Abstract
TOPIIA topoisomerases are required for the regulation of DNA topology by DNA cleavage and re-ligation and are important targets of antibiotic and anticancer agents. Humans possess two TOPIIA paralogue genes (TOP2A and TOP2B) with high sequence and structural similarity but distinct cellular functions. Despite their functional and clinical relevance, the evolutionary history of TOPIIA is still poorly understood. Here we show that TOPIIA is highly conserved in Metazoa. We also found that TOPIIA paralogues from jawed and jawless vertebrates had different origins related with tetraploidization events. After duplication, TOP2B evolved under a stronger purifying selection than TOP2A, perhaps promoted by the more specialized role of TOP2B in postmitotic cells. We also detected genetic signatures of positive selection in the highly variable C-terminal domain (CTD), possibly associated with adaptation to cellular interactions. By comparing TOPIIA from modern and archaic humans, we found two amino acid substitutions in the TOP2A CTD, suggesting that TOP2A may have contributed to the evolution of present-day humans, as proposed for other cell cycle-related genes. Finally, we identified six residues conferring resistance to chemotherapy differing between TOP2A and TOP2B. These six residues could be targets for the development of TOP2A-specific inhibitors that would avoid the side effects caused by inhibiting TOP2B. Altogether, our findings clarify the origin, diversification and selection pressures governing the evolution of animal TOPIIA.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
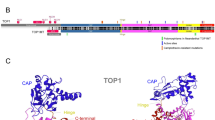
Topoisomerases are important enzymes that support cell viability and chromosome topology. They have the ability to cut, shuffle and reconnect DNA strands by adding or removing DNA supercoils and disentangle DNA segments (Champoux 2001; Wang 1996). Type IIA topoisomerases transiently cleave two strands of DNA and include bacterial and archaeal gyrase, bacterial topoisomerase IV and eukaryotic topoisomerase II. Some viruses also encode for type II topoisomerases, such as some Nucleocytoplasmic Large DNA Viruses (NCLDV) and T4-like bacterioviruses (Forterre et al. 2007; Gadelle et al. 2003; Schoeffler and Berger 2008). Type II topoisomerases have been identified in early diverging lineages of eukaryotes, such as kinetoplastid protozoans, Giardia lamblia and Plasmodium falciparum (Chakraborty and Majumder 1987; Cheesman et al. 1994; De et al. 2005; Strauss and Wang 1990). Previous studies have shown that most eukaryotes have a single type IIA topoisomerase (TOPIIA), with the notable exception of vertebrates that have two paralogues, topoisomerase IIα (TOP2A) and topoisomerase IIβ (TOP2B) (Austin and Fisher 1990; Drake et al. 1987). In humans, TOP2A is encoded by the TOP2A gene on chromosome 17q21-22 (Tsai-Pflugfelder et al. 1988) and TOP2B by TOP2B gene on chromosome 3p24 (Austin et al. 1993; Jenkins et al. 1992; Tan et al. 1992). Both proteins display similar structures (Fig. 1) and biochemical activities but have different biological roles (Cornarotti et al. 1996; Drake et al. 1989; Leontiou et al. 2003; Marsh et al. 1996; Wang 2002).
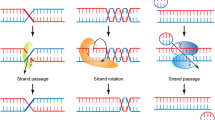
TOPIIA activity relies on a mechanism that involves the controlled association and dissociation of three subunit-dimerization interfaces, or ‘gates’, termed the N-gate, DNA-gate and C-gate, which guide the physical movement of one DNA duplex through another (Berger et al. 1996; Cabral et al. 1997; Roca et al. 1996; Roca and Wang 1992, 1994; Wigley et al. 1991). TOPIIA can also be described as including three structural domains (Fig. 1): an N-terminal ATPase domain (NTD), a central catalytic DNA-binding/cleavage domain (CD) and a C-terminal domain (CTD). The N-terminal gate (ATPase domain) is composed by two elements: an ATP binding domain of the GHKL superfamily of proteins, including the Bergerat fold (Dutta and Inouye 2000), and an adjacent domain called the transducer (Classen et al. 2003; Corbett and Berger 2003; Wigley et al. 1991) that is thought to transmit signals (Corbett and Berger 2003; Kingma et al. 2000) from the N-gate to the DNA-gate (where DNA is bound and cleaved). The DNA-gate is formed by a divalent metal-binding TOPRIM domain and a WHD similar to that typified by the catabolite activation protein (Aravind et al. 1998; Berger et al. 1996; Gajiwala and Burley 2000; McKay and Steitz 1981). The WHD contains the catalytic tyrosine and cooperates with the TOPRIM fold to cleave DNA, generating a pair of 5′cuts staggered 4 bp apart from each other on opposite strands (Liu et al. 1983; Morrison and Cozzarelli 1979; Sander and Hsieh 1983). Adjacent to the WHD is a fold termed the “shoulder” or “tower” (Cabral et al. 1997), which also participates in DNA binding (Dong and Berger 2007). The third dimerization interface, the C-gate, is a coiled-coil element capped at its distal end with a small globular domain that extends from the Tower. This interface is well conserved and serves as the primary dimer interface of the protein (Berger et al. 1996; Cabral et al. 1997; Corbett et al. 2005; Fass et al. 1999; Laponogov et al. 2007).
TOP2A and TOP2B have very similar structures resulting from a high degree of sequence homology (~ 70 to 80%) (Austin et al. 2018; Austin et al. 1993; Jenkins et al. 1992). The main differences reside in the C-terminal region, which is responsible for their different biological roles (Kozuki et al. 2017; Linka et al. 2007). The complete CTD crystal structure has not been determined, but secondary structure prediction suggests that it is structurally disordered (Broeck et al. 2021). The TOP2A CTD seems to act in the preferential relaxation of positive supercoils, whereas the equivalent region of TOP2B does not reveal a supercoil preference (McClendon et al. 2005). The C-terminal region contains nuclear localization signals and undergo extensive post-translational modifications (Lane et al. 2013; Lotz and Lamour 2020).
TOP2A is highly expressed during mitosis, being essential in proliferating cells and assists in chromosome segregation and replication (Akimitsu et al. 2003; Ali and Abd Hamid 2016; Cuvier and Hirano 2003; Grue et al. 1998; Niimi et al. 2001; Ye et al. 2010). On the other hand, TOP2B appears dispensable for cell proliferation, but regulates gene expression and is associated with developmental and differentiation events, in particular nerve growth and brain development (Austin et al. 2018; Bollimpelli et al. 2017; Ju et al. 2006; Lyu et al. 2006; Lyu and Wang 2003; McNamara et al. 2008; Tiwari et al. 2012; Yang et al. 2000). Mutations in TOP2B have been associated with B-cell immunodeficiency (Broderick et al. 2019; Papapietro et al. 2020), hearing loss (Xia et al. 2019) and intellectual disability (Lam et al. 2017). For instance, mice lacking TOP2A fail to develop beyond the 4–8-cell stage, whilst those without TOP2B exhibit a perinatal death due to defects in neuronal development (Akimitsu et al. 2003; Lyu and Wang 2003; Yang et al. 2000).
In addition to their vital cellular functions, TOPIIA are a target for some of the most active anticancer agents (Nitiss 2009). TOP2A is responsible for the anticancer effects of TOP2 inhibitors due to its high activity in proliferating cells. TOP2 poisons (e.g. etoposide, doxorubicin) increase the levels of TOP2–DNA covalent complexes, resulting in double-strand DNA breaks that can cause cell death. TOP2B is believed to be responsible for most of the secondary malignancies and cardiotoxicity caused by TOP2-targeting drugs due to its similar structure to TOP2A (Azarova et al. 2007; Chen et al. 2012; Haffner et al. 2010). Moreover, several mutations in TOP2A confer resistance to anticancer drugs (Nitiss and Beck 1996; Wu et al. 2011).
TOPIIA structure and function have been well studied in model organisms and humans, but often without including a comprehensive evolutionary analysis. Here, we address this limitation by providing a detailed evolutionary study of TOPIIA in animals. We provide a comprehensive phylogeny of TOPIIA in animals, including a detailed view of the duplication event that originated the TOP2A and TOP2B paralogues. We also identified the most conserved protein domains of functional relevance and assessed selective pressures governing the evolution of these important topoisomerases.
Materials and Methods
TOPIIA Sequences
We obtained TOPIIA protein sequences using the Ortho DB v10 (https://www.orthodb.org), which is a comprehensive catalogue of putative orthologues from more than 400 metazoan species (Kriventseva et al. 2019). We also used the protein–protein BLAST (blastp) to retrieve sequences from phyla that were not found in the Ortho DB, by using as query TOPIIA sequences from close phylogenetic groups. We excluded repeated sequences from the same species that showed 100% identity, which most likely represented different entries of the same sequence in the databases. We also removed short sequences with less than half of the average of TOPIIA length from further analyses as they can represent partial protein sequences derived from poorly assembled genomes. In some cases, contigs do not cover the complete genomic region where the protein is encoded, resulting in partial protein sequences. By this reason, some species lack one of the paralogues in our dataset, although it may be present in their genomes.
Denisovan and Neanderthal TOP2A and TOP2B sequences were downloaded from the UCSC Genome Browser (http://genome.ucsc.edu/) (Kent et al. 2002). We downloaded all BAM reads for tracks Denisova and Neanderthal Cntgs matching the Human Mar. 2006 (NCBI36/hg18) chr17:35,798,322-35,827,695 (TOP2A) and chr3:25,614,479-25,680,835 (TOP2B). The BAM reads from each track were then reassembled against the human TOP2A (NC_000017.11) and TOP2B (NC_000003.12) reference sequences using Geneious v2020.2.4 (http://www.geneious.com). We only considered a polymorphic position in archaic hominids when: (1) at least two reads overlap in that position; (2) the polymorphism represents more than 75% of all the reads and (3) the polymorphism is not at the end of a read.
TOPIIA Multiple Sequence Alignments
We aligned the TOPIIA protein sequences with MAFFT version 7 (Katoh et al. 2019). The following three multiple alignments of protein sequences were used in subsequent analyses: Metazoa (n = 389), Chordata TOP2A (n = 105) and Chordata TOP2B (n = 125). The conservation across the alignments was estimated using the percentage of pairwise identity (PI) calculated in the Geneious program. Note that PI constitutes the average percent identity calculated by comparing the base pairs at every site.
TOPIIA coding domain sequences (CDS) from chordates were obtained from the Ensembl Genome Server (Hunt et al. 2018), as multiple sequence alignments of TOP2A (ENSG00000131747) and TOP2B (ENSG00000077097) human orthologues. The orthologues were organized in four different alignments for either TOP2A or TOP2B: Chordata (n = 159), Actinopteri (n = 51), Aves (n = 11) and Mammalia (n = 86). For coherence, we included the same species in the alignments of TOP2A and TOP2B.
The Ensembl server includes two long transcripts for TOP2B: TOP2B-201 (ENST00000264331.9) with 5814 nucleotides and 1626 amino acids and TOP2B-204 (ENST00000435706.6) with 5389 nucleotides and 1621 amino acids. Here we used the longest transcript (TOP2B-201) and resulting protein sequence unless stated otherwise.
The sequence alignments are available at Mendeley Data (https://data.mendeley.com//datasets/h2xfj5fsxw/1).
Phylogenetic Analyses
The phylogenetic tree with all metazoan species (n = 389), considering Arabidopsis thaliana as outgroup, was obtained from the corresponding multiple alignment of protein sequences. We reconstructed a maximum likelihood (ML) phylogenetic tree with PhyML 3.0 (Guindon et al. 2010), implemented in the ATGC bioinformatics platform (http://www.atgc-montpellier.fr). The JTT+G+I substitution model of protein evolution was selected with the Smart Model Selection (SMS) v1.8.4 method implemented in PhyML (Lefort et al. 2017), under the Akaike Information Criterion (AIC). The branch support was evaluated with aBayes method (Anisimova et al. 2011). We analysed the duplication events in chordates with a phylogenetic tree based on the alignment of 34 CDS from Cephalochordata, Tunicata and Vertebrata species, considering Acanthaster planci (Echinodermata) as outgroup. Again, the ML tree was reconstructed with the ATGC bioinformatics platform, under the GTR+G+I substitution model and bootstrap based on 100 replicates. The resulting phylogenetic trees were edited with FigTree v1.4.3 (http://tree.bio.ed.ac.uk/software/figtree) and TreeViewer v1.2.2 (https://treeviewer.org).
Evaluation of Selection
We evaluated molecular adaptation in TOP2A and TOP2B protein-coding sequence alignments with the nonsynonymous/synonymous substitution rate ratio (dN/dS) (Del Amparo et al. 2021; Jeffares et al. 2015). We started by identifying the best-fitting substitution model of DNA evolution with jModelTest2 (Darriba et al. 2012) and reconstructed a ML phylogenetic tree under the selected substitution model. Next, we estimated dN/dS under a ML method considering the reconstructed phylogenetic tree with the well-established evolutionary framework Hyphy (Kosakovsky Pond and Frost 2005; Kosakovsky Pond et al. 2020). In particular, we applied the single-likelihood ancestor counting (SLAC) method, which provides dN/dS estimation with accuracy similar to that obtained with other likelihood-based methods (Kosakovsky Pond and Frost 2005). We identified global (entire sequence) genetic signatures of selection and positively selected sites (PSSs). The difference dN − dS was also used to evaluate selection at the site codon level.
TOPIIA Protein Structure
We used the TOP2A domains and other regions previously described (Broeck et al. 2021) and we considered them also in TOP2B by aligning the human reference sequences of both proteins. The TOP2A protein structure with PDB (Protein Data Bank) (Berman et al. 2000) code 6ZY7 (Broeck et al. 2021) and TOP2B structure with code 5ZAD (Sun et al. 2018) were obtained with Mol* (Sehnal et al. 2021) and RCSB PDB. The structures were coloured according to the protein domains and main regions.
Results and Discussion
TOPIIA Proteins are Conserved Across Metazoa
The phylogenetic tree built with TOPIIA protein sequences placed Cnidaria at the root of Metazoa, which was expected considering that it was the only phylum in our dataset not belonging to Bilateria (Fig. 2, Supplementary Fig. S1). In addition to Cnidaria, our dataset included eight Protostomia and 13 Deuterostomia phyla. Our phylogeny supports the split of Protostomia in Ecdysozoa (those exhibiting moulting) and Spiralia or Lophotrochozoa (those having lophophores and trochophore larvae), although only Spiralia was retrieved as a monophyletic group. Within Ecdysozoa, our results did not support the existence of Cycloneuralia, a clade including Scalidophora (represented here by Priapulida) and Nematoida (represented here by Nematoda) (Dunn et al. 2008). Indeed, our phylogenetic tree placed Nematoida more related to Panarthropoda (represented here by Tardigrada and Arthropoda) than to Scalidophora (Priapulida) (Campbell et al. 2011; Pisani et al. 2013). Spiralia formed a well-supported monophyletic group including Annelida, Mollusca, Brachiopoda and Platyhelminthes. The relationships within Spiralia are poorly resolved and often a matter of debate (Dunn et al. 2014). The clustering of Brachiopoda with Platyhelminthes supports the hypothesis that Lophophorata, organisms with a rake-like feeding structure (represented here by Brachiopoda), forms a separate clade from Trochozoa, a group defined by trochophore larvae, including at least Annelida and Mollusca (Dunn et al. 2014; Nesnidal et al. 2013) (Fig. 2).
Phylogenetic analysis of TOPIIA in Metazoa. Maximum likelihood (ML) phylogenetic tree built with an alignment of 389 TOPIIA protein sequences from metazoans and considering Arabidopsis thaliana as outgroup. The branch support was estimated with aBayes, shown on the internal nodes. The scale bar indicates substitutions per site. Major monophyletic clades are collapsed for visualization purposes. The complete tree can be accessed in the supplementary material
The Deuterostomia clade includes a well-supported monophyletic group with Hemichordata and Echinodermata, which together form Ambulacraria. Within the Chordata, cephalochordates and tunicates diverged first and vertebrates formed a monophyletic group with the two paralogues TOP2A and TOP2B in different branches. In order to have a better resolution within Chordata, we built a phylogeny with all available TOPIIA CDS sequences from basal chordates (Fig. 3A). In the CDS tree, cephalochordates diverged first, with urochordates and vertebrates forming a sister group known as Olfactores, an arrangement that has now been widely accepted (Delsuc et al. 2006; Putnam et al. 2008; Satoh et al. 2014). We found particularly long branches in tunicates that suggests a high rate of molecular evolution, as previously noted for other genes (Delsuc et al. 2006; Tsagkogeorga et al. 2010). In particular, the extremely long branch of Oikopleura dioica (Fig. 3A) supports the claim that it is the fastest evolving metazoan recorded so far (Berna and Alvarez-Valin 2014; Denoeud et al. 2010).
Evolutionary history of TOPIIA in basal chordates. A Maximum likelihood (ML) phylogenetic tree built with an alignment of 34 TOPIIA coding domain sequences (CDS) from Cephalochordata, Tunicata and Vertebrata, considering Acanthaster planci (Echinodermata) as outgroup. The branch support was estimated with 100 bootstrap cycles, shown on the internal nodes. The scale bar indicates substitutions per site. The putative occurrence of two rounds of tetraploidization (1R and 2R) is indicated. B Pairwise identity between Petromyzon marinus and Homo sapiens paralogues. C Hypothetical succession of events to explain the presence of duplicated genes in Cyclostomata and Gnathostomata. The Cyclostomata paralogues originated during the 1R event, the Gnathostomata lost one of the duplicated genes from 1R and the TOP2A and TOP2B paralogues originated in the 2R event
TOPIIA Genes Duplicated Independently in Cyclostomata and Gnathostomata
We extensively searched for TOPIIA sequences in all available chordate genomes, and only found cases of TOPIIA paralogue genes within vertebrates. It is therefore evident that the formation of TOPIIA paralogues is related with vertebrate-specific genome duplication events. There is now convincing evidence that early vertebrate evolution is characterized by two rounds of tetraploidization (known as 1R and 2R), whose timing is still a topic of debate (Ohno 2013; Smith and Keinath 2015; Van de Peer et al. 2009). Irrespectively of when tetraploidization events occurred, our analyses suggest that the formation of paralogues in jawless vertebrates (Cyclostomata) and in jawed vertebrates (Gnathostomata) were independent events. Although with weak bootstrap values (~ 60%), our analyses always placed Cyclostomata paralogues in a different group from Gnathostomata TOP2A and TOP2B paralogues (Fig. 3A). In fact, Cyclostomata TOPIIA paralogues cannot even be classified as TOP2A or TOP2B, as they equally diverged from Gnathostomata paralogues, as shown by the similar pairwise sequence identities between Petromyzon marinus and Homo sapiens paralogues (Fig. 3B).
There is a growing consensus that all vertebrates share the first tetraploidization event (1R), but only jawed vertebrates had a second whole genome duplication (2R) (Aase-Remedios and Ferrier 2021; Nakatani et al. 2021; Simakov et al. 2020), following previous studies (Escriva et al. 2002; Stadler et al. 2004). If that is the case, the most parsimonious succession of events to explain TOPIIA paralogues was as follows: (a) Cyclostomata paralogues originated during the 1R event; (b) Gnathostomata lost one of the duplicated genes from 1R and (c) Gnathostomata TOP2A and TOP2B originated in the 2R event (Fig. 3C). This scenario only assumes a single gene loss to explain the observed phylogeny. Other scenarios are less parsimonious by requiring two or more gene loss/duplication events (Supplementary Fig. S2). For example, if 1R duplicated genes were retained in Gnathostomata at the time of 2R, two of the four resulting copies had to be subsequently lost. Nonetheless, further studies on early vertebrates will better elucidate these evolutionary events.
Non-synonymous Changes in TOP2A and TOP2B Amongst Modern and Archaic Humans
Neanderthals and Denisovans are the closest relatives to modern humans, diverging from the modern human lineage early in the Middle Pleistocene (Green et al. 2010; Reich et al. 2010). The recent shared ancestry explains the low genetic divergence amongst modern and archaic humans. We identified four polymorphisms amongst human, Denisovan and Neanderthal TOPIIA coding sequences (Table 1). The polymorphic positions were observed in the Transducer region of the ATPase domain (n = 1) and CTD (n = 3). The mutation in the TOP2A Transducer (position 267) occurred in the Denisovan lineage (A>G) without replacing the amino acid, perhaps the only possible type of mutation considering that this site is invariable in the 389 species analysed here. Two missense mutations were found in the TOP2A CTD. A T>C mutation in the human lineage replaced aspartate by glycine at position 1386. A C>A mutation in the Neanderthal lineage changed alanine by serine in position 1515 (Table 1). In both cases, the replacement was between amino acids with different charges or polarities. Such replacements may not affect the protein structure as the CTD is believed to be disordered. However, it is possible that such substitutions may impact the CTD interactions with other cellular components, raising the possibility that it may have contributed to the evolution of present-day humans. For example, position 1515 is within the chromatin tether domain (ChT), shown to be essential for TOP2A to interact robustly with chromosomes in mitosis (Lane et al. 2013). The only polymorphism detected in TOP2B occurs also in the CTD in the human lineage without replacing the amino acid. We found that all the polymorphic positions in the CTD are not conserved (< 50% pairwise identity), as expected considering the high variability of this protein region.
Very few amino acid changes become fixed in modern humans when comparing with archaic humans. For example, only 78 of those substitutions were described in the original publication of the Neandertal genome (Green et al. 2010). A recent survey, using data from different Neanderthal and Denisovan genomes, identified 571 genes with human-specific amino acid-changes (Kuhlwilm and Boeckx 2019). In fact, human TOP2A was identified as the protein with the largest number of interactions with other proteins (n = 53), suggesting that it may operate as an interaction hub in modifications of the cell division complex (Kuhlwilm and Boeckx 2019). Experimental studies exploring the influence of these changes on the cell cycle machinery could evaluate this intriguing hypothesis.
TOP2A and TOP2B Evolved Under Strong Purifying Selection But a Few Sites were Positively Selected
We tested for the strength and mode of selection acting on TOP2A and TOP2B using dN/dS in chordates (Table 2). We found molecular signatures of purifying selection in both genes (dN/dS < 0.3), indicative of a strong selective pressure to conserve both TOP2A and TOP2B. It has been suggested that paralogues are subject to weaker purifying selection than single-copy genes (Kondrashov et al. 2002; Scannell and Wolfe 2008). In this concern, dN/dS values for TOP2A and TOP2B are higher than those estimated for Topoisomerase III Beta (TOP3B) in chordates (dN/dS < 0.1) (Moreira et al. 2021), which suggests that functional constraints were relaxed during the functional divergence to TOP2A and TOP2B. Moreover, the presence of (at least partially) redundant gene copies may have permitted the accumulation of previously forbidden deleterious mutations, which can explain the higher dN/dS values.
Paralogues may exhibit asymmetric rates of sequence evolution (Conant and Wagner 2003; Scannell and Wolfe 2008; Van de Peer et al. 2001). The strength of purifying selection was stronger in TOP2B (i.e. dN/dS = 0.156 on Chordata) than in TOP2A (e.g. dN/dS = 0.238 on Chordata), which suggests that TOP2B is under stronger functional constraints. Moreover, TOP2A displayed a higher nucleotide diversity and substitution rate than TOP2B (Table 2), which can also be noted in its longer branches in the reconstructed phylogenetic trees (Figs. 2, 3A). The specific activity of TOP2B in nerve growth and brain development (Lyu and Wang 2003; Yang et al. 2000) could impose relevant constrains on molecular evolution by the need of interacting with different partners and chemical environments. For example, it has been suggested that the role of TOP2B in the organism development involves the activation and repression of specific developmental genes (e.g. Myt1l, Cacna2d1, Syt1, Kcnd2) in association with diverse proteins (Lyu et al. 2006; Lyu and Wang 2003; Tiwari et al. 2012).
We detected signatures of positive selection in three TOP2A and two TOP2B sites in mammals (Table 2). Three of these positively selected sites (PSSs) are placed in the CTD, which is believed to conform specificity to the different activities of TOP2A and TOP2B (Kozuki et al. 2017; Linka et al. 2007). The TOP2A position 928 in the Tower domain was also found under positive selection (dN − dS = 4.832). The highest dN − dS values were obtained for the TOP2B position 28 in the ATPase domain in both the Chordata and Mammalia datasets (Table 2). The ATPase domain binds to ATP for a nucleotide-actuated protein dimerization gate through which DNA duplexes are passed. Considering the domains where the PSSs are located, we believe that these sites accumulated nonsynonymous substitutions over time to improve TOP2A and TOP2B interaction and cellular functions, although it should be evaluated with experimental studies.
Low Conservation in Putative Regulatory Regions of the C-terminal Domain (CTD)
Human TOP2A and TOP2B proteins had an overall pairwise identity of 66.6% (Fig. 4A, B). The DNA Binding/Cleavage domain was slightly more conserved (80.2%) than the ATPase domain (77.3%), whilst the paralogues mainly diverged in the CTD (28.8%). In this concern, the alignment of TOPIIA sequences of metazoans clearly showed the contrast between the poorly conservated CTD and the other well-conserved protein domains (Fig. 4C). The most conserved regions were the WHD (84.8%) and TOPRIM (76.9%) domains (Fig. 4D), where conservation can be explained by their critical role in interacting with DNA (Aravind et al. 1998; Gajiwala and Burley 2000; McKay and Steitz 1981; Roca et al. 1996). The same domains stand out as the most conserved when comparing TOP2A with TOP2B (Fig. 4D). All the other TOPIIA domains displayed a pairwise identity of around 50–70%, excepting the CTD linker (32.9%) and CTD (21.9%), which are almost impossible to align when using all metazoans. The CTD regulates nuclear localization and protein–protein interactions, which could have differentially evolved amongst species. Moreover, the CTD is believed to be a disordered region, which is known to evolve faster than well-structured regions (Brown et al. 2011). Overall, we found that TOP2B domains are more conserved than TOP2A domains (Fig. 4D), in agreement with the strong purifying selection observed in TOP2B (Table 2). We found poor conservation in the ending CTD 30 amino acids (positions 1502–1531) that constitute the ChT domain (Lane et al. 2013). The ChT domain had a 24.8% pairwise identity in Metazoa whilst a 55.7% in TOP2A Chordata. Similarly, the bipartite nuclear localisation signal (NLS) near the TOP2A CTD end (positions 1454–1497) (Mirski et al. 1997) was found poorly conserved in Metazoa (24.9%) and Chordata TOP2A (49.9%).
TOPIIA diversity and structural organization. A Identity plot for the alignments of human TOP2A and TOP2B reference sequences. The identical positions are shown in green bars and the different positions are shown in yellow bars. Highlighted are the positions polymorphic in Denisovan or Neanderthal (green bars), associated with resistance to anticancer drugs (red bars) and positively selected (grey bars). TOP2B mutations resulting in disease are indicated by black bars. B Percentage of pairwise identity between human TOP2A and TOP2B protein complete sequence and domains. C Identity plot for the alignment of 389 TOPIIA protein sequences from metazoan species. The most conserved positions are indicated with brown bars, the less conserved with red bars. The main protein domains are included. D TOPIIA conservation across Metazoa. The percentage of pairwise identity was calculated for the full TOPIIA protein and domains in three different alignments, TOP2A (chordates), TOP2B (chordates) and all metazoans (Color figure online)
High Conservation of the Linker Connecting the ATPase and the TOPRIM Domains
We found that the linker connecting the ATPase and the TOPRIM domains (Fig. 5A) is well conserved in both TOP2A (92.7%) and TOP2B (96.4%), but more variable (65.4%) when all metazoans are compared (Fig. 4D). Two insertions of several amino acids are observed in Trichuris suis and Habropoda laboriosa. This linker forms an alpha helix with 29 amino acids connecting the N-gate to the DNA-gate (Figs. 1, 5B). Broeck et al. (2021) identified four highly conserved residues (positions 414, 417, 418 and 425) in this linker and tested different mutants to assess their contribution to the allosteric regulation of the human TOP2A. Our dataset of metazoan TOPIIA sequences confirmed that the positions 414, 417 and 418 were highly conserved (> 93%), but the position 425 showed a moderate level of conservation (70.8%; Fig. 5C). Indeed, positions 431 (99%), 409 (92.5%), 419 (84.5%) and 407 (82.4%) were also conserved, suggesting that they may play an important role in connections between the ATPase and TOPRIM domains. These residues could be tested in future experiments on allosteric regulation of TOP2A.
Structure and diversity of TOPIIA linker regions. A Identity plot and sequence logo for the linker joining the N-gate to DNA-gate. The most conserved positions are indicated by brown bars and the less conserved by red bars. B Cartoon representation of the linker region in the human TOP2A structure. C Percentage of pairwise identity per site for the linker region obtained from the alignment of 389 animal species. From D to F, we show the same information as from A to C but for the C-terminal domain (CTD) linker (Color figure online)
It has been suggested that the CTD linker (Fig. 5D, E) can structurally favour the curvature of the G-segment, stimulating DNA cleavage and facilitating strand passage (Broeck et al. 2021). Our analyses show that the CTD linker is poorly conserved in Metazoa, with several insertions and deletions (Fig. 4C, D). A better conservation was observed in Chordata TOP2A and TOP2B, but still much lower than other domains. In the Metazoa alignment, only the CTD linker region near the CTD is relatively conserved, with a few sites showing a pairwise identity above 75% (Fig. 5F). A few lysines (K) stand out as relatively conserved (Fig. 5F). The TOP2A position 1213 is phosphorylated during mitosis and contributes to localization of the protein to the centromere (Ishida et al. 2001). This position is moderately conserved in Metazoa (57.1%) and Chordata TOP2A (68.5%), but completely conserved in TOP2B (100%), which suggests that it is fundamental for the localization of TOP2B.
TOP2B Mutations Associated with Disease Occur in Conserved Sites and Replace Amino Acids with Different Physicochemical Properties
Alterations in topoisomerases have been associated with neurodegenerative and immune disorders and cancer (Pommier et al. 2016). TOP2A is essential for life; therefore, mutations that significantly affect its activity in relaxing topological stress are expected to be lethal. In fact, human disorders caused by mutations in TOP2A are rare. For example, the Human Gene Mutation Database (http://www.hgmd.cf.ac.uk/) only reports a gross deletion associated with congenital heart disease (Glessner et al. 2014). On the other hand, TOP2B is not embryonic lethal and acts particularly in postmitotic cells. Perhaps because of this, a few cases of inherited TOP2B mutations associated with diseases have been reported (Broderick et al. 2019; Erdős et al. 2021; Lam et al. 2017; Papapietro et al. 2020; Xia et al. 2019). We identified six TOP2B mutations in the literature related with impaired B-cell development and function, hearing loss or neurodevelopmental disease (Table 3). A mutation at position 63 replaced a histidine by tyrosine in the Bergerat fold of the ATP binding domain, in an invariable site across Chordates. The mutation replaces a positively charged (histidine) by a neutral (tyrosine) amino acid. A total of four mutations were observed in the TOPRIM domain, all of them in highly conserved residues (PI > 95.2%). The heterozygous mutations affecting the TOPRIM were shown as partially dominant loss-of-function mutations (Broderick et al. 2019) and affect essential sites for the catalytic activity of the TOP2B. For example, the alanine to proline replacement at position 490 is predicted to destabilize an alfa helix within the TOPRIM domain (Papapietro et al. 2020). The Ser>Leu and Gly>Ser replacements change polar and nonpolar amino acids.
A single mutation (position 1618) was described in the TOP2B CTD (Table 3). The residue is poorly conserved (PI of 54.9%). The mutation occurs near the end of the TOP2B sequence in a region that is homologous to the TOP2A ChT domain (Lane et al. 2013) that facilitates stable binding to chromatin. Still, it remains to be determined if TOP2B presents a similar domain and if the identified mutation could affect its activity. Overall, TOP2B replacement mutations resulting in disease are found at conserved amino acid positions, a pattern also observed in other genes (Miller and Kumar 2001). The high conservation of several TOP2B residues suggests that other undetected mutations could cause similar diseases.
Six Residues Conferring Resistance to TOP2 Poisons Differ Amongst TOP2A and TOP2B, and Can be Used to Develop Paralogue-Specific Drugs
TOPIIA is a molecular target of several important classes of anticancer drugs, whose efficiency can be affected by mutations in critical protein sites (Delgado et al. 2018; Nitiss 2009). We analysed 27 amino acid replacements previously shown to confer resistance to anticancer drugs (Beck et al. 1993; Gilroy et al. 2006; Leontiou et al. 2006; Leontiou et al. 2007; Vassetzky et al. 1995), all of them located in the DNA Binding/Cleavage domain (Table 4). A total of six out of the 27 sites represented different amino acids in TOP2A and TOP2B sequences (positions 450, 480, 762, 763, 908 and 909 in TOP2A). These six residues are amongst the most variable sites in Chordata TOP2A. However, we did not find the same pattern in TOP2B, with only the position 929 being variable amongst chordates. TOP2B is believed to be responsible for undesirable side effects of anticancer chemotherapy by leading to therapy-related leukaemia (Azarova et al. 2007; Cowell et al. 2012). Therefore, we believe that future works could explore these six variable sites to design TOP2A-specific anticancer drugs with less undesirable side effects caused by interfering with TOP2B (Wu et al. 2013). With the exceptions mentioned above, positions conferring resistance to chemotherapy were well conserved (Table 4). This pattern is expected because target sites of TOP2 poisons should disrupt functionally relevant protein sites, which therefore are under strong negative selection. Nevertheless, such sites can vary in cases of resistance to TOP2 poisons and still allow functional topoisomerases, perhaps only possible in the specific environment of cancer cells under different selective pressures.
Conclusion
Our results suggest that the long-term evolution of TOPIIA is primarily driven by strong purifying selection, which also explains the high levels of sequence conservation. TOP2B is under stronger selective constraints than TOP2A, which may be explained by the specialized role of TOP2B in the genetic programming of postmitotic cells that impose additional constrains to its evolution. The TOPIIA phylogeny lead us to conclude that Cyclostomata TOPIIA paralogues have evolved independently from jawed vertebrates. Therefore, jawless vertebrates are a good model to uncover the role of additional TOPIIA genes in vertebrate evolution. Our study also identified two missense mutations in the TOP2A CTD when comparing modern and archaic humans that may have contributed to the evolution of human-specific features. We found that almost all mutations related with resistance to chemotherapy or causing diseases occur in conserved sites of the TOP2B ATPase and DNA Binding/Cleavage domains, including their linker region. Therefore, we recommend that these domains should be included in the screening of undiagnosed diseases, particularly considering the multiple roles of TOP2B in cells. Similarly, we provide a list of residues that could be a good target to design TOP2A-specific anticancer drugs that would avoid the undesirable side effects caused by interfering with TOP2B. Overall, our study provides important insights into the evolution of TOPIIA in animals and represents a valuable resource for future functional studies of topoisomerases.
Data Availability
The sequence alignments are available at Mendeley Data, https://data.mendeley.com//datasets/h2xfj5fsxw/1 NB. The dataset has been published under embargo, so the files will become available after publication.
Code Availability
Not applicable.
References
Aase-Remedios ME, Ferrier DEK (2021) Improved understanding of the role of gene and genome duplications in chordate evolution with new genome and transcriptome sequences. Front Ecol Evol 9:429
Akimitsu N et al (2003) Enforced cytokinesis without complete nuclear division in embryonic cells depleting the activity of DNA topoisomerase IIα. Genes Cells 8:393–402
Ali Y, Abd Hamid S (2016) Human topoisomerase II alpha as a prognostic biomarker in cancer chemotherapy. Tumor Biol 37:47–55
Anisimova M, Gil M, Dufayard J-F, Dessimoz C, Gascuel O (2011) Survey of branch support methods demonstrates accuracy, power, and robustness of fast likelihood-based approximation schemes. Syst Biol 60:685–699
Aravind L, Leipe DD, Koonin EV (1998) Toprim—a conserved catalytic domain in type IA and II topoisomerases, DnaG-type primases, OLD family nucleases and RecR proteins. Nucleic Acids Res 26:4205–4213
Austin CA, Fisher LM (1990) Isolation and characterization of a human cDNA clone encoding a novel DNA topoisomerase II homologue from HeLa cells. FEBS Lett 266:115–117
Austin CA, Sng J-H, Patel S, Fisher LM (1993) Novel HeLa topoisomerase II is the IIβ isoform: complete coding sequence and homology with other type II topoisomerases. Biochem Biophys Acta 1172:283–291
Austin CA et al (2018) TOP2B: the first thirty years. Int J Mol Sci 19:2765
Azarova AM, Lyu YL, Lin C-P, Tsai Y-C, Lau JY-N, Wang JC, Liu LF (2007) Roles of DNA topoisomerase II isozymes in chemotherapy and secondary malignancies. Proc Natl Acad Sci USA 104:11014–11019
Beck WT, Danks MK, Wolverton JS, Kim R, Chen M (1993) Drug resistance associated with altered DNA topoisomerase II. Adv Enzym Regul 33:113–116
Berger JM, Gamblin SJ, Harrison SC, Wang JC (1996) Structure and mechanism of DNA topoisomerase II. Nature 379:225–232
Berman HM et al (2000) The protein data bank. Nucleic Acids Res 28:235–242
Berna L, Alvarez-Valin F (2014) Evolutionary genomics of fast evolving tunicates. Genome Biol Evol 6:1724–1738
Bollimpelli VS, Dholaniya PS, Kondapi AK (2017) Topoisomerase IIβ and its role in different biological contexts. Arch Biochem Biophys 633:78–84
Broderick L et al (2019) Mutations in topoisomerase IIβ result in a B cell immunodeficiency. Nat Commun 10:1–15
Broeck AV, Lotz C, Drillien R, Haas L, Bedez C, Lamour V (2021) Structural basis for allosteric regulation of Human Topoisomerase IIα. Nat Commun 12:1–13
Brown CJ, Johnson AK, Dunker AK, Daughdrill GW (2011) Evolution and disorder. Curr Opin Struct Biol 21:441–446
Cabral JHM, Jackson AP, Smith CV, Shikotra N, Maxwell A, Liddington RC (1997) Crystal structure of the breakage–reunion domain of DNA gyrase. Nature 388:903–906
Campbell LI et al (2011) MicroRNAs and phylogenomics resolve the relationships of Tardigrada and suggest that velvet worms are the sister group of Arthropoda. Proc Natl Acad Sci USA 108:15920–15924
Chakraborty AK, Majumder HK (1987) Decatenation of kinetoplast DNA by an ATP-dependent DNA topoisomerase from the kinetoplast hemoflagellate Leishmania donovani. Mol Biochem Parasitol 26:215–224
Champoux JJ (2001) DNA topoisomerases: structure, function, and mechanism. Annu Rev Biochem 70:369–413
Cheesman S, McAleese S, Goman M, Johnson D, Horrocks P, Ridley RG, Kilbey BJ (1994) The gene encoding topoisomerase II from Plasmodium falciparum. Nucleic Acids Res 22:2547–2551
Chen W, Qiu J, Shen Y (2012) Topoisomerase IIα, rather than IIβ, is a promising target in development of anti-cancer drugs. Drug Discov Ther 6:230–237
Classen S, Olland S, Berger JM (2003) Structure of the topoisomerase II ATPase region and its mechanism of inhibition by the chemotherapeutic agent ICRF-187. Proc Natl Acad Sci USA 100:10629–10634
Conant GC, Wagner A (2003) Asymmetric sequence divergence of duplicate genes. Genome Res 13:2052–2058
Corbett KD, Berger JM (2003) Structure of the topoisomerase VI-B subunit: implications for type II topoisomerase mechanism and evolution. EMBO J 22:151–163
Corbett KD, Schoeffler AJ, Thomsen ND, Berger JM (2005) The structural basis for substrate specificity in DNA topoisomerase IV. J Mol Biol 351:545–561
Cornarotti M, Tinelli S, Willmore E, Zunino F, Fisher LM, Austin CA, Capranico G (1996) Drug sensitivity and sequence specificity of human recombinant DNA topoisomerases IIalpha (p170) and IIbeta (p180). Mol Pharmacol 50:1463–1471
Cowell IG et al (2012) Model for MLL translocations in therapy-related leukemia involving topoisomerase IIβ-mediated DNA strand breaks and gene proximity. Proc Natl Acad Sci USA 109:8989–8994
Cuvier O, Hirano T (2003) A role of topoisomerase II in linking DNA replication to chromosome condensation. J Cell Biol 160:645–655
Darriba D, Taboada GL, Doallo R, Posada D (2012) jModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9:772–772
De H, Wen JF, Chen WQ, Lu SQ, De Dong X (2005) Identification, characteristic and phylogenetic analysis of type II DNA topoisomerase gene in Giardia lamblia. Cell Res 15:474–482
Del Amparo R, Branco C, Arenas J, Vicens A, Arenas M (2021) Analysis of selection in protein-coding sequences accounting for common biases. Brief Bioinform. https://doi.org/10.1093/bib/bbaa431
Delgado JL, Hsieh C-M, Chan N-L, Hiasa H (2018) Topoisomerases as anticancer targets. Biochem J 475:373–398
Delsuc F, Brinkmann H, Chourrout D, Philippe H (2006) Tunicates and not cephalochordates are the closest living relatives of vertebrates. Nature 439:965–968
Denoeud F et al (2010) Plasticity of animal genome architecture unmasked by rapid evolution of a pelagic tunicate. Science 330:1381–1385
Dong KC, Berger JM (2007) Structural basis for gate-DNA recognition and bending by type IIA topoisomerases. Nature 450:1201–1205
Drake FH et al (1987) Purification of topoisomerase II from amsacrine-resistant P388 leukemia cells. Evidence for two forms of the enzyme. J Biol Chem 262:16739–16747
Drake FH, Hofmann GA, Bartus HF, Mattern MR, Crooke ST, Mirabelli CK (1989) Biochemical and pharmacological properties of p170 and p180 forms of topoisomerase II. Biochemistry 28:8154–8160
Dunn CW et al (2008) Broad phylogenomic sampling improves resolution of the animal tree of life. Nature 452:745–749
Dunn CW, Giribet G, Edgecombe GD, Hejnol A (2014) Animal phylogeny and its evolutionary implications. Annu Rev Ecol Evol Syst 45:371–395
Dutta R, Inouye M (2000) GHKL, an emergent ATPase/kinase superfamily. Trends Biochem Sci 25:24–28
Erdős M, Lányi Á, Balázs G, Casanova J-L, Boisson B, Maródi L (2021) Inherited TOP2B mutation: possible confirmation of mutational hotspots in the TOPRIM domain. J Clin Immunol 41:817–819
Escriva H, Manzon L, Youson J, Laudet V (2002) Analysis of lamprey and hagfish genes reveals a complex history of gene duplications during early vertebrate evolution. Mol Biol Evol 19:1440–1450
Fass D, Bogden CE, Berger JM (1999) Quaternary changes in topoisomerase II may direct orthogonal movement of two DNA strands. Nat Struct Biol 6:322–326
Forterre P, Gribaldo S, Gadelle D, Serre M-C (2007) Origin and evolution of DNA topoisomerases. Biochimie 89:427–446
Gadelle D, Filée J, Buhler C, Forterre P (2003) Phylogenomics of type II DNA topoisomerases. BioEssays 25:232–242
Gajiwala KS, Burley SK (2000) Winged helix proteins. Curr Opin Struct Biol 10:110–116
Gilroy KL, Leontiou C, Padget K, Lakey JH, Austin CA (2006) mAMSA resistant human topoisomerase IIβ mutation G465D has reduced ATP hydrolysis activity. Nucleic Acids Res 34:1597–1607
Glessner JT et al (2014) Increased frequency of de novo copy number variants in congenital heart disease by integrative analysis of single nucleotide polymorphism array and exome sequence data. Circ Res 115:884–896
Green RE et al (2010) A draft sequence of the Neandertal genome. Science 328:710–722
Grue P et al (1998) Essential mitotic functions of DNA topoisomerase IIα are not adopted by topoisomerase IIβ in human H69 cells. J Biol Chem 273:33660–33666
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Haffner MC et al (2010) Androgen-induced TOP2B-mediated double-strand breaks and prostate cancer gene rearrangements. Nat Genet 42:668–675
Hunt SE et al (2018) Ensembl variation resources. Database 2018:bay 119
Ishida R et al (2001) Mitotic specific phosphorylation of serine-1212 in human DNA topoisomerase IIα. Cell Struct Funct 26:215–226
Jeffares DC, Tomiczek B, Sojo V, dos Reis M (2015) A beginners guide to estimating the non-synonymous to synonymous rate ratio of all protein-coding genes in a genome. Methods Mol Biol 1201:65–90. https://doi.org/10.1007/978-1-4939-1438-8_4
Jenkins JR et al (1992) Isolation of cDNA clones encoding the β isozyme of human DNA topoisomerase II and localisation of the gene to chromosome 3p24. Nucleic Acids Res 20:5587–5592
Ju B-G, Lunyak VV, Perissi V, Garcia-Bassets I, Rose DW, Glass CK, Rosenfeld MG (2006) A topoisomerase IIß-mediated dsDNA break required for regulated transcription. Science 312:1798–1802
Katoh K, Rozewicki J, Yamada KD (2019) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20:1160–1166
Kent WJ, Sugnet CW, Furey TS, Roskin KM, Pringle TH, Zahler AM, Haussler D (2002) The human genome browser at UCSC. Genome Res 12:996–1006
Kingma P, Osheroff N, Bjergbaek L, Nielsen IS, Wang Y, Westergaard O, Andersen AH (2000) Communication between the ATPase and cleavage/religation domains of human topoisomerase IIα. J Biol Chem 275:13041–13048
Kondrashov FA, Rogozin IB, Wolf YI, Koonin EV (2002) Selection in the evolution of gene duplications. Genome Biol 3:1–9
Kosakovsky Pond SL, Frost SD (2005) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22:1208–1222
Kosakovsky Pond SL et al (2020) HyPhy 2.5—a customizable platform for evolutionary hypothesis testing using phylogenies. Mol Biol Evol 37:295–299
Kozuki T et al (2017) Roles of the C-terminal domains of topoisomerase IIα and topoisomerase IIβ in regulation of the decatenation checkpoint. Nucleic Acids Res 45:5995–6010
Kriventseva EV, Kuznetsov D, Tegenfeldt F, Manni M, Dias R, Simão FA, Zdobnov EM (2019) OrthoDB v10: sampling the diversity of animal, plant, fungal, protist, bacterial and viral genomes for evolutionary and functional annotations of orthologs. Nucleic Acids Res 47:D807–D811
Kuhlwilm M, Boeckx C (2019) A catalog of single nucleotide changes distinguishing modern humans from archaic hominins. Sci Rep 9:1–14
Lam C-W, Yeung W-L, Law C-Y (2017) Global developmental delay and intellectual disability associated with a de novo TOP2B mutation. Clin Chim Acta 469:63–68
Lane AB, Giménez-Abián JF, Clarke DJ (2013) A novel chromatin tether domain controls topoisomerase IIα dynamics and mitotic chromosome formation. J Cell Biol 203:471–486
Laponogov I et al (2007) Breakage-reunion domain of Streptococcus pneumoniae topoisomerase IV: crystal structure of a Gram-positive quinolone target. PLoS ONE 2:e301
Lefort V, Longueville J-E, Gascuel O (2017) SMS: smart model selection in PhyML. Mol Biol Evol 34:2422–2424
Leontiou C, Lightowlers R, Lakey JH, Austin CA (2003) Kinetic analysis of human topoisomerase IIα and β DNA binding by surface plasmon resonance. FEBS Lett 554:206–210
Leontiou C, Lakey JH, Lightowlers R, Turnbull RM, Austin CA (2006) Mutation P732L in human DNA topoisomerase IIβ abolishes DNA cleavage in the presence of calcium and confers drug resistance. Mol Pharmacol 69:130–139
Leontiou C et al (2007) Differential selection of acridine resistance mutations in human DNA topoisomerase IIβ is dependent on the acridine structure. Mol Pharmacol 71:1006–1014
Linka RM, Porter AC, Volkov A, Mielke C, Boege F, Christensen MO (2007) C-terminal regions of topoisomerase II α and II β determine isoform-specific functioning of the enzymes in vivo. Nucleic Acids Res 35:3810–3822
Liu LF, Rowe TC, Yang L, Tewey KM, Chen GL (1983) Cleavage of DNA by mammalian DNA topoisomerase II. J Biol Chem 258:15365–15370
Lotz C, Lamour V (2020) The interplay between DNA topoisomerase 2α post-translational modifications and drug resistance. Cancer Drug Resist. https://doi.org/10.20517/cdr.2019.114
Lyu YL, Wang JC (2003) Aberrant lamination in the cerebral cortex of mouse embryos lacking DNA topoisomerase IIβ. Proc Natl Acad Sci USA 100:7123–7128
Lyu YL, Lin C-P, Azarova AM, Cai L, Wang JC, Liu LF (2006) Role of topoisomerase IIβ in the expression of developmentally regulatedgenes. Mol Cell Biol 26:7929–7941
Marsh KL et al (1996) Amsacrine-promoted DNA cleavage site determinants for the two human DNA topoisomerase II isoforms α and β. Biochem Pharmacol 52:1675–1685
McClendon AK, Rodriguez AC, Osheroff N (2005) Human topoisomerase IIα rapidly relaxes positively supercoiled DNA: implications for enzyme action ahead of replication forks. J Biol Chem 280:39337–39345
McKay DB, Steitz TA (1981) Structure of catabolite gene activator protein at 2.9 Å resolution suggests binding to left-handed B-DNA. Nature 290:744–749
McNamara S, Wang H, Hanna N, Miller WH Jr (2008) Topoisomerase IIβ negatively modulates retinoic acid receptor α function: a novel mechanism of retinoic acid resistance. Mol Cell Biol 28:2066–2077
Miller MP, Kumar S (2001) Understanding human disease mutations through the use of interspecific genetic variation. Hum Mol Genet 10:2319–2328
Mirski SE, Gerlach JH, Cummings HJ, Zirngibl R, Greer PA, Cole SP (1997) Bipartite nuclear localization signals in the C terminus of human topoisomerase IIα. Exp Cell Res 237:452–455
Moreira F, Arenas M, Videira A, Pereira F (2021) Molecular evolution of DNA topoisomerase III beta (TOP3B) in metazoa. J Mol Evol 89:384–395
Morrison A, Cozzarelli NR (1979) Site-specific cleavage of DNA by E. coli DNA gyrase. Cell 17:175–184
Nakatani Y, Shingate P, Ravi V, Pillai NE, Prasad A, McLysaght A, Venkatesh B (2021) Reconstruction of proto-vertebrate, proto-cyclostome and proto-gnathostome genomes provides new insights into early vertebrate evolution. Nat Commun 12:1–14
Nesnidal MP et al (2013) New phylogenomic data support the monophyly of Lophophorata and an Ectoproct-Phoronid clade and indicate that Polyzoa and Kryptrochozoa are caused by systematic bias. BMC Evol Biol 13:1–13
Niimi A, Suka N, Harata M, Kikuchi A, Mizuno S (2001) Co-localization of chicken DNA topoisomerase IIα, but not β, with sites of DNA replication and possible involvement of a C-terminal region of α through its binding to PCNA. Chromosoma 110:102–114
Nitiss JL (2009) Targeting DNA topoisomerase II in cancer chemotherapy. Nat Rev Cancer 9:338–350
Nitiss J, Beck W (1996) Antitopoisomerase drug action and resistance. Eur J Cancer 32:958–966
Ohno S (2013) Evolution by gene duplication. Springer Science & Business Media, New York
Papapietro O et al (2020) Topoisomerase 2β mutation impairs early B-cell development. Blood 135:1497–1501
Pisani D, Carton R, Campbell LI, Akanni WA, Mulville E, Rota-Stabelli O (2013) An overview of arthropod genomics, mitogenomics, and the evolutionary origins of the arthropod proteome. In: Minelli A, Boxshall G, Fusco G (eds) Arthropod biology and evolution. Springer, Berlin, Heidelberg. https://doi.org/10.1007/978-3-642-36160-9_3
Pommier Y, Sun Y, Shar-yin NH, Nitiss JL (2016) Roles of eukaryotic topoisomerases in transcription, replication and genomic stability. Nat Rev Mol Cell Biol 17:703–721
Putnam NH et al (2008) The amphioxus genome and the evolution of the chordate karyotype. Nature 453:1064–1071
Reich D et al (2010) Genetic history of an archaic hominin group from Denisova Cave in Siberia. Nature 468:1053–1060
Roca J, Wang JC (1992) The capture of a DNA double helix by an ATP-dependent protein clamp: a key step in DNA transport by type II DNA topoisomerases. Cell 71:833–840
Roca J, Wang JC (1994) DNA transport by a type II DNA topoisomerase: evidence in favor of a two-gate mechanism. Cell 77:609–616
Roca J, Berger JM, Harrison SC, Wang JC (1996) DNA transport by a type II topoisomerase: direct evidence for a two-gate mechanism. Proc Natl Acad Sci USA 93:4057–4062
Sander M, Hsieh T-S (1983) Double strand DNA cleavage by type II DNA topoisomerase from Drosophila melanogaster. J Biol Chem 258:8421–8428
Satoh N, Rokhsar D, Nishikawa T (2014) Chordate evolution and the three-phylum system. Proc R Soc B 281:20141729
Scannell DR, Wolfe KH (2008) A burst of protein sequence evolution and a prolonged period of asymmetric evolution follow gene duplication in yeast. Genome Res 18:137–147
Schoeffler AJ, Berger JM (2008) DNA topoisomerases: harnessing and constraining energy to govern chromosome topology. Q Rev Biophys 41:41–101
Sehnal D et al (2021) Mol* viewer: modern web app for 3D visualization and analysis of large biomolecular structures. Nucleic Acids Res. https://doi.org/10.1093/nar/gkab314
Simakov O et al (2020) Deeply conserved synteny resolves early events in vertebrate evolution. Nat Ecol Evol 4:820–830
Smith JJ, Keinath MC (2015) The sea lamprey meiotic map improves resolution of ancient vertebrate genome duplications. Genome Res 25:1081–1090
Stadler PF, Fried C, Prohaska SJ, Bailey WJ, Misof BY, Ruddle FH, Wagner GP (2004) Evidence for independent Hox gene duplications in the hagfish lineage: a PCR-based gene inventory of Eptatretus stoutii. Mol Phylogenet Evol 32:686–694
Strauss PR, Wang JC (1990) The TOP2 gene of Trypanosoma brucei: a single-copy gene that shares extensive homology with other TOP2 genes encoding eukaryotic DNA topoisomerase II. Mol Biochem Parasitol 38:141–150
Sun L-Y, Zhu L-W, Tang Y-J (2018) Increasing the distance between two monomers of topoisomerase IIβ under the action of antitumor agent 4 β-sulfur-(benzimidazole) 4′-demethylepipodophyllotoxin. Sci Rep 8:1–9
Tan K, Dorman TE, Falls KM, Chung TD, Mirabelli CK, Crooke ST, Mao J-i (1992) Topoisomerase IIα and topoisomerase IIβ genes: characterization and mapping to human chromosomes 17 and 3, respectively. Cancer Res 52:231–234
Tiwari VK et al (2012) Target genes of Topoisomerase IIβ regulate neuronal survival and are defined by their chromatin state. Proc Natl Acad Sci USA 109:E934–E943
Tsagkogeorga G, Turon X, Galtier N, Douzery EJ, Delsuc F (2010) Accelerated evolutionary rate of housekeeping genes in tunicates. J Mol Evol 71:153–167
Tsai-Pflugfelder M et al (1988) Cloning and sequencing of cDNA encoding human DNA topoisomerase II and localization of the gene to chromosome region 17q21-22. Proc Natl Acad Sci USA 85:7177–7181
Van de Peer Y, Taylor JS, Braasch I, Meyer A (2001) The ghost of selection past: rates of evolution and functional divergence of anciently duplicated genes. J Mol Evol 53:436–446
Van de Peer Y, Maere S, Meyer A (2009) The evolutionary significance of ancient genome duplications. Nat Rev Genet 10:725–732
Vassetzky YS, Alghisi GC, Gasser SM (1995) DNA topoisomerase II mutations and resistance to anti-tumor drugs. BioEssays 17:767–774
Wang JC (1996) DNA topoisomerases. Annu Rev Biochem 65:635–692
Wang JC (2002) Cellular roles of DNA topoisomerases: a molecular perspective. Nat Rev Mol Cell Biol 3:430–440
Watterson G (1975) On the number of segregating sites in genetical models without recombination. Theor Popul Biol 7:256–276
Wigley DB, Davies GJ, Dodson EJ, Maxwell A, Dodson G (1991) Crystal structure of an N-terminal fragment of the DNA gyrase B protein. Nature 351:624–629
Wu C-C et al (2011) Structural basis of type II topoisomerase inhibition by the anticancer drug etoposide. Science 333:459–462
Wu C-C, Li Y-C, Wang Y-R, Li T-K, Chan N-L (2013) On the structural basis and design guidelines for type II topoisomerase-targeting anticancer drugs. Nucleic Acids Res 41:10630–10640
Xia W et al (2019) Mutations in TOP 2B cause autosomal-dominant hereditary hearing loss via inhibition of the PI 3K-Akt signalling pathway. FEBS Lett 593:2008–2018
Yang X, Li W, Prescott ED, Burden SJ, Wang JC (2000) DNA topoisomerase IIβ and neural development. Science 287:131–134
Ye J et al (2010) TRF2 and Apollo cooperate with topoisomerase 2α to protect human telomeres from replicative damage. Cell 142:230–242
Funding
This work was partially supported by a research grant to FM (SFRH/BD/131584/2017) from Fundação para a Ciência e a Tecnologia. MA was supported by the Spanish Ministerio de Ciencia e Innovación through the Grants [RYC-2015-18241] and [PID2019-107931GA-I00].
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
None to declare.
Additional information
Handling editor: David Alvarez-Ponce.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Moreira, F., Arenas, M., Videira, A. et al. Evolutionary History of TOPIIA Topoisomerases in Animals. J Mol Evol 90, 149–165 (2022). https://doi.org/10.1007/s00239-022-10048-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-022-10048-2