected by BCA protein assay kit (Beyotime, Jiangsu, China). The chemical fluorescence intensity of samples was measured by a multifunctional microplate reader SpectraMax M5 (Molecular Devices, Sunnyvale, CA, USA) and the ATP concentration was calculated according to the standard curve.
Total RNA Isolation and Real-Time Quantitative PCR
Total RNA was extracted by Trizol reagent (Biosharp, Guangzhou, China). The quality and concentration of RNA were determined by NanoDrop 2000 (Thermo Scientific, Waltham, MA, USA) and agarose gel electrophoresis. An amount of 1 μg total RNA was reverse transcribed into cDNA using a cDNA synthesis kit (Vazyme, Nanjing, China). The steady-state mRNA levels were detected by Real-Time quantitative PCR based on SYBR green detection (Vazyme, Nanjing, China) with quantitative thermal cycler (MyiQTM 2 Two Color Quantity PCR Detection System, Bio-Rad, Hercules, CA, USA). Primers are listed in Table S1, supplementary files. Calculations for relative expression were performed using the 2−∆∆Ct method.
Statistical Analysis
All the experiments were presented at least three times with a minimum of three repeats. Statistical analysis was conducted with SPSS 16.0 software. Results were performed as means ± standard error (SE). The data were presented by one-way analysis of variance (ANOVA) and followed by Tukey’s multiple range tests. In each analysis, a value of P less than 0.05 was considered to indicate the statistical significance.
Results
Hypoxia Reduces Cell Viability and Induces ROS Production
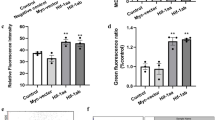
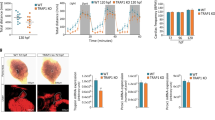
The ability to tolerate hypoxia varies greatly among different cell types. To determine the viability of cells under different hypoxia stress, a CCK-8 assay was performed after the EPCs were treated with different oxygen levels (0.3%, 1%, 3%, 10%, 20%). Morphologically, the shape and density of EPCs were severely affected after being treated in acute hypoxia (0.3%, 1%, 3% O2) for 24 h, and over 50% of cells died in 0.3% O2 by the end of 48 h, whereas cells under 10% O2 were not inhibited (Fig. 1A). We also examined mitochondrial ROS generation following a 24 h exposure of cells to varying oxygen concentrations. In comparison to normoxia, ROS production in cells exhibited a significant increase under 0.3%, 1%, and 3% hypoxia. However, ROS production remained unchanged at 10% O2 (Fig. 1B). The CCK-8 assay results (Fig. 1C) were in accordance with the cell morphology. The cell viability was not affected by 3% and 1% O2 after 12 h but sharply declined in 0.3% O2 group. After 24 h, the viability of all the cells suffered from acute hypoxia and viability was inhibited. However, cell viability under 10% O2 was relatively high, reaching a peak at 24 h. The viability remained high over 48 h, which may have been due to the stress response to the hypoxia. These findings indicate that the viability of EPCs is influenced by hypoxia in a manner dependent on both time and oxygen concentration. In light of these results, we opted for the 10% O2 treatment for our subsequent experiments.
Cell state and ROS generation under various degree of hypoxic stress after 24 h. A Severe morphological changes in EPCs treated with 3% O2, 1% O2, and 0.3% O2 were observed. Conversely, shape and density were well maintained under 10% O2. B ROS generation varied in EPCs treated with different oxygen concentrations for 24 h. Higher fluorescence represents increased ROS production. C Cell viability declined sharply in the 0.3% O2 group after 12 h and was affected under 3% and 1% O2 after 24 h. The cells in the 10% O2 group did not show compromised viability (n = 3 per group)
HIF-1αA Promotes ROS Production and Affects Mitochondrial Membrane Potential of EPCs
We next investigated how the two HIF-1α paralogs affect the stability of EPCs. Prior to that, we assessed the transfection efficiency of HIF-1αA and HIF-1αB. In our experiment, we observed that the fraction of positively transfected cells ranges from 40 to 50%, as determined by counting (Table S2, supplementary files). The Dual Luciferase Reporter Assay demonstrated significantly higher luciferase activity in cells transfected with either HIF-1αA or HIF-1αB comparing to control groups (Figure S1A). Notably, as the luciferase activity of the negative control (Myc vector) exceeded that of the blank control, we further conducted an examination of the mRNA levels of HIF-1αA and HIF-1αB in each experimental group to address the concerns. The results revealed a pronounced increase in the expression levels of HIF-1αA and HIF-1αB in the cells transfected with their respective counterparts (Figure S1B). ROS generation in cells transfected with HIF-1αA increased significantly, while the ROS level in cells transfected with HIF-1αB was not significantly affected, indicating that HIF-1αA and HIF-1αB were functionally different in regulating ROS production (Fig. 2A, B). The promotion of ROS can induce further damage to mitochondria; therefore, the MMPs of the transfected EPCs were investigated. The JC-1 staining showed that the MMP was downregulated only by HIF-1αA (Fig. 2C, D), indicating that the HIF-1αA induces depolarization of the MMP by promoting ROS production, which can lead to decreased stability of mitochondria.
ROS production and mitochondrial membrane potential regulated by HIF-1αA and HIF-1αB. ROS production and mitochondrial membrane potential were measured after 24 h incubation under 10% O2 (n = 3 per group). A ROS production measured by flow cytometry. B ROS production following transfection with either HIF-1αA or HIF-1αB. Significant differences between treatments are marked with different letters (P < 0.05). C Mitochondrial membrane potential was assessed under a Leica TCS SP confocal microscope using JC-1 Green and JC-1 Red. D Red/green florescence ratio assessed mitochondrial membrane potential in cells with HIF-1αA and HIF-1αB. Significant differences between treatments are marked with different letters (P < 0.05)
Diverged Roles of HIF-1αA and HIF-1αB in Apoptosis and Mitophagy Regulation
The ROS production and depolarization of mitochondrial membrane are closely related to mitophagy and cell apoptosis. We therefore next investigated the roles of HIF-1αA and HIF-1αB in regulating mitophagy and apoptosis under hypoxia. The AnnexinV-FITC/PI staining revealed a significant promotion of apoptosis when HIF-1αA was over-expressed, while no remarkable changes were observed with HIF-1αB over-expression (Fig. 3A, B). For mitophagy experiments, the co-localization of mitochondria and lysosomes were investigated 24 h after the EPCs were transfected with HIF-1αA or HIF-1αB, respectively. We observed a noteworthy rise in the co-localization of mitochondria and lysosomes in cells transfected with HIF-1αB (Fig. 3C), indicating that the over-expression of HIF-1αB enhances mitophagy. A slight degree of co-localization was also noted in other groups, potentially attributable to the presence of endogenous HIF-1αB.
Apoptosis and mitophagy of the EPCs induced by HIF-1αA and HIF-1αB. A Apoptosis in EPCs using flow cytometry. B Cell apoptosis following HIF-1αA or HIF-1αB transfection. C Mitophagy following HIF-1αA and HIF-1αB transfection was detected by mitochondria and lysosomes co-localization assays. Mitophagy was induced in EPCs transfected with HIF-1αB. Significant differences between treatments are marked with different letters (P < 0.05)
HIF-1αB Mediates the Expression of Key Genes Related to Mitochondrial Autophagy
Since HIF-1αB was the dominant paralog that regulated mitophagy, we next investigated how over-expression of HIF-1αA and HIF-1αB regulated mitophagy-related genes. We first measured the expression of genes involved in the classic PINK1-Parkin mitophagy pathway, including pink1, parkin, vdac1, optn, ambra1, and sqstm1. Both pink1 and optn were not affected by HIF-1αA or HIF-1αB over-expression. Conversely, the expression of parkin was strongly up-regulated by HIF-1αB. The transcript level of vdac1, a crucial substrate of parkin that regulates mitophagy, was also up-regulated together with sqstm1, an autophagic adaptor that is essential for mitochondrial clearance. Intriguingly, a second autophagic adaptor, ambra1, was downregulated by HIF-1αB over-expression only. These results indicate that the PINK1-Parkin mitophagy pathway was regulated by HIF-1αB under hypoxic stress (Fig. 4).
We also investigated how HIF-1αA and HIF-1αB regulated bnip3 and nix expression, as each of these transcripts are key to cell apoptosis and mitophagy. Our results revealed divergent regulation by HIF-1αA and HIF-1αB. Transcript levels for bnip3, together with beclin1, were significantly up-regulated by HIF-1αA, while nix was strongly up-regulated by HIF-1αB. Transcripts for bcl-2, an apoptosis inhibitor, were also up-regulated, but only by HIF-1αB. Over-expression of HIF-1αA promoted the expression of foxo3, while both HIF-1αA and HIF-1αB had suppressive effects on mTOR regulation (Fig. 5). Both Ulk1 and Ulk2 are two important initiators to regulate the induction of autophagy, and the expression level of these genes was strongly affected by HIF-1αA. HIF-1αB also exerted the effects on the two initiators, and in cells transfected with HIF-1αB, transcript levels of ulk1 were remarkably increased; however, this was not observed for ulk2.
Intriguingly, a crucial hypoxia-sensitive mitophagy-involved gene, fundc1, was up-regulated by HIF-1αA but not HIF-1αB, while lc3, a key protein contributing to major steps of autophagy, was only regulated by HIF-1αB under hypoxic stress (Fig. 5). Taken together, although the mitophagy process was mainly regulated by HIF-1αB, HIF-1αA also contributed to the regulation of genes important for mitophagy under hypoxia.
Divergent Impact of HIF-1αA and HIF-1αB on Mitochondrial Oxidative Phosphorylation-Related Genes and ATP Production
We next investigated the impact of HIF-1αA and HIF-1αB over-expression of mitochondrial oxidative phosphorylation of EPCs 24 h after hypoxic stress. We chose representative genes that encode subunits from each mitochondrial complex, one mitochondrial gene, and one nuclear gene (except for Complex II, which is encoded by four nuclear genes).
For Complex I, the expression levels of mtnd1 and ndufs1 were measured, and the results showed a remarkable increase of ndufs1 in the cells transfected with HIF-1αB (Fig. 6). For Complex II, we measured sdha and sdhb expression levels following transfection. There was a significant increase of sdha in the cells with HIF-1αB over-expression (Fig. 6). For Complex III, the transcription levels of cytochrome b (mtcyb) and uqcrc1 were determined. The expression of mtcyb was up-regulated by HIF-1αA, while that of the uqcrc1 was significantly up-regulated by HIF-1αB (Fig. 6). For Complex IV, the expression levels of coxI and coxIV were measured, and it was revealed that HIF-1αB up-regulated the level of coxIV, but neither HIF-1αA nor HIF-1αB exerted effects on the mitochondrial gene coxI (Fig. 6). These results suggest that HIF-1αA and HIF-1αB have different roles in regulating mitochondrial complex synthesis.
Lastly, the karyogenes from each complex were primarily affected by HIF-1αB, which may further alter ATP production. Therefore, ATP content was measured, and the results revealed that under hypoxia, the production of ATP was driven by HIF-1αB (Fig. 7).
Discussion
For most cell types, hypoxia can lead to the decrease of proliferation. However, cell proliferation can be maintained in certain cell populations when subjected to hypoxic stress, which is one of the common pathological traits in tumor cells, and this adaptive response is also crucial for maintenance of stem cell populations in the face of oxygen insufficiency (Hubbi and Semenza 2015). How mammalian cells sense oxygen levels to coordinate diverse biological outputs during hypoxia has been well demonstrated in recent years. Perhaps the best studied mechanism-mediating responses in low oxygen environments are hypoxia-inducible factors (HIFs). HIFs are stabilized by hypoxia and control the expression of several genes. For fish, however, the exact roles of HIF paralogs under hypoxia are not fully characterized.
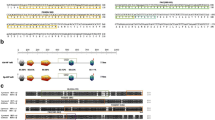
The epithelioma papulosum cyprini (EPC) cell line is derived from the epithelial cells found in the skin of the fathead minnow (Pimephales promelas), a freshwater fish belonging to the Cyprinidae family, same as zebrafish. HIF-1α exhibits a remarkable degree of conservation, as evidenced by prior research indicating that zebrafish HIF-1α can successfully recruit HIF-β and operate effectively even within human cells. (Chen et al. 2020; Guan et al. 2014). Consequently, it is reasonable to anticipate that zebrafish HIF-1α can readily recruit HIF-β and function effectively in EPCs. Zebrafish HIF-1αA consists of 717 amino acids, while HIF-1αB comprises 776 amino acids. These two paralogs exhibit a substantial sequence similarity within their binding domains, encompassing the basic helix-loop-helix (bHLH) and Per-Arnt-Sim (PAS) domains, the oxygen-dependent degradation domain (ODDD), and the C-terminal transactivation domain (C-TAD) (see Figure S2). Notably, a deletion is present within the conserved N-terminal ODDD hydroxylation motif (LXXLAP), and similar deletions have been observed in other cyprinid fish species (Guan et al. 2014). This deletion is believed to potentially reduce the protein's sensitivity to oxygen levels (Rytkonen et al. 2013). The N-TAD, working in conjunction with the C-TAD, plays a pivotal role in influencing the repertoire of target genes activated by HIF-1. The coexistence of both N-TAD and C-TAD empowers HIF-1 to selectively trigger a specific subset of genes in response to hypoxia. Consequently, variations in the amino acid sequences within the transactivation domains (TADs) of these two paralogs may contribute to the divergence in their target gene profiles.
EPCs are not hypoxia-tolerable, and their viability can be severely depleted under acute hypoxia. Our experiments showed that the survival rate of the cells under hypoxia is time- and oxygen-concentration-dependent. Nevertheless, cells can usually improve their ability to adapt to hypoxia to a certain degree and improve their viability through self-regulation, thus promoting survival in a short time (Lee et al. 2020). Given the functions of HIF-1α, the increased viability of cells under 10% O2 may be due to the function of endogenous HIF-1αs.
Under hypoxic stress, mitochondria can become uncoupled because there is not enough O2 to serve as electron receptors during oxidative phosphorylation. As a result, free electrons are increased in the form of ROS. The accumulation of ROS will oxidize and damage macromolecules in cells, which will lead to elevated mitochondrial membrane permeability, the release of cytochrome C from mitochondria, the activation of apoptosis-inducing factors, and subsequently apoptosis (Ruth and Zvulun 2011). In most physiological or pathological cases, a change of MMP is an early response of apoptosis. Studies show that there is decline of MMP is one of the important factors leading to apoptosis (Yang et al. 1997). Our results revealed that the ROS production in EPC cells is oxygen-concentration-dependent (Fig. 1B). Studies have shown that the increase in mitochondrial ROS caused by hypoxia could extend the lifespan of multiple types of cells in vitro (Bell et al. 2007; Liu et al. 2005; Schulz et al. 2007). This response, however, must be finely tuned as overproduction of ROS can eventually result in damage leading to cell death. HIF-1α regulates the generation of ROS by unbalancing the electron flow in the Electron Transport Chain. The specialized functions of HIF-αA and HIF-αB isoforms in cyprinids have been the subject of investigation regarding their distinct roles in the regulation of mitochondrial biogenesis and oxidative phosphorylation. Research findings indicate that, when stabilized by CoCl2, both HIF-αA and HIF-αB isoforms exhibit the capacity to enhance mitochondrial biogenesis. HIF-1αA and HIF-1αB have partially overlapped target gene pools. They have similar functions in promoting mitochondrial biogenesis, but different roles in other aspects like oxidative phosphorylation, the tricarboxylic acid cycle, and the maintenance of mitochondrial stability (Chen et al. 2020). HIF-α is unstable when oxygen is sufficient. The CoCl2 -cultivated experiment stabilizes HIF-α under normoxic conditions, but the cells do not experience hypoxia. As a result, it remains unclear how HIF-1αA and HIF-1αB function in response to hypoxia. Our results demonstrate that HIF-1αA maintained ancestral gene function of regulating ROS under hypoxia, which may be beneficial to both HIF-1αA and HIF-1αB because intracellular ROS can inhibit the activity of prolyl hydroxylase to prevent the degradation of HIF-1α (Li et al. 2014). Our study also illustrated that over-expression of HIF-1αA causes the collapse of MMP, an outcome of excessive accumulation of ROS, and increases apoptosis under hypoxic stress. These results suggested that HIF-1αA can induce apoptosis through the promotion of ROS and mitochondrial depolarization when cells are subjected to hypoxic stress. This ability appears to be lost for HIF-1αB after duplication of the ancestral gene.
Mitochondrial autophagy, or mitophagy, is a crucial strategy for cells to prolong viability under hypoxia or other environmental stressors. In this study, only HIF-1αB significantly promoted the co-localization of mitochondria and lysosomes, suggesting a prominent role in the induction of mitophagy. However, both HIF-1αA and HIF-1αB were functionally involved in different mitophagy pathways regulation at the gene level. The PINK-Parkin pathway, usually mediates mitophagy under cellular starvation. Its activation is not hypoxia dependent, but it can be strongly affected by hypoxia and ROS. In our study, the expression of Parkin was up-regulated by HIF-1αB. Two autophagic adaptors Vdac1 and Sqstm1, each important in the PINK-Parkin-mediated clearance of mitochondria (Geisler et al. 2010), were also increased by HIF-1αB. In addition, Ambra1 has emerged as a scaffold protein regulating autophagy in multiple ways. The down-regulation of Ambra1 by HIF-1αB may be due to its negative regulation to autophagy by promoting proteasomal degradation (Cianfanelli and Cecconi 2015). However, studies also demonstrate its positive role in inducing mitophagy regardless of Parkin and Sqstm1 (Antonioli et al. 2015; Strappazzon et al. 2015). These results suggested that HIF-1αB was predominantly responsible for mitophagy induction through the PINK-Parkin pathway under oxygen deficiency.
Hypoxia can activate mitophagy in different cell types through Bnip3L/Nix, Bcl-2/Bnip3, and Fundc1 (Liu et al. 2012a; Zhou et al. 2011). Our data revealed that over-expression of HIF-1αA induced expression of multiple genes involved in these pathways including Bnip3, Fundc1, Beclin1, and Foxo3 (Fig. 5); these proteins act as intermediates in autophagy and apoptosis. For instance, Beclin1, an ortholog of the Atg6 in yeast, is essential for localization of autophagic proteins to a pre-autophagosomal structure (He and Levine 2010), and this function is closely related to the function of Bcl-2, an anti-apoptotic protein and inhibitor of mitochondrial dysfunction during apoptosis, which is transcriptionally controlled by HIF-1αB. The dissociation of Beclin1 from Bcl-2 is essential for its autophagic activity (Maiuri et al. 2007). Foxo3 is a transcriptional factor responsible for rapid induction of autophagy in cells by transactivating multiple autophagy-related genes. Studies demonstrate that Foxo3 is a substrate and is consumed by basal autophagic degradation, which confers low sensitivity to apoptosis. If autophagy is inhibited, cells will activate pro-apoptotic genes such as Bbc3 via the accumulation of Foxo3 becoming sensitized to apoptotic signals (Fitzwalter and Thorburn 2018; Thorburn et al. 2014). Therefore, HIF-1αA possesses dual functions in regulating mitochondrial autophagy and cell death, and to some extent, it plays more of a role than HIF-1αB in promoting apoptosis. Conversely, HIF-1αB appears to be primarily responsible for the induction of mitophagy, preventing apoptosis via transactivating Bcl-2. Autophagy and apoptosis are two important cellular processes with complex and intersecting protein networks and the transition from “self-eating” to “self-killing” relies on the extent to which stress can be tolerated in terms of hypoxia and ROS overproduction. Therefore, our study shows that the timing of these processes is finely tuned by the divergent function of HIF-1αA and HIF-1αB.
Since the mitochondrial ETC is the largest consumer of intracellular O2 for the generation of ATP, it can be expected that changes in O2 supply will affect the activity mitochondrial complexes in the ETC. The results revealed functional divergence and target shifting of HIF-1αA and HIF-1αB (Fig. 6), suggesting that the protein from one copy targets mitochondrial genes while the other primarily targeted nuclear genes that encodes subunits of the complexes. Complex IV (also known as cytochrome c oxidase, COX) is the last complex within the ETC, providing 4 electrons to O2 to generate water. Since Complex IV possesses a high affinity for O2, the ETC can function at near anoxic levels (Cooper 1990), cells can maintain their ATP levels during hypoxia, which slows accumulation of intermediate ROS. Our study suggests that HIF-1αB inherited a role in promoting ATP generation from its ancestral gene. We propose this is the reason HIF-1αB does not facilitate ROS production.
Conclusions
Subfunctionalization results in an increase in the preservation of duplicated gene copies, which are essential to the generation of evolutionary novelty. The two HIF-1α duplicates were specifically retained in cyprinid fish. Our study unraveled their diverged function in regulating ROS and ATP generation, mitochondrial autophagy, and apoptosis. The distinct collaboration between the two HIF-1α copies has unveiled precise regulation of mitophagy and apoptosis in response to hypoxia exposure.
References
Antonioli M, Albiero F, Fimia GM, Piacentini M (2015) AMBRA1-regulated autophagy in vertebrate development. Int J Dev Biol 59:109–117
Bell EL, Klimova TA, Eisenbart J, Schumacker PT, Chandel NS (2007) Mitochondrial reactive oxygen species trigger hypoxia-inducible factor-dependent extension of the replicative life span during hypoxia. Mol Cell Biol 27:5737–5745
Chen G, Han Z, Feng D, Chen Y, Chen L, Wu H, Huang L, Zhou C, Cai X, Fu C et al (2014) A regulatory signaling loop comprising the PGAM5 phosphatase and CK2 controls receptor-mediated mitophagy. Mol Cell 54:362–377
Chen J, Guan L, Zou M, He S, Li D, Chi W (2020) Specific cyprinid HIF isoforms contribute to cellular mitochondrial regulation. Sci Rep 10:17246
Cianfanelli V, Cecconi F (2015) AMBRA1: when autophagy meets cell proliferation. Autophagy 11:1705–1707
Cooper CE (1990) The steady-state kinetics of cytochrome c oxidation by cytochrome oxidase. Biochim Biophys Acta 1017:187–203
Daido S, Kanzawa T, Yamamoto A, Takeuchi H, Kondo Y, Kondo S (2004) Pivotal role of the cell death factor BNIP3 in ceramide-induced autophagic cell death in malignant glioma cells. Cancer Res 64:4286–4293
Fitzwalter BE, Thorburn A (2018) FOXO3 links autophagy to apoptosis. Autophagy 14:1467–1468
Geisler S, Holmstrom KM, Skujat D, Fiesel FC, Rothfuss OC, Kahle PJ, Springer W (2010) PINK1/Parkin-mediated mitophagy is dependent on VDAC1 and p62/SQSTM1. Nat Cell Biol 12:119–131
Guan L, Chi W, Xiao W, Chen L, He S (2014) Analysis of hypoxia-inducible factor alpha polyploidization reveals adaptation to Tibetan Plateau in the evolution of schizothoracine fish. BMC Evol Biol 14:192
Hamacher-Brady A, Brady NR, Logue SE, Sayen MR, Jinno M, Kirshenbaum LA, Gottlieb RA, Gustafsson AB (2007) Response to myocardial ischemia/reperfusion injury involves Bnip3 and autophagy. Cell Death Differ 14:146–157
Han JE, Lim PW, Na CM, Choi YS, Lee JY, Kim Y, Park HW, Moon HE, Heo MS, Park HR et al (2017) Inhibition of HIF1alpha and PDK induces cell death of glioblastoma multiforme. Exp Neurobiol 26:295–306
He C, Levine B (2010) The beclin 1 interactome. Curr Opin Cell Biol 22:140–149
Hubbi ME, Semenza GL (2015) Regulation of cell proliferation by hypoxia-inducible factors. Am J Physiol Cell Physiol 309:C775-782
Huifang W, Lei L, Quan C (2015) Selective removal of mitochondria via mitophagy: distinct pathways for different mitochondrial stresses. Biochim Biophys Acta 1853:2784
Jiang H, Guo R, Powell-Coffman JA (2001) The Caenorhabditis elegans hif-1 gene encodes a bHLH-PAS protein that is required for adaptation to hypoxia. Proc Natl Acad Sci U S A 98:7916–7921
Kanzawa T, Zhang L, Xiao L, Germano IM, Kondo Y, Kondo S (2005) Arsenic trioxide induces autophagic cell death in malignant glioma cells by upregulation of mitochondrial cell death protein BNIP3. Oncogene 24:980–991
Kung-Chun Chiu D, Pui-Wah Tse A, Law CT, Ming-Jing Xu I, Lee D, Chen M, Kit-Ho Lai R, Wai-Hin Yuen V, Wing-Sum Cheu J, Wai-Hung Ho D et al (2019) Hypoxia regulates the mitochondrial activity of hepatocellular carcinoma cells through HIF/HEY1/PINK1 pathway. Cell Death Dis 10:934
Lee P, Chandel NS, Simon MC (2020) Cellular adaptation to hypoxia through hypoxia inducible factors and beyond. Nat Rev Mol Cell Biol 21:268–283
Lemasters JJ (2005) Selective mitochondrial autophagy, or mitophagy, as a targeted defense against oxidative stress, mitochondrial dysfunction, and aging. Rejuvenation Res 8:3–5
Li YN, Xi MM, Guo Y, Hai CX, Yang WL, Qin XJ (2014) NADPH oxidase-mitochondria axis-derived ROS mediate arsenite-induced HIF-1alpha stabilization by inhibiting prolyl hydroxylases activity. Toxicol Lett 224:165–174
Liu X, Jiang N, Hughes B, Bigras E, Shoubridge E, Hekimi S (2005) Evolutionary conservation of the clk-1-dependent mechanism of longevity: loss of mclk1 increases cellular fitness and lifespan in mice. Genes Dev 19:2424–2434
Liu L, Feng D, Chen G, Chen M, Zheng Q, Song P, Ma Q, Zhu C, Wang R, Qi W et al (2012a) Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat Cell Biol 14:177–185
Liu L, Feng D, Chen G, Chen M, Zheng Q, Song P, Ma Q, Zhu C, Wang R, Qi W et al (2012b) Mitochondrial outer-membrane protein FUNDC1 mediates hypoxia-induced mitophagy in mammalian cells. Nat Cell Biol 14:177
Liu M, Fan Y, Li D, Han B, Meng Y, Chen F, Liu T, Song Z, Han Y, Huang L et al (2021) Ferroptosis inducer erastin sensitizes NSCLC cells to celastrol through activation of the ROS-mitochondrial fission-mitophagy axis. Mol Oncol 15:2084–2105
Loenarz C, Coleman ML, Boleininger A, Schierwater B, Holland PW, Ratcliffe PJ, Schofield CJ (2011) The hypoxia-inducible transcription factor pathway regulates oxygen sensing in the simplest animal, Trichoplax adhaerens. EMBO Rep 12:63–70
Maiuri MC, Zalckvar E, Kimchi A, Kroemer G (2007) Self-eating and self-killing: crosstalk between autophagy and apoptosis. Nat Rev Mol Cell Biol 8:741–752
Majmundar AJ, Wong WJ, Simon MC (2010) Hypoxia-inducible factors and the response to hypoxic stress. Mol Cell 40:294
Nathan C, Cunningham-Bussel A (2013) Beyond oxidative stress: an immunologist’s guide to reactive oxygen species. Nat Rev Immunol 13:349–361
Ruth S-S, Zvulun E (2011) Regulation of autophagy by ROS: physiology and pathology. Trends Biochem Sci 36:30
Rytkonen KT, Williams TA, Renshaw GM, Primmer CR, Nikinmaa M (2011) Molecular evolution of the metazoan PHD-HIF oxygen-sensing system. Mol Biol Evol 28:1913–1926
Rytkonen KT, Akbarzadeh A, Miandare HK, Kamei H, Duan C, Leder EH, Williams TA, Nikinmaa M (2013) Subfunctionalization of cyprinid hypoxia-inducible factors for roles in development and oxygen sensing. Evolution 67:873–882
Sandoval H, Thiagarajan P, Dasgupta SK, Schumacher A, Prchal JT, Chen M, Wang J (2008) Essential role for Nix in autophagic maturation of erythroid cells. Nature 454:232–235
Sangbin L, Hao L, Luciana MDS, Ritu A, Zixing L et al (2016) Immunoregulatory protein B7–H3 reprograms glucose metabolism in cancer cells by ROS-mediated stabilization of HIF1α. Cancer Res 76:2231
Schofield CJ, Ratcliffe PJ (2004) Oxygen sensing by HIF hydroxylases. Nat Rev Mol Cell Biol 5:343–354
Schulz TJ, Zarse K, Voigt A, Urban N, Birringer M, Ristow M (2007) Glucose restriction extends Caenorhabditis elegans life span by inducing mitochondrial respiration and increasing oxidative stress. Cell Metab 6:280–293
Semenza GL (2014) Oxygen sensing, hypoxia-inducible factors, and disease pathophysiology. Annu Rev Pathol 9:47–71
Sowter HM, Ratcliffe PJ, Watson P, Greenberg AH, Harris AL (2001) HIF-1-dependent regulation of hypoxic induction of the cell death factors BNIP3 and NIX in human tumors. Cancer Res 61:6669–6673
Strappazzon F, Nazio F, Corrado M, Cianfanelli V, Romagnoli A, Fimia GM, Campello S, Nardacci R, Piacentini M, Campanella M et al (2015) AMBRA1 is able to induce mitophagy via LC3 binding, regardless of PARKIN and p62/SQSTM1. Cell Death Differ 22:517
Sylvie C, Silke O, Bettina W, Sven D, Michael T, Peter R, Jan D (2017) Phospho-ubiquitin-PARK2 complex as a marker for mitophagy defects. Autophagy 13:201
Tanaka A, Cleland MM, Xu S, Narendra DP, Suen DF, Karbowski M, Youle RJ (2010) Proteasome and p97 mediate mitophagy and degradation of mitofusins induced by Parkin. J Cell Biol 191:1367–1380
Thorburn J, Andrysik Z, Staskiewicz L, Gump J, Maycotte P, Oberst A, Green DR, Espinosa JM, Thorburn A (2014) Autophagy controls the kinetics and extent of mitochondrial apoptosis by regulating PUMA levels. Cell Rep 7:45–52
Vincow ES, Merrihew G, Thomas RE, Shulman NJ, Beyer RP, MacCoss MJ, Pallanck LJ (2013) The PINK1-Parkin pathway promotes both mitophagy and selective respiratory chain turnover in vivo. Proc Natl Acad Sci USA 110:6400–6405
Webb JD, Coleman ML, Pugh CW (2009) Hypoxia, hypoxia-inducible factors (HIF), HIF hydroxylases and oxygen sensing. Cell Mol Life Sci 66:3539–3554
Wu X, Gong L, Xie L, Gu W, Wang X, Liu Z, Li S (2021) NLRP3 deficiency protects against intermittent hypoxia-induced neuroinflammation and mitochondrial ros by promoting the pink1-parkin pathway of mitophagy in a murine model of sleep apnea. Front Immunol 12:628168
Xu J, Wang L, Zhang L, Zheng F, Wang F, Leng J, Wang K, Heroux P, Shen HM, Wu Y et al (2021) Mono-2-ethylhexyl phthalate drives progression of PINK1-parkin-mediated mitophagy via increasing mitochondrial ROS to exacerbate cytotoxicity. Redox Biol 38:101776
Yang J, Liu X, Bhalla K, Kim CN, Ibrado AM, Cai J, Peng TI, Jones DP, Wang X (1997) Prevention of apoptosis by Bcl-2: release of cytochrome c from mitochondria blocked. Science 275:1129–1132
Yoshii SR, Kishi C, Ishihara N, Mizushima N (2011) Parkin mediates proteasome-dependent protein degradation and rupture of the outer mitochondrial membrane. J Biol Chem 286:19630–19640
Zeb A, Choubey V, Gupta R, Kuum M, Safiulina D, Vaarmann A, Gogichaishvili N, Liiv M, Ilves I, Tamm K et al (2021) A novel role of KEAP1/PGAM5 complex: ROS sensor for inducing mitophagy. Redox Biol 48:102186
Zepeda Andrea B, Pessoa Adalberto, Castillo Rodrigo L, Figueroa Carolina A, Pulgar Victor M, Farías Jorge G (2013) Cellular and molecular mechanisms in the hypoxic tissue: role of HIF-1 and ROS. Cell Biochem Funct 31:451
Zhang J, Ney PA (2009) Role of BNIP3 and NIX in cell death, autophagy, and mitophagy. Cell Death Differ 16:939–946
Zhou R, Yazdi AS, Menu P, Tschopp J (2011) A role for mitochondria in NLRP3 inflammasome activation. Nature 469:221–225
Acknowledgements
We thank Xiaomin Feng and Wenwen Zhang for their helpful suggestions and assistance to this study.
Funding
This work was supported by the Professorial and Doctoral Scientific Research Foundation of Huizhou University (2020JB065), Team of Guangdong Provincial Department of Education: Marine shellfish ecological breeding and disease prevention and control innovation (2021kcxtd057), and Curriculum Ideology and Politics Foundation of Huizhou University (X-KCSZ2021004).
Author information
Authors and Affiliations
Contributions
WC cointributed to conceptualization, methodology, data curation, and writing—original draft preparation. JF contributed to software and formal analysis. WC and CJM contributed to investigation. . CJM contributed to writing—review & editing.JW and LZ contributed to funding acquisition .
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Handling editor: John Bracht.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Chi, W., Fu, J., Martyniuk, C.J. et al. Post-Subfunctionalization Functions of HIF-1αA and HIF-1αB in Cyprinid Fish: Fine-Tuning Mitophagy and Apoptosis Regulation Under Hypoxic Stress. J Mol Evol 91, 780–792 (2023). https://doi.org/10.1007/s00239-023-10138-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-023-10138-9