Abstract
Bilateria exhibit whole-body handedness in internal structure. This left–right polarity is evolutionarily conserved with virtually no reversed extant lineage, except in molluscan Gastropoda. Phylogenetically independent snail groups contain both clockwise-coiled (dextral) and counterclockwise-coiled (sinistral) taxa that are reversed from each other in bilateral handedness as well as in coiling direction. Within freshwater Hygrophila, Lymnaea with derived dextrality have diaphanous related formin (diaph) gene duplicates, while basal sinistral groups possess one diaph gene. In terrestrial Stylommatophora, dextral Bradybaena also have diaph duplicates. Defective maternal expression of one of those duplicates gives rise to sinistral hatchlings in Lymnaea and handedness-mixed broods in Bradybaena, through polarity change in spiral cleavage of embryos. These findings led to the hypothesis that diaph duplication was crucial for the evolution of dextrality by reversal. The present study discovered that diaph duplication independently occurred four times and its duplicate became lost twice in gastropods. The dextrality of Bradybaena represents the ancestral handedness conserved across gastropods, unlike the derived dextrality of Lymnaea. Sinistral lineages recurrently evolved by reversal regardless of whether diaph had been duplicated. Amongst the seven formin gene subfamilies, diaph has most thoroughly been conserved across eukaryotes of the 14 metazoan phyla and choanoflagellate. Severe embryonic mortalities resulting from insufficient expression of the duplicate in both of Bradybaena and Lymnaea also support that diaph duplicates bare general roles for cytoskeletal dynamics other than controlling spiralian handedness. Our study rules out the possibility that diaph duplication or loss played a primary role for reversal evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Bilateria possess a characteristic body plan with the left–right axis. Their body architecture may externally be left–right symmetric by and large. On the other hand, their internal structure frequently exhibits bilateral asymmetry such as seen in the organ arrangement. In this whole-body asymmetry, virtually no left–right reversed taxa are known (Wood 2005), indicating the presence of evolutionary constraint against reversal across bilaterian groups except for one: molluscan gastropods. Diverse snail groups with internal fertilization uniquely present both clockwise-coiled (dextral) and counterclockwise-coiled (sinistral) taxa in many phylogenetically independent clades (Vermeij 1975; Okumura et al. 2008; Ponder et al. 2020a).
The aperture of the dextral shell is to the right when the apex is up and to the left in sinistral snails (Fig. 1a). In most gastropod groups, their coiling direction reflects the polarity of bilateral asymmetry in the body plan of Bilateria (Ponder et al. 2020a). The location of their genital orifice, situated at the body side instead of the midplane unlike typical bilaterians, is thus also reversed between dextral and sinistral gastropods (Fig. 1b, c). Therefore, physical genital mismatch interferes with copulation between the reversed mutant and the wild-type, resulting in positive frequency-dependent selection against reversal in gastropods with internal fertilization (Johnson 1982; Gittenberger 1988; Asami et al. 1998). Because of this mechanism, reversed individuals are rarely found in wild populations. Their lineages/groups mostly remain without population fixation for the reversal, i.e., reversal evolution. Reversed taxa are unknown in marine gastropods with external fertilization, in which stringent survival selections against reversal can be in effect under large effective population size (Vermeij 1975; Utsuno et al. 2011).
Left–right handedness of gastropod snails in coiling direction (secondary asymmetry) and bilateral (primary) asymmetry. a Dextral (left in figure) and sinistral shells of a chirally dimorphic Cuban tree snail Liguus vittatus. b Dextral (left) and sinistral siblings produced by a racemic mutant parent in a dextral Asian terrestrial snail Bradybaena similaris. c Dextral (left) and sinistral siblings produced by a racemic mutant parent in a sinistral Asian clausiliid terrestrial snail Euphaedusa tau. Arrows indicate genital orifice locations reversed between the dextral and sinistral
Typical difficulties of gastropod breeding in captivity with rarely available mutants and a characteristic inheritance mode, maternal inheritance (Toyama 1913; Sturtevant 1923), historically hampered studies to determine the genetic basis of gastropod handedness (Boycott and Diver 1923; Boycott et al. 1930). Most results of crossing experiment supported inheritance of whole-body handedness at one nuclear locus in four phylogenetically independent groups of hermaphroditic pulmonates (Murray and Clarke 1976; Asami et al. 2008; Utsuno and Asami 2010; Utsuno et al. 2010). It deserves attention, however, that the wild-type handedness was contrasted with the mutant phenotype in most cases. At least in freshwater pulmonate (hygrophilid) snails, early spiral cleavages proceed in largely opposite direction between dextral and sinistral taxa (Crampton 1894; Camey and Verdonk 1970; Raven 1972; Meshcheryakov and Beloussov 1975). This reversal of spiral cleavage in early development between the wild types with opposite handedness, however, remains unverified in other gastropod clades, for similar technical reasons that hinder breeding.
A racemic mutant line of terrestrial dextral Bradybaena similaris produces handedness-mixed broods from which both dextral and sinistral progeny consistently hatch (Fig. 1b, Utsuno and Asami 2010). Another racemic mutant strain has been established in sinistral clausiliid Euphaedusa tau (Fig. 1c, Utsuno et al. 2010). Similarly mixed broods were observed in crossing experiments with dextral mutants of sinistral Alinda biplicata (Degner 1952) and in chirally dimorphic Partula snails (Murray and Clarke 1976). In freshwater gastropods, handedness-mixed broods were also found to be produced by genetically fixed strains of dextral Lymnaea peregra (Freeman and Lundelius 1982) as well as in wild populations of an operculate dextral Campeloma (van Cleave 1936). These findings illuminate the persistence of an evolutionarily conserved genetic system for development of the wild-type handedness, for which gastropod populations are ordinarily fixed to be either dextral or sinistral (Hierck et al. 2005; Okumura et al. 2008; Utsuno et al. 2011).
Bilaterally reversed gastropods evolved through population fixation for the reversal in this conserved developmental system, in which early embryos undergo spiral cleavages (Lesoway and Henry 2019; Martín-Durán and Marlétaz 2020). This evolutionary process has been accelerated through the course of lineage radiation in the land pulmonate clade Stylommatophora. The gene responsible for the reversal has acted as a speciation gene that establishes sexual isolation by reversal through population fixation under given conditions of terrestrial environment (Gittenberger 1988; Orr 1991; van Batenburg and Gittenberger 1996; Ueshima and Asami 2003; Hoso et al. 2010; Richards et al. 2017). The frequency of chiral speciation was estimated as once every 4.6–3.3 MY in the door snail family Clausiliidae (Gittenberger et al. 2012).
In the case of a handedness mutant strain of L. stagnalis, established with reversed (sinistral) individuals found in a natural pond (Hierck et al. 2005; Asami et al. 2008), the sinistral phenotype of hatching individuals was the outcome of frameshift mutation at one of the diaphanous-related formin (diaph) gene duplicates (Davison et al. 2016; Kuroda et al. 2016). Insufficient expression of the diaph duplicate results in loss of maternal handedness determination in the racemic strain of terrestrial B. similaris, which thus produces handedness-mixed broods (Noda et al. 2019). These cases indicate that the diaph duplicate bares an indispensable role for development of the wild-type dextrality at least in pulmonate clades.
Within a freshwater clade, composed of Lymnaeidae and two sinistral families Physidae and Planorbidae, duplication of the diaph gene probably occurred in the ancestor of Lymnaeidae, because other sinistral groups have a single diaph gene in genome (Davison et al. 2016). Dextral B. similaris of the terrestrial stylommatophoran clade also possesses diaph duplicates (Noda et al. 2019). Whether recurrent reversal evolution occurred depending on diaph duplication has, therefore, arisen as a critical question to be answered. However, few studies examined how the molecular evolution of diaph proceeded during the course of gastropod cladogenesis. It thus remains unknown whether the wild-type sinistrality of those reversed lineages could have originated in the evolutionary change of diaph. Here, we show that the reversal of bilateral handedness recurrently evolved without depending on the dynamic molecular evolution of diaph that occurred in Gastropoda.
Methods
Molluscan Diaph Sequences
Diaph sequences of molluscan species were assembled using available RNA sequence reads and whole-genome sequences deposited in NCBI public database. Reads similar to those sequences of diaph were extracted by tblastn and manually assembled. Sequences were extended with reads found by blastn with the edge sequence of the first assembles as queries to reconstruct whole-protein coding regions.
Molecular Phylogenetic Analysis
We carried out molecular phylogenetic analysis of diaph using Mrbayes version 3.2.6 (Ronquist et al. 2012). Amino acid sequences were first aligned by ClustalW version 2.1.0 and modified properly considering with exon–intron structure. JTT model was adopted as an evolutionary model according to estimation by MEGA 7.0.9 (Kumar et al. 2016). We reconstructed the basian and maximum likelihood trees of diaph based on its whole-protein sequences as well as on FH2 domain sequences by MEGA 7.0.9. To examine the legitimacy of evolutionary loss of one of diaph duplicates, we conducted tblastn searches in whole-genome sequence data of Littorina saxatilis (SRX4809132) and Potamopyrgus antipodarum (SRX2550281). Sequences of DIAPH proteins from Rapana venosa (diaph-Hyp1 and diaph-Hyp2), L. saxatilis and P. antipodarum were used for the query of tblsatn.
Protein Domain and Exon–Intron Structure Analysis
Domain composition of DIAPH proteins was examined by using Pfam version 32.0 (Finn et al., 2014) as previously described with slight modifications (Noda and Satoh 2008). Exon–intron structure was compared using reconstructed diaph transcript sequences and reads of whole-genome sequencing by NGS. All introns’ 5' and 3' splice sites, i.e. intron boundaries, in protein coding regions were identified.
Formin Family Genes Survey
To examine the extent of evolutionary gene conservation across the animal phyla, gene models of 24 animals and a choanoflagellate as an out group in NCBI database were surveyed by tblastn with Formin family proteins of humans as queries. Annotation of formin genes was carried out as previously described with blast bidirectional hit search, Pfam domain analysis and molecular phylogenetic analysis (Noda 2011).
Results
Molecular Phylogeny of Diaph in Gastropoda
Taking advantage of the high quantity and quality RNA and DNA sequence data of molluscs especially gastropods deposited in NCBI, we assembled diaph-related sequences from 67 species in 51 families of the phylum Mollusca (Ponder et al. 2020b; Supplementary Table S1). Our results show that all of these molluscs including three bivalves and one cephalopod have at least one diaph copy, which is probably highly conserved. We found that two copies of diaph are present in the genome of 21 of 63 gastropods included, suggesting the occurrence of duplication during gastropod evolutionary radiation. To substantiate our re-examination of molluscan diaph genes and thus to determine evolutionary points where the duplication occurred in the animal tree of gastropods, we established the molecular phylogeny of diaph using the entire amino-acid sequences (Fig. 2; Supplementary Fig. S1 for ML tree) and FH2 domain sequences (Fig. S2 for Bayesian inference tree; Fig. S3 for ML tree). Almost the same tree topologies were obtained by the four methods. We plotted diaph duplication and loss events detected by this analysis on the phylogeny of gastropod groups (Fig. 3).
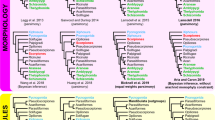
Molecular phylogeny of diaphanous-related formin (diaph) genes in gastropods, reconstructed by Bayesian method. Red circles show the four independent occurrences of duplication. Blue circles indicate duplicates that are expressed in sinistral taxa. Posterior probabilities higher than 0.8 and bootstrap values of maximum likelihood tree which supports the topologies of bayesian tree are shown on nodes, for which diaph sequences of three bivalve species were used as the outgroup
Diaph Duplication and Loss
Our diaph phylogeny showed that duplications have independently occurred in at least four phylogenetic lineages of the class Gastropoda. Within the Panpulmonata clade, the gene has been duplicated at least twice independently, in the ancestors of the superorder Stylommatophora (diaph-a and diaph-b) (Fig. 2, clade E) and the hygrophilid family Lymnaeidae (dia1 and dia2) (Fig. 2, a class of within clade D). In addition, two duplications occurred in the ancestors of Hypsogastropoda (diaph-Hyp1 and diaph-Hyp2) (Fig. 2, clade B) and Ampullarioidea (diaph-Amp1 and diaph-Amp2) (Fig. 2, clade C).
One of the duplicates (diaph-Hyp2) has been lost twice within the Hypsogastropoda clade; in the ancestor of Truncatelloidea (Potamopyrgus antipodarum and Oncomelania hupensis) (Fig. 2, clade H) and in that of Littorinoidea and Naticoidea (Littorina saxatilis, Echinolittorina malaccana and Neverita didyma) (Fig. 2, clade F). In theory there is a risk to interpret the technical overlooking of one of those duplicates as the loss in the genome. We thus conducted tblastn search in genome sequencing data from Littorina saxatilis and Potamopyrgus antipodarum. A plenty of reads were counted for diaph-Hyp1. However, no diaph-Hyp2 like reads were found (Table 1). Our results therefore support the loss of diaph-Hyp2 in the ancestors of these species.
Reversal Does not Correlate with Diaph Duplication or Loss
We found that independent events of diaph duplication or loss in Gastropoda accompanied no case of reversal, except one that may have occurred after duplication in the ancestor of freshwater Lymnaeidae. Many lineages of Stylommatophora have retained dextrality after diaph became duplicated in their dextral ancestor. Similarly, in Caenogastropoda, diaph duplication has preceded in the dextral ancestor of Ampullarioidea as well as in that of Hypsogastropoda, which had been followed by no case of sinistrality evolution.
One of diaph duplicated paralogs has been lost at least twice in the clade of hypsogastropods, which also did not accompany the evolution of reversal in the following lineage. Moreover, the two events of reversal in pulmonates independently happened without accompanying diaph duplication or loss. The sinistral clade has been derived from the dextral ancestor of freshwater Hygrophila with no duplication. In parallel, in terrestrial Stylommatophora, the sinistrality of Clausiliidae, represented by Euphaedusa tau and Alinda biplicata in the present study, has evolved with no loss of diaph duplicates. These lines of evidence indicate that left–right reversals in Gastropoda have evolved without addition or reduction of the diaph paralog.
Domain Composition and Exon–Intron Structure
Modification in DIAPH protein domain composition might have been involved in the evolution of left–right reversal. We confirmed that DIAPH protein did not, however, differ in domain composition among three molluscan classes: Gastropoda, Bivalvia and Cephalopoda (Fig. 4). From the N- to C-terminal the protein contains domains of formin homology 2, diaphanous FH3, diaphanous GTPase-binding, and DRF autoregulatory, regardless of the taxon examined. This has similarly been reported for other animals (Schönichen and Geyer 2010).
DIAPH protein domain structure in three molluscan classes. Magenta, formin homology 2 domains; cyan, diaphanous FH3 domain; green, diaphanous GTPase-binding domain; and red, DRF autoregulatory domain. Vertical broken lines show exon–intron boundaries of DIAPH in gastropods. Red arrowheads indicate difference in exon–intron boundaries between bivalves/cephalopods and gastropods. The top ruler indicates amino acid positions in the primary sequence
On the other hand, molluscan diaph genes vary in exon–intron structure among the classes. There are 24 introns within the protein coding region of gastropod diaph. The bivalve diaph contains 23 introns without the one corresponding to the 8th gastropod intron (Fig. 4). Cephalopod genomes lack the intron corresponding to the 21st in gastropods and possess the 25th intron with no correspondence to the gastropod or bivalve introns. However, these changes in intron–exon structure of diaph genes are unlikely to affect the domain composition of DIAPH protein (Fig. 4).
Conservation of Formin Family Genes in Metazoans
Diaph is a member of one of seven subfamilies that compose the formin gene family (Schönichen and Geyer 2010). To assess the relative extent of evolutionary conservation, we examined whether a gene of each subfamily is present in the genome of 24 animal species that represent clades across 14 metazoan phyla, including a sponge (Amphimedon queenslandica) and a placozoan (Trichoplax adhaerens). The genome of a choanoflagellate (Monosiga brevicollis) was used as an outgroup.
We found that the seven gene subfamilies strikingly differ in the extent of evolutionary conservation across these clades of eukaryotes (Fig. 5). The diaph subfamily is consistently retained by all the organisms examined. FHOD is also commonly present in all the animal genomes including those of ecdysozoan and spiralian parasites (Necator americanus, Schistosoma mansoni, and Intoshia linei). On the other hand, we did not find the subfamilies of INF in a platyhelminth (Macrostomum lignano) and of DAAM in the spiralian parasite or the placozoan. FMNL is missing in five animal species of the spiralian, cnidarian and placozoan. Five ecdysozoan and spiralian species also lack FMN. The subfamily GRID2IP conspicuously differs from the diaph subfamily in its absence over nine species, including the ecdysozoan, spiralian, chidarian, placozoan and choanoflagellate. These results show that DIAPH proteins are most thoroughly conserved among the formin subfamilies regardless of whether bilateral handedness develops through spiral cleavage.
Discussion
Our molecular phylogeny of the diaph gene disclosed that it has gone through at least four events of duplication and two events of loss after duplication phylogenetically independently from one another in Gastropoda, in which left–right reversed lineages recurrently evolved. We found no sign of evolutionary correlation between diaph duplication and whole-body reversal.
Gastropoda radiated from the dextral ancestor with a single diaph homolog. Our results show that two independent events of sinistrality evolution by reversal gave rise to sinistral clade including Physidae and Planorbidae in freshwater Hygrophila and the mostly sinistral clausiliid clades in terrestrial Stylommatophora, regardless of whether their dextral ancestors had one or two diaph copies. Terrestrial dextral Bradybaena and freshwater dextral Lymnaea possess two diaph copies that originated in different events of duplication within pulmonates. The dextrality of the former represents the ancestral handedness of Gastropoda conserved since before diaph duplication, unlike the latter exhibits dextrality that was recovered by reversal from sinistrality.
We demonstrated that DIAPH proteins are expressed in animals of the 14 metazoan phyla as well as in a choanoflagellate. The diaph gene only exhibits such a thorough evolutionary conservation among the seven formin gene subfamilies. This finding corroborates that DIAPH proteins bare a variety of indispensable functions for cytoskeletal processes generally across eukaryotes, including Deuterostomia and Ecdysozoa which develop bilateral asymmetry in their distinctive modes with no spiral cleavage (Bogdan et al. 2013). In a fruit fly and round worm of the latter clade, diaph mutants go through defects in cytokinesis to be lethal (Castrillon and Wasserman 1994; Severson et al. 2002). Similarly, in both dextral Lymnaea (Hierck et al. 2005; Asami et al. 2008) and Bradybaena (Utsuno and Asami 2010), insufficient maternal expression at only one diaph despite having been duplicated produces the reversed progeny that barely survives to the veliger stage (Utsuno et al. 2011; Davison et al. 2016; Kuroda et al. 2016; Noda et al. 2019). These cases exemplify that both of diaph duplicates have to be expressed not to be eliminated by internal selection. This indicates that the protein products of both of diaph duplicates hold general roles for development as cellular machinery components (Bogdan et al. 2013) other than determining the wild type handedness of spiral cleavage (Martín-Durán & Marlétaz 2020), which gastropods uniquely reversed (Vermeij 1975; Lesoway and Henry 2019; Ponder et al. 2020a).
Sinistrals develop with dextrals in the same uterus in natural populations of a dextral freshwater operculate snail of Campeloma with ovoviviparity, but undergo survival selection against after birth (van Cleave 1936). Handedness mutant strains of sinistral ovoviviparous clausiliids Alinda biplicata (Degner 1952) and Euphaedusa tau (Utsuno et al. 2010) also produce both dextral and sinistral offspring, in which at least the latter strain has no mutation in amino acid sequence or expression of either one of diaph duplicates (Noda et al. unpublished data). These lines of currently available knowledge support that diaph is neither a primary determinant of spiralian handedness nor has been responsible for the evolution of reversal in ovoviviparous gastropods.
In terrestrial Stylommatophora, all of extant taxa examined possess two diaph copies in the genome, regardless of their handedness, indicating that one duplication independently occurred in their dextral ancestor. Bradybaena similaris is an example of dextral species in the stylommatophoran family Camaenidae, in which decline of maternal expression of a diaph duplicate causes failure in embryonic metamorphosis by and large (Noda et al. 2019). This indicates that the loss-of-function mutation of a diaph duplicate could not have been responsible for sinistrality, for which populations have become fixed repeatedly within the Camaenidae clade (Hoso et al. 2010; Chen et al. 2021). In chirally dimorphic Partula suturalis of the family Partulidae, there were populations that were fixed for dextrality and those for sinistrality (Johnson et al. 1993). Crossing experiments showed that their sinistrality was dominant to dextrality in maternal inheritance (Murray and Clarke 1976). Thus, the sinistral handedness of this species could not have been the outcome of expression loss of a duplicate gene. Moreover, dextral lineages have also repeatedly evolved by recovering the ancestral handedness within derived sinistral clades, for example in Camaenidae (Ueshima and Asami 2003), Clausiliidae (Gittenberger et al. 2012; Fehér et al. 2013; Páll‐Gergely et al. 2019) and Dyakiidae (Jirapatrasilp et al. 2021). These reversals to the sinistral and back to the dextral in the Stylommatophora could not have occurred repeatedly if the expression loss and recovery of a diaph duplicate were responsible for sinistrality and dextrality, respectively.
We therefore conclude that the wild-type sinistrality of gastropods represents a unique example of a recurrently evolved novelty, loss of which recovers the ancestral dextrality, instead of the phenotype resulting from loss-of-function mutation of a diaph duplicate. Our findings direct further empirical studies to unravel the genetic key determinant of the wild-type handedness in dextral and sinistral lineages that evolved through population fixation for the reversal of bilateral handedness in Gastropoda.
Data availability
Data necessary to re-create the figures can be found on our Unit site: https://osit.jp.group.mgu.
References
Asami T, Ohbayashi K, Cowie RH (1998) Evolution of mirror images by sexually asymmetric mating behaviour in hermaphroditic snails. Am Nat 152:225236. https://doi.org/10.1086/286163
Asami T, Gittenberger E, Falkner G (2008) Whole-body enantiomorphy and maternal inheritance of chiral reversal in the pond snail Lymnaea stagnalis. J Hered 99:552–557. https://doi.org/10.1093/jhered/esn032
Bogdan S, Schultz J, Grosshans J (2013) Formin’ cellular structures: physiological roles of Diaphanous (Dia) in actin dynamics. Commun Integr Biol 6:e27634. https://doi.org/10.4161/cib.27634
Boycott AE, Diver C (1923) On the inheritance of sinistrality in Limnaeaperegra. Proc Roy Soc Lond B 95:207–213. https://doi.org/10.1098/rspb.1923.0033
Boycott AE, Diver C, Garstang SL, Turner FM (1930) The inheritance of sinistrality in Limnaeaperegra (Mo1lusca, Pulmonata). Phil Trans r Soc Lond b 219:51–131. https://doi.org/10.1098/rspb.1923.0033
Camey T, Verdonk N (1970) The early development of the snail Biomphalariaglabrata (Say) and the origin of the head organs. Neth J Zool 20:93–121. https://doi.org/10.1163/002829670x00097
Castrillon DH, Wasserman SA (1994) Diaphanous is required for cytokinesis in Drosophila and shares domains of similarity with the products of the limb deformity gene. Development 120:3367–3377. https://doi.org/10.1242/dev.120.12.3367
Chen ZY, Lyu ZT, Wu M (2021) Systematic revision of Stegodera Martens, 1876 (Gastropoda, Stylommatophora, Camaenidae), with description of a new genus. ZooKeys 1059:1. https://doi.org/10.3897/zookeys.1059.68385
Crampton HE Jr (1894) Reversal of cleavage in a sinistral gastropod. Ann N Y Acad Sci 8:167–169. https://doi.org/10.1111/j.1749-6632.1894.tb55419
Davison A, McDowell GS, Holden JM, Johnson HF, Koutsovoulos GD, Liu MM, Hulpiau P, Van Roy F, Wade CM, Banerjee R, Yang F, Chiba S, Davey JW, Jackson DJ, Levin M, Blaxter ML (2016) Formin is associated with left-right asymmetry in the pond snail and the frog. Curr Biol 26:654–660. https://doi.org/10.1016/j.cub.2015.12.071
Degner E (1952) Der erbgang der invergion bei Laciniaria biplicata MTG. Mtteil Hamburg Zool Mus Inst 51:3–61
Fehér Z, Németh L, Nicoară A, Szekeres M (2013) Molecular phylogeny of the land snail genus Alopia (Gastropoda: Clausiliidae) reveals multiple inversions of chirality. Zool J Linn Soc 167:259–272. https://doi.org/10.1111/zoj.12002
Finn RD, Bateman A, Clements J, Coggill P, Eberhardt RY, Eddy SR, Heger A, Hetherington K, Holm L, Mistry J, Sonnhammer EL, Tate J, Punta M (2014) Pfam: the protein families database. Nucleic Acids Res 42:D222–D230. https://doi.org/10.1093/nar/gkt1223
Freeman G, Lundelius JW (1982) The developmental genetics of dextrality and sinistrality in the gastropod Lymnaeaperegra. Roux Arch Dev Biol 191:69–83. https://doi.org/10.1007/BF00848443
Gittenberger T (1988) Sympatric speciation in snails; a largely neglected model. Evolution 42:826–828. https://doi.org/10.1111/j.1558-5646.1988.tb02502.x
Gittenberger E, Hamann TD, Asami T (2012) Chiral speciation in terrestrial pulmonate snails. PLoS ONE 7:e34005. https://doi.org/10.1371/journal.pone.0034005
Hierck BP, Witte B, Poelmann RE, Gittenberger-de Groot AC, Gittenberger E (2005) Chirality in snails is determined by highly conserved asymmetry genes. J Moll Stud 71:192–195. https://doi.org/10.1093/mollus/eyi023
Hoso M, Kameda Y, Wu SP, Asami T, Kato M, Hori M (2010) A speciation gene for left-right reversal in snails results in anti-predator adaptation. Nat Commun 1:133. https://doi.org/10.1038/ncomms1133
Jirapatrasilp P, Tongkerd P, Jeratthitikul E, Liew TS, Pholyotha A, Sutcharit C, Panha S (2021) Molecular phylogeny of the limacoid snail family Dyakiidae in Southeast Asia, with the description of a new genus and species. Zool J Linn Soc 193:250–280. https://doi.org/10.1093/zoolinnean/zlaa129
Johnson MS (1982) Polymorphism for direction of coil in Partula suturalis: Behavioural isolation and positive frequency dependent selection. Heredity 49:145–151. https://doi.org/10.1038/hdy.1982.80
Johnson MS, Murray J, Clarke B (1993) The ecological genetics and adaptive radiation of Partula on Moorea. Oxford Surv Evol Biol 9:167–238
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kuroda R, Fujikura K, Abe M, Hosoiri Y, Asakawa S, Shimizu M, Umeda S, Ichikawa F, Takahashi H (2016) Diaphanous gene mutation affects spiral cleavage and chirality in snails. Sci Rep 6:34809. https://doi.org/10.1038/srep34809
Lesoway MP, Henry JQ (2019) Twisted Shells, spiral cells, and asymmetries: evo-devo lessons learned from gastropods. In: Nuno de la Rosa L, Müller G (eds) Evolutionary Developmental Biology. Springer, pp 1–18
Martín-Durán JM, Marlétaz F (2020) Unravelling spiral cleavage. Development 147:181081. https://doi.org/10.1242/dev.181081
Meshcheryakov VN, Beloussov LV (1975) Asymmetrical rotations of blastomeres in early cleavage of gastropoda. Wilehm Roux Arch Dev Biol 177:193–203. https://doi.org/10.1007/bf00848080
Murray J, Clarke B (1976) Supergenes in polymorphic land snails. II. Partula Suturalis. Heredity 37:271–282. https://doi.org/10.1038/hdy.1976.87
Noda T (2011) The maternal genes Ci-p53/p73-a and Ci-p53/p73-b regulate zygotic ZicL expression and notochord differentiation in Ciona intestinalis embryos. Dev Biol 360:216–229. https://doi.org/10.1016/j.ydbio.2011.08.029
Noda T, Satoh N (2008) A comprehensive survey of cadherin superfamily gene expression patterns in Ciona intestinalis. Gene Expr Patterns 8:349–356. https://doi.org/10.1016/j.gep.2008.01.004
Noda T, Satoh N, Asami T (2019) Heterochirality results from reduction of maternal diaph expression in a terrestrial pulmonate snail. Zool Lett 5:2. https://doi.org/10.1186/s40851-018-0120-0
Okumura T, Utsuno H, Kuroda J, Gittenberger E, Asami T, Matsuno K (2008) The development and evolution of left-right asymmetry in invertebrates: lessons from Drosophila and snails. Dev Dyn 237:3497–3515. https://doi.org/10.1002/dvdy.21788
Orr HA (1991) Is single-gene speciation possible? Evolution 45:764–769. https://doi.org/10.2307/2409927
Páll-Gergely B, Szekeres M, Fehér Z, Asami T, Harl J (2019) Evolution of a dextral lineage by left–right reversal in Cristataria (Gastropoda, Pulmonata, Clausiliidae). J Zool Sys Evol Res 57:520–526. https://doi.org/10.1111/jzs.12277
Ponder WF, Lindberg DR, Ponder JM. 2020a Biology and Evolution of the Mollusca. Volume 1. CRC Press.
Ponder WF, Lindberg DR, Ponder JM. 2020b Biology and Evolution of the Mollusca. Volume 2. CRC Press.
Raven CP (1972) Determination of the direction of spiral coiling in Lymnaeaperegra. Acta Morph Neerl Scand 10:165–178
Richards PM, Morii Y, Kimura K, Hirano T, Chiba S, Davison A (2017) Single-gene speciation: mating and gene flow between mirorr-image snails. Evol Lett 1:282–291. https://doi.org/10.1002/evl3.31
Ronquist F, Teslenko M, van der Mark P, Ayres DL, Darling A, Höhna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) MrBayes 3.2: efficient Bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61:539–542. https://doi.org/10.1093/sysbio/sys029
Schönichen A, Geyer M (2010) Fifteen formins for an actin filament: a molecular view on the regulation of human formins. Biochim Biophys Acta 1803:152–163. https://doi.org/10.1016/j.bbamcr.2010.01.014
Severson AF, Baillie DL, Bowerman B (2002) A Formin homology protein and a profilin are required for cytokinesis and Arp2/3-independent assembly of cortical microfilaments in C. elegans. Curr Biol 12:2066–2075. https://doi.org/10.1016/S0960-9822(02)01355-6
Sturtevant AH (1923) Inheritance of direction of coiling in Limnaea. Science 58:269–270. https://doi.org/10.1126/science.58.1501.26
Toyama K (1913) Maternal inheritance and mendelism. J Genet 2:351–405. https://doi.org/10.1007/BF01837304
Ueshima R, Asami T (2003) Single-gene speciation by left-right reversal. Nature 425:679. https://doi.org/10.1038/425679a
Utsuno H, Asami T (2010) Maternal Inheritance of racemism in the terrestrial snail Bradybaenasimilaris. J Hered 101:11–19. https://doi.org/10.1093/jhered/esp058
Utsuno H, Kasem S, Fukuda H, Asami T (2010) Genetic basis of racemism and ease of interchiral mating in a clausiliid species of snails. Moll Res 30:37–47
Utsuno H, Asami T, Van Dooren TJ, Gittenberger E (2011) Internal selection against the evolution of left–right reversal. Evolution 65:2399–2411. https://doi.org/10.1111/j.1558-5646.2011.01293.x
van Cleave HJ (1936) Reversal of symmetry in Campeloma rufum, a fresh-water snail. Am Nat 70:567–573. https://doi.org/10.1086/280697
van Batenburg FHD, Gittenberger E (1996) Ease of fixation of a change in coiling: computer experiments on chirality in snails. Heredity 76:278–286. https://doi.org/10.1038/hdy.1996.41
Vermeij GJ (1975) Evolution and distribution of left-handed and planispiral coiling in snails. Nature 254:419–420. https://doi.org/10.1038/254419a0
Wood WB (2005) The left–right polarity puzzle: determining embryonic handedness. PLoS Biol 3:e292. https://doi.org/10.1371/journal.pbio.0030292
Acknowledgements
We are grateful to Adriana Gittenberger-de Groot for inspiring discussion, Rei Ueshima, Masatoyo Okamoto and Hiroshi Fukuda for help to establish racemic strains, David Maceira for help in the field, Victor Misaka for help in literature survey and Kanako Hisata for help in manuscript preparation.
Funding
The work was supported in part by JSPS KAKENHI grants to T.A. (17H01673, 17H01448, 19K22450), and OIST Internal Research Fund to N.S. (Marine Genomics Unit).
Author information
Authors and Affiliations
Contributions
TN, TA conceived the experiment, TN carried out bioinformatic analysis, NS, TN, TA, EG carefully finalized the manuscript. All authors gave final approval for publication and agreed to be held accountable for the work performed therein.
Corresponding author
Ethics declarations
Conflict of interest
We declare no competing interests.
Additional information
Handling editor: Arturo Becerra.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Noda, T., Satoh, N., Gittenberger, E. et al. Left–Right Reversal Recurrently Evolved Regardless of Diaphanous-Related Formin Gene Duplication or Loss in Snails. J Mol Evol 91, 721–729 (2023). https://doi.org/10.1007/s00239-023-10130-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-023-10130-3