Abstract
Transthyretin was discovered in the 1940s, named after its ability to bind thyroid hormones and retinol. In the genomic era, transthyretins were found to be part of a larger family with homologs of no obvious function, then called transthyretin-related proteins. Thus, it was proposed that the transthyretin gene could be the result of gene duplication of an ancestral of this newly identified homolog, later found out to be an enzyme involved in uric acid degradation, then named HIUase (5-hydroxy-isourate hydrolase). Here, we sought to re-enact the evolutionary history of this protein family by reconstructing, from a phylogeny inferred from 123 vertebrate sequences, three ancestors corresponding to key moments in their evolution—before duplication; the common transthyretin ancestor after gene duplication and the common ancestor of Eutheria transthyretins. Experimental and computational characterization showed the reconstructed ancestor before duplication was unable to bind thyroxine and likely presented the modern HIUase reaction mechanism, while the substitutions after duplication prevented that activity and were enough to provide stable thyroxine binding, as confirmed by calorimetry and x-ray diffraction. The Eutheria transthyretin ancestor was less prone to characterization, but limited data suggested thyroxine binding as expected. Sequence/structure analysis suggests an early ability to bind the Retinol Binding Protein. We solved the X-ray structures from the two first ancestors, the first at 1.46 resolution, the second at 1.55 resolution with well-defined electron density for thyroxine, providing a useful tool for the understanding of structural adaptation from enzyme to hormone distributor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Transthyretins
Thyroid hormones (TH) are fundamental for body growth, development, and metabolism. The circulating TH in plasma is thyroxine (T4), which through action of deiodinases, is converted into its active form, triiodothyronine (T3) (Köhrle 2007). While the main TH carrier protein in blood plasma is the thyroxine binding globulin (TBG), followed by transthyretin (TTR) and albumin, in cerebrospinal fluid (CSF) TTR is the main T4 carrier, accounting for 80% of its transport (Hagen and Elliott 1973).
Discovered in 1942 as a protein present in CSF and in human plasma (Seibert and Nelson 1942; Kabat et al. 1942), TTR was initially called pre-albumin due to its migration pattern in electrophoresis gel, running faster than albumin. Years later it was found to be involved in transport of TH and retinol (vitamin A1) and its name was then changed to transthyretin, which reflects its then known functionalities. TTR was first described in Eutheria (placental mammals) as a T4 carrier protein, but in other vertebrates it has a higher affinity for the active form of this hormone (T3), a feature first observed in a bullfrog homolog (Yamauchi et al. 1993). The TTR crystallographic structure was initially determined in the seventies (Blake et al. 1971, 1974), and the Protein Data Bank (PDB) currently presents hundreds of transthyretin structures from various vertebrates, while it appears to be absent in invertebrates.
In addition to carrying THs, TTR also participates in the retinol transport through the formation of the TTR-RBP (retinol binding protein) complex (Kanai et al. 1968; Monaco et al. 1995), thus avoiding loss of retinol and RBP in kidneys via glomerular filtration (Monaco 2009). Furthermore, some studies have shown that transthyretin can bind to retinoic acid (Smith et al. 1994), flavonoids (Lueprasitsakul et al. 1990) and associate with high-density lipoprotein (HDL) through binding to apolipoprotein A-I (apoA-I), which is also considered a substrate for possible TTR peptidase activity. This would characterize it as a zinc-dependent metalloprotease (Liz et al. 2012) made possible by the structural changes observed upon metal binding (De Palmieri et al. 2010).
Another transthyretin function is related to sequestration of the Aβ peptide in CSF, forming stable complexes with TTR (Schwarzman et al. 1994). Although there are other proteins capable of binding to the Aβ peptide, TTR is the main protein that acts on β-amyloid peptide modulation in CSF, preventing amyloidosis and Alzheimer's disease (Riisøen 1988; Schwarzman et al. 1994). Other roles in nervous system include participation in sensor-motor function and improvement in nerve regeneration (Riisøen 1988), increase in exploratory capacity in TTR knockout mice and reduction of depression behaviour (Sousa et al. 2004), neuropeptide processing (Nunes et al. 2006), and maintenance of memory-related processes during aging (Brouillette and Quirion 2008).
Mutations in TTR can cause several types of diseases, from hyperthyroxinemia caused by increased thyroxine affinity to neurodegenerative diseases related to the formation of amyloid fibrils (Refetoff et al. 1996; Saraiva 2001). Such diseases include senile systemic amyloidosis (SSA), familial amyloid cardiomyopathy (FAC), leptomeningeal amyloidosis (LA) and familial amyloid polyneuropathy (FAP).
Three-Dimensional Structure
Transthyretin is a homotetrameric protein with a molecular mass of approximately 55 KDa, each of the four subunits having around 14 KDa. In human TTR, each monomer is composed of 127 amino acids arranged in eight β-strands, named A-H, which are connected by seven loops and divided into two β-sheets, forming a β-sandwich. The two β-sheets are composed of four antiparallel β-strands called DAGH and CBEF, and the structure also presents a small α-helix composed of nine residues between β-bands E and F (Blake et al. 1974).
TTR dimers are composed of two pairs of β-sheets, one internal called DAGHH'G'A'D' and one external called CBEFF'E'B'C' (Blake et al. 1974). Each monomer interacts predominantly by hydrogen bonds between the adjacent H–H 'and F–F' β-strands, forming the dimer. Formation of the tetrameric structure occurs through interaction of two dimers, AB and CD, predominantly by hydrophobic contact between residues located in two loops: A–B and G–H (Blake et al. 1974).
Tetramerization of transthyretin leads to formation of a broad hydrophobic channel through the protein, approximately 8 Å in diameter and 50 Å in length (Blake and Oatley 1977). In this channel there is one site of thyroid hormone binding on each end. However, under physiological conditions, only one of the sites is occupied due to a negative cooperativity between the sites (Ferguson et al. 1975; Neumann et al. 2001).
Transthyretin-Related Proteins
In 2000, transthyretin-like sequences were found in open reading frames from Caenorhabditis elegans and several microorganisms (Prapunpoj et al. 2000). Subsequently, it was proposed that the TTR gene could be the result of a gene duplication event of an ancestral of this newly identified homolog, which have been transthyretin-related protein (TRP). To test this hypothesis, TTR and TRP sequences were analysed and it was demonstrated that they shared motifs that could be mapped in structurally and functionally important regions of TTR (Hennebry et al. 2006b).
In 2001, a role for TRPs in purine catabolism was proposed: Schultz et al. generated a series of mutants of Bacillus subtilis with a mutation in TRP gene, which is located near the uricase gene (Schultz et al. 2001). These bacteria were then cultured in medium containing only uric acid as a source of nitrogen and the bacteria with this mutation had a low proliferation rate compared to wild bacteria, evidencing a possible role of TRP in a purine degradation pathway. In 2006, the conversion mechanism of 5-hydroxyisourate (5-HIU) to 2-oxo-4-hydroxy-4-carboxy-5-ureidoimidazoline (OHCU) was elucidated for Bacillus subtilis (Jung et al. 2006) and TRP was renamed HIUase (5-hydroxy-isourate hydrolase).
Like TTR, each HIUase monomer is composed of the same eight anti-parallel β-strands (A-H) and a small α-helix (Hennebry et al. 2006a). Two monomers form dimers by means of interactions between the main chain of amino acids in the H and F ß-strands. The tetramer is formed by hydrophobic interactions between the AB and GH loops of two dimers, leading to the formation of a channel similar to that of TTR. In addition to high structural similarity, they have considerable identity values. Human transthyretin and HIUase from snapper fish, for example, exhibit 48% identity (Hennebry 2009).
In the proposed mechanism for HIUase, a water molecule donates a proton to a histidine (His14) located at the channel entrance. After being deprotonated, it makes a nucleophilic attack on the C6 carbon of 5-hydroxy-isourate. The oxyanion resulting from nucleophilic attack is negatively charged and stabilized by Arg49. An electron rearrangement in the oxyanion leads to ring rupture at the C6-N1 bond, probably by extracting a proton from the guanidine group of Arg49. The catalytic cycle is completed by a proton transfer from His14 to Arg49, re-establishing the original configuration of the enzyme site. Site-directed mutagenesis studies confirms the crucial role of His14 and Arg49 residues for the enzyme, since mutants lost their catalytic activity (Jung et al. 2006). The same happens for Tyr118Phe mutant, suggesting that this residue may be interacting with 5-HIU’s O8, probably stabilizing 5-HIU position during catalysis, and may also be related to OHCU’s O2 tautomerization (Jung et al. 2006). We now know that four residues are fundamental to catalytic activity of this enzyme: His14, Arg49, His105, all located at the entrance of the channel, and residue Tyr118, positioned further to the bottom, closing the channel (Jung et al. 2006). On the same year, Hennebry et al. have independently shown by site directed mutagenesis upon their publication of the Salmonella dublin structure, that mutations on residues His6, His95 and Tyr108 also abolished catalytic activity.
Reconstructing the History of HIUases and Transthyretins
Evidence indicates that the TTR gene would have originated during the emergence of vertebrates, from the duplication of an ancestral of modern HIUases—enzymes present from bacteria to vertebrates (Ramazzina et al. 2006; Zanotti et al. 2006; Hennebry 2009; Kratzer et al. 2014). Given the same exon/intron structure of HIUase and TTR coding genes, following gene duplication, point mutations in the first coding exon could have led to changes in the peptide that in HIUase signals to peroxisomes, so that in TTR it signalled for cellular secretion (Zanotti et al. 2009). Such a proposition would agree with the fact that TTR expression occurs mainly in the liver, where the degradation of urate happens (Stevenson et al. 2010). However, there are known cases on which the first exon is not detected (Hennebry 2009), and so another possibility would be that the exon 1 region could be replaced by other peptide signal encoding exon by exon shuffling.
Subsequently, a series of mutations would have dramatically altered the protein function at the same time that the structure remained largely unchanged. Although only a few substitutions at the active site would supposedly be sufficient for conversion of HIUase to TTR, only after a fine tuning resulting from selection of several subsequent mutations has the thyroid hormone binding function been definitively established (Zanotti et al. 2006; Romero and Arnold 2009; Cendron et al. 2011).
Nonetheless, selective pressure would apparently have acted on the length and composition of the N-terminal portion of TTR by a series of individual base mutations, resulting in a shift of the intron 1 / exon 2 border in the 3 'direction (Aldred et al. 1997). This would have led to a gradual change in primary structure and, consequently, changed affinity of TTR from T3 to T4 (Chang et al. 1999; Eneqvist et al. 2004; Prapunpoj et al. 2006; Morgado et al. 2008; Prapunpoj and Leelawatwattana 2009; Kasai et al. 2018).
Resurrecting Ancient Proteins
In this work, we aim to re-enact the history of modern transthyretins by experimentally characterizing transthyretins ancestors in key moments in their evolution. Ancestral sequence reconstruction studies consist of synthesis, expression and experimental characterization of hypothetical ancestral proteins inferred from large alignments of contemporary homologous protein sequences using phylogenetic methods (Pauling et al. 1963; Thornton 2004; Perez-Jimenez et al. 2011; Morrow et al. 2017). This approach has been benchmarked experimentally (Randall et al. 2016) and has been successfully applied in many different cases (Risso et al. 2013; Kratzer et al. 2014; Wilson et al. 2015). This includes those similar to HIUase / TTR (Huang et al. 2012), having demonstrated, for example, that the structure of certain proteins is preserved for billions of years (Ingles-Prieto et al. 2013). Therefore, the ancestral protein reconstruction method is ideal for experimental studies of molecular evolution and divergence of protein function. Here, we describe the sequence reconstruction of three ancestors of modern transthyretins and characterize them to understand how they evolved over time.
Results
We synthesized genes based on ancestral sequence reconstruction data of three different timepoints in HIUase-TTR evolution as shown in Fig. 1. The first protein expressed from such genes, to be referred to as HIUase-TTR ancestor, corresponds to the sequence reconstruction of the node immediately before gene duplication. The second protein, named TTR ancestor is a reconstruction of the node right after duplication, corresponding to the ancestor of all contemporary transthyretins. Finally, the Eutheria TTR ancestor is a reconstruction of the ancestral node of the clade containing all Eutheria transthyretins. All the three nodes of phylogenetic tree taken for ancestral sequences reconstruction have support (posterior probability) of 100% (the original consensus tree file including support values for all nodes is included as supporting material).
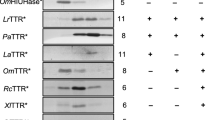
Phylogeny used for ancestral sequence reconstruction. Nodes marked in circles labelled H–T, T and ET refer to the points used for ancestral reconstruction for HIUase-TTR, TTR Ancestor and Eutheria TTR Ancestor. Unlabelled nodes marked with black circles show posterior probabilities above 90%. Background colours refer to the division between vertebrate HIUases (blue) and transthyretins (yellow) (Color figure online)
A comparison of the three reconstructed ancestors and human transthyretin can be seen in the multiple sequence alignment shown in Fig. 2.
Multiple sequence alignment for HIUase and TTR proteins. Sequences include human transthyretin (hTTR, Uniprot: P02766), the reconstructed ancestors for Eutheria transthyretins (TTR_Eutheria), all transthyretins (AncTTR) and the transthyretin/HIUase ancestor before the gene duplication event (HIUase-TTR). Positions marked in asterisks (*) refer to key positions for HIUase function. Numbers on top of the sequences refer to the HIUase-TTR ancestor
To study the three-dimensional structure of TTR ancestors and their binding to T4, the three proteins were expressed in E. coli, purified, crystallized, subjected to x-ray diffraction and isothermal titration calorimetry (ITC) experiments (see Methods).
The HIUase-TTR ancestor crystal diffracted to 1.46 Å. In Fig. 3a and b, its three-dimensional structure is superposed to that of human transthyretin (PDB: 2ROX). On a cartoon representation, which considers the backbones only, the two proteins seem to have changed little over such a long period of time. However, changes in side chains modify the available volume in the channel (being bulkier in transthyretin as seen in Fig. 3c and d), which means less space for a molecule like T4. Indeed, we detected no T4 binding to the HIUase-TTR ancestor from the ITC experiments and attempts to crystallize the protein with T4 yielded no electron density inside the tunnel. The overall structure of the HIUase-TTR ancestor is similar to modern HIUases, especially on the binding pocket, as can be seen in its superposition to proteins such as the HIUase from S. dublin (Hennebry et al. 2006a), as shown in Fig. 3e. A superposition of thyroxine as bound to human transthyretin to H–T (Fig. 3f), which is incompatible with this binding pocket, further supports the hypothesis that the ability of thyroid hormone binding was not available before the lineage of proteins derived from gene duplication.
a HIUase-TTR ancestor (salmon) superposed to human TTR (yellow). b Same as A upon a 180° rotation through the vertical axis. c Available volume in the HIUase-TTR tunnel represented as cyan superposed spheres, calculated using CAVER (Jurcik et al. 2018). d Same as C for human transthyretin. e HIUase-TTR ancestor (salmon) superposed with the HIUase from S. Dublin (green). f The HIUase-TTR binding pocket superposed with thyroxine as positioned in human TTR. Figure prepared with PyMOL (DeLano 2007; Schrödinger 2010) (Color figure online)
As opposed to the HIUase-TTR ancestor, the TTR ancestor has shown T4 binding in isothermal titration calorimetry (ITC) experiments as seen in Fig. 4. Its adjusted KD was 2.33 μM, i.e., larger than that observed for human transthyretin (0.36 μM).
The TTR ancestor was also subjected to crystallization trials in the presence of T4. The best available crystal diffracted to 1.55 Å and this time an interpretable electron density was observed in both tunnel sides, which could be modelled as T4 molecules. A superposition of the TTR ancestor to human transthyretin can be seen in Fig. 5.
Attempts to characterize the Eutheria TTR ancestor were also made. This construct expressed very poorly in E. coli and was not prone to crystallization. Although it was possible to obtain an ITC profile indicating binding to T4, results were not sufficiently reproducible in different conditions. For this reason we included such results as supplementary material only.
Docking
To analyze the potential interactions among 5-HIU and the protein binding sites, we docked 5-HIU and the high energy intermediate (HIU_HEI) of the hydrolysis reaction against three protein structures: Bacillus subtilis PucM (PDB 2H0F), HIUase-TTR and TTR ancestor. Analysis of the best scoring binding modes suggests a very good fit between 5-HIU and the binding sites of PucM and HIUase-TTR, as indicated by good shape complementarity and the presence of many polar interactions involving all polar atoms of 5-HIU (Fig. 6). For PucM, hydrogen bonds were predicted for residues His14, Arg49, His105, Tyr 118 and Ser121 (PDB 2H0F numbering) and the same structurally aligned residues from HIUase-TTR (Supplementary Table S1). Among those, site-directed mutagenesis with a homologous protein indicated that His14, His105 and Tyr118 play an important role in catalysis (Hennebry et al. 2006a). The best scoring binding modes were very conserved for 5-HIU and HIU_HEI, with additional hydrogen bonds involving the tetrahedral OH present in HIU_HEI and Arg49. This is in agreement with the reaction mechanism previously proposed by Jung et al., who hypothesized that the oxyanion formed during the reaction could be stabilized by the active site arginine (Jung et al. 2006).
Predicted binding modes for 5-HIU and HIU_HEI. Poses predicted in the binding site of: PucM (PDB 2H0F) for 5-HIU (a) and HIU_HEI (b); HIUase-TTR for -5HIU (c) and HIU_HEI (d); and TTR ancestor, for 5-HIU (e) and HIU_HEI (f). Docking was performed with the program Glide SP precision. Hydrogen bonds are represented as dashed lines. Figures prepared with PyMOL (DeLano 2007; Schrödinger 2010)
On the other hand, poor chemical complementarity was observed between 5-HIU and TTR ancestor (Fig. 6). This is reflected in worse total docking scores when compared to the results for the other proteins (Supplementary Table S2). Only four hydrogen bonds were predicted between ligand and residues Ser120 and Thr122, leaving most polar atoms of 5-HIU and HIU_HEI stranded. This result was expected, considering the low identity among this binding site and the other two proteins considered in this study. Key differences are observed, such as the absence of the catalytic histidine involved in the cleavage of 5-HIU. Other important residues, such as Arg49 and Tyr118, which are predicted to stabilize the binding of 5-HIU in HIUase-TTR, are also replaced by residues with different physicochemical properties in TTR ancestor.
Discussion
The distribution of transthyretin homologs among different taxa shows that it is a very old protein family, earlier than the divergence of eukaryotes and prokaryotes (Prapunpoj et al. 2000; Eneqvist et al. 2003). Context analysis of these genes and experimental characterization of their resulting proteins suggest that their earliest function was in uric acid catabolism. This was present until very recently in the human lineage, when primates lost the uricase gene, which was followed by the loss of the HIUase genes (Keebaugh and Thomas 2010).
However, one of the paralogs that resulted from gene duplication in chordates followed a process of neofunctionalization as commonly seen in events of duplication followed by divergence (Hennebry 2009). Our results show that the early substitutions after duplication were already sufficient for that protein to be a fully functioning thyroxine distributor.
The third known function of modern transthyretins, i.e. cryptic protease, was proposed to have developed much later in TTR evolution (Liz et al. 2012). In 2010, we observed that the alpha-helix between loop E and loop F, which is involved in holo-RBP recognition, undergoes a conformational change upon binding to zinc (De Palmieri et al. 2010). This finding enabled Liz and colleagues to explain why the protease activity of transthyretin is cryptic: this change makes possible the rearrangement of the proposed catalytic residues (His88, His90 and Glu92) in order to make a functional active site (Liz et al. 2012). They have also noted that the residues necessary for catalysis are present in humans and some other primates while most other organisms lack His90—from our reconstruction, that position is occupied by a tyrosine at least until Eutheria transthyretins.
It has been recently observed that the transthyretin from Crocodylus porosus also presents peptidase activity, although lower than human TTR (Leelawatwattana et al. 2016). This study has not assessed the role of zinc as in the former analysis from Liz and colleagues, but the authors noted that a tyrosine could also bind zinc, and that the protein terminals would also be involved in catalysis. A later publication from the same group showed proteolytic activity also in chicken transthyretin (Tola et al. 2019). Even though these studies on non-primate transthyretins did not include mutagenesis experiments to map which are the residues necessary for catalysis, they show that it is possible to present a significant activity without His90. Further experiments and investigation on other proteins would be necessary in order to assess if the lack of His90 is compensated by the presence of other residue(s), whether protease function evolved independently and if it could be or not cryptic in different taxa.
For the specific mechanism shown for human TTR (Liz et al. 2012), on which metal binding triggers a conformational change related to the formation of an active site, one could analyse the evolution of metal binding site residues in transthyretin. Three zinc binding sites have been found on human transthyretin (De Palmieri et al. 2010a): the one making up the proposed catalytic triad (Liz et al. 2012), a second one formed by Cys10 and His56 and a third one by Glu72/Asp74/His31. Those residues were all present in the TTR ancestral except for His31, where there is a lysine at least until the Eutheria TTR.
Yamauchi and Kasai have recently proposed that metal binding may be an ancient property in the HIUase/TTR family, by observing the ability of different members to bind to Ni-affinity columns in the absence of His-tags, as well as the observation of Zn2+ in different crystal structures (Yamauchi and Kasai 2018). As observed in that study, histidine binding residues observed in vertebrate and invertebrate HIUases are not the same ones seen in the zinc binding sites we previously described in human TTR (De Palmieri et al. 2010a), but they are present in the HIUase-TTR ancestor reconstructed in this work. The authors propose, based on their observations that T3 binding to different transthyretins is zinc-dependant, that such zinc-binding histidines would be related to transthyretin being a high-affinity thyroid hormone distributor in early vertebrate evolution. This hypothesis is in accordance with our reconstructed sequences—five histidines mentioned in the study appear in the Ancestral TTR sequence substituting Thr1, Pro55 and Gln101 (see Fig. 1) in the reconstructed HIUase-TTR ancestor, as well as inserted between residues 102–103 and in the C-terminal extension, but the ones in the N-terminal and C-terminal are lost in the reconstructed Eutheria ancestor already, and only the one substituting Pro55 is still present in human transthyretin.
Conclusions
In this article, we reconstructed the evolutionary history of the HIUase/transthyretin protein family, starting before the gene duplication that gave rise to modern transthyretins. By reconstructing ancestors in three evolutionary timepoints, we assessed how the various functions observed in this protein family likely arose over time, using various experimental and computational methodologies.
Depending on the species, the differences between modern HIUases and the HIUase-TTR ancestor reconstructed in this study varied greatly. The HIUase from Japanese rice fish (UniProt: A0A3B3I7P0_ORYLA) is 97.5% identical to the reconstructed ancestor, while the closest primate sequence found in a BLAST search (Altschul et al. 1990), that of tufted capuchin (RefSeq: XP_032140684) changed almost half of its residues, showing only 55.83% identity to HIUase-TTR. The closest structurally characterized HIUase is the one from zebrafish, at 69.23% identity. Yet, structural analysis and docking support an identical binding and catalytic mechanism for the HIUase-TTR ancestral.
Changes after gene duplication were far more dramatic. The TTR ancestral and HIUase-TTR share only 53.5% identity, with a longer C-terminal replacing the typical HIUase YRGS sequence. Yet these substitutions enabled it to bind to T4 very early in evolution, as shown by calorimetry and x-ray crystallography. A further 35 substitutions over a 124-residues overlap separates the TTR ancestral to its most likely sequence in Eutheria, which differs from human transthyretin by 15 additional substitutions. The substitution rate was clearly higher for transthyretins when compared to HIUases—while it is possible to find modern HIUases which are 97% identical to the HIUase-TTR ancestral (i.e., the most likely sequence before the gene duplication), modern transthyretins are at most around 77% identical to the TTR ancestral (those coming mostly from birds).
Interestingly, our analysis suggests that the other functions found in transthyretins, binding to the retinol binding protein and presenting peptidase activity, could have appeared earlier in evolution and are worth investigating. Finally, our X-ray structures from reconstructed proteins in this family provide a useful tool for investigating the evolution of this protein family.
Methods
Phylogeny and Ancestral Sequence Reconstruction
Extant sequences were obtained from the UniProt database (UniProt Consortium 2018) through advanced search by protein family (family:"transthyretin family") and taxonomy (taxonomy:"Vertebrata [7742]"). In order to minimize redundancy, only representative sequences from UniRef90 were used. Finally, manual screening was performed to remove excessively long sequences or fragments. The remaining 123 sequences, 74 being transthyretins and 49 HIUases, were aligned using the M-coffee algorithm (Wallace et al. 2006). Alignment filtering was performed by TrimAl Automated 1 method (Capella-Gutiérrez et al. 2009). From the multiple sequence alignment, a phylogenetic reconstruction was performed using MrBayes (Ronquist and Huelsenbeck 2003) over 1,000,000 iterations with JTT as the best model according to Prottest (Abascal et al. 2005; Darriba et al. 2011). Support values were estimated as Bayesian posterior probabilities. The resulting phylogenetic tree was rooted by the midpoint method and, along with the alignment, was used as input to obtain candidate sequences for the reconstructed ancestral proteins by FastML (Ashkenazy et al. 2012) with branch lengths optimization, Gamma distribution with 8 discrete categories and JTT as amino acid replacement matrix.
Heterologous Expression and Purification
The nucleotide sequences for the ancestral proteins were synthesized and cloned into expression vector pET28a-TEV by GenScript, which includes codons for a 6xHis tag and a TEV protease site for its removal. Plasmids were transformed by electroporation into electrocompetent bacteria E. coli BL21 (DE3) or BL21 (DE3) PLysS. These were cultured in LB medium with antibiotic kanamycin, for E. coli BL21 (DE3), and kanamycin + chloramphenicol for BL21 (DE3) PlysS, and expression was performed by addition of isopropyl β-D-1 thiogalactopyranoside during the log phase of bacterial proliferation. The theoretical molecular weight of the proteins was obtained using ProtParam (Gasteiger et al. 2003), available on the ExPaSy Proteomics Server ("http://www.expasy.org"). Expression was confirmed by SDS-PAGE, in which it was possible to observe a band with the expected molecular mass only in the cell extract at the end of the expression.
For large scale purification, cells were lysed by sonication, centrifugated, and the supernatant was loaded onto an affinity chromatography column containing immobilized nickel (His-Trap HP—5 mL—GE Healthcare) coupled to a AKTA pure system (GE Healthcare). The purity and integrity of the eluted samples was evaluated by applying them to a polyacrylamide gel, which was subsequently stained with Coomassie Brilliant Blue R250. The eluted samples were incubated with the TEV protease to remove the polyhistidine tag and subjected to another affinity chromatography step to separate the TEV protease (which also has a polyhistidine tag) from the recombinant proteins. This was followed by molecular exclusion chromatography. The concentration of the recombinant proteins was obtained by absorbance at 280 nm using the molar extinction coefficient obtained by amino acid sequence analysis in the ExPaSy Proteomics Server.
X-ray Crystallography
The HIUase-TTR and TTR ancestor crystal structures were elucidated by SAD (Single Anomalous Dispersion) phasing using, respectively, an iodine derivative crystal (Iod-HIUase-TTR) prepared according to the quick cryo soaking approach (Dauter et al. 2000; Nagem et al. 2003) and a native iodine containing T4/TTR ancestor complex crystal (Nat-T4/TTR ancestor). Briefly, crystals were transferred for a few seconds to a cryogenic solution (crystallization mother liquor with 10% ethylene glycol with or without 100–200 mM NaI) before freezing and data acquisition. All datasets were collected at the Brazilian Synchrotron Light Source MX2 beamline (Guimarães et al. 2009). X-ray diffraction statistics for all datasets are shown in Table 1.
The HIUase-TTR structure was elucidated in a two-step procedure. Initially, an Iod-HIUase-TTR incomplete model at 1.70 Å resolution was automatically built using the Phenix Program Suite (Adams et al. 2010) and later completed by manual building using Coot (Emsley and Cowtan 2004). Later, this model and the Nat-HIUase-TTR dataset were used for Molecular Replacement phasing, and the native model was refined to a final Rfactor/Rfree of 16.1/18.2. The T4/TTR ancestor complex preliminary model was also automatically built using the Phenix Program Suite. It was completed and refined to a final Rfactor/Rfree of 17.2/18.5 after a few cycles of manual and TLS refinement in Coot and Phenix, respectively. Refinement statistics for the final models are also reported in Table 1. The Nat-HIUase-TTR and the Nat-T4/TTR ancestor final models, and their respective datasets, were deposited at the Protein Data Bank under the accession codes 7KCN and 7KJJ, respectively.
As usual with ligand bound transthyretin structures, one of the binding sites usually presents poorer electron density compared to the other. This makes the determination of the ligand binding mode more difficult, as transthyretin commonly crystallizes as a dimer in the asymmetric unit, with a two-fold axis passing through the tunnel where both binding sites lie, and therefore the electron density represents the combination of two binding modes rotated by a 180° angle. The orientation of the two rings can be very easily determined for both sites, mostly due to the electron density of the iodine atoms which are unequivocal even at higher σ omit maps, while the orientation of the amino acid group is less clear for one of the sites (see supplementary materials for electron density maps).
Isothermal Titration Calorimetry
T4 binding to the ancestral proteins was assessed using isothermal titration calorimetry (ITC). A HIUase from Herbaspirillum seropedicae (Matiollo et al. 2009) was used as negative control, and human TTR as a positive control. All proteins were dialysed against a buffer containing 25 mM Tris pH 8,0, 100 mM KCl, 1 mM EDTA then centrifuged at 10,000 rpm for 10 min at 4 °C. T4 (100 μM) was titrated in a cell containing the proteins at 12 μM in a VP-ITC (Malvern) microcalorymeter. Experiments were made at 25 °C with a preliminary injection of 2 μM followed by 5 or 10 μM injections in 250 s intervals. The heat of dilution was obtained from independent experiments and subtracted from each titration. Thermodynamic parameters were obtained using Microcal Origin 7 (OriginLab Corporation).
Ligand and Protein File Preparation
All steps were performed in Maestro version 11.6.13, Release 2018-2 (Schrödinger, LLC, New York, USA). The HIU enantiomer predicted by Pipolo (Pipolo et al. 2011) and the respective high energy intermediate state for the reaction were prepared with LigPrep 3.6 (Schrödinger, LLC, New York, USA). Tridimensional structures were generated from SMILES strings, employing the OPLS3e force field and with the protonation state predicted by Epik, at pH 7.0. Protein files were prepared with the Protein Preparation Wizard, with the addition of hydrogens, removal of waters and optimization of hydrogen positions, based on protonation states predicted with PropKa at pH 7.0.
Docking
Grids were prepared with the Receptor Grid Preparation tool (Schrödinger, LLC, New York, USA). Grid centers were defined based on four residues from the binding site of each protein: His14 and His105 from chains B and D, for PucM (PDB 2H0F), HIUase; His13 and His104 from chains A and B for HIUase-TTR; Lys18 and Leu112 from chains B and D for TTR ancestor. Grids had inner box dimensions of 10 Å and outer box dimensions of 30 Å in each axis. Docking was performed with Glide 6.9 (Schrödinger, LLC, New York, USA), (Friesner et al. 2004) with the following parameters: scaling of van der Waals radii by a scaling factor of 0.8, with charge cutoff = 0.15; SP precision; application of Epik state penalties for docking scores; enhance planarity of conjugated pi groups. Up to 5 poses were saved for each ligand. After docking, results were analyzed in PyMOL (DeLano 2007; Schrödinger 2010).
References
Abascal F, Zardoya R, Posada D (2005) ProtTest: selection of best-fit models of protein evolution. Bioinformatics 21:2104–2105. https://doi.org/10.1093/bioinformatics/bti263
Adams PD, Afonine PV, Bunkóczi G et al (2010) PHENIX : a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr Sect D Biol Crystallogr 66:213–221. https://doi.org/10.1107/S0907444909052925
Aldred AR, Prapunpoj P, Schreiber G (1997) Evolution of shorter and more hydrophilic transthyretin N-termini by stepwise conversion of exon 2 into intron 1 sequences (shifting the 3’ splice site of intron 1). Eur J Biochem 246:401–409. https://doi.org/10.1111/j.1432-1033.1997.t01-1-00401.x
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. https://doi.org/10.1016/S0022-2836(05)80360-2
Ashkenazy H, Penn O, Doron-Faigenboim A et al (2012) FastML: a web server for probabilistic reconstruction of ancestral sequences. Nucleic Acids Res. https://doi.org/10.1093/nar/gks498
Blake CCF, Oatley SJ (1977) Protein–DNA and protein–hormone interactions in prealbumin: a model of the thyroid hormone nuclear receptor? Nature 268:115–120. https://doi.org/10.1038/268115a0
Blake CCF, Swan IDA, Rerat C et al (1971) An X-ray study of the subunit structure of prealbumin. J Mol Biol 61:217–224. https://doi.org/10.1016/0022-2836(71)90218-X
Blake CC, Geisow MJ, Swan ID et al (1974) Structure of human plasma prealbumin at 2–5 A resolution. A preliminary report on the polypeptide chain conformation, quaternary structure and thyroxine binding. J Mol Biol 88:1–12. https://doi.org/10.1016/0022-2836(74)90291-5
Brouillette J, Quirion R (2008) Transthyretin: a key gene involved in the maintenance of memory capacities during aging. Neurobiol Aging 29:1721–1732. https://doi.org/10.1016/j.neurobiolaging.2007.04.007
Capella-Gutiérrez S, Silla-Martínez JM, Gabaldón T (2009) trimAl: a tool for automated alignment trimming in large-scale phylogenetic analyses. Bioinformatics 25:1972–1973. https://doi.org/10.1093/bioinformatics/btp348
Cendron L, Ramazzina I, Percudani R et al (2011) Probing the evolution of hydroxyisourate hydrolase into transthyretin through active-site redesign. J Mol Biol 409:504–512. https://doi.org/10.1016/j.jmb.2011.04.022
Chang L, Munro SLA, Richardson SJ, Schreiber G (1999) Evolution of thyroid hormone binding by transthyretins in birds and mammals. Eur J Biochem 259:534–542. https://doi.org/10.1046/j.1432-1327.1999.00076.x
Darriba D, Taboada GL, Doallo R, Posada D (2011) ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27:1164–1165. https://doi.org/10.1093/bioinformatics/btr088
Dauter Z, Dauter M, Rajashankar KR (2000) Novel approach to phasing proteins: derivatization by short cryo-soaking with halides. Acta Crystallogr D Biol Crystallogr 56:232–237
De Palmieri LC, Lima LMTR, Freire JBB et al (2010a) Novel Zn2+-binding sites in human transthyretin: Implications for amyloidogenesis and retinol-binding protein recognition. J Biol Chem 285:31731–31741. https://doi.org/10.1074/jbc.M110.157206
DeLano WL (2007) PyMOL: An Open-Source Molecular Graphics Tool
Emsley P, Cowtan K (2004) Coot: model-building tools for molecular graphics. Acta Crystallogr Sect D Biol Crystallogr 60:2126–2132. https://doi.org/10.1107/S0907444904019158
Eneqvist T, Lundberg E, Nilsson L et al (2003) The transthyretin-related protein family. Eur J Biochem 270:518–532
Eneqvist T, Lundberg E, Karlsson A et al (2004) High resolution crystal structures of piscine transthyretin reveal different binding modes for triiodothyronine and thyroxine. J Biol Chem 279:26411–26416. https://doi.org/10.1074/jbc.M313553200
Ferguson RN, Edelhoch H, Saroff HA et al (1975) Negative cooperativity in the binding of thyroxine to human serum prealbumin. Biochemistry 14:282–289. https://doi.org/10.1021/bi00673a014
Friesner RA, Banks JL, Murphy RB et al (2004) Glide: a new approach for rapid, accurate docking and scoring. 1. Method and assessment of docking accuracy. J Med Chem 47:1739–1749. https://doi.org/10.1021/jm0306430
Gasteiger E, Gattiker A, Hoogland C et al (2003) ExPASy: the proteomics server for in-depth protein knowledge and analysis. Nucleic Acids Res 31:3784–3788. https://doi.org/10.1093/nar/gkg563
Guimarães BG, Sanfelici L, Neuenschwander RT et al (2009) The MX2 macromolecular crystallography beamline: a wiggler X-ray source at the LNLS. J Synchrotron Radiat 16:69–75. https://doi.org/10.1107/S0909049508034870
Hagen GA, Elliott WJ (1973) Transport of thyroid hormones in serum and cerebrospinal fluid. J Clin Endocrinol Metab 37:415–422. https://doi.org/10.1210/jcem-37-3-415
Hennebry SC (2009) Evolutionary changes to transthyretin: structure and function of a transthyretin-like ancestral protein. FEBS J 276:5367–5379. https://doi.org/10.1111/j.1742-4658.2009.07246.x
Hennebry SC, Law RHP, Richardson SJ et al (2006a) The crystal structure of the transthyretin-like protein from Salmonella dublin, a prokaryote 5-hydroxyisourate hydrolase. J Mol Biol 359:1389–1399. https://doi.org/10.1016/j.jmb.2006.04.057
Hennebry SC, Wright HM, Likic VA, Richardson SJ (2006b) Structural and functional evolution of transthyretin and transthyretin-like proteins. Proteins Struct Funct Bioinforma 64:1024–1045. https://doi.org/10.1002/prot.21033
Huang R, Hippauf F, Rohrbeck D et al (2012) Enzyme functional evolution through improved catalysis of ancestrally nonpreferred substrates. Proc Natl Acad Sci USA 109:2966–2971. https://doi.org/10.1073/pnas.1019605109
Ingles-Prieto A, Ibarra-Molero B, Delgado-Delgado A et al (2013) Conservation of protein structure over four billion years. Structure 21:1690–1697. https://doi.org/10.1016/j.str.2013.06.020
Jung D-K, Lee Y, Park SG et al (2006) Structural and functional analysis of PucM, a hydrolase in the ureide pathway and a member of the transthyretin-related protein family. Proc Natl Acad Sci USA 103:9790–9795. https://doi.org/10.1073/pnas.0600523103
Jurcik A, Bednar D, Byska J et al (2018) CAVER Analyst 2.0: analysis and visualization of channels and tunnels in protein structures and molecular dynamics trajectories. Bioinformatics 34:3586–3588. https://doi.org/10.1093/bioinformatics/bty386
Kabat EA, Moore DH, Landow H (1942) An electrophoretic study of the protein components in cerebrospinal fluid and their relationship to the serum proteins 1. J Clin Investig 21:571–577. https://doi.org/10.1172/jci101335
Kabsch W (2010) XDS. Acta Crystallogr Sect D Biol Crystallogr 66:125–132. https://doi.org/10.1107/S0907444909047337
Kanai M, Raz A, Goodman DS (1968) Retinol-binding protein: the transport protein for vitamin A in human plasma. J Clin Investig 47:2025–2044. https://doi.org/10.1172/JCI105889
Kasai K, Nishiyama N, Yamauchi K (2018) Molecular and thyroid hormone binding properties of lamprey transthyretins: the role of an N-terminal histidine-rich segment in hormone binding with high affinity. Mol Cell Endocrinol 474:74–88. https://doi.org/10.1016/j.mce.2018.02.012
Keebaugh AC, Thomas JW (2010) The evolutionary fate of the genes encoding the purine catabolic enzymes in hominoids, birds, and reptiles. Mol Biol Evol 27:1359–1369. https://doi.org/10.1093/molbev/msq022
Köhrle J (2007) Thyroid hormone transporters in health and disease: advances in thyroid hormone deiodination. Best Pract Res Clin Endocrinol Metab 21:173–191. https://doi.org/10.1016/j.beem.2007.04.001
Kratzer JT, Lanaspa MA, Murphy MN et al (2014) Evolutionary history and metabolic insights of ancient mammalian uricases. Proc Natl Acad Sci USA 111:3763–3768. https://doi.org/10.1073/pnas.1320393111
Leelawatwattana L, Praphanphoj V, Prapunpoj P (2016) Proteolytic activity of Crocodylus porosus transthyretin protease and role of the terminal polypeptide sequences. ScienceAsia 42:190–200. https://doi.org/10.2306/scienceasia1513-1874.2016.42.190
Liz MA, Leite SC, Juliano L et al (2012) Transthyretin is a metallopeptidase with an inducible active site. Biochem J 443:769–778. https://doi.org/10.1042/BJ20111690
Lueprasitsakul W, Alex S, Fang SL et al (1990) Flavonoid administration immediately displaces thyroxine (T4) from serum transthyretin, increases serum free t4and decreases serum thyrotropin in the rat. Endocrinology 126:2890–2895. https://doi.org/10.1210/endo-126-6-2890
Matiollo C, Vernal J, Ecco G et al (2009) A transthyretin-related protein is functionally expressed in Herbaspirillum seropedicae. Biochem Biophys Res Commun 387:712–716. https://doi.org/10.1016/j.bbrc.2009.07.094
Monaco HL (2009) The transthyretin—retinol-binding protein complex. Recent advances in transthyretin evolution, structure and biological functions. Springer, Berlin, pp 123–142
Monaco HL, Rizzi M, Coda A (1995) Structure of a complex of two plasma proteins: transthyretin and retinol-binding protein. Science 268:1039–1041. https://doi.org/10.1126/science.7754382
Morgado I, Melo EP, Lundberg E et al (2008) Hormone affinity and fibril formation of piscine transthyretin: the role of the N-terminal. Mol Cell Endocrinol 295:48–58. https://doi.org/10.1016/j.mce.2008.06.010
Morrow JM, Castiglione GM, Dungan SZ et al (2017) An experimental comparison of human and bovine rhodopsin provides insight into the molecular basis of retinal disease. FEBS Lett 591:1720–1731
Nagem RAP, Polikarpov I, Dauter Z (2003) Phasing on rapidly soaked ions. Methods Enzymol 374:120–137. https://doi.org/10.1016/S0076-6879(03)74005-1
Neumann P, Cody V, Wojtczak A (2001) Structural basis of negative cooperativity in transthyretin. Acta Biochim Pol 48:867–875
Nunes AF, Saraiva MJ, Sousa MM (2006) Transthyretin knockouts are a new mouse model for increased neuropeptide Y. FASEB J 20:166–168. https://doi.org/10.1096/fj.05-4106fje
Pauling L, Zuckerkandl E, Henriksen T, Lövstad R (1963) Chemical paleogenetics. Molecular “Restoration Studies” of extinct forms of life. Acta Chem Scand 17 supl.:9–16. https://doi.org/10.3891/acta.chem.scand.17s-0009
Perez-Jimenez R, Inglés-Prieto A, Zhao Z-M et al (2011) Single-molecule paleoenzymology probes the chemistry of resurrected enzymes. Nat Struct Mol Biol 18:592–596. https://doi.org/10.1038/nsmb.2020
Pipolo S, Percudani R, Cammi R (2011) Absolute stereochemistry and preferred conformations of urate degradation intermediates from computed and experimental circular dichroism spectra. Org Biomol Chem 9:5149–5155. https://doi.org/10.1039/c1ob05433c
Prapunpoj P, Leelawatwattana L (2009) Evolutionary changes to transthyretin: structure-function relationships. FEBS J 276:5330–5341. https://doi.org/10.1111/j.1742-4658.2009.07243.x
Prapunpoj P, Yamauchi K, Nishiyama N et al (2000) Evolution of structure, ontogeny of gene expression, and function of Xenopus laevis transthyretin. Am J Physiol Regul Integr Comp Physiol 279:R2026–R2041. https://doi.org/10.1152/ajpregu.2000.279.6.R2026
Prapunpoj P, Leelawatwatana L, Schreiber G, Richardson SJ (2006) Change in structure of the N-terminal region of transthyretin produces change in affinity of transthyretin to T4 and T3. FEBS J 273:4013–4023. https://doi.org/10.1111/j.1742-4658.2006.05404.x
Ramazzina I, Folli C, Secchi A et al (2006) Completing the uric acid degradation pathway through phylogenetic comparison of whole genomes. Nat Chem Biol 2:144–148. https://doi.org/10.1038/nchembio768
Randall RN, Radford CE, Roof KA et al (2016) An experimental phylogeny to benchmark ancestral sequence reconstruction. Nat Commun 7:12847. https://doi.org/10.1038/ncomms12847
Refetoff S, Marinov VS, Tunca H et al (1996) A new family with hyperthyroxinemia caused by transthyretin Val109 misdiagnosed as thyrotoxicosis and resistance to thyroid hormone–a clinical research center study. J Clin Endocrinol Metab 81:3335–3340. https://doi.org/10.1210/jcem.81.9.8784093
Riisøen H (1988) Reduced prealbumin (transthyretin) in CSF of severely demented patients with Alzheimer’s disease. Acta Neurol Scand 78:455–459. https://doi.org/10.1111/j.1600-0404.1988.tb03687.x
Risso VA, Gavira JA, Mejia-Carmona DF et al (2013) Hyperstability and substrate promiscuity in laboratory resurrections of Precambrian β-lactamases. J Am Chem Soc 135:2899–2902. https://doi.org/10.1021/ja311630a
Romero PA, Arnold FH (2009) Exploring protein fitness landscapes by directed evolution. Nat Rev Mol Cell Biol 10:866–876. https://doi.org/10.1038/nrm2805
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19:1572–1574. https://doi.org/10.1093/bioinformatics/btg180
Saraiva MJM (2001) Transthyretin mutations in hyperthyroxinemia and amyloid diseases. Hum Mutat 17:493–503. https://doi.org/10.1002/humu.1132
Schrödinger L (2010) The PyMOL Molecular Graphics System, Version~1.3r1
Schultz AC, Nygaard P, Saxild HH (2001) Functional analysis of 14 genes that constitute the purine catabolic pathway in Bacillus subtilis and evidence for a novel regulon controlled by the PucR transcription activator. J Bacteriol 183:3293–3302. https://doi.org/10.1128/JB.183.11.3293-3302.2001
Schwarzman AL, Gregori L, Vitek MP et al (1994) Transthyretin sequesters amyloid beta protein and prevents amyloid formation. Proc Natl Acad Sci 91:8368–8372. https://doi.org/10.1073/pnas.91.18.8368
Seibert FB, Nelson JW (1942) Electrophoretic study of the blood protein response in tuberculosis. J Biol Chem 143:29–38
Smith TJ, Davis FB, Deziel MR et al (1994) Retinoic acid inhibition of thyroxine binding to human transthyretin. Biochim Biophys Acta Gen Subj 1199:76–80. https://doi.org/10.1016/0304-4165(94)90099-X
Sousa JC, Grandela C, Fernández-Ruiz J et al (2004) Transthyretin is involved in depression-like behaviour and exploratory activity. J Neurochem 88:1052–1058. https://doi.org/10.1046/j.1471-4159.2003.02309.x
Stevenson WS, Hyland CD, Zhang J-G et al (2010) Deficiency of 5-hydroxyisourate hydrolase causes hepatomegaly and hepatocellular carcinoma in mice. Proc Natl Acad Sci USA 107:16625–16630. https://doi.org/10.1073/pnas.1010390107
Thornton JW (2004) Resurrecting ancient genes: experimental analysis of extinct molecules. Nat Rev Genet 5:366–375
Tola AJ, Leelawatwattana L, Prapunpoj P (2019) The catalytic kinetics of chicken transthyretin towards human Aβ1–42. Comp Biochem Physiol Part C Toxicol Pharmacol 226:108610. https://doi.org/10.1016/j.cbpc.2019.108610
UniProt Consortium T (2018) UniProt: the universal protein knowledgebase. Nucleic Acids Res 46:2699–2699. https://doi.org/10.1093/nar/gky092
Wallace IM, O’Sullivan O, Higgins DG, Notredame C (2006) M-Coffee: combining multiple sequence alignment methods with T-Coffee. Nucleic Acids Res 34:1692–1699. https://doi.org/10.1093/nar/gkl091
Wilson C, Agafonov RV, Hoemberger M et al (2015) Kinase dynamics. Using ancient protein kinases to unravel a modern cancer drug’s mechanism. Science 347:882–886. https://doi.org/10.1126/science.aaa1823
Yamauchi K, Kasai K (2018) Sequential molecular events of functional trade-offs in 5-hydroxyisourate hydrolase before and after gene duplication led to the evolution of transthyretin during chordate diversification. J Mol Evol 86:457–469. https://doi.org/10.1007/s00239-018-9858-4
Yamauchi K, Kasahara T, Hayashi H, Horiuchi R (1993) Purification and characterization of a 3, 5, 3′-l- triiodothyronine-specific binding protein from bullfrog tadpole plasma: a homolog of mammalian transthyretin. Endocrinology 132:2254–2261. https://doi.org/10.1210/endo.132.5.8477670
Zanotti G, Cendron L, Ramazzina I et al (2006) Structure of zebra fish HIUase: insights into evolution of an enzyme to a hormone transporter. J Mol Biol 363:1–9. https://doi.org/10.1016/j.jmb.2006.07.079
Zanotti G, Ramazzina I, Cendron L et al (2009) Vertebrate 5-hydroxyisourate hydrolase identification, function, structure, and evolutionary relationship with transthyretin. Recent advances in transthyretin evolution, structure and biological functions. Springer, Berlin, pp 95–108
Acknowledgements
We thank Hernan Terenzi and Debora Foguel for kindly providing plasmids for the proteins used as experimental controls (H. seropedicae HIUase and human transthyretin, respectively). Funding: this work was supported by CNPq (grant 457851/2014-7) and CAPES (grant 051/2013). LB is a fellow researcher of CNPq.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Belinda Chang.
Accession Numbers: PDB: 7KCN, 7KJJ.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Carrijo de Oliveira, L., Figueiredo Costa, M.A., Gonçalves Pedersolli, N. et al. Reenacting the Birth of a Function: Functional Divergence of HIUases and Transthyretins as Inferred by Evolutionary and Biophysical Studies. J Mol Evol 89, 370–383 (2021). https://doi.org/10.1007/s00239-021-10010-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-021-10010-8