Abstract
Understanding the origin and early evolution of proteins is important for unveiling how the RNA world developed into an RNA–protein world. Because the composition of organic molecules in the Earth’s primitive environment was plausibly not as diverse as today, the number of different amino acids used in early protein synthesis is likely to be substantially less than the current 20 proteinogenic residues. In this study, we have explored the thermal stability and RNA binding of ancestral variants of the ribosomal protein uS8 constructed from a reduced-alphabet of amino acids. First, we built a phylogenetic tree based on the amino acid sequences of uS8 from multiple extant organisms and used the tree to infer two plausible amino acid sequences corresponding to the last bacterial common ancestor of uS8. Both ancestral proteins were thermally stable and bound to an RNA fragment. By eliminating individual amino acid letters and monitoring thermal stability and RNA binding in the resulting proteins, we reduced the size of the amino acid set constituting one of the ancestral proteins, eventually finding that convergent sequences consisting of 15- or 14-amino acid alphabets still folded into stable structures that bound to the RNA fragment. Furthermore, a simplified variant reconstructed from a 13-amino-acid alphabet retained affinity for the RNA fragment, although it lost conformational stability. Collectively, RNA-binding activity may be achieved with a subset of the current 20 amino acids, raising the possibility of a simpler composition of RNA-binding proteins in the earliest stage of protein evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
All extant organisms use the same DNA–RNA–protein system known as the “central dogma of molecular biology”, wherein DNA contains the genetic information and proteins work as functional molecules. Therefore, many biologists believe that all extant organisms are descendants of a single common ancestor – in other words, the hypothesis first predicted by Darwin (Darwin 1859). While not all biologists necessarily believe in the single common ancestor hypothesis (Doolittle 1999; Kandler 1995; Woese and Fox 1977), all modern organisms share significantly similar mechanisms for replication and expression of genetic information – a fact that supports the existence of a single ancestor because it seems unlikely that such similar mechanisms were established many times independently. It should be kept in mind that the common ancestor is not the “oldest”, but the “most recent” common ancestor of all extant organisms, which is often referred to as the last universal common ancestor (LUCA). This differentiation is necessary because the oldest ancestor that appeared on primitive Earth might have diversified over several million years (Cornish-Bowden and Cardenas 2017). Most of the subsequent primitive organisms might have become extinct for various reasons and only LUCA might survive (Nisbet and Sleep 2001). One study has suggested that LUCA was an anaerobic, autotrophic microorganism with a metabolic system for nitrogen fixation and carbonate fixation using hydrogen as an electron acceptor (Weiss et al. 2016), while another has suggested that LUCA could already synthesize proteins using 20 types of amino acids, similar to extant organisms (Mat et al. 2008).
Regardless of the exact nature of LUCA, the origins of life, which emerged earlier, remain a long-running controversy because, in extant organisms, the nucleic acid polymers (i.e., DNA and RNA) carry the information for the amino acid sequences of proteins, while proteins play a central role in the replication of nucleic acid polymers. More than 50 years ago, Rich proposed the idea that RNA served as both a genetic and functional molecule in the primitive environment (Rich 1962), while Gilbert coined the ‘RNA world hypothesis’ in 1986 (Gilbert 1986). Much experimental evidence now supports the hypothesis that life on Earth began with an RNA molecule (Guerrier-Takada et al. 1983; Kruger et al. 1982; Nissen et al. 2000).
In extant organisms, proteins are generally composed from 20 types of l-amino acid, which the organisms obtain by two primary means: intracellular synthesis by the metabolic system if the corresponding amino acid synthesis pathway is available; or acquisition from the external environment if it is not. It can be reasonably assumed that primitive proteins were synthesized using only amino acids available in the environment before innovation of the corresponding intracellular amino acid synthetic pathways (Cleaves 2010; Shibue et al. 2018). To our best knowledge, no experimental evidence supports the plausibility that all 20 of the current proteinogenic amino acids were present at concentrations sufficient for synthesizing primitive proteins in the environment 4 billion years ago. However, classic Miller–Urey experiments simulating the hypothetical early Earth environment suggest that the synthesis of organic compounds from inorganic substances was possible (Ferus et al. 2017; Weber and Miller 1981; Miller 1953). As well as various other compounds, those experiments yielded many amino acids, of which only 10 of the 20 current proteinogenic amino acids were present. The 10 amino acids were also found in samples returned from the asteroid Ryugu by the Hayabusa2 spacecraft (Nakamura et al. 2022). Further clues to the amino acids present on early Earth have been obtained from the Murchison meteorite, which is rich in organic compounds (Wolman et al. 1972). The meteorite contains more than 70 amino acids, but only eight are essential for current protein synthesis (Cronin 1989; Cronin and Pizzarello 1983). Thus, it can be reasonably assumed that only a subset of the current 20 amino acids was present in sufficient amount in primitive Earth’s environment. Other evidence supports the idea that proteins with fewer than 20 amino acids were synthesized in the early stages of evolution before the emergence of LUCA (Akanuma et al. 2002; Angyan et al. 2014; Cornell et al. 2019; Giacobelli et al. 2022; Longo et al. 2013; Shibue et al. 2018; Solis 2019; Yagi et al. 2021).
It is possible that early proteins and RNA served as mutual cofactors and scaffolds, necessitating strong interactions (Lupas and Alva 2017; Vázquez-Salazar and Lazcano 2018). Because ribose, which forms the backbone of RNA, and its analog are quickly decomposed at high temperatures (Larralde et al. 1995), Miller and Lazcano pointed out that the earliest life was unlikely to thrive in a high-temperature environment (Miller and Lazcano 1995). Conversely, Pearce and colleagues predicted that a hot early environment on Earth (50–80 °C) would favor rapid nucleotide synthesis as compared with a warm early environment (5–35 °C) (Pearce et al. 2017). Geochemical evidence suggests that early Earth underwent catastrophic meteoritic bombardment (Chyba 1990); therefore, the presumed early environment was likely to be hotter and more unstable in terms of climate change than today (Knauth and Lowe 1978; Robert and Chaussidon 2006). This unstable environment would be fatal to single-stranded RNA structures and highly likely to cause RNA inactivation and even breakage. Therefore, the role of the earliest proteins might have been to stabilize the RNA molecules that are hypothesized to play a central role in the RNA world (Shibue et al. 2018).
By comparing a large number of extant homologous protein sequences, ancestral protein reconstruction can resurrect the proteins of past organisms (Akanuma and Yamagishi 2016; Gaucher et al. 2010; Merkl and Sterner 2016; Rouet et al. 2017; Thornton 2004; Wheeler et al. 2016). Many studies have used this approach, for example, to understand the evolution of ethanol production and consumption in yeast (Thomson et al. 2005) and the trajectory of ligand-specific changes in hormone receptors (Bridgham et al. 2006, 2009; Harms and Thornton 2013; Ortlund et al. 2007), and to estimate ancient biosphere temperature (Akanuma et al. 2013; Garcia et al. 2017; Gaucher et al. 2008, 2003).
To further explore the subset of amino acids present in early proteins, here we have applied the reconstruction method to the ribosomal protein uS8 (named according to the new system for ribosomal proteins; Ban et al. 2014), which directly interacts with the central domain of 16S rRNA (Wiener et al. 1988). uS8 is a 130-residue protein essential for organization of the central domain of the small subunit of the ribosome (Collatz et al. 1976). Its deletion prevents other ribosomal proteins from assembly on the 30S small subunit, resulting in a significant loss of protein synthesis (Allmang et al. 1994; Shimojo et al. 2020). First, we inferred two potential ancestral sequences of uS8 using the information contained in a predictive phylogenetic tree of the amino acid sequences of extant uS8 proteins, and characterized the resulting ancestral uS8 proteins in terms of thermal stability and RNA-binding properties. Next, by eliminating one amino acid letter at a time from the ancestral uS8 sequence, we identified amino acids that are not essential for RNA binding, and used this information to create simplified uS8 variants lacking multiple types of amino acid to derive a minimal set of amino acids essential for a stable uS8 variant with RNA-binding activity. Lastly, we compared this minimal set with amino acids that have been identified as plausibly abundant in the prebiotic environment by earlier geochemical studies.
Materials and Methods
Phylogenetic Tree Building and Ancestral Amino Acid Sequence Inference
A BlastP search (Altschul et al. 1997) of the inhouse KF database v.1.2, which contains all protein sequences of 804 organisms (Furukawa et al. 2017), was performed to construct a dataset of uS8 amino acid sequences. The amino acid sequence of Thermus thermophilus uS8 (accession numbers: AAB25287) was used as a query sequence because it is one of the most well-studied uS8 proteins. Methanococcus maripaludis uS8 (WP_011171358) was used as a query sequence to retrieve archaeal and eukaryote sequences. Duplicate identical amino acid sequences were removed and the remaining sequences were used as the primary dataset. Individual sequence datasets for Bacteria, Archaea and Eukaryotes were aligned independently using MAFFT ver.7.3 (Katoh and Standley 2013). Amino acid sequences annotated as proteins other than uS8 were removed from the alignment. Sequences with a long insertion (> 50 amino acids) relative to T. thermophilus and M. maripaludis uS8 proteins were also removed. The alignment was then manually corrected by referring to secondary structure information and the known tertiary structures of uS8 proteins from T. thermophilus (PDB code: 1QD7), Bacillus anthracis (PDB code: 4PDB) and Methanocaldococcus jannaschii (PDB code: 1I6U).
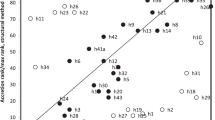
Conserved regions in the final alignment were selected via the automated1 mode, gappyout mode, and no gaps mode of trimAl (Capella-Gutierrez et al. 2009). IQ-TREE v. 1.6.9 (Nguyen et al. 2015), in conjunction with the LG + R8 amino acid substitution model, was used to build a phylogenetic tree (Figure. S1). ModelFinder (Kalyaanamoorthy et al. 2017) selected LG + R8 as the optimal amino acid substitution model. We removed eukaryotic sequences from the dataset, as well as some prokaryotic sequences that might cause long-branch attraction, and recalculated the tree. Again, IQ-TREE v. 1.6.9 (Nguyen et al. 2015) was used in conjunction with the LG + R9 amino acid substitution model, which was selected as the optimal amino acid substitution model by ModelFinder (Kalyaanamoorthy et al. 2017). We also build other trees with LG + R7, LG + R8, and LG + R10 amino acid substitution models. The four resulting trees all showed a pectinate shape topology with a few sequences branching directly near the basal position of the tree (Fig. S2-1–4), probably due to long-branch attraction. We therefore used a site-heterogeneous mixture model (CAT) as an alternative amino acid substitution model (Lartillot and Philippe 2004) because this model is expected to suppress long-branch attraction artefacts (Lartillot et al. 2007). We build six more phylogenetic trees using IQ-TREE in conjunction with an LG + C10 + F + G, LG + C20 + F + G, LG + C30 + F + G, LG + C40 + F + G, LG + C50 + F + G, or LG + C60 + F + G amino acid substitution model (Fig. S2-5–10). Among the six resulting trees, the tree built with the LG + C30 + F + G model showed the best log likelihood score, although a possible long-branch attraction artefact (branch leading to Gold_HGW-Goldbacteria-1_PKL91838.1) was still observed (Fig. 1, Fig. S2-7 and Fig. S3). Using the phylogenetic tree built with the LG + C30 + F + G model and either IQ-TREE or CodeML in PAML (Yang 2007), we inferred two ancestral uS8 sequences (named I_Bac and P_Bac, respectively; Fig. 2 and Fig. S4) that might correspond to the last bacterial common ancestor. GASP (Edwards and Shields 2004) was used to estimate the location of gaps in the ancestral sequences. The amino acid sequences of I_Bac and P_Bac are available in fasta format (Supplementary Data 1).
Amino acid sequence comparison of the two bacterial ancestral uS8 proteins. Residues that differ between the two ancestral sequences are shown in bold. Boxes indicate plausible RNA-binding residues predicted using the structure of the B. anthracis uS8–RNA complex (PDB code: 4PDB) as a guide (see Fig. S9)
Construction of Expression Plasmids for Ancestral uS8 and Simplified Variants
Nucleotide sequences encoding the last bacterial common ancestral uS8 were generated by reverse-translating the inferred ancestral amino acid sequences. Codon usage was optimized for an Escherichia coli expression system. The nucleotide sequences were artificially synthesized by Eurofins Genomics and cloned into the NdeI-BamHI site of plasmid pET23a( +) (Merck).
The genes encoding uS8 from T. thermophilus and B. anthracis were artificially synthesized by Eurofins Genomics and cloned into the NdeI-XhoI site of plasmid pET23a( +) (Merck), which expressed the proteins as a C-terminally His-tagged form.
The genes encoding simplified P_Bac variants were also synthesized by Eurofins Genomics and cloned into the NdeI-BamHI site of plasmid pET23a( +), except for those encoding simplified P_Bac variants lacking cysteine, phenylalanine, threonine or tryptophan, which were synthesized by the splicing-by-overlap-extension PCR method (Horton et al. 1993). The mutated genes were PCR-amplified in a reaction mixture containing 1 × PCR buffer for KOD Plus DNA polymerization (Toyobo), 1 mM MgSO4, 0.2 mM each of the dNTPs, 0.25 μM each of the synthetic oligonucleotides, 1.0 unit of KOD Plus DNA polymerase, and the expression plasmid for P_Bac as the template DNA. The PCR conditions were 95 °C for 3 min, followed by 25 cycles of 95 °C for 30 s, 55 °C for 30 s, and 68 °C for 1 min. The PCR product was digested with NdeI and BamHI (New England Biolabs), and cloned into the NdeI-BamHI site of pET23a( +). The genes encoding simplified P_Bac variants devoid of multiple types of amino acid were artificially synthesized by Eurofins Genomics and cloned into the NdeI-XhoI site of plasmid pET23a( +) (Merck) to produce the protein as a C-terminally His-tagged form.
Overexpression of uS8 Proteins
E. coli C41 (DE3) pLysS (Lucigen) was transformed with the expression plasmids for the bacterial ancestral proteins and simplified variants. Transformants were spread on Luria–Bertani (LB) medium plate supplemented with ampicillin (150 μg/ml) and grown overnight at 37 °C. For protein production, one colony was inoculated into 2 ml of LB liquid medium containing ampicillin (150 μg/ml) and shaken at 37 °C for 15 h. Next, 2 ml of this culture was added to 200 ml of LB medium containing ampicillin (150 μg/ml) and shaken at 37 °C for 3 h. Isopropyl β-D-thiogalactopyranoside (final concentration, 1 mM) was added and incubation was continued at 30 °C for 18 h. Finally, the cells were harvested by centrifugation at 5,000 g for 10 min, the supernatant was removed, and the cells were stored at –20 °C.
Purification of Proteins
To purify the bacterial ancestral proteins and the simplified variants lacking a single type of amino acid, each cell pellet was resuspended in 10 ml of 20 mM Tris–HCl, pH 6.8, 800 mM NaCl, disrupted by sonication, and then centrifuged at 18,000×g for 20 min at 4 °C. The supernatant was heat-treated at 70 °C for 20 min to precipitate proteins originating from E. coli, which were removed by centrifugation at 18,000×g for 20 min at 4 °C. The resulting supernatant was diluted with 20 mM Tris–HCl, pH 6.8, to a NaCl concentration of 250 mM and then passed through HiTrap-SP FF (Cytiva). Fractions containing uS8 were recovered, dialyzed against 20 mM Tris–HCl, pH 8.8, 250 mM NaCl, and then passed through HiTrap-SP FF again.
To purify the simplified variants lacking multiple types of amino acid, each cell pellet was resuspended in 10 ml of 20 mM Tris–HCl, pH 7.5, 800 mM NaCl, 30 mM imidazole and disrupted by sonication. After centrifugation at 18,000×g for 20 min at 4 °C, the supernatant was passed through a HisTrap HP column (Cytiva). T. thermophilus uS8 was purified by a similar method.
B. anthracis uS8 was purified under denaturing conditions because the protein collected in inclusion bodies. The cell pellet was resuspended in 10 ml of 20 mM Tris–HCl, pH 7.5, 800 mM NaCl, 30 mM imidazole, and disrupted by sonication. The soluble protein fraction was removed by centrifugation at 18,000×g for 20 min at 4 °C. Insoluble protein was dissolved in 10 ml of 20 mM Tris–HCl, pH 7.5, 800 mM NaCl, 1 mM dithiothreitol, 30 mM imidazole containing 7.0 M urea, and passed through a HisTrap HP column (Cytiva). The solution containing B. anthracis uS8 was step-wise dialyzed against 20 mM Tris–HCl, pH 7.5, 800 mM NaCl containing 7.0 M, 5.0 M, 3.0 M, 1.0 M and 0.5 M urea. Lastly, the protein solution was dialyzed against 20 mM Tris–HCl, pH 7.5, 800 mM NaCl twice, and the protein molecules that remained insoluble were removed by centrifugation at 18,000×g for 20 min at 4 °C.
The purity of each protein was > 95% as judged by SDS–polyacrylamide gel electrophoresis (SDS-PAGE) followed by Coomassie Brilliant Blue staining (Fig. S5). Protein concentrations were determined by measuring the A280 values of the samples as described by Pace et al. (1995) because all proteins analyzed in this study contained either tyrosine or tryptophan, or both residues.
Circular Dichroism Measurement
Circular dichroism (CD) measurements were carried out using a J-1100 CD spectropolarimeter (Jasco) equipped with a programmable temperature controller. Proteins were diluted to 20 μM in 20 mM potassium phosphate buffer (pH 7.6), 200 mM NaCl, and placed in a quartz glass cell with a 0.1-cm path length. Far-UV CD spectra were recorded from 200 to 250 nm at 25 °C.
Temperature-induced unfolding of the proteins was measured in duplicate by monitoring the change in ellipticity at 222 nm. Proteins were diluted to 20 μM in 20 mM potassium phosphate buffer, pH 7.6, 200 mM NaCl. The temperature was increased at a rate of 1.0 °C/min. A pressure-proof cell compartment was used to prevent the solutions from bubbling and evaporating at high-temperature.
RNA-Binding Assays
Interactions between an RNA fragment and I_Bac, P_Bac and some variants of P_Bac were examined quantitatively using a BLItz System with a streptavidin sensor chip (FORTEBIO/Zartorius Japan) at 25 °C. We used a previously reported RNA fragment selected for binding to B. anthracis uS8 by an in vitro aptamer selection method (Davlieva et al. 2014). The sequence (5’-GGG AUG CUC AGU GAU CCU UCG GGA UAU CAG GGC AUC CC-3’) with a 5’ biotin modification was artificially synthesized by Eurofins Genomics. The sensor chip was washed with running buffer (20 mM Tris–HCl, pH 7.5, 800 mM NaCl, and 50 μM BSA), placed in RNA (50 μM) solution diluted with running buffer, and washed with buffer again. The sensor chip was then placed in uS8 (2.0 or 10 μM) solution diluted with running buffer, and binding of uS8 to RNA captured on the sensor chip was measured. Lastly, the sensor chip was placed in the running buffer to measure the dissociation of uS8 from the RNA fragment. Rate constants for association (ka) and dissociation (kd), and dissociation constant (KD) were calculated by the BLItz System built-in software.
Interactions between the RNA fragment and P_Bac variants lacking a single type of amino acid were examined by pull-down assay. The RNA fragment was captured with Magnosphere MS300/Streptavidin (SR Life Sciences) magnet beads. In brief, 1 µL of 50 μM RNA solution was incubated with 100 µL of magnet beads (pre-treated according to the manufacturer’s protocol) for 15 min at 4 °C with agitation. The beads were precipitated by the magnetic stand, the supernatant was removed, and the RNA-bound beads were washed twice with 20 mM Tris–HCl, pH 7.5. The beads were resuspended in 100 µL of 20 mM Tris–HCl, pH 7.5, 2 mM MgCl2, 0.1% Tween 20, 800 mM NaCl, an equal volume of a solution containing 3.0 µM uS8 and 30 µM BSA was added, and the suspension was incubated for 15 min at 4 °C with agitation. The beads were then precipitated by the magnetic stand, the supernatant was removed, and the beads were washed three times with 20 mM Tris–HCl, pH 7.5, 800 mM NaCl, 0.1% Tween 20. Lastly, the beads were resuspended in 10 µL of ultrapure water, uS8 was dissociated by boiling for 20 min in 2% SDS, and the samples were analyzed by SDS-PAGE. The interaction of the variants with RNA-free magnetic beads was also analyzed as a negative control.
Results and Discussion
Phylogenetic Tree Building and Ancestral Amino Acid Sequence Inference
The first step in reconstructing an ancestral sequence is to perform a multiple sequence alignment using the amino acid sequences of the target protein from multiple living organisms. The alignment is then used to build a phylogenetic tree by a modeling approach, such as maximum–likelihood (ML) (Yang et al. 1995) or Bayesian (Yang and Rannala 1997) modelling. In this study, an ML method was used because ML is reported to be relatively accurate in the reconstruction of ancestral sequences (Hanson-Smith et al. 2010).
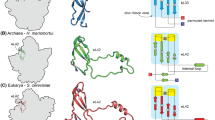
The multiple sequence alignment included the amino acid sequences of the ribosomal protein uS8 from 582 bacterial, 140 archaeal and 138 eukaryotic species. Although there were many more sequences from bacteria than from archaea or eukaryotes, this was not considered an issue because the primary aim at this stage was to infer the sequence of the bacterial common ancestor. In the resulting phylogenetic tree, built using the ML program IQ-TREE (Nguyen et al. 2015), the bacteria and archaeal sequences are clearly divided into their own monophyletic groups, while eukaryotic sequences are found among the archaeal sequences (Fig. S1). Our phylogenetic tree supports the two-domain hypothesis of all modern life, which proposes that eukaryotes emerged within the archaeal domain (Cox et al. 2008; Raymann et al. 2015; Rivera and Lake 1992; Williams et al. 2013), and is consistent with a recently reported tree built using the concatenated sequences of ribosomal proteins (Hug et al. 2016). By contrast, a phylogenetic tree based on small subunit ribosomal RNA sequences showed a monophyletic status of Bacteria, Archaea, and Eukarya (Woese et al. 1990), with Eukarya located as a sister group of Archaea. That tree, together with various molecular phylogenetic studies and phylogenomic studies support the three-domain hypothesis of all modern life (Ciccarelli et al. 2006; Fournier and Gogarten 2010; Harris et al. 2003; Rinke et al. 2013; Yutin et al. 2008).
Because the eukaryotic sequences are unlikely to influence estimation of the sequence at the deepest node, we removed them from the dataset, as well as some prokaryotic sequences that might cause long-branch attraction, and recalculated the tree. In the final tree built from 527 bacterial and 124 archaeal sequences (Fig. 1 and Fig. S3), the two major domains form two distinct monophyletic groups.
To define the root of the phylogenetic tree, we needed to include sequences that diverged from uS8 before LUCA as an outgroup. However, no such sequences were identified in our Blast search. Therefore, the sequence at the deepest bacterial node was inferred from the tree by treating the archaeal sequences as an outgroup. We used two programs to predict the sequence of the last bacterial common ancestor: the sequence predicted by CODEML in PAML (Yang 2007) was named P_Bac; and that predicted by IQ-TREE (Nguyen et al. 2015) was named I_Bac (Fig. 2). The amino acid sequences of P_Bac and I_Bac were very similar (121 of 130 residues are identical). Furthermore, most of the identical residues in P_Bac and I_Bac had an a posteriori probability of higher than 0.9 (Fig. S4); therefore, the inclusion of these residues seems likely to be correct. In contrast, the residues that differed between the two sequences had a relatively low a posteriori probability.
Thermal Stabilities of P_Bac and I_Bac
Genes encoding the two inferred ancestral amino acid sequences were artificially synthesized and the encoded proteins were individually expressed in E. coli and purified. Ellipticity at 222 nm was monitored as a function of temperature to generate the temperature-induced unfolding curves of the two ancestral proteins. As shown in Fig. 3, both P_Bac and I_Bac were less thermostable than T. thermophilus uS8, but still had very high unfolding mid-point temperatures (Tm, ~ 85 °C for both proteins), comparable to those of extant thermophilic proteins.
Thermal denaturation of T. thermophilus and ancestral uS8 proteins. Change in ellipticity at 222 nm was monitored as a function of temperature for T. thermophilus uS8 (dotted line), P_Bac (solid line), and I_Bac (dashed line). The temperature was increased at a rate of 1.0 °C/min. The samples comprised 20 μM protein in 20 mM potassium phosphate (pH 7.6), 200 mM NaCl. Each experiment was conducted in duplicate, which produced identical melting profiles within experimental error. The plots have been normalized with respect to the baseline of the native and denatured states
The high thermal stability of bacterial ancestral uS8 is consistent with observations of other reconstructed ancestral proteins (Akanuma et al. 2013; Busch et al. 2016; Butzin et al. 2013; Gaucher et al. 2008; Gumulya et al. 2018). There is often a direct correlation between the unfolding temperature of a protein and the optimal environmental temperature of its host organism (Akanuma et al. 2013; Gromiha et al. 1999). Furthermore, most reconstructed ancestral proteins are very thermostable, suggesting that ancestral organisms such as the last bacterial common ancestor, last archaeal common ancestor, and LUCA were thermophilic or hyperthermophilic. The high thermal stabilities of the two reconstructed bacterial ancestral ribosomal uS8 proteins also support the idea that the last bacterial common ancestor was a thermophilic organism that thrived in a high-temperature environment. We note, however, that the environmental temperature of primitive organisms remains under debate, and a non-thermophilic ancestry of life is supported by computational studies focusing on the environmental temperatures experienced by ancient life (Boussau et al. 2008; Galtier et al. 1999; Groussin et al. 2013).
It should also be noted that an accurate tree is not always obtained and ancestral sequences cannot be reconstructed with absolute certainty, although the techniques used to infer ancestral sequences have greatly improved in the past decade. Therefore, any implications derived from the tree and the ancestral sequence are hard to verify. Williams et al. proposed that the high thermal stabilities observed for ancestral proteins might be related to the inherent nature of the ancestral sequence reconstructions (Williams et al. 2006). They asserted that an inaccurately reconstructed sequence would result in an overestimation of its thermostability. Furthermore, Tawfik and coworkers have suggested that a high environmental temperature may not have been the only factor requiring the high thermodynamic stability of ancestral proteins (Trudeau et al. 2016). They considered that the stability of ancestral proteins might have been driven by high oxidative pressure and radiation levels, the absence of cellular osmolytes and/or chaperones, or the low fidelity of the transcription–translation machinery.
Other studies based on ancestral sequence reconstruction have also connected ancient proteins to early environments. For example, Schopf and colleagues reconstructed proteins from phototrophic species, which suggested that there has been a general cooling of the Earth’s photic zone from the Archean Eon to the present. In addition, Kaçar and colleagues resurrected a Precambrian-age, ancestral RuBisCO gene from extant cyanobacteria (Kędzior et al. 2022) and found that the carbon isotope signatures of the engineered cyanobacteria being cultured under potential Precambrian environments fell within modern ranges. Therefore, uniformitarian assumptions of carbon isotope signatures over geologic time might be justified, but with an important caveat because the modern organism and its proteins might have influenced the ancestral RuBisCO phenotype. Garcia and Kaçar also warned of the pitfalls of facilely interpreting paleophenotype models and data (Garcia and Kaçar 2019).
Nevertheless, even if the inferred sequence is not the correct ancestral sequence and the high thermostability of the reconstructed proteins does not reflect a high-temperature environment of the ancient organism, proteins with high thermostability remain suitable as starting molecules to simplify amino acid usage. Therefore, one of the ancestral uS8 proteins was chosen as the scaffold on which to reduce the size of the amino acid alphabet.
Interaction Between Ancestral uS8 and RNA
The interaction of the two wild-type and two ancestral uS8 proteins with biotinylated synthetic RNA captured on a streptavidin sensor chip was quantitatively analyzed in the presence of 800 mM NaCl and 50 μM BSA to suppress nonspecific adsorption of the protein. We used an RNA fragment selected by the systematic evolution of ligands using an exponential enrichment (SELEX) technique (Davlieva et al. 2014). We note here that the interactions of protein residues with SELEX-generated RNA are not the same as those in natural protein–RNA interactions. The sensor chip provides real-time data on molecular interactions: when protein molecules interact with the RNA on the sensor chip, the binding signal shifts in the positive direction. Here, the binding signal sharply increased after the RNA-bound chip was exposed to both ancestral uS8 solutions, suggesting the formation of a protein–RNA complex on the sensor chip (Fig. 4). Upon changing the solution to protein-free buffer, the binding signal slightly decreased, indicating the dissociation of some protein molecules from RNA.
Interaction of wild-type uS8 proteins, ancestral uS8 proteins and reduced-alphabet variants with RNA. RNA binding of T. thermophilus uS8 (yellow), B. anthracis uS8 (brown), P_Bac (magenta), I_Bac (blue), P_Bac-15 (green), P_Bac-14 (cyan), P_Bac-13N (purple) and P_Bac-13M (grey) was measured at a protein concentration of 2.0 μM (A) or 10 μM (B) using a streptavidin sensor chip (BLItz system). The association and dissociation curves of P_Bac-13L were almost identical to those of P_Bac-13M and have been therefore omitted. The running buffer was 20 mM Tris–HCl, pH 7.5, 800 mM NaCl, and 50 μM BSA. The RNA fragment used was previously selected for binding to B. anthracis uS8 (Davlieva et al. 2014) (Color figure online)
The BLItz System’s built-in software generated ka, kd and KD values from the association and dissociation curves (Table 1). The KD values (33 × 10–7 M and 14 × 10–7 M at protein concentrations of 10 μM and 2.0 μM, respectively) observed for B. anthracis uS8 were more than 10 times weaker than that (1.1 × 10–7 M) previously reported for the binding of B. anthracis uS8 to an RNA fragment with a similar but not identical sequence (Davlieva et al. 2014). This difference may be due to the following two reasons: (i) the previous study measured binding to free RNA, whereas we measured with RNA immobilized to a sensor chip; (ii) the previous study measured RNA–protein binding in a moderate salt concentration (150 mM potassium acetate), whereas our measurement was performed in high salt (800 mM sodium chloride) and the high salt concentration may have somewhat inhibited the binding of B. anthracis uS8 to RNA. In contrast, the binding of T. thermophilus uS8 to the RNA fragment exhibited much better KD values of 6.2 × 10–8 M and 3.5 × 10–8 M at protein concentrations of 10 μM and 2.0 μM, respectively.
I_Bac showed KD values of 8.4 × 10–8 M and 5.5 × 10–8 M at protein concentrations of 10 μM and 2.0 μM, respectively, while P_Bac showed KD values of 35 × 10–8 M (10 μM) and 27 × 10–8 M (2.0 μM); thus, I_Bac showed 4–5-fold stronger binding. The KD values observed for I_Bac were similar to those reported for the binding of T. thermophilus uS8 to the RNA fragment.
Effect of Eliminating One Amino Acid Letter on the Stability and RNA Binding of P_Bac
In a first step toward identifying a minimal set of amino acids that would retain the stability and RNA-binding properties of uS8, we individually eliminated each type of amino acid from the inferred uS8 ancestral protein. For this experiment, we used P_Bac because, based on the a posteriori probability, the accuracy of the residues in P_Bac was higher than that in I_Bac (Fig. S4). We constructed 19 simplified variants of P_Bac, the sequences of which each lacked one amino acid letter. Because P_Bac contains no histidine residues, each variant comprised an 18-amino-acid alphabet. In each variant, amino acids were replaced with the “second-best” ancestral residue; in other words, ancestral amino acids with a posteriori probability < 1.0 were replaced by the amino acid that showed second highest probability. Ancestral amino acids with a posteriori probability of 1.0 were replaced by the amino acid that was second most frequent at the corresponding position in the multiple sequence alignment used for tree building and ancestral sequence inference. Completely conserved residues were replaced by physicochemically similar amino acids. For constructing variants lacking methionine, the N-terminal residue was not taken into account. The amino acid sequences of P_Bac and its 19 variants are given in Supplementary Data 1 and aligned in Fig. S6.
The variants that lacked glycine, glutamate, isoleucine or lysine appeared to be insoluble and could not be subjected to further analysis. It seems likely that the presence of glycine, glutamate, isoleucine and lysine is crucial for the proper folding and/or thermodynamic stability of the protein. The other 15 variants were successfully expressed and recovered as soluble forms.
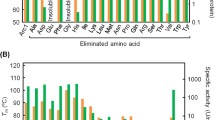
We measured the temperature-induced unfolding of each protein by monitoring the change in ellipticity at 222 nm as a function of temperature (Fig. S7). For each protein, the duplicate measurements gave identical unfolding curves within experimental error (data not shown). The unfolding curves of P_Bac and its variants showed a single transition (Fig. S7), the midpoint of which was used to compare the thermal stabilities of the proteins and identify which types of amino acid are important for protein stability. We also assessed the interaction between the variants and an RNA fragment by a magnetic beads-based pull-down assay (Fig. S8).
Table 2 summarizes the unfolding midpoint temperature and the RNA-binding ability of the simplified variants, showing that the elimination of some amino acid letters from the sequence of P_Bac exerts a large effect on its stability and/or RNA-binding ability. For example, elimination of arginine, tyrosine, proline, alanine or serine resulted in a lower unfolding temperature and loss of RNA-binding activity. Among these residues, the hydroxyl groups on the side chains of serine at positions 105 and 107 are predicted to be closely involved in RNA-binding via the formation of hydrogen bonds with the RNA molecule (Fig. S9). In addition, valine was found to be crucial for RNA binding but not for thermal stability. Elimination of methionine, asparagine or leucine lowered the unfolding temperature but did not affect RNA-binding activity, while elimination of glutamine moderately reduced the unfolding temperature. In contrast, the remaining four amino acids (phenylalanine, tryptophan, cysteine, threonine) could be eliminated from the sequence of P_Bac without compromising its stability or RNA-binding activity. Based on the structure of the B. anthracis uS8 and RNA complex (Davlieva et al. 2014), methionine, asparagine, leucine, phenylalanine, tryptophan, cysteine and threonine are unlikely to be involved in RNA binding (Fig. 2 and Fig. S9). In contrast, the side chain of glutamine at position 56 may form hydrogen bonds with the RNA molecule (Fig. S9). Nevertheless, replacement of the glutamine residue by lysine did not affect RNA binding (Table 2 and Fig. S8). Thus, the various types of amino acid do not contribute equally to the stability and RNA binding of P_Bac; in particular, these findings suggested that phenylalanine, tryptophan, cysteine and threonine might be eliminated in combination to produce more simplified variants of ancestral uS8.
Construction of Simplified P_Bac Variants Lacking Multiple Amino Acid Letters
Next, we tested whether four or more types of amino acid could be eliminated in combination without substantial loss of stability or RNA-binding activity. We first eliminated phenylalanine, tryptophan, cysteine and threonine from the sequence of P_Bac by replacing them with “second-best” ancestral amino acids. The resulting protein, P_Bac-15 (Supplementary Data 1; Fig. S6), was reasonably thermally stable (Tm = 84 °C; Fig. 5A) and its RNA-binding activity was significant (Fig. 5B). We further eliminated glutamine from P_Bac-15, thus producing P_Bac-14 (Supplementary Data 1; Fig. S6). The thermal stability of P_Bac-14 (Tm = 86 °C) was comparable to that of P_Bac and P_Bac-15 (Fig. 5A), and its RNA-binding activity was also significant (Fig. 5B). These findings indicate that a reduced amino acid alphabet, comprising only 14-amino acid letters, is sufficient to achieve high thermal stability and strong RNA binding in the ribosomal protein uS8.
Functional analysis of P_Bac-15 and P_Bac-14. A Thermal denaturation curves of P_Bac-15 (solid line) and P_Bac-14 (dashed line). The unfolding midpoint temperature is indicated. B RNA-binding assay. The binding buffer was 20 mM Tris–HCl (pH 7.5), 800 mM NaCl, 0.1% Tween 20. The wash buffer was 20 mM Tris–HCl (pH 7.5), 2 mM MgCl2, 0.1% Tween 20 containing 800 mM NaCl. Plus and minus signs above the gels represent the presence and absence of RNA, respectively
We further eliminated methionine, asparagine or leucine from P_Bac-14 by replacing them with “second-best” ancestral amino acids to produce P_Bac-13M, P_Bac-13N and P_Bac-13L, respectively (Supplementary Data 1; Fig. S6). It should be noted that methionine, leucine and asparagine are not directly involved in RNA binding (Fig. 2 and Fig. S9). The far-UV CD spectra of P_Bac-13M and P_Bac-13N (Fig. 6) indicated the presence of significant secondary structure in these variants; however, the ellipticities were smaller than that of P_Bac. The smaller ellipticity may reflect a reduced secondary structure content; alternatively, it might indicate that a certain percentage of protein molecules did not fold correctly even at room temperature. The far-UV CD spectrum of P_Bac-13L indicated that this variant did not contain significant secondary structure. In the temperature-induced unfolding experiment, none of the three variants (P_Bac-13M, P_Bac-13N and P_Bac-13L) showed a cooperative secondary structure unfolding transition under the conditions used (Fig. S10). Therefore, Tm values could not be determined for these three proteins.
We also measured the interaction of P_Bac-15, P_Bac-14 and P_Bac-13N with an RNA fragment using a streptavidin sensor chip, which determined the association and dissociation curves for P_Bac-15, P_Bac-14 and P_Bac-13N (Fig. 4). At protein concentrations of 10 μM and 2.0 μM, the KD values of P_Bac-15 were both 4.8 × 10–7 M, which is similar to those of P_Bac (Table 1). Furthermore, the KD values of P_Bac-14 were 1.9 × 10–6 M (10 μM) and 2.0 × 10–6 M (2.0 μM), indicating that this simplified uS8 variant also interacted with the RNA fragment significantly, albeit with weaker binding than P_Bac and P_Bac-15. Unexpectedly, the assay indicated that P_Bac-13N also bound to RNA to some extent, although the shift in binding signal was much smaller than that observed for the ancestral uS8 protein, P_Bac-15 and P_Bac-14. Correspondingly, the KD values of P_Bac-13N were 3.3 × 10–6 M and 2.1 × 10–6 M at protein concentrations of 10 μM and 2.0 μM, respectively, similar to those of P_Bac-14 (Table 1). No change in binding signal was observed on exposure of the RNA-bound chip to P_Bac-13M or P_Bac-13L solution (Fig. 4), showing that these two simplified proteins did not bind to the RNA fragment.
Implications for the Amino Acid Repertoire in Primitive RNA-Binding Proteins
The amino acid repertoire used in primordial protein synthesis must be closely related to the origin and early evolution of the genetic code. The ‘frozen accident’ and other theories commonly predict that, by gradually incorporating new amino acids into the repertoire, the modern genetic code has progressively evolved from a primitive, simpler one involving a subset of the current 20 proteinogenic amino acids (Baumann and Oro 1993; Crick 1968; Eigen and Schuster 1977; Higgs 2009; Ikehara et al. 2002; Johnson and Wang 2010; Wong 1975). Several studies have proposed that the functionality of proteins would have been increased by the amino acids added later to the proteinogenic amino acid repertoire. For example, the sidechains of subsequently added amino acids might have had higher chemical reactivity (Granold et al. 2018), and expanded the chemistry space in terms of size, charge and hydrophobicity (Ilardo and Freeland 2014). They might also have increased protein function (Francis 2013), or both protein structure and function to improve the fitness of primitive organisms (Muller et al. 2013). Consistent with these ideas, Trifonov proposed an all-encompassing order for amino acid emergence (G/A, V/D, P, S, E/L, T, R, N, K, Q, I, C, H, F, M, Y, W; Trifonov 2000). Recently, Mayer-Bacon and Freeland examined how the current set of 20 proteinogenic amino acids is distributed throughout extant life in terms of quantitative measures (Mayer-Bacon and Freeland 2021), showing that the remarkable distributions of volume, hydrophobicity and charge (pKa) become far more obscure when comparing a prebiotically plausible subset of amino acids with a much smaller subset of prebiotically plausible alternatives detected in meteorites. Lastly, Masel and colleagues performed integrated phylostratigraphy across 435 organisms with full genome sequences, observing that trends in amino acid usage among ancient domains reflect the order in which the amino acids were incorporated into the genetic code (James et al. 2021). They suggested that amino acid usage in the extant descendants of ancient sequences may reflect the availability of the amino acids when the sequences first emerged.
In our experiments to explore the properties of uS8 variants constructed from a reduced set of amino acids, elimination of histidine, phenylalanine, tryptophan, cysteine, threonine, glutamine, or methionine from the ancestral uS8 protein P_Bac had little effect on thermal stability or RNA-binding activity. Notably, these types of amino acid, with the exception of threonine, were presumably incorporated into protein synthesis at a relatively late stage of evolution (Jordan et al. 2005; Trifonov 2000).
The ancestral uS8 variants lacking glutamate, glycine, isoleucine or lysine were insoluble when they were expressed using the recombinant E. coli expression system. It is possible that these variants could not form adequate tertiary structures. In our previous experiment, in which the size of the amino acid set constituting an ancestral nucleoside kinase was systematically reduced, the two variants lacking either glycine or glutamate seemed to be insoluble (Shibue et al. 2018). Therefore, glutamate and glycine – considered members of the plausible prebiotically available amino acid set and presumably incorporated into protein synthesis at a relatively early stage of evolution – may be necessary for ensuring a stable conformation of proteins in general.
Because RNA is negatively charged, it is reasonable to predict that amino acids with positively charged side chains, such as lysine and arginine, will be important for RNA binding by proteins. Indeed, elimination of arginine from ancestral uS8 did result in loss of RNA-binding activity. As mentioned above, however, elimination of lysine from the ancestral uS8 led to no detectable level of expression in E. coli. It seems unlikely that lysine and arginine were synthesized in the prebiotic environment (McDonald and Storrie-Lombardi 2010) and therefore the positively charged amino acids were plausibly unavailble for the earliest protein synthesis. One hypothesis for this discrepancy is that amino acids with a simpler positively charged side chain, such as ornithine and 2,4-diaminobutyrate, may have been used instead of lysine and arginine in the synthesis of primitive proteins. For example, Tawfik and coworkers suggested that the first nucleic acid-binding proteins may have arisen from short simple sequences containing ornithine, which is not used in extant proteins (Longo et al. 2020). Alternatively, metal ions might have mediated the binding of proteins to RNA. In this regard, Hlouchová and coworkers generated a genetic library encoding modified amino acid sequences of the C-terminal domain of ribosomal protein uL11 by combining 10 types of amino acid without a positively charged side chain (Giacobelli et al. 2022). After selection for RNA binding by a mRNA display method, they obtained a uL11 variant in which glutamate residues, instead of positively charged amino acids, facilitated binding to the phosphate groups of RNA via Mg2+ ions. However, the possibility that either lysine or arginine or both were abiotically synthesized in some way and available in the primordial environment cannot be completely ruled out. For example, Sutherland and colleagues have reported an abiotic synthesis pathway for a precursor to arginine (Patel et al. 2015).
We note that our study has some limitations. First, our sequence-wide individual substitutions ignored any epistasis effect between residues: while all residues of one kind were substituted across the alphabet, we note that compensatory mutations would be likely to occur in nature throughout evolution. Second, we reduced the alphabet for a reconstructed ancestral protein, and it remains to be tested whether mutating wild-type ribosomal proteins would have the same impact. In the future, we will conduct studies along these lines to explore further the implications for evolution of the amino acid alphabet.
In conclusion, construction of a phylogenetic tree based on uS8 amino acid sequences from representative extant organisms enabled us to infer two amino acid sequences corresponding to the last bacterial common ancestor of uS8; the proteins reconstructed from the sequences were thermally stable and bound to an RNA fragment. Among a series of elimination variants, the most simplified sequence variant (P_Bac-13N), lacking seven amino acid letters, was still able to bind to the RNA fragment. Collectively, our findings show that the full set of 20 proteinogenic amino acids is not necessarily essential to create an RNA-binding protein, raising the possibility that primitive RNA-binding proteins in the early stage of evolution were made from a reduced amino acid set. It is impossible, however, to assert definitively that the amino acids excluded from our simplified uS8 protein were not used in the synthesis of primitive proteins. Moreover, even if not all of the current 20 proteinogenic amino acids were available, other non-proteinogenic amino acids might have existed in the primitive environment. In short, it cannot be entirely ruled out that, before the emergence of LUCA, protein synthesis involved more than 20 amino acids, which were subsequently ‘standardized’ to the current set at an early stage of evolution leading to LUCA. In that case, the early single genetic code might have specified multiple amino acids unambiguously in the primitive translation system.
References
Akanuma S et al (2013) Experimental evidence for the thermophilicity of ancestral life. Proc Natl Acad Sci USA 110:11067–11072. https://doi.org/10.1073/pnas.1308215110
Akanuma S, Kigawa T, Yokoyama S (2002) Combinatorial mutagenesis to restrict amino acid usage in an enzyme to a reduced set. Proc Natl Acad Sci USA 99:13549–13553. https://doi.org/10.1073/pnas.222243999
Allmang C, Mougel M, Westhof E, Ehresmann B, Ehresmann C (1994) Role of conserved nucleotides in building the 16S rRNA binding site of E. coli ribosomal protein S8. Nucleic Acids Res 22:3708–3714. https://doi.org/10.1093/nar/22.18.3708
Altschul SF, Madden TL, Schäffer AA, Zhang J, Zhang Z, Miller W, Lipman DJ (1997) Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res 25:3389–3402. https://doi.org/10.1093/nar/25.17.3389
Angyan AF, Ortutay C, Gaspari Z (2014) Are proposed early genetic codes capable of encoding viable proteins? J Mol Evol 78:263–274. https://doi.org/10.1007/s00239-014-9622-3
Ban N et al (2014) A new system for naming ribosomal proteins. Curr Opin Struct Bio 24:165–169. https://doi.org/10.1016/j.sbi.2014.01.002
Baumann U, Oro J (1993) Three stages in the evolution of the genetic code. Biosystems 29:133–141
Boussau B, Blanquart S, Necsulea A, Lartillot N, Gouy M (2008) Parallel adaptations to high temperatures in the Archaean eon. Nature 456:942–945. https://doi.org/10.1038/nature07393
Bridgham JT, Carroll SM, Thornton JW (2006) Evolution of hormone-receptor complexity by molecular exploitation. Science 312:97–101. https://doi.org/10.1126/science.1123348
Bridgham JT, Ortlund EA, Thornton JW (2009) An epistatic ratchet constrains the direction of glucocorticoid receptor evolution. Nature 461:515–519. https://doi.org/10.1038/nature08249
Busch F, Rajendran C, Heyn K, Schlee S, Merkl R, Sterner R (2016) Ancestral tryptophan synthase reveals functional sophistication of primordial enzyme complexes. Cell Chem Biol 23:709–715. https://doi.org/10.1016/j.chembiol.2016.05.009
Butzin NC, Lapierre P, Green AG, Swithers KS, Gogarten JP, Noll KM (2013) Reconstructed ancestral Myo-inositol-3-phosphate synthases indicate that ancestors of the thermococcales and thermotoga species were more thermophilic than their descendants. PLoS ONE 8:e84300. https://doi.org/10.1371/journal.pone.0084300
Capella-Gutierrez S, Silla-Martinez JM, Gabaldon T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Chyba CF (1990) Impact delivery and erosion of planetary oceans in the early inner solar system. Nature 343:129–133. https://doi.org/10.1038/343129a0
Ciccarelli FD, Doerks T, von Mering C, Creevey CJ, Snel B, Bork P (2006) Toward automatic reconstruction of a highly resolved tree of life. Science 311:1283–1287. https://doi.org/10.1126/science.1123061
Cleaves HJ 2nd (2010) The origin of the biologically coded amino acids. J Theor Biol 263:490–498. https://doi.org/10.1016/j.jtbi.2009.12.014
Collatz E, Wool IG, Lin A, Stöffler G (1976) The isolation of eukaryotic ribosomal proteins. the purification and characterization of the 40 S ribosomal subunit proteins S2, S3, S4, S5, S6, S7, S8, S9, S13, S23/S24, S27, and S28. J Biol Chem 251:4666–4672
Cornell CE et al (2019) Prebiotic amino acids bind to and stabilize prebiotic fatty acid membranes. Proc Natl Acad Sci USA 116:17239–17244. https://doi.org/10.1073/pnas.1900275116
Cornish-Bowden A, Cardenas ML (2017) Life before LUCA. J Theor Biol 434:68–74. https://doi.org/10.1016/j.jtbi.2017.05.023
Cox CJ, Foster PG, Hirt RP, Harris SR, Embley TM (2008) The archaebacterial origin of eukaryotes. Proc Natl Acad Sci USA 105:20356–20361. https://doi.org/10.1073/pnas.0810647105
Crick FH (1968) The origin of the genetic code. J Mol Biol 38:367–379
Cronin JR (1989) Origin of organic compounds in carbonaceous chondrites. Adv Space Res 9:59–64. https://doi.org/10.1016/0273-1177(89)90364-5
Cronin JR, Pizzarello S (1983) Amino acids in meteorites. Adv Space Res 3:5–18
Darwin C (1859) On the origin of species by means of natural selection, or the preservation of favoured races in the struggle for life. John Murray, London
Davlieva M, Donarski J, Wang J, Shamoo Y, Nikonowicz EP (2014) Structure analysis of free and bound states of an RNA aptamer against ribosomal protein S8 from Bacillus anthracis. Nucleic Acids Res 42:10795–10808. https://doi.org/10.1093/nar/gku743
Doolittle WF (1999) Phylogenetic classification and the universal tree. Science 284:2124–2129
Edwards RJ, Shields DC (2004) GASP: gapped ancestral sequence prediction for proteins. BMC Bioinformatics 5:123. https://doi.org/10.1186/1471-2105-5-123
Eigen M, Schuster P (1977) The hypercycle. a principle of natural self-organization. part a: emergence of the hypercycle. Naturwissenschaften 64:541–565
Ferus M et al (2017) Formation of nucleobases in a miller-urey reducing atmosphere. Proc Natl Acad Sci USA 114:4306–4311. https://doi.org/10.1073/pnas.1700010114
Fournier GP, Gogarten JP (2010) Rooting the ribosomal tree of life. Mol Biol Evol 27:1792–1801. https://doi.org/10.1093/molbev/msq057
Francis BR (2013) Evolution of the genetic code by incorporation of amino acids that improved or changed protein function. J Mol Evol 77:134–158. https://doi.org/10.1007/s00239-013-9567-y
Furukawa R, Nakagawa M, Kuroyanagi T, Yokobori SI, Yamagishi A (2017) Quest for ancestors of eukaryal cells based on phylogenetic analyses of aminoacyl-tRNA synthetases. J Mol Evol 84:51–66. https://doi.org/10.1007/s00239-016-9768-2
Galtier N, Tourasse N, Gouy M (1999) A nonhyperthermophilic common ancestor to extant life forms. Science 283:220–221
Garcia AK, Kaçar B (2019) How to resurrect ancestral proteins as proxies for ancient biogeochemistry. Free Radic Biol Med 140:260–269. https://doi.org/10.1016/j.freeradbiomed.2019.03.033
Garcia AK, Schopf JW, Yokobori SI, Akanuma S, Yamagishi A (2017) Reconstructed ancestral enzymes suggest long-term cooling of earth’s photic zone since the Archean. Proc Natl Acad Sci USA 114:4619–4624. https://doi.org/10.1073/pnas.1702729114
Gaucher EA, Thomson JM, Burgan MF, Benner SA (2003) Inferring the palaeoenvironment of ancient bacteria on the basis of resurrected proteins. Nature 425:285–288. https://doi.org/10.1038/nature01977
Gaucher EA, Govindarajan S, Ganesh OK (2008) Palaeotemperature trend for precambrian life inferred from resurrected proteins. Nature 451:704–707. https://doi.org/10.1038/nature06510
Gaucher EA, Kratzer JT, Randall RN (2010) Deep phylogeny–how a tree can help characterize early life on earth. Cold Spring Harb Perspect Biol 2:a002238. https://doi.org/10.1101/cshperspect.a002238
Giacobelli VG et al (2022) In vitro evolution reveals noncationic protein-RNA interaction mediated by metal ions. Mol Biol Evol 39:msac032. https://doi.org/10.1093/molbev/msac032
Gilbert W (1986) Origin of life: the RNA world. Nature 319:618–618. https://doi.org/10.1038/319618a0
Granold M, Hajieva P, Toşa MI, Irimie FD, Moosmann B (2018) Modern diversification of the amino acid repertoire driven by oxygen. Proc Natl Acad Sci USA 115:41–46. https://doi.org/10.1073/pnas.1717100115
Gromiha MM, Oobatake M, Sarai A (1999) Important amino acid properties for enhanced thermostability from mesophilic to thermophilic proteins. Biophys Chem 82:51–67
Groussin M, Boussau B, Charles S, Blanquart S, Gouy M (2013) The molecular signal for the adaptation to cold temperature during early life on earth. Biol Lett 9:20130608
Guerrier-Takada C, Gardiner K, Marsh T, Pace N, Altman S (1983) The RNA moiety of ribonuclease P is the catalytic subunit of the enzyme. Cell 35:849–857. https://doi.org/10.1016/0092-8674(83)90117-4
Gumulya Y et al (2018) Engineering highly functional thermostable proteins using ancestral sequence reconstruction. Nat Catal 1:878–888. https://doi.org/10.1038/s41929-018-0159-5
Hanson-Smith V, Kolaczkowski B, Thornton JW (2010) Robustness of ancestral sequence reconstruction to phylogenetic uncertainty. Mol Biol Evol 27:1988–1999. https://doi.org/10.1093/molbev/msq081
Harms MJ, Thornton JW (2013) Evolutionary biochemistry: revealing the historical and physical causes of protein properties. Nat Rev Genet 14:559–571. https://doi.org/10.1038/nrg3540
Harris JK, Kelley ST, Spiegelman GB, Pace NR (2003) The genetic core of the universal ancestor. Genome Res 13:407–412. https://doi.org/10.1101/gr.652803
Higgs PG (2009) A four-column theory for the origin of the genetic code: tracing the evolutionary pathways that gave rise to an optimized code. Biol Direct 4:16. https://doi.org/10.1186/1745-6150-4-16
Horton RM, Ho SN, Pullen JK, Hunt HD, Cai Z, Pease LR (1993) Gene splicing by overlap extension. Methods Enzymol 217:270–279
Hug LA et al (2016) A new view of the tree of life. Nat Microbiol 1:16048. https://doi.org/10.1038/nmicrobiol.2016.48
Ikehara K, Omori Y, Arai R, Hirose A (2002) A novel theory on the origin of the genetic code: a GNC-SNS hypothesis. J Mol Evol 54:530–538. https://doi.org/10.1007/s00239-001-0053-6
Ilardo MA, Freeland SJ (2014) Testing for adaptive signatures of amino acid alphabet evolution using chemistry space. J Syst Chem 5:1. https://doi.org/10.1186/1759-2208-5-1
James JE, Willis SM, Nelson PG, Weibel C, Kosinski LJ, Masel J (2021) Universal and taxon-specific trends in protein sequences as a function of age. Elife. https://doi.org/10.7554/eLife.57347
Johnson DB, Wang L (2010) Imprints of the genetic code in the ribosome. Proc Natl Acad Sci USA 107:8298–8303. https://doi.org/10.1073/pnas.1000704107
Jordan IK, Kondrashov FA, Adzhubei IA, Wolf YI, Koonin EV, Kondrashov AS, Sunyaev S (2005) A universal trend of amino acid gain and loss in protein evolution. Nature 433:633–638. https://doi.org/10.1038/nature03306
Kalyaanamoorthy S, Minh BQ, Wong TKF, von Haeseler A, Jermiin LS (2017) ModelFinder: fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–589. https://doi.org/10.1038/nmeth.4285
Kandler O (1995) Cell wall biochemistry in archaea and its phylogenetic implications. J Biol Phys 20:165–169. https://doi.org/10.1007/bf00700433
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Kędzior M, Garcia AK, Li M, Taton A, Adam ZR, Young JN, Kaçar B (2022) Resurrected Rubisco suggests uniform carbon isotope signatures over geologic time. Cell Rep 39:110726. https://doi.org/10.1016/j.celrep.2022.110726
Knauth LP, Lowe DR (1978) Oxygen isotope geochemistry of cherts from the onverwacht group (3.4 billion years), transvaal, south africa, with implications for secular variations in the isotopic composition of cherts. Earth Planet Sci Lett 41:209–222. https://doi.org/10.1016/0012-821X(78)90011-0
Kruger K, Grabowski PJ, Zaug AJ, Sands J, Gottschling DE, Cech TR (1982) Self-splicing RNA: autoexcision and autocyclization of the ribosomal RNA intervening sequence of Tetrahymena. Cell 31:147–157. https://doi.org/10.1016/0092-8674(82)90414-7
Larralde R, Robertson MP, Miller SL (1995) Rates of decomposition of ribose and other sugars: implications for chemical evolution. Proc Natl Acad Sci USA 92:8158–8160
Lartillot N, Philippe H (2004) A bayesian mixture model for across-site heterogeneities in the amino-acid replacement process. Mol Biol Evol 21:1095–1109. https://doi.org/10.1093/molbev/msh112
Lartillot N, Brinkmann H, Philippe H (2007) Suppression of long-branch attraction artefacts in the animal phylogeny using a site-heterogeneous model. BMC Evol Biol 7(Suppl 1):S4. https://doi.org/10.1186/1471-2148-7-s1-s4
Longo LM et al (2020) Primordial emergence of a nucleic acid-binding protein via phase separation and statistical ornithine-to-arginine conversion. Proc Natl Acad Sci USA 117:15731–15739. https://doi.org/10.1073/pnas.2001989117
Longo LM, Lee J, Blaber M (2013) Simplified protein design biased for prebiotic amino acids yields a foldable, halophilic protein. Proc Natl Acad Sci USA 110:2135–2139. https://doi.org/10.1073/pnas.1219530110
Lupas AN, Alva V (2017) Ribosomal proteins as documents of the transition from unstructured (poly)peptides to folded proteins. J Struct Biol 198:74–81. https://doi.org/10.1016/j.jsb.2017.04.007
Mat WK, Xue H, Wong JT (2008) The genomics of LUCA. Front Biosci 13:5605–5613. https://doi.org/10.2741/3103
Mayer-Bacon C, Freeland SJ (2021) A broader context for understanding amino acid alphabet optimality. J Theor Biol 520:110661. https://doi.org/10.1016/j.jtbi.2021.110661
McDonald GD, Storrie-Lombardi MC (2010) Biochemical constraints in a protobiotic earth devoid of basic amino acids: the “BAA(-) world.” Astrobiology 10(10):989–1000. https://doi.org/10.1089/ast.2010.0484
Merkl R, Sterner R (2016) Ancestral protein reconstruction: techniques and applications. Biol Chem 397:1–21. https://doi.org/10.1515/hsz-2015-0158
Miller SL (1953) A production of amino acids under possible primitive earth conditions. Science 117:528–529
Miller SL, Lazcano A (1995) The origin of life–did it occur at high temperatures? J Mol Evol 41:689–692
Muller MM et al (2013) Directed evolution of a model primordial enzyme provides insights into the development of the genetic code. PLoS Genet 9:e1003187. https://doi.org/10.1371/journal.pgen.1003187
Nakamura E et al (2022) On the origin and evolution of the asteroid Ryugu: a comprehensive geochemical perspective. Proc Jpn Acad Ser B 98:227–282. https://doi.org/10.2183/pjab.98.015
Nguyen LT, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Nisbet EG, Sleep NH (2001) The habitat and nature of early life. Nature 409:1083–1091. https://doi.org/10.1038/35059210
Nissen P, Hansen J, Ban N, Moore PB, Steitz TA (2000) The structural basis of ribosome activity in peptide bond synthesis. Science 289:920–930. https://doi.org/10.1126/science.289.5481.920
Ortlund EA, Bridgham JT, Redinbo MR, Thornton JW (2007) Crystal structure of an ancient protein: evolution by conformational epistasis. Science 317:1544–1548. https://doi.org/10.1126/science.1142819
Pace CN, Vajdos F, Fee L, Grimsley G, Gray T (1995) How to measure and predict the molar absorption coefficient of a protein. Protein Sci 4:2411–2423. https://doi.org/10.1002/pro.5560041120
Patel BH, Percivalle C, Ritson DJ, Duffy CD, Sutherland JD (2015) Common origins of RNA, protein and lipid precursors in a cyanosulfidic protometabolism. Nat Chem 7:301–307. https://doi.org/10.1038/nchem.2202
Pearce BKD, Pudritz RE, Semenov DA, Henning TK (2017) Origin of the RNA world: the fate of nucleobases in warm little ponds. Proc Natl Acad Sci USA 114:11327–11332. https://doi.org/10.1073/pnas.1710339114
Raymann K, Brochier-Armanet C, Gribaldo S (2015) The two-domain tree of life is linked to a new root for the Archaea. Proc Natl Acad Sci USA 112:6670–6675. https://doi.org/10.1073/pnas.1420858112
Rich A (1962) On the problems of evolution and biochemical information transfer. In: Kasha M, Pullman B (eds) Horizons in Biochemistry. Academic Press, NY, pp 103–126
Rinke C et al (2013) Insights into the phylogeny and coding potential of microbial dark matter. Nature 499:431–437. https://doi.org/10.1038/nature12352
Rivera MC, Lake JA (1992) Evidence that eukaryotes and eocyte prokaryotes are immediate relatives. Science 257:74–76
Robert F, Chaussidon M (2006) A palaeotemperature curve for the precambrian oceans based on silicon isotopes in cherts. Nature 443:969–972. https://doi.org/10.1038/nature05239
Rouet R et al (2017) Structural reconstruction of protein ancestry. Proc Natl Acad Sci USA 114:3897–3902. https://doi.org/10.1073/pnas.1613477114
Shibue R, Sasamoto T, Shimada M, Zhang B, Yamagishi A, Akanuma S (2018) Comprehensive reduction of amino acid set in a protein suggests the importance of prebiotic amino acids for stable proteins. Sci Rep 8:1227. https://doi.org/10.1038/s41598-018-19561-1
Shimojo M, Amikura K, Masuda K, Kanamori T, Ueda T, Shimizu Y (2020) In vitro reconstitution of functional small ribosomal subunit assembly for comprehensive analysis of ribosomal elements in E. coli. Commun Biol 3:142. https://doi.org/10.1038/s42003-020-0874-8
Solis AD (2019) Reduced alphabet of prebiotic amino acids optimally encodes the conformational space of diverse extant protein folds. BMC Evol Biol 19:158. https://doi.org/10.1186/s12862-019-1464-6
Thomson JM, Gaucher EA, Burgan MF, De Kee DW, Li T, Aris JP, Benner SA (2005) Resurrecting ancestral alcohol dehydrogenases from yeast. Nat Genet 37:630–635. https://doi.org/10.1038/ng1553
Thornton JW (2004) Resurrecting ancient genes: experimental analysis of extinct molecules. Nat Rev Genet 5:366–375. https://doi.org/10.1038/nrg1324
Trifonov EN (2000) Consensus temporal order of amino acids and evolution of the triplet code. Gene 261:139–151
Trudeau DL, Kaltenbach M, Tawfik DS (2016) On the potential origins of the high stability of reconstructed ancestral proteins. Mol Biol Evol 33:2633–2641. https://doi.org/10.1093/molbev/msw138
Vázquez-Salazar A, Lazcano A (2018) Early life: embracing the RNA world. Curr Biol 28:R220–R222. https://doi.org/10.1016/j.cub.2018.01.055
Weber AL, Miller SL (1981) Reasons for the occurrence of the twenty coded protein amino acids. J Mol Evol 17:273–284
Weiss MC, Sousa FL, Mrnjavac N, Neukirchen S, Roettger M, Nelson-Sathi S, Martin WF (2016) The physiology and habitat of the last universal common ancestor. Nat Microbiol 1:16116. https://doi.org/10.1038/nmicrobiol.2016.116
Wheeler LC, Lim SA, Marqusee S, Harms MJ (2016) The thermostability and specificity of ancient proteins. Curr Opin Struct Biol 38:37–43. https://doi.org/10.1016/j.sbi.2016.05.015
Wiener L, Schüler D, Brimacombe R (1988) Protein binding sites on E. coli 16S ribosomal RNA; RNA regions that are protected by proteins S7, S9 and S19, and by proteins S8, S15 and S17. Nucleic Acids Res 16:1233–1250. https://doi.org/10.1093/nar/16.4.1233
Williams PD, Pollock DD, Blackburne BP, Goldstein RA (2006) Assessing the accuracy of ancestral protein reconstruction methods. PLoS Comput Biol 2:e69. https://doi.org/10.1371/journal.pcbi.0020069
Williams TA, Foster PG, Cox CJ, Embley TM (2013) An archaeal origin of eukaryotes supports only two primary domains of life. Nature 504:231–236. https://doi.org/10.1038/nature12779
Woese CR, Fox GE (1977) The concept of cellular evolution. J Mol Evol 10:1–6
Woese CR, Kandler O, Wheelis ML (1990) Towards a natural system of organisms: proposal for the domains Archaea, Bacteria, and Eucarya. Proc Natl Acad Sci USA 87:4576–4579
Wolman Y, Haverland WJ, Miller SL (1972) Nonprotein amino acids from spark discharges and their comparison with the murchison meteorite amino acids. Proc Natl Acad Sci USA 69:809–811. https://doi.org/10.1073/pnas.69.4.809
Wong JT (1975) A co-evolution theory of the genetic code. Proc Natl Acad Sci USA 72:1909–1912
Yagi S et al (2021) Seven amino acid types suffice to create the core fold of RNA polymerase. J Am Chem Soc 143:15998–16006. https://doi.org/10.1021/jacs.1c05367
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591. https://doi.org/10.1093/molbev/msm088
Yang Z, Rannala B (1997) Bayesian phylogenetic inference using DNA sequences: a Markov Chain Monte Carlo method. Mol Biol Evol 14:717–724
Yang Z, Kumar S, Nei M (1995) A new method of inference of ancestral nucleotide and amino acid sequences. Genetics 141:1641–1650
Yutin N, Makarova KS, Mekhedov SL, Wolf YI, Koonin EV (2008) The deep archaeal roots of eukaryotes. Mol Biol Evol 25:1619–1630. https://doi.org/10.1093/molbev/msn108
Akanuma S, Yamagishi A (2016) A strategy for designing thermostable enzymes by reconstructing ancestral sequences possessed by ancient life. Biotechnology of Extremophiles: Advances and Challenges, (ed Rampelotto PH), Springer, cham
Acknowledgements
The authors are grateful to Dr. Ryutaro Furukawa for assisting the phylogenetic analysis and ancestral sequence inference. This work was supported by JSPS KAKENHI (Grant Number 21H01200) and the Astrobiology Center Program of the National Institutes of Natural Sciences (Grant Number AB031007).
Funding
Japan Society for the Promotion of Science,21H01200,Satoshi Akanuma,National Institutes of Natural Sciences,AB031007,Satoshi Akanuma
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Belinda Chang .
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Springer Nature or its licensor (e.g. a society or other partner) holds exclusive rights to this article under a publishing agreement with the author(s) or other rightsholder(s); author self-archiving of the accepted manuscript version of this article is solely governed by the terms of such publishing agreement and applicable law.
About this article
Cite this article
Zhao, F., Akanuma, S. Ancestral Sequence Reconstruction of the Ribosomal Protein uS8 and Reduction of Amino Acid Usage to a Smaller Alphabet. J Mol Evol 91, 10–23 (2023). https://doi.org/10.1007/s00239-022-10078-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-022-10078-w