Abstract
Preventing and controlling epidemics caused by vector-borne viruses are particularly challenging due to their diverse pool of hosts and highly adaptive nature. Many vector-borne viruses belong to the Flavivirus genus, whose members vary greatly in host range and specificity. Members of the Flavivirus genus can be categorized to four main groups: insect-specific viruses that are maintained solely in arthropod populations, mosquito-borne viruses and tick-borne viruses that are transmitted to vertebrate hosts by mosquitoes or ticks via blood feeding, and those with no-known vector. The mosquito-borne group encompasses the yellow fever, dengue, and West Nile viruses, all of which are globally spread and cause severe morbidity in humans. The Flavivirus genus is genetically diverse, and its members are subject to different host-specific and vector-specific selective constraints, which do not always align. Thus, understanding the underlying genetic differences that led to the diversity in host range within this genus is an important aspect in deciphering the mechanisms that drive host compatibility and can aid in the constant arms-race against viral threats. Here, we review the phylogenetic relationships between members of the genus, their infection bottlenecks, and phenotypic and genomic differences. We further discuss methods that utilize these differences for prediction of host shifts in flaviviruses and can contribute to viral surveillance efforts.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The Flavivirus genus consists of more than 90 species that exhibit high variability in host range and specificity (Schoch et al. 2020). Some of these are significant public-health pathogens, such as dengue virus, zika virus, yellow fever virus, West Nile virus, and Japanese encephalitis virus. Dengue virus alone is responsible for about 390 million infections each year, and currently only limited antiviral treatments are available for its control (Thomas and Yoon 2019; Troost and Smit 2020). According to the World Health Organization, the number of reported dengue cases increased by over eightfold in the last two decades, with over 4.2 million reported cases in 2019 (World Health Organization 2020).
All flaviviruses are enveloped, positive-sense, and single-stranded RNA (ssRNA) viruses, with small genomes, ranging from 9 to 13 Kb, and consisting of a 3’ and 5’ untranslated regions (UTRs) wrapping a single open-reading frame (ORF). This reading frame is translated into a polyprotein, from which both structural and non-structural proteins are derived. The structural region, located at the N terminus of the polyprotein, contains the capsid protein, followed by a pre-membrane glycoprotein which protects the virion from pH changes in host environment, and an envelope glycoprotein which acts as a receptor for binding and entry to host cells. The non-structural region consists of the proteins NS1, NS2A, NS2B, NS3, NS4A, NS4B, and NS5 (Chambers et al. 1991). These non-structural proteins are responsible for various functions in the virus, ranging from proteolysis, genomic replication, and evasion from the host immune system (Chong et al. 2019).
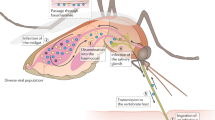
The Flavivirus genus can be categorized into four (not necessarily monophyletic) groups according to their host range and transmission mode: insect-specific flavivirus (ISF), mosquito-borne flavivirus (MBFV), tick-borne flavivirus (TBFV), and no-known arthropod vector (NKV). While ISFs only infect invertebrate hosts and NKVs primarily infect vertebrate hosts, MBFVs, and TBFVs can replicate in both. In accordance with the variation in their host-tropism, flaviviruses from these different groups also vary in their mode of transmission between infected individuals. ISFs are usually transmitted vertically, from arthropod mother to offspring (Halbach et al. 2017), while NKVs were shown to transmit mainly horizontally between their vertebrate hosts (Blitvich and Firth 2017). In contrast, MBFVs and TBFVs, which possess dual-host tropism, undergo alternating transmission cycles between their insect vectors and their vertebrate hosts (Franz et al. 2015). As such, successful transmission of vector-borne viruses from their arthropod vectors to their vertebrate hosts entails the infection of both the midgut and salivary gland of the insect (Brackney and Armstrong 2016; Miesen et al. 2016). Aside from transmission to vertebrates via insect bites, vertebrates can be infected through ingestion of the insect (Dobson et al. 1995), consumption of dairy and meat of infected vertebrates (Swanepoel et al. 1985; Offerdahl et al. 2016) and blood transfusions (Leiby and Gill 2004).
Vector-borne viruses exhibit greater host plasticity, infecting up to three times the number of host taxonomic groups, compared to non-vector-borne viruses (Kreuder Johnson et al. 2015). This property could be attributed to the increased opportunities for effective contact across diverse animal hosts enabled by vector-borne transmission. Indeed, vectors provide an effective bridge for transmission of disease from wild animals that do not normally contact humans, thereby increasing the zoonosis frequency of vector-borne viruses. Furthermore, vector-borne flaviviruses demonstrate highly variable host plasticity, with some infecting only closely related hosts while others infect a wide range of vertebrate taxa (Weaver and Barrett 2004). For example, the vector-borne West Nile virus can infect a wide range of vertebrate hosts, from humans and horses to birds (e.g., Turdus merula), while the vector-borne yellow fever virus was only isolated from primates (Mihara et al. 2016; Michel et al. 2019). These differences in host range make the Flavivirus genus a compelling target for studying the viral traits that control host range and specificity. Such knowledge can be utilized for early detection of outbreaks through identification of potential host species and monitoring of viral threats (Bicca-Marques and de Freitas 2010).
The evolutionary basis for the adaptability of vector-borne viruses to multiple hosts has been previously explored. In this context, three hypotheses consider the fitness landscape of vector-borne viruses in the context of their multiple hosts (reviewed in Novella et al. 2012). The first hypothesis, termed the tradeoff hypothesis, proposes that vector-borne viruses maintain adequate fitness to their multiple hosts in exchange for lower adaptation capabilities towards any one of them (Wilson and Yoshimura 1994; Kassen 2002). The second hypothesis suggests that vector-borne viruses do occasionally reach an optimal peak that is shared across multiple hosts. The third hypothesis argues that the evolutionary constraints of vector-borne viruses are mainly determined by their vector host due to its key role in transmission to vertebrates. Accordingly, viruses reside in the optimal fitness peak within the vector, rather than in that of the vertebrate host. All three hypotheses are supported by different experimental studies (e.g., Ciota et al. 2008; Deardorff et al. 2011). The discrepancy across these studies could be attributed to differences in the methods utilized for measuring fitness and for manipulating the passage of vector-borne viruses through their different hosts. For example, the quasispecies theory proposes that viral fitness should be measured while considering the variation across multiple viral individuals within a population, rather than virus individuals (Domingo et al. 2012). This variation, viewed as mutant clouds, is induced by a population of interacting viruses and enables the expression of phenotypic variance and virus adaptability (Eigen 1996; Wilke 2005).
In addition to intrinsic viral characteristics, host traits and environmental factors may also influence viral capability to replicate and persist within hosts. Thus, the identification of macro-ecological attributes that facilitate viral persistence can contribute to our understanding of the variation, prevalence, and propensity of viruses towards specific hosts. Several recent studies have utilized machine learning and explanatory models to predict virus-host associations based on traits of viruses and their respective hosts (Olival et al. 2017; Babayan et al. 2018; Pandit et al. 2018; Mollentze and Streicker 2020; Albery et al. 2020). Yet, the choice of the traits used for predictions and the underlying causes for their contribution to the prediction accuracy are not fully understood. In addition, empirical validation of such predictions is still largely missing. In this review, we first examine the phylogenetic relationships within the Flavivirus genus and point to the multiple potential shifts in host tropism. We then discuss infection barriers and phenotypic and genomic differences between members of the Flavivirus genus. Finally, we review emerging methods that partially utilize this knowledge, together with a range of viral and host traits, to predict potential hosts shifts within the Flavivirus genus and other viral genera.
Phylogenetic Relationships and Evolution Among Flaviviruses
Taking advantage of many complete genomic sequences of flaviviruses available at that time, Moureau et al. (2015) established the phylogenetic relationships within the Flaviviridae family in the context of vector-host relationships. In agreement with previous studies (Cook et al. 2012; Marklewitz et al. 2015), the ISF clade was placed externally to all other clades, suggesting that the ancestral state at the root of the Flavivirus genus was limited to infecting invertebrates. Based on that phylogeny, at least five major transitions in host compatibility were observed (Fig. 1 in Moureau et al. 2015): Two leading to the TBFV and NKV clades, and another to a large clade, dominated by mosquito-borne viruses. Two additional transitions were observed within this latter clade. First, three NKV viruses (sokuluk, Entebbe bat and yokose viruses) that were later denoted NKV-like viruses, were clustered within the MBFV group. This suggests that the ancestors of these NKV-like viruses were able to replicate in mosquitos, and this ability has been lost at the present lineages (Cook and Holmes 2006). Despite the different categorization of the Flavivirus groups of species, the phylogeny exhibited in Ochsenreiter et al. (2019), which was also reconstructed based on the complete polyprotein sequences of several flaviviruses, also supports the paraphyly of MBFVs with the three NKV-like viruses. Second, seven previously unknown ISF viruses, later denoted ISF-like viruses, which do not exhibit an ability to replicate in vertebrate cells, were clustered together with MBFVs such as dengue and yellow fever viruses, (Huhtamo et al. 2009, 2014; Junglen et al. 2009). Thus, these viruses may have lost the genetic trait that enabled them to infect vertebrate hosts.
Phylogenetic tree of 83 Flavivirus species. The tree is colored according to host-tropism groups: insect-specific (ISF) in gray, no-known vector (NKV) in blue, tick-borne (TBFV) in green, mosquito-borne (MBFV) in red, NKV-like in orange, and ISF-like in yellow. Twelve species with available NS5 sequence data but whose classification is unknown or whose sequences are unreliable were pruned from the tree.
Following the accumulation of sequences from additional Flavivirus species, we have re-examined the phylogenetic relationships within Flavivirus based on the NS5 protein sequences (Fig. 1), as opposed to the longer and more variable polyprotein sequences that were used in Moureau et al. (2015). The vital functionality of NS5 in viral replication (Murray et al. 2008; Bollati et al. 2010) and suppression of the host immune response (Lin et al. 2006) leads to high genetic conservation, making it an appealing target for serving as a phylogenetic marker as well as for antiviral treatments. Sequences of different Flavivirus species were collected from the Virus Pathogen Resource database (Pickett et al. 2012). These were supplemented with several sequences of species that were present in Moureau et al. (2015) but absent from the Virus Pathogen Resource database. A single-representative NS5 protein sequence was selected for each species using CD-HIT version 4.6 (Li and Godzik 2006). The sequences were then aligned with MAFFT version 7.471 (Katoh et al. 2002). The phylogenetic tree was reconstructed using IQTree version 2.0.3 (Minh et al. 2020) given an amino-acid substitution model customized for Flavivirus evolution (Le and Vinh 2020), coupled with a four-category Gamma distribution modeling rate variation across sites, with bootstrap analysis consisting of 1000 repeats. The resulting tree was rooted using minimal ancestor deviation as implemented in Python (Tria et al. 2017). The accessions of the selected sequences (Supplementary text S1), the alignment (Supplementary text S2), the resulting phylogeny (Supplementary text S3), and the classification of species to host tropism (Supplementary text S4) are available in the supplementary materials.
The reconstructed phylogeny consists of 95 species, 31 of which are absent from the phylogeny reconstructed by Moureau et al. (2015). There is a general agreement between the two phylogenies, despite differences in the reconstruction strategies employed: The two phylogenies differ with respect to the taxonomic level (species versus strains), sequence marker (polyprotein ORF versus NS5 protein), substitution model [WAG (Whelan and Goldman 2001) versus FLAVI (Le and Vinh 2020)], and phylogenetic reconstruction method (Bayesian versus maximum likelihood). In the phylogeny obtained here, all five transitions identified previously could be detected, although the exact placement of the NKV-like clade varied between the two phylogenies. In the phylogeny reconstructed here, an additional transition from a putative MBFV ancestor to the ISF-like nounane and barkedji viruses could be identified. This transition suggests the possibility that more than one back-transition from dual-host tropism to single-host tropism have occurred, although this additional transition is supported by a weak bootstrap value (0.32) and could be an artifact of the relatively short sequence length of the NS5 protein. Additional putative transitions of this type have been implicated. For example, the rabensburg virus, which is considered as an ISF, is closely related to the mosquito-borne West Nile virus (Shah-Hosseini et al. 2014; Elrefaey et al. 2020), but its NS5 sequence is missing from our compilation and thus not included in our phylogeny. A promising future research direction would verify whether these back-transitions have occurred and, if so, what are the genetic modifications that have led to the loss of dual-host tropism in these lineages.
Host Barriers of ISFs Preventing Infection of Vertebrates
The placement of the root in the Flavivirus phylogeny reconstructed here, as well as in those previously inferred (Cook et al. 2012; Marklewitz et al. 2015; Moureau et al. 2015; Ochsenreiter et al. 2019), suggests that vector-borne flaviviruses are the descendants of ISF lineages that later acquired dual-host tropism. Detecting the underlying genomic transitions that led to the acquisition of these abilities in vector-borne viruses could contribute to the identification of viral species which are more likely to acquire such abilities in the future and consequently widen their range of hosts. For an in-depth review of the bottlenecks of mosquito-specific viruses to enter and replicate in vertebrate cells, see Halbach et al. (2017). Below we point to their main findings and review additional research concerning TBFVs.
Higher temperature in vertebrates compared to invertebrates was shown to act as a replication bottleneck in mosquito-specific (Jerzak et al. 2008). For example, studies on the insect-specific rabensburg virus, which is closely related to the mosquito-borne West Nile virus, showed that acquisition of the ability to replicate at higher temperatures can lead to vertebrate host compatibility (Aliota and Kramer 2012; Aliota et al. 2012; Ngo et al. 2019). However, studies on other ISFs indicate that additional factors, other than the ability to replicate at higher temperatures, are needed for successful transmission of ISFs to vertebrate hosts (Huhtamo et al. 2014).
Inherent vertebrate factors can also limit their infection by ISFs. Such factors can be either virus Agonist that are required for the virus to complete its replicative cycle in the infected cell (Junglen et al. 2017), or virus antagonist, of which the most widely recognized are interferons that suppress viral replication. It appears that certain proteins in vector-borne flaviviruses, including NS1 and NS5, interact with either of these host factor types to enable successful infection of vertebrate cells (Grant et al. 2016; Best 2017; Xia et al. 2018). The inability of ISFs to infect vertebrates suggests that these proteins interact differently with the host factors in ISFs, thereby limiting viral infection. This altered functionality of ISF proteins enabled the development of a vaccine against the MBFV chikungunya virus based on a recombinant insect-specific virus carrying the chikungunya virus structural proteins (Erasmus et al. 2017).
Invertebrate factors may also play a role in host restriction of ISFs through tissue-specific immune response. Successful infection of vertebrate hosts via insect bites requires the presence of viral particles in the insect salivary gland. However, the insect immune response in the salivary gland acts as a barrier to many ISFs, preventing their transmission to vertebrates (Blitvich and Firth 2015; Hall-Mendelin et al. 2016). While the mechanisms underlying suppression of this tissue-specific immune response by vector-borne viruses are not well understood, evidence indicates that mosquito-borne viruses overcome this response through neutrilizing agnets. For example, Kent et al. (2010) showed that co-infecting mosquitoes with the insect-specific culex Flavivirus and the mosquito-borne West Nile virus enabled succesful infection of the salivary gland by both. This suggests that the ability of the West Nile virus to infect the salivary gland involves neutralization of some unknown mosquito factors that would otherwise prevent infection by ISFs. Tick-borne viruses face similar tissue-specific barriers in the infected tick (Nuttall 2014). Interstingly, the tick salivary gland was shown to contain active molecules that aid tick-borne viruses in coping with the vertebrate immune response following its transmission to vertebrates (Hermance and Thangamani 2015; Kotál et al. 2015).
Proteome variation between mosquito-borne and mosquito-specific flaviviruses may also explain differences in host compatibility. Aside from fixed proteome differences, dynamic variation can be the result of programmed ribosomal frameshifting (PRF)—A phenomenon in which viruses harbor sequences that induce a proportion of translating ribosomes to shift by one nucleotide position to a new reading frame, thus, producing a ‘transframe’ fusion protein (Firth and Brierley 2012). For example, a PRF product, termed NS1', has been observed in several MBFVs (Moureau et al. 2015) and was reported to play a role in the inhibition of the vertebrate interferon-mediated immune response (Zhou et al. 2018) and in increased transmission efficiency from mosquitoes to vertebrates (Melian et al. 2014). Another PRF that is unique to ISFs, results in a modified NS2A/NS2B-coding region, termed Fairly Interesting Flavivirus ORF (FIFO) (Firth et al. 2010). The function of this PRF product is still unknown. Surely, additional research is required to better understand the prevalence of other PRFs, whether they are also present in TBFVs, and their possible contribution in mediating host compatibly.
Finally, UTRs of flaviviruses include conserved, and often duplicated, RNA structural elements that were shown to be involved in various stages of the viral life cycle (Filomatori et al. 1995; Alvarez et al. 2005; Manzano et al. 2011; Brinton and Basu 2015; Ng et al. 2017). Unique structures were found in the different Flavivirus groups, suggesting that their presence could be associated with host adaptation (Davies and Pedersen 2008; Villordo et al. 2015, 2016; Pallarés et al. 2020). Ochsenreiter et al. (2019) performed a comparative analysis of the 3’UTR structures and detected a consistent 3’UTR architectural organization across TBFVs, and inconsistent organization in ISF and NKV, suggesting that the organization detected in TBFVs may confer their dual-host tropism.
Genome Composition of Flavivirus
Due to their alternating replication in invertebrate and vertebrate hosts, vector-borne flaviviruses face strong selective constraints that could limit reaching optimal fitness in either of them. These cumulative constraints are likely stronger than the ones that operate on single-host viruses. The fitness landscape of vector-borne viruses is constantly fluctuating according to their current host, leading to time-averaged adaptation (Wilke 2001). At the quasispecies level, these fluctuating environments are expected to affect the size of the mutant cloud and the pattern of mutation accumulation, reflecting the genetic variation within the virus population and its ability for adaptation. However, this does not always seem to be the case (Novella et al. 2012).
Molecular differences between dual-host and single-host viruses may also be observed at the genome composition level, reflected by variation in the relative frequencies of pairs of nucleotides and codons compared to their expected frequencies across genomic sequences, termed dinucleotide and codon usage, respectively. Lobo et al. (2009) explored the differential patterns of dinucleotide and codon usage across different Flaviviridae groups and their hosts. Using extracted ORF sequences from the complete genomes of 39 Flaviviridae species and the complete mRNA sequences of 9 vertebrates and 4 invertebrates, the authors discovered that the transcriptomes of all vertebrate-infecting flaviviruses display a dinucleotide usage pattern similar to that of vertebrates. Specifically, they found a bias against CpG and CpA in these groups, which was in accordance with the one observed in vertebrates. In vertebrates, this bias could be explained by cytosine methylation and deamination of the DNA in regions that are rich with CpG and CpA, a mechanism that is absent from insects (Bird 2007). In vertebrate-infecting flaviviruses, this bias can be explained as either mimicry of the host genomic composition (Kandimalla et al. 2003) or evasion from host immune system components, such as the Zinc-finger antiviral protein (ZAP) that recognize CpG containing viral RNA sequences (Takata et al. 2017; Odon et al. 2019; Meagher et al. 2019; Luo et al. 2020). A similar bias was not observed in ISFs. Indeed, Colmant et al. (2021) showed that the insect-specific binjari and Hidden Valley viruses were able to replicate in ZAP-knockout human cells, leading to the conclusion that ZAP is as an important barrier in prevention of ISF infection in vertebrates. Lobo et al. (2009) also observed a bias against UpA that was consistent across all viral groups. UpA is known to trigger vertebrate immune response of the RNase L enzyme (Han et al. 2004), but no such response has been recorded in insects. However, low UpA abundance can be explained in ISFs by the mimicry of dinucleotide usage of the insect transcriptome which is underabundant in UpA-containing tRNAs (Sexton and Ebel 2019). In general, UpA avoidance in both vertebrates and invertebrates can be triggered by avoidance of TATA box, TAA and TAG stop codons, or transcribed UA-rich RNA regions that lead to mRNA decay (Karlin and Mrázek 1997). At last, the biases against CpG and UpA appear to be compensated by a positive bias towards UpG in all the examined groups.
Several studies conducted a codon usage analysis to examine whether co-evolutionary signals are expressed at the codon level as well. In accordance with their dinucleotide analysis, Lobo et al. (2009) found a bias against CpG containing codons in vertebrate-infecting viruses and in their vertebrate hosts, but not in ISFs. However, no depletion in CpA containing codons was found, as might be expected from the detected dinucleotide underabundance. The authors also found depletion in UpA-containing codons in both insects and ISFs, which suggested mimicry of host codon usage. Notably, the analysis of Lobo et al. (2009) indicated that genome composition in vector-borne flaviviruses is influenced mostly by their vertebrate hosts—A finding that contradicts previous hypotheses that considered the insect vectors as the dominant driver of vector-borne viral evolution (see Introduction).
A recent study by Di Paola et al. (2018) investigated the codon adaptation of flaviviruses to their hosts using the codon adaptation index (CAI) (Sharp et al. 1986; Sharp and Li 1987), based on extracted ORFs from the complete genomic sequences of 205 flaviviruses and the codon usage tables of three vertebrate species and three invertebrate species. As expected, the CAIs to human genes were found to be higher in vector-borne flaviviruses compared to those of ISFs. The authors also detected significantly higher CAI to vertebrate genes in MBFVs compared to that of TBFVs, suggesting more rapid adaptation in the former group. The higher adaptation to humans in vector-borne viruses may be associated with their increased translation and replication capabilities in vertebrates (Andersen et al. 2015; Cugola et al. 2016). However, codon usage biases have not been directly associated with increased replication rates (Vasilakis et al. 2009; Shin et al. 2015). Interestingly, Di Paola et al. (2018) also collected sequences of West Nile viral genomes from the time of its recent emergence out of Africa and into Europe (2010–2014). The authors observed increased CAI of the virus to human-housekeeping genes between 2012 and 2014, compared to previous years. This increase was correlated with increased levels of infection, suggesting an association between codon usage adaptation and high infection efficacy. The increase in CAI during the time of the outbreak of the West Nile virus in Europe could potentially be attributed to increased exposure to humans. Thus, increasing similarity of codon usage between humans and vector-borne viruses that are not yet adapted to human hosts could possibly serve as an indicator of emerging zoonosis.
Notably, the studies described above showed resemblance in dinucleotide bias and codon usage between flaviviruses and their hosts. Di Giallonardo et al. (2017) hypothesized that such characteristics in viral species are more closely associated with their evolutionary history rather than with their hosts. This hypothesis is supported by the observation that dinucleotide bias in viruses simply reflects background mutation pressure (Wright 1990; Jenkins and Holmes 2003). To test their hypothesis, Di Giallonardo et al. predicted the host group and the viral taxonomic classifications based on the odds ratios of all 16 dinucleotides of 29,310 viral sequences from 20 families spanning a variety of hosts, some of which belonged to the Flavivirus genus. The prediction sensitivity was much higher for viral taxonomic classification as compared to the prediction of host groups (true positive prediction rate of 0.71 compared to 0.33). Moreover, exclusion of dual-host viruses from the analysis did not result in improved sensitivity of host prediction. However, the accuracy of host prediction based on dinucleotide information was much higher for the Flaviviridae family (true positive prediction rate of 0.76) and even more so when focusing on Flaviviridae viruses from the “vector-borne” host category. This suggests that vector-borne flaviviruses carry a unique genomic compositional signature, and this may be utilized for predicting virus-host associations in this group.
Methods for Prediction of Host Shifts and Compatibility
Identification of potential host shifts and zoonotic viral species can aid in the ongoing arms-race against viral threats by enabling risk-based allocations of research and surveillance effort. In this section, we discuss recent studies that have attempted to tackle these challenges, some of which focused on Flavivirus and others that examined a wider range of viruses.
Pandit et al. (2018) divided 35 Flavivirus species to several groups based on their known primary hosts and collected 29 traits of their hosts. Based on these traits, the authors trained a machine learning model for each group of flaviviruses to predict the identity of hosts that it may infect. The models corresponding to dengue and Japanese encephalitis viruses predicted 139 and 388 novel hosts for the respective viruses. The host traits that contributed most to the predictive model were those related to the geographical range of the species, body mass, and a feature that accounted for the biases in research efforts (i.e., the number of PubMed hits). In the model corresponding to the group of zika and Japanese encephalitis viruses, host metabolic rate was also determined as an important feature. This study demonstrated that geographical and physiological traits of the hosts can serve as important predictors of potential host shifts in flaviviruses. Interestingly, the importance of some features varied across models trained for different Flavivirus groups, which could indicate the presence of variation in the mechanisms conferring host adaptation in these different groups.
Other studies have built prediction models based on viral traits rather than those of the hosts. Many of these studies focused on genomic traits that have been suggested to indicate adaptation to hosts. For example, Babayan et al. (2018) trained gradient boosting models on features describing the phylogenetic relatedness between viruses as well as various genomic compositional biases of ssRNA viruses to predict for a given virus: (1) its host, (2) weather it can be transmitted via an arthropod vector, and if so (3) its transmission vector. The models produced predictions of high accuracy as assessed using cross-validation (0.84, 0.97, and 0.91 for the respective three categories). The models also predicted the hosts of 36 viruses in which hosts have not been verified empirically and arthropod vector compatibility for 17 viruses that have not been classified as vector borne thus far. Finally, the models predicted the vector class (i.e., midge, mosquito, sandfly, tick) of 31 vector-borne viruses whose vectors are currently unknown. However, some of these predictions were found to be contradictory to current evidence. For example, the model for reservoir hosts predicted Pterobats and Vestbats as the hosts of the MERS virus, while evidence suggest that its main host is in fact Artiodactyl (Ghai et al. 2021). An interesting future development could combine features that are based on traits of both the host and the virus for predicting probable virus-host associations. Furthermore, additional features could be integrated into the learning framework, strengthening the predictive power, including those that are related to transmission bottlenecks such as the host body temperature, virus transmission mode, identity of duplicated molecular secondary structures in the viral 3’UTR, and the identity of different products of programmed ribosomal frameshifting (Fig. 2).
Host and virus factors that can be utilized for predictions of virus-host associations. The figure depicts the factors discussed within the scope of this review that can be used as features in machine learning models for predictions of virus-host associations in Flavivirus. Factors that characterize different hosts are shown at the left side while factors that characterize viruses are shown at the right. These features are fed into machine learning methods to predict various virus-host associations, such as probable hosts or dual-host tropism. The four Flavivirus host-tropism groups listed in the figure are: mosquito-borne (MBFV), tick-borne (TBFV), insect-specific (ISF), and no-known vector (NKV).
While computational methods provide the opportunity to detect the host range of novel viruses, empirical validation of their results can be challenging. Isolation and identification of viruses from hosts can be obtained in several manners, including those that are based on serological samples using enzyme-linked immunosorbent assays (ELISA) and immunofluorescence assays (Gubler et al. 1984; Innis et al. 1989), or via molecular techniques that involve nucleic acid amplification using polymerase chain reaction (PCR) (or reverse transcriptase PCR in the case of RNA viruses) (Guarneri et al. 2001) or nucleic acid hybridization and microarrays. While molecular methods are considered more sensitive (Lanciotti et al. 2000; Lanciotti and Kerst 2001), they may fail to detect viruses that produce low and short-lived viremias, such as the West Nile virus (Murray et al. 2011). Molecular methods can also be applied to archival samples to characterize epidemics over time and to retrospectively diagnose viral diseases (Frisbie et al. 2004; Decaro et al. 2013).
The above experimental detection methods rely on the prior knowledge of the identity of the viruses that are expected to reside in a sample of interest and are thus limited to viruses that are well described. To overcome this limitation, detection methods based on metagenomic data were developed. Such methods use data collected from environmental samples and animal tissues and do not rely on viral amplification in cell culture (Delwart 2007). Rather, these methods entail the use of computational tools for the assembly and identification of viral sequences from high-throughput metagenomic sequence data (Cholleti et al. 2018). Viral metagenomic analysis has provided means for discovery of many novel viruses. For example, Li et al. (2015) identified 112 novel RNA viruses across 70 arthropod species. In another large-scale analysis of metagenomic samples extracted from 220 invertebrate species, Shi et al. (2016) discovered 1445 novel RNA viruses. Still, the use of metagenomic-based methods entails several challenges. First, assembly that is based on viral reference genomes may fail to detect viral sequences that have diverged from the reference, while a reference-free de-novo assembly is more susceptible to produce ambiguous and chimeric sequences of closely related viruses (Domingo et al. 2012). Furthermore, classification of viral sequences is based on similarity searches and is thus limited by the variety of annotated viral genomes in existing databases (Simmonds 2015). A promising future development could utilize the abundance of available metagenomic data from different sources for the simultaneous analysis of multiple viral metagenomes using a pan-genomics approach, similar to approaches for microbial communities profiling (Zhong et al. 2021). This will enable distinguishing between genuine viral species and those that are a product of specific modifications in a metagenomic sample and are not likely to survive. Such developments would also enable the identification of viruses with wide host range and the detection of conserved genetic regions that confer wide host compatibility.
Conclusions
In the past few years, important progress has been made in our understanding of Flavivirus evolution, including better resolution of the phylogenetic relationships and co-evolutionary patterns at the genomic level between viruses and their hosts. Ongoing advances in predictive models that utilize artificial intelligence techniques are expected to play a key role in research efforts aimed at deciphering the differences between distinct Flavivirus groups and mechanisms that induce compatibility to their different host types. To date, learning methods, trained on viral traits, demonstrated great potential for the prediction of novel vectors and hosts based on cross-validation. However, empirical support of novel virus-host associations is largely missing and their result can often be biased due to imbalanced data or increased error due to the “curse of dimensionality” (Agany et al. 2020). It is expected that models that utilize a combination of features, spanning both viral and host traits, would provide greater predictive power. Such anticipated developments should contribute greatly to the ongoing battle against viral threats and would focus viral surveillance efforts on host species and viral species with high predicted potential of transmission to humans.
Data Availability
The list of collected accessions and sequences, alignment, and tree are available in the supplementary materials.
References
Agany DDM, Pietri JE, Gnimpieba EZ (2020) Assessment of vector-host-pathogen relationships using data mining and machine learning. Comput Struct Biotechnol J 18:1704–1721. https://doi.org/10.1016/j.csbj.2020.06.031
Albery GF, Eskew EA, Ross N, Olival KJ (2020) Predicting the global mammalian viral sharing network using phylogeography. Nat Commun 11:2260. https://doi.org/10.1038/s41467-020-16153-4
Aliota MT, Jones SA, Dupuis AP et al (2012) Characterization of Rabensburg virus, a Flavivirus closely related to West Nile virus of the Japanese encephalitis antigenic group. PLoS ONE. https://doi.org/10.1371/journal.pone.0039387
Aliota MT, Kramer LD (2012) Replication of West Nile virus, Rabensburg lineage in mammalian cells is restricted by temperature. Parasites Vectors 5:293. https://doi.org/10.1186/1756-3305-5-293
Alvarez DE, De Lella Ezcurra AL, Fucito S, Gamarnik AV (2005) Role of RNA structures present at the 3′UTR of dengue virus on translation, RNA synthesis, and viral replication. Virology 339:200–212. https://doi.org/10.1016/j.virol.2005.06.009
Andersen KG, Shapiro BJ, Matranga CB et al (2015) Clinical sequencing uncovers origins and evolution of lassa virus. Cell 162:738–750. https://doi.org/10.1016/j.cell.2015.07.020
Babayan SA, Orton RJ, Streicker DG (2018) Predicting reservoir hosts and arthropod vectors from evolutionary signatures in RNA virus genomes. Science 362:577–580. https://doi.org/10.1126/science.aap9072
Best SM (2017) The many faces of the Flavivirus NS5 protein in antagonism of Type I interferon signaling. J Virol. https://doi.org/10.1128/jvi.01970-16
Bicca-Marques JC, de Freitas DS (2010) The role of monkeys, mosquitoes, and humans in the occurrence of a yellow fever outbreak in a fragmented landscape in south brazil: protecting howler monkeys is a matter of public health. Trop Conserv Sci 3:78–89. https://doi.org/10.1177/194008291000300107
Bird A (2007) Perceptions of epigenetics. Nature 447:396–398. https://doi.org/10.1038/nature05913
Blitvich B, Firth A (2015) Insect-specific Flaviviruses: a systematic review of their discovery, host range, mode of transmission, superinfection exclusion potential and genomic organization. Viruses 7:1927–1959. https://doi.org/10.3390/v7041927
Blitvich B, Firth A (2017) A review of Flaviviruses that have no known arthropod vector. Viruses 9:154. https://doi.org/10.3390/v9060154
Bollati M, Alvarez K, Assenberg R et al (2010) Structure and functionality in Flavivirus NS-proteins: perspectives for drug design. Antiviral Res 87:125–148. https://doi.org/10.1016/j.antiviral.2009.11.009
Brackney DE, Armstrong PM (2016) Transmission and evolution of tick-borne viruses. Curr Opin Virol 21:67–74. https://doi.org/10.1016/j.coviro.2016.08.005
Brinton MA, Basu M (2015) Functions of the 3′ and 5′ genome RNA regions of members of the genus Flavivirus . Virus Res 206:108–119. https://doi.org/10.1016/j.virusres.2015.02.006
Chambers TJ, Grakoui A, Rice CM (1991) Processing of the yellow fever virus nonstructural polyprotein: a catalytically active NS3 proteinase domain and NS2B are required for cleavages at dibasic sites. J Virol 65:6042–6050. https://doi.org/10.1128/jvi.65.11.6042-6050.1991
Cholleti H, Berg M, Hayer J, Blomström A-L (2018) Vector-borne viruses and their detection by viral metagenomics. Infect Ecol Epidemiol 8:1553465. https://doi.org/10.1080/20008686.2018.1553465
Chong HY, Leow CY, Abdul Majeed AB, Leow CH (2019) Flavivirus infection—A review of immunopathogenesis, immunological response, and immunodiagnosis. Virus Res 274:197770
Ciota AT, Lovelace AO, Jia Y et al (2008) Characterization of mosquito-adapted West Nile virus. J Gen Virol 89:1633–1642. https://doi.org/10.1099/vir.0.2008/000893-0
Colmant AMG, Hobson-Peters J, Slijkerman TAP et al (2021) Insect-specific Flavivirus replication in mammalian cells is inhibited by physiological temperature and the zinc-finger antiviral protein. Viruses 13:573. https://doi.org/10.3390/v13040573
Cook S, Holmes EC (2006) A multigene analysis of the phylogenetic relationships among the Flaviviruses (Family: Flaviviridae) and the evolution of vector transmission. Arch Virol 151:309–325. https://doi.org/10.1007/s00705-005-0626-6
Cook S, Moureau G, Kitchen A et al (2012) Molecular evolution of the insect-specific Flaviviruses. J Gen Virol 93:223–234. https://doi.org/10.1099/vir.0.036525-0
Cugola FR, Fernandes IR, Russo FB et al (2016) The Brazilian zika virus strain causes birth defects in experimental models. Nature 534:267–271. https://doi.org/10.1038/nature18296
Davies TJ, Pedersen AB (2008) Phylogeny and geography predict pathogen community similarity in wild primates and humans. Proc R Soc B Biol Sci 275:1695–1701. https://doi.org/10.1098/rspb.2008.0284
Deardorff ER, Fitzpatrick KA, Jerzak GVS et al (2011) West Nile virus experimental evolution in vivo and the trade-off hypothesis. PLoS Pathog 7:e1002335. https://doi.org/10.1371/journal.ppat.1002335
Decaro N, Mari V, Lucente MS et al (2013) Detection of a Hobi-like virus in archival samples suggests circulation of this emerging pestivirus species in Europe prior to 2007. Vet Microbiol 167:307–313. https://doi.org/10.1016/j.vetmic.2013.09.006
Delwart EL (2007) Viral metagenomics. Rev Med Virol 17:115–131. https://doi.org/10.1002/rmv.532
Di Giallonardo F, Schlub TE, Shi M, Holmes EC (2017) Dinucleotide composition in animal RNA viruses is shaped more by virus family than by host species. J Virol. https://doi.org/10.1128/jvi.02381-16
Di Paola N, De Melo Freire CC, De Andrade Zanotto PM (2018) Does adaptation to vertebrate codon usage relate to Flavivirus emergence potential? PLoS ONE 13:e0191652. https://doi.org/10.1371/journal.pone.0191652
Dobson A, Jones L, Gould E (1995) Persistence and transmission of tick-borne viruses: ixodes ricinus and louping-Ill virus in red grouse populations. Parasitology 111:S49–S58. https://doi.org/10.1017/S0031182000075818
Domingo E, Sheldon J, Perales C (2012) Viral quasispecies evolution. Microbiol Mol Biol Rev 76:159–216. https://doi.org/10.1128/mmbr.05023-11
EIGEN M, (1996) On the nature of virus quasispecies. Trends Microbiol 4:216–218. https://doi.org/10.1016/0966-842X(96)20011-3
Elrefaey AM, Abdelnabi R, Rosas ALR et al (2020) Understanding the mechanisms underlying host restriction of insect-specific viruses. Viruses 12:1–19. https://doi.org/10.3390/v12090964
Erasmus JH, Auguste AJ, Kaelber JT et al (2017) A chikungunya fever vaccine utilizing an insect-specific virus platform. Nat Med 23:192–199. https://doi.org/10.1038/nm.4253
Filomatori CV, Lodeiro MF, Alvarez DE et al (1995) A 5 RNA element promotes dengue virus RNA synthesis on a circular genome. Sullivan Ahlquist 20:2238–2249. https://doi.org/10.1101/gad.1444206
Firth AE, Blitvich BJ, Wills NM et al (2010) Evidence for ribosomal frameshifting and a novel overlapping gene in the genomes of insect-specific Flaviviruses. Virology 399:153–166. https://doi.org/10.1016/j.virol.2009.12.033
Firth AE, Brierley I (2012) Non-canonical translation in RNA viruses. J Gen Virol 93:1385–1409. https://doi.org/10.1099/vir.0.042499-0
Franz A, Kantor A, Passarelli A, Clem R (2015) Tissue barriers to arbovirus infection in mosquitoes. Viruses 7:3741–3767. https://doi.org/10.3390/v7072795
Frisbie B, Tang YW, Griffin M et al (2004) Surveillance of childhood influenza virus infection: what is the best diagnostic method to use for archival samples? J Clin Microbiol 42:1181–1184. https://doi.org/10.1128/JCM.42.3.1181-1184.2004
Ghai RR, Carpenter A, Liew AY et al (2021) Animal reservoirs and hosts for emerging alphacoronaviruses and betacoronaviruses. Emerg Infect Dis 27:1015–1022. https://doi.org/10.3201/eid2704.203945
Grant A, Ponia SS, Tripathi S et al (2016) Zika virus targets human STAT2 to inhibit type i interferon signaling. Cell Host Microbe 19:882–890. https://doi.org/10.1016/j.chom.2016.05.009
Guarneri T, Rossetti L, Giraffa G (2001) Rapid identification of Lactobacillus brevis using the polymerase chain reaction. Lett Appl Microbiol 33:377–381. https://doi.org/10.1046/j.1472-765X.2001.01014.x
Gubler DJ, Kuno G, Sather GE et al (1984) Mosquito cell cultures and specific monoclonal antibodies in surveillance for dengue viruses. Am J Trop Med Hyg 33:158–165. https://doi.org/10.4269/ajtmh.1984.33.158
Halbach R, Junglen S, van Rij RP (2017) Mosquito-specific and mosquito-borne viruses: evolution, infection, and host defense. Curr Opin Insect Sci 22:16–27. https://doi.org/10.1016/j.cois.2017.05.004
Hall-Mendelin S, McLean BJ, Bielefeldt-Ohmann H et al (2016) The insect-specific Palm Creek virus modulates West Nile virus infection in and transmission by Australian mosquitoes. Parasites Vectors 9:1–10. https://doi.org/10.1186/s13071-016-1683-2
Han JQ, Wroblewski G, Xu Z et al (2004) Sensitivity of hepatitis C virus RNA to the antiviral enzyme ribonuclease L is determined by a subset of efficient cleavage sites. J Interferon Cytokine Res 24:664–676. https://doi.org/10.1089/jir.2004.24.664
Hermance ME, Thangamani S (2015) Tick saliva enhances powassan virus transmission to the host, influencing its dissemination and the course of disease. J Virol 89:7852–7860. https://doi.org/10.1128/jvi.01056-15
Huhtamo E, Cook S, Moureau G et al (2014) Novel Flaviviruses from mosquitoes: mosquito-specific evolutionary lineages within the phylogenetic group of mosquito-borne Flaviviruses. Virology 464–465:320–329. https://doi.org/10.1016/j.virol.2014.07.015
Huhtamo E, Putkuri N, Kurkela S et al (2009) Characterization of a novel Flavivirus from mosquitoes in Northern Europe that Is related to mosquito-borne Flaviviruses of the tropics. J Virol 83:9532–9540. https://doi.org/10.1128/jvi.00529-09
Innis BL, Nisalak A, Nimmannitya S et al (1989) An enzyme-linked immunosorbent assay to characterize dengue infections where dengue and Japanese encephalitis co-circulate. Am J Trop Med Hyg 40:418–427. https://doi.org/10.4269/ajtmh.1989.40.418
Jenkins GM, Holmes EC (2003) The extent of codon usage bias in human RNA viruses and its evolutionary origin. Virus Res 92:1–7. https://doi.org/10.1016/S0168-1702(02)00309-X
Jerzak GVS, Brown I, Shi PY et al (2008) Genetic diversity and purifying selection in West Nile virus populations are maintained during host switching. Virology 374:256–260. https://doi.org/10.1016/j.virol.2008.02.032
Junglen S, Kopp A, Kurth A et al (2009) A New Flavivirus and a new vector: characterization of a novel Flavivirus isolated from uranotaenia mosquitoes from a tropical rain forest. J Virol 83:4462–4468. https://doi.org/10.1128/jvi.00014-09
Junglen S, Korries M, Grasse W et al (2017) Host range restriction of insect-specific Flaviviruses occurs at several levels of the viral life cycle. Sphere. https://doi.org/10.1128/msphere.00375-16
Kandimalla ER, Zhu FG, Bhagat L et al (2003) Toll-like receptor 9: modulation of recognition and cytokine induction by novel synthetic CpG DNAs. Biochem Soc Trans 31:654–658. https://doi.org/10.1042/BST0310654
Karlin S, Mrázek J (1997) Compositional differences within and between eukaryotic genomes. Proc Natl Acad Sci 94:10227–10232. https://doi.org/10.1073/pnas.94.19.10227
Kassen R (2002) The experimental evolution of specialists, generalists, and the maintenance of diversity. J Evol Biol 15:173–190
Katoh K, Misawa K, Kuma KI, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30:3059–3066. https://doi.org/10.1093/nar/gkf436
Kent RJ, Crabtree MB, Miller BR (2010) Transmission of West Nile virus by Culex quinquefasciatus say infected with culex Flavivirus izabal. PLoS Negl Trop Dis 4:e671. https://doi.org/10.1371/journal.pntd.0000671
Kotál J, Langhansová H, Lieskovská J et al (2015) Modulation of host immunity by tick saliva. J Proteom 128:58–68. https://doi.org/10.1016/j.jprot.2015.07.005
Kreuder Johnson C, Hitchens PL, Smiley Evans T et al (2015) Spillover and pandemic properties of zoonotic viruses with high host plasticity. Sci Rep 5:14830. https://doi.org/10.1038/srep14830
Lanciotti RS, Kerst AJ, Nasci RS et al (2000) Rapid detection of West Nile virus from human clinical specimens, field-collected mosquitoes, and avian samples by a TaqMan reverse transcriptase-PCR assay. J Clin Microbiol 38:4066–4071. https://doi.org/10.1128/jcm.38.11.4066-4071.2000
Lanciotti RS, Kerst AJ (2001) Nucleic acid sequence-based amplification assays for rapid detection of West Nile and St Louis Encephalitis viruses. J Clin Microbiol 39:4506–4513. https://doi.org/10.1128/JCM.39.12.4506-4513.2001
Le TK, Vinh LS (2020) FLAVI: an amino acid substitution model for Flaviviruses. J Mol Evol 88:445–452. https://doi.org/10.1007/s00239-020-09943-3
Leiby DA, Gill JE (2004) Transfusion-transmitted tick-borne infections: a cornucopia of threats. Transfus Med Rev 18:293–306. https://doi.org/10.1016/j.tmrv.2004.07.001
Li W, Godzik A (2006) Cd-hit: a fast program for clustering and comparing large sets of protein or nucleotide sequences. Bioinformatics 22:1658–1659. https://doi.org/10.1093/bioinformatics/btl158
Li CX, Shi M, Tian JH et al (2015) Unprecedented genomic diversity of RNA viruses in arthropods reveals the ancestry of negative-sense RNA viruses. Elife. https://doi.org/10.7554/eLife.05378
Lin R-J, Chang B-L, Yu H-P et al (2006) Blocking of interferon-induced Jak-Stat signaling by Japanese encephalitis Virus NS5 through a protein tyrosine phosphatase-mediated mechanism. J Virol 80:5908–5918. https://doi.org/10.1128/jvi.02714-05
Lobo FP, Mota BEF, Pena SDJ et al (2009) Virus-host coevolution: Common patterns of nucleotide motif usage in Flaviviridae and their hosts. PLoS ONE. https://doi.org/10.1371/journal.pone.0006282
Luo X, Wang X, Gao Y et al (2020) Molecular mechanism of RNA recognition by zinc-finger antiviral protein. Cell Rep 30:46-52.e4. https://doi.org/10.1016/j.celrep.2019.11.116
Manzano M, Reichert ED, Polo S et al (2011) Identification of cis-acting elements in the 3′-untranslated region of the dengue virus type 2 RNA that modulate translation and replication. J Biol Chem 286:22521–22534. https://doi.org/10.1074/jbc.M111.234302
Marklewitz M, Zirkel F, Kurth A et al (2015) Evolutionary and phenotypic analysis of live virus isolates suggests arthropod origin of a pathogenic RNA virus family. Proc Natl Acad Sci 112:7536–7541. https://doi.org/10.1073/pnas.1502036112
Meagher JL, Takata M, Gonçalves-Carneiro D et al (2019) Structure of the zinc-finger antiviral protein in complex with RNA reveals a mechanism for selective targeting of CG-rich viral sequences. Proc Natl Acad Sci 116:24303–24309. https://doi.org/10.1073/pnas.1913232116
Melian EB, Hall-Mendelin S, Du F et al (2014) Programmed ribosomal frameshift alters expression of West Nile virus genes and facilitates virus replication in birds and mosquitoes. PLoS Pathog 10:e1004447. https://doi.org/10.1371/journal.ppat.1004447
Michel F, Sieg M, Fischer D et al (2019) Evidence for West Nile virus and Usutu virus infections in wild and resident birds in Germany, 2017 and 2018. Viruses 11:674. https://doi.org/10.3390/v11070674
Miesen P, Joosten J, van Rij RP (2016) PIWIs go viral: arbovirus-derived pirnas in vector mosquitoes. PLOS Pathog 12:e1006017. https://doi.org/10.1371/journal.ppat.1006017
Mihara T, Nishimura Y, Shimizu Y et al (2016) Linking virus genomes with host taxonomy. Viruses. https://doi.org/10.3390/v8030066
Minh BQ, Schmidt HA, Chernomor O et al (2020) IQ-TREE 2: new models and efficient methods for phylogenetic inference in the genomic era. Mol Biol Evol 37:1530–1534. https://doi.org/10.1093/molbev/msaa015
Mollentze N, Streicker DG (2020) Viral zoonotic risk is homogenous among taxonomic orders of mammalian and avian reservoir hosts. Proc Natl Acad Sci 117:9423–9430. https://doi.org/10.1073/pnas.1919176117
Moureau G, Cook S, Lemey P et al (2015) New insights into Flavivirus evolution, taxonomy and biogeographic history, extended by analysis of canonical and alternative coding sequences e0117849. PLoS ONE 10:e0117849. https://doi.org/10.1371/journal.pone.0117849
Murray CL, Jones CT, Rice CM (2008) Architects of assembly: roles of Flaviviridae non-structural proteins in virion morphogenesis. Nat Rev Microbiol 6:699–708. https://doi.org/10.1038/nrmicro1928
Murray KO, Walker C, Gould E (2011) The virology, epidemiology, and clinical impact of West Nile virus: a decade of advancements in research since its introduction into the Western Hemisphere. Epidemiol Infect 139:807–817
Ng W, Soto-Acosta R, Bradrick S et al (2017) The 5′ and 3′ untranslated regions of the Flaviviral genome. Viruses 9:137. https://doi.org/10.3390/v9060137
Ngo KA, Rose JT, Kramer LD, Ciota AT (2019) Adaptation of Rabensburg virus (RBGV) to vertebrate hosts by experimental evolution. Virology 528:30–36. https://doi.org/10.1016/j.virol.2018.11.015
Novella IS, Presloid JB, Smith SD, Wilke CO (2012) Specific and nonspecific host adaptation during arboviral experimental evolution. J Mol Microbiol Biotechnol 21:71–81. https://doi.org/10.1159/000332752
Nuttall PA (2014) Tick-borne viruses. In: Biol. Ticks, Vol. 2, eds D. E. Sonenshine R. M. Roe (Oxford University Press, New York, NY). https://global.oup.com/academic/product/biology-of-ticks-volume-2-9780199744060?cc=illang=en&lang=en&
Ochsenreiter R, Hofacker IL, Wolfinger MT (2019) Functional RNA structures in the 3’UTR of tick-borne, insect-specific and no-known-vector Flaviviruses. Viruses 11:298. https://doi.org/10.3390/v11030298
Odon V, Fros JJ, Goonawardane N et al (2019) The role of ZAP and OAS3/RNAseL pathways in the attenuation of an RNA virus with elevated frequencies of CpG and UpA dinucleotides. Nucleic Acids Res 47:8061–8083. https://doi.org/10.1093/nar/gkz581
Offerdahl DK, Clancy NG, Bloom ME (2016) Stability of a tick-borne Flavivirus in milk. Front Bioeng Biotechnol. https://doi.org/10.3389/fbioe.2016.00040
Olival KJ, Hosseini PR, Zambrana-Torrelio C et al (2017) Host and viral traits predict zoonotic spillover from mammals. Nature 546:646–650. https://doi.org/10.1038/nature22975
Pallarés HM, Costa Navarro GS, Villordo SM et al (2020) Zika virus subgenomic Flavivirus RNA generation requires cooperativity between duplicated RNA structures that are essential for productive infection in human cells. J Virol 94:343–363. https://doi.org/10.1128/jvi.00343-20
Pandit PS, Doyle MM, Smart KM et al (2018) Predicting wildlife reservoirs and global vulnerability to zoonotic Flaviviruses. Nat Commun 9:1–10. https://doi.org/10.1038/s41467-018-07896-2
Pickett BE, Sadat EL, Zhang Y et al (2012) ViPR: an open bioinformatics database and analysis resource for virology research. Nucleic Acids Res. https://doi.org/10.1093/nar/gkr859
Schoch CL, Ciufo S, Domrachev M et al (2020) NCBI taxonomy: a comprehensive update on curation, resources and tools. Database. https://doi.org/10.1093/database/baaa062
Sexton NR, Ebel GD (2019) Effects of arbovirus multi-host life cycles on dinucleotide and codon usage patterns. Viruses 11:643
Shah-Hosseini N, Chinikar S, Ataei B et al (2014) Phylogenetic analysis of West Nile virus genome. Iran Emerg Infect Dis 20:1419–1421
Sharp PM, Li WH (1987) The codon adaptation index-a measure of directional synonymous codon usage bias, and its potential applications. Nucleic Acids Res 15:1281–1295. https://doi.org/10.1093/nar/15.3.1281
Sharp PM, Tuohy TMF, Mosurski KR (1986) Codon usage in yeast: cluster analysis clearly differentiates highly and lowly expressed genes. Nucleic Acids Res 14:5125–5143. https://doi.org/10.1093/nar/14.13.5125
Shi M, Lin XD, Tian JH et al (2016) Redefining the invertebrate RNA virosphere. Nature 540:539–543. https://doi.org/10.1038/nature20167
Shin YC, Bischof GF, Lauer WA, Desrosiers RC (2015) Importance of codon usage for the temporal regulation of viral gene expression. Proc Natl Acad Sci 112:14030–14035. https://doi.org/10.1073/pnas.1515387112
Simmonds P (2015) Methods for virus classification and the challenge of incorporating metagenomic sequence data. J Gen Virol 96:1193–1206. https://doi.org/10.1099/jgv.0.000016
Swanepoel R, Shepherd AJ, Leman PA, Miller GB (1985) A common-source outbreak of Crimean-Congo haemorrhagic fever on a dairy farm. South Afr Med J 68:635–637. https://doi.org/10.10520/AJA20785135_6416
Takata MA, Gonçalves-Carneiro D, Zang TM et al (2017) CG dinucleotide suppression enables antiviral defence targeting non-self RNA. Nature 550:124–127. https://doi.org/10.1038/nature24039
Thomas SJ, Yoon IK (2019) A review of Dengvaxia®: development to deployment. Hum Vaccines Immunother 15:2295–2314. https://doi.org/10.1080/21645515.2019.1658503
Tria FDK, Landan G, Dagan T (2017) Phylogenetic rooting using minimal ancestor deviation. Nat Ecol Evol 1:193. https://doi.org/10.1038/s41559-017-0193
Troost B, Smit JM (2020) Recent advances in antiviral drug development towards dengue virus. Curr Opin Virol 43:9–21. https://doi.org/10.1016/j.coviro.2020.07.009
Vasilakis N, Deardorff ER, Kenney JL et al (2009) Mosquitoes put the brake on arbovirus evolution: experimental evolution reveals slower mutation accumulation in mosquito than vertebrate cells. PLoS Pathog 5:e1000467. https://doi.org/10.1371/journal.ppat.1000467
Villordo SM, Carballeda JM, Filomatori CV, Gamarnik AV (2016) RNA structure duplications and Flavivirus host adaptation. Trends Microbiol 24:270–283. https://doi.org/10.1016/j.tim.2016.01.002
Villordo SM, Filomatori CV, Sánchez-Vargas I et al (2015) Dengue virus RNA structure specialization facilitates host adaptation. PLOS Pathog 11:e1004604. https://doi.org/10.1371/journal.ppat.1004604
Weaver SC, Barrett ADT (2004) Transmission cycles, host range, evolution and emergence of arboviral disease. Nat Rev Microbiol 2:789–801. https://doi.org/10.1038/nrmicro1006
Whelan S, Goldman N (2001) A general empirical model of protein evolution derived from multiple protein families using a maximum-likelihood approach. Mol Biol Evol 18:691–699. https://doi.org/10.1093/oxfordjournals.molbev.a003851
Wilke C (2001) Dynamic fitness landscapes in molecular evolution. Phys Rep 349:395–446. https://doi.org/10.1016/S0370-1573(00)00118-6
Wilke CO (2005) Quasispecies theory in the context of population genetics. BMC Evol Biol. https://doi.org/10.1186/1471-2148-5-44
Wilson DS, Yoshimura J (1994) On the coexistence of specialists and generalists. Am Nat 144:692–707. https://doi.org/10.1086/285702
World health Organization W (2020) Dengue and severe dengue. World health Organization, Geneva
Wright F (1990) The “effective number of codons” used in a gene. Gene 87:23–29. https://doi.org/10.1016/0378-1119(90)90491-9
Xia H, Luo H, Shan C et al (2018) An evolutionary NS1 mutation enhances Zika virus evasion of host interferon induction. Nat Commun 9:1–13. https://doi.org/10.1038/s41467-017-02816-2
Zhong C, Chen C, Wang L, Ning K (2021) Integrating pan-genome with metagenome for microbial community profiling. Comput Struct Biotechnol J 19:1458–1466. https://doi.org/10.1016/j.csbj.2021.02.021
Zhou D, Jia F, Li Q et al (2018) Japanese encephalitis virus NS1′ protein antagonizes interferon beta production. Virol Sin 33:515–523. https://doi.org/10.1007/s12250-018-0067-5
Funding
This study was supported in part by a fellowship to KH from the Edmond J. Safra Center for Bioinformatics at Tel Aviv University and by the Koret-UC Berkeley-Tel Aviv University Initiative in Computational Biology and Bioinformatics to IM.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The author declares no conflict of interest.
Additional information
Handling editor: David Liberles
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Halabi, K., Mayrose, I. Mechanisms Underlying Host Range Variation in Flavivirus: From Empirical Knowledge to Predictive Models. J Mol Evol 89, 329–340 (2021). https://doi.org/10.1007/s00239-021-10013-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-021-10013-5