Abstract
Severe Acute Respiratory Syndrome Coronavirus-2 is a zoonotic virus with a possible origin in bats and potential transmission to humans through an intermediate host. When zoonotic viruses jump to a new host, they undergo both mutational and natural selective pressures that result in non-synonymous and synonymous adaptive changes, necessary for efficient replication and rapid spread of diseases in new host species. The nucleotide composition and codon usage pattern of SARS-CoV-2 indicate the presence of a highly conserved, gene-specific codon usage bias. The codon usage pattern of SARS-CoV-2 is mostly antagonistic to human and bat codon usage. SARS-CoV-2 codon usage bias is mainly shaped by the natural selection, while mutational pressure plays a minor role. The time-series analysis of SARS-CoV-2 genome indicates that the virus is slowly evolving. Virus isolates from later stages of the outbreak have more biased codon usage and nucleotide composition than virus isolates from early stages of the outbreak.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Background
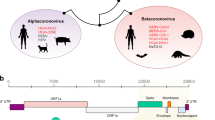
Emerging viral diseases are major threat to the public health systems across the globe (Berry et al. 2015). Coronavirus infectious disease 19(COVID-19) is the latest addition in the growing list of emerging and re-merging diseases with significant financial and health impacts (Zaki et al. 2012; Rota et al. 2003; Zhu et al. 2020; Carroll et al. 2015; Faria et al. 2016). COVID-19 is the third coronavirus outbreak in two decades. The viruses included in Coronaviridae family are found in fish, birds and mammals (Kahn and McIntosh 2005; Fehr and Perlman 2015). The human coronaviruses were first characterized more than fifty years ago and were found to be associated with respiratory infections in humans (Paules et al. (2020)). The infections were of a lesser concern until turn of the century when Severe Acute Respiratory Syndrome Coronavirus (SARS-CoV) emerged in southern China and infected more than 8000 people worldwide, killing 774 of them (Peiris et al. (2003); Guan et al. 2003; Drosten et al. 2003). This was followed by another relatively low-profile outbreak of the Middle East Respiratory Syndrome Coronavirus (MERS-CoV) in Middle East in 2012–2013, causing over 2500 laboratory-confirmed MERS-CoV infections and 866 associated deaths. A third outbreak of highly pathogenic Severe Acute Respiratory Syndrome Coronavirus-2 (SARS-CoV-2) was first reported from Hubei province of China in late 2019, which is liable for severe human infection with a facile human to human transmission (Hui et al. 2019; Tang et al. 2020). The SARS-CoV-2 is a betacoronavirus, which is genetically different from SARS-CoV and MERS-CoV but closely related to bat coronavirus RaTG13 found in the bats from Yunnan province of China (Zhu et al. (2020); Zhou et al. 2020). Bats were also suspected as primary reservoir of SARS-CoV and MERS-CoV, which transmitted to humans through civet cats and dromedary camels, respectively (Guan et al. 2003; Reusken et al. 2013; Li et al. 2005; Memish et al. 2013). Bats serve as reservoirs for several zoonotic viruses, including rabies, Marburg, Nipah, Hendra and probably Ebola viruses (Smith and Wang 2013; Lau et al. 2005; Poon et al. 2005).
When a zoonotic virus jump from one specie to another, it mutates rapidly to cope with host's cellular environment, particularly the translational machinery that will allow a virus to complete its replication cycle and host immune response that will prevent its replication (Lahouassa et al. 2012; Stremlau et al. 2004; Campos et al. 2017; Chassey et al. 2013). During replication, the coding regions of viral genomes undergo both mutational and natural selective pressures that result in non-synonymous and synonymous changes (Sanjuan and Domingo-Calap 2016; Elena and Sanjuan 2005; Peck and Lauring 2018). The viral diversity and evolution are shaped by the intrinsic differences in the fidelity of polymerase, host-encoded mutation rate modifiers and mutation buffering mechanisms that mitigate the deleterious effects of mutations. A time-series analysis of simple measures such as nucleotide composition, codon usage bias (ENC) and codon adaptation index (CAI) could reveal the directional changes in the genome composition after invasion from its natural host and allow the analysis of convergence or divergence of viral codon adaptation towards or away from that of the host over time (Wada et al. 2016).
Several studies have been conducted to understand the emergence and evolution of SARS-CoV-2. The nucleotide composition of SARS-CoV-2 genome was found to be biased and reflected in its codon and amino acid usage patterns (Tort et al. 2020). Most high-frequency codons are A- or T-ending, while the low frequency and rare codons are G- or C-ending (Kandeel et al. 2020). The codon usage pattern of SARS-CoV-2 is affected by both mutational bias and natural selection (Dilucca et al. 2020). SARS-CoV-2 structural and non-structural genes are evolving at varying pace. The integral membrane proteins M and E tend to evolve slowly by accumulating nucleotide mutations on their corresponding genes, while replicase (ORF1ab) and structural proteins S and N tend to evolve faster in comparison to M and E proteins (Dilucca et al. 2020). The codon usage patterns of SARS-CoV-2 orf1ab, spike and nucleocapsid genes were found to be more similar to bat RaTG13 virus, while the codon usage pattern of membrane protein is more similar to Pangolin P1E virus (Gu et al. 2020). In a phylogenetic network analysis of 160 complete SARS-CoV-2 genomes, Forster et al. found three central variants distinguished by amino acid changes, with different geographic distributions. The B type is the most common type in East Asia, while the ancestral A and C types are found in significant proportions in Europe and Americas (Forster et al. 2020). Van Dorp et al. have identified 198 recurrent mutations in 7,666 SARS-CoV-2 genome, with nearly 80% of the recurrent mutations producing non-synonymous changes at the protein level (Dorp et al. 2020). To build upon the foundation laid by the above-mentioned studies, we have analyzed 10,114 SARS-CoV-2 genomes to gain more insights into evolution of this virus. The nucleotide composition and the codon usage biases of protein coding genes were analyzed to identify the evolutionary forces that contributed the most in shaping the codon usage patterns. Time-series analysis was performed on different codon usage indices to estimate virus adaptation and directional changes in SARS-CoV-2 genome during the course of the outbreak. Our data suggest that SARS-CoV-2 has not achieved a codon usage bias that balances the selective and mutational pressures on the synonymous codon usage. The SARS-CoV-2 genome is slowly evolving. Virus isolates from later stages of the outbreak have more biased codon usage and nucleotide composition than virus isolates from early days of the outbreak. Codon usage pattern of SARS-CoV-2 is diverging away from the constitutively expressed human genes.
Materials and Methods
Sequence Data
10,114 SARS-CoV-2 genomes with > 29,000 base pair length, < 1% non-synonymous mutations and < 0.05 unique amino acid mutations were retrieved from the GISAID Initiative EpiCoV platform (https://www.epicov.org/epi3/frontend#49972e) (Table S1). Sequences were aligned with the local version of MAFFT software (https://mafft.cbrc.jp) using default parameters. For comparative genomic analysis, one bat coronavirus (MN996532, Bat-CoV-RaTG13) sequence, two full-length bat SARS like coronavirus sequences (MG772933, Bat SLCoV-ZC45 and MG772934, Bat SLCoV-ZXC21) and one SARS-CoV sequence (NC_004718, SARS-CoV Tor2) were retrieved from the Virus Pathogen Resource database (https://www.viprbrc.org/brc). For predicting statistical properties of coding and non-coding sequences of each genome, conserved coding sequences were concatenated into a single open reading frame, whereas untranslated regions (5′ and 3′) were merged and analyzed as a single sequence.
Compositional Properties
Base composition, including overall frequency of mononucleotides (G%, C%, A% and U%), frequency of mononucleotides at third wobble position (G3s, C3s, U3s and A3s) and frequency of G + C at three codon positions (GC1, GC2 and GC3) were calculated using the program Codon W 1.4.4 (http://sourceforge.net/projects/codonw/). The ratio of observed and expected frequencies of 16 dinucleotides were calculated using default parameter in SSE software package (Simmonds 2012) and used for designation of over-representation (> 1.235) or under-representation (< 0.765) in terms of relative abundance compared with a random association of mononucleotides.
Codon Usage Bias in SARS-CoV-2 Genome
Relative synonymous codon usage (RSCU) is one of the most widely used parameters for examining the overall synonymous codon usage variation among genes (Sharp and Li 1986). The RSCU values of SARS-CoV-2 and other coronaviruses listed in table S2 were calculated using following equation: RSCUj(x) = (fjx /Ejx), where fjx is the observed frequency of codon j in the genome x and Ejx is the expected frequency of the codon j. Expected values are calculated by counting the total number of synonymous codons for a given amino acid in the sequence divided by the number of existing codons that code for it. The codon usage frequencies of Rhinolophus affinis were retrieved from High-performance Integrated Virtual Environment-Codon Usage Tables (HIVE-CUT) database (Athey et al. 2017), while codon usage table representing human house-keeping genes is described elsewhere (Paola et al. 2018).
Codon Adaptation Index (CAI)
The CAI is a measure of the adaptiveness of synonymous codon usage bias of a gene towards the codon usage of highly expressed genes. To assess the SARS-CoV-2 adaptation to its hosts, the CAI values for all SARS-CoV genes were calculated using Homo sapiens house-keeping genes codon usage table and Rhinolophus affinis codon usage table.
Effective Number of Codons (ENC)
The ENC is a measure of the total number of different codons present in a sequence and shows the bias from equal use of all synonymous codons for a given amino acid (Novembre 2002; Wright 1990). The ENC value range from 20 to 61, with low ENC values indicating higher codon usage bias and vice versa. The sequences in which ENC values are < 30 are highly expressed, while those with > 55 are poorly expressed genes (Sau et al. 2005; Sharp and Cowe 1991). The ENC values for SARS-CoV-2 genome and individual genes were calculated using EMBOSS (Carver and Bleasby 2003).
ENC-Plot Mapping Analysis
The ENC plot (a plot of ENC vs. GC3s) provides a visual display of the main features of codon usage patterns for several genes. Expected ENC values for all GC3 compositions (0–1.0) were calculated using the equation ENCexp = 2 + S + (29/S2 + (1 − S)2) (Wright 1990), where S is the frequency of G + C at third-codon position. When the corresponding points fall near the expected neutral curve, mutational bias is the main factor affecting the observed codon diversity. However, when the corresponding points fall considerably below the expected curve, the observed codon bias is mainly affected by natural selection.
Neutrality Plot Mapping Analysis
The neutrality plot is an analytical method used to examine the relative influence of mutation pressure and natural selection on the codon usage pattern (Sueoka 1988). Among three codon positions, second-codon position is the most functionally constrained codon position, while third-codon position is the least functionally constrained. A nucleotide substitution in the second-codon position causes a non-synonymous change in the coding sequence. Most nucleotide substitutions in the third-codon position and some nucleotide substitutions in the first-codon position are synonymous (Bofkin and Goldman 2007). Theoretically, mutation should occur randomly at any of three codon positions if there is no external pressure. A neutrality plot was generated, where GC3 was compared to the average GC content at the first- and second-codon positions (GC1,2) with the Pearson product-moment correlation coefficient(r). In the plot, the regression coefficient against GC3 is regarded as the mutation-selection equilibrium coefficient.
Principal Component Analysis
The principal component analysis (PCA) is a type of multivariate statistical method used to identify major trends of variation in synonymous codon usage among genes/genomes. PCA is an orthogonal linear transformation procedure that transforms some correlated variable (RSCU) into a smaller number of uncorrelated variables called principal components. To investigate the major trends of codon usage variation, the PCA was performed on the RSCU values of 10,114 SARS-CoV-2 genomes. Factor retention was determined using a scree plot and the proportion of variance. First three principal components with high eigenvalues values (7.71, 5.37 and 4.74, respectively) and 30.22% cumulative variation were ranked as major axes. Correlation analysis based on Spearman’s rank correlation (with a level of significance of p < 0.05 or p < 0.01) was used to reveal correlations between various codon usage indices and major axes of PCA.
Results
Composition Analysis of SARS-CoV-2 Genome
To study molecular evolution of SARS-CoV-2, we analyzed genomic nucleotide sequences of 10,114 isolates for which the isolation/reporting dates were given. The genome of SARS-CoV-2 was found to be AT(U) rich with ~ 62% mean A + T(U) contents (Fig. 1a). The G + C contents of the coding and non-coding sequences are 38% and 41.3%, respectively, indicating more biased nucleotide composition in the coding regions of the genome. Similar compositional bias was observed in bat-CoV-RaTG13, bat SLCoV-ZC45, bat SLCoV-ZXC21 and SARS-CoV genomes (Fig. 1b & c). Among individual SARS-CoV-2 genes, the nucleocapsid (N) and matrix (M) genes have least biased nucleotide composition with 52.8% and 57.4% AT(U) contents, respectively. The ORF6 has the most biased nucleotide composition with ~ 72% AT(U) contents. All other genes have AT(U) contents in the range of 60–65% (Fig. 1d). Analysis of AT(U)/GC nucleotides distribution at three codon positions revealed a progressively declining trends in G/C use at second- and third-codon positions. The A/T(U) were preferably used at third synonymous codon position in the SARS-CoV-2 genome (Fig. S1).
Whole genome comparative nucleotide composition analysis of SARS-CoV-2, SARS-CoV (SARS-CoV-Tor2), bat coronavirus (Bat-CoV-RaTG13) and bat SARS like coronaviruses (Bat SLCoV-ZXC21 and Bat SLCoV-ZXC21). a Boxplot diagram depicting the GC/AT(U) contents in coding and non-coding (NC) regions of SARS-CoV-2 genome. b, c Boxplot depicting the A/T(U) and G/C contents of Bat-CoV, Bat_SLCoV, SARS-CoV and SARS-CoV-2 genomes. d GC/AT(U) contents (mean ± SD) of SARS-CoV-2 genes
SARS-CoV-2 Codon Usage Analysis
To explore the variation of synonymous codon usage in the SARS-CoV-2 genome, the RSCU values of 59 synonymous codons were calculated (Fig. 2, Table S2). SARS-CoV-2 strongly preferred A- and U-ending codons with 25/26 preferred codon (RSCU > 1.0) in the SARS-CoV-2 genome being A/U-ending. Among preferred codon, twelve codons AGA(R), GGU (G), GCU(A), UCU(S), GUU(V), CCU(P), ACU(T), CUU (L), UCA (S), ACA(T) and UUA(L) have RSCU value > 1.6 and hence were considered over-represented. The G/C-ending codons are suppressed in SARS-CoV-2 genome with 20/21 under-represented codons (RSCU < 0.6) being G/C-ending. The A/U-ending codons were used as a preferred codon for all 18 amino acids encoded by synonymous codons in the SARS-CoV-2 genome. Further analysis of amino acids encoded by four or more synonymous codons showed that approximately 78% of amino acids in SARS-CoV-2 proteins are encoded by A/U-ending codons. For example, 67.47% of leucine, 71.39 of valine, 73.54% of arginine, 79.30% of glycine, 82% of alanine, 84.50% of serine, 85.60% of threonine and 88.50% of proline are encoded by A/U-ending codons.
Principal Component Analysis of Codon Usage
To identify the major trends of variation in synonymous codon usage, PCA was implemented for all 10,114 sequences. The first three principal components in the PCA accounted for 13.02%, 9.11% and 8.0% of the total variation, indicating that first three principal axes are the major explanatory axes for interpreting codon usage variation (Fig. 3a). The distribution of SARS-CoV-2 isolates based on RSCU values on first and second principal axes resulted in segregation of SARS-CoV-2 isolates into five overlapping clusters along first and second principal axes (Fig. 3b). A vast majority of US isolates clustered in the upper-right and lower-left quadrants of the plot. The European isolates segregated into three clusters, two along the second principal axis and third in the upper-left quadrant of the plot. Furthermore, each cluster contains a significant number of isolates from other geographic regions as well, suggesting multiple parallel transmission events. A highly significant positive correlation of first two principal axes (f1 and f2) with U% (f1 = 0.411, f2 = 0.187; p < 0.0001), and negative correlation with A%(f1 = -0.149, f2 = -0.168; p < 0.0001) and C% (f1 = -0.379, f2 = -0.111; p < 0.0001), indicates that compositional constraints (especially U and C nucleotides) affected the codon usage bias of the SARS-CoV-2 genome (Table S3).
The principal component analysis of RSCU of SARS-CoV-2. a The scree plot of the eigenvalues of the first 40 PCs and cumulative variance plot from principal component analysis of the SARS-CoV-2 RSCU values. b A plot of the values of the Axis1(13.072%) and the Axis2 (9.11%) of all SARS-CoV-2 strains in principal component analysis
Codon Adaptation Index
CAI quantifies the similarity between the synonymous codon usage of a gene and the synonymous codon frequency of a reference set. As inferred from the RSCU values, the codon usage pattern of SARS-CoV-2 is very similar to other coronaviruses but mostly antagonistic to both of its hosts, with only one preferred codon for arginine (AGA) common between SARS-CoV-2 and Homo sapiens and five preferred codons common with Rhinolophus Affinis (Table S2). To gain further insights into the SARS-CoV-2 adaptation to specific hosts, CAI analysis was performed using Homo sapiens and Rhinolophus Affinis codon usage tables as references. As shown in Fig. 4a, all SARS-CoV-2 genes are predicted to be expressed at high levels in Homo sapiens and Rhinolophus Affinis. The mean CAI for SARS-CoV-2 genes ranged from 0.652 to 0.766 against Homo sapiens codon usage reference and 0.633–0.731 against Rhinolophus Affinis codon usage reference. With exception of E gene, which has the lowest CAI (0.65 and 0.63, respectively) in both hosts, all structural and non-structural genes have high CAI (> 0.70) in both hosts.
The Analysis of Codon Usage Bias in the SARS-CoV-2 Genes
The genomic composition of SARS-CoV-2 was found to be relatively stable and conserved among different isolates. The mean ENC value of SARS-CoV-2 coding sequence is 45.372 ± 0.02, which is similar to bat-CoV-RaTG13 but slightly lower than other bat-SLCoVs or SARS-CoV (Fig. 4b). Analysis of individual SARS-CoV-2 genes showed substantial variations in the ENC of different genes (Fig. 4b). The ORF10, Matrix, Nucleocapsid and ORF3 of the SARS-CoV-2 have low codon usage bias with mean ENC values of 57.80 ± 0.00, 54.17 ± 0.14, 53.22 ± 0.13, and 51.66 ± 0.02, respectively. However, ORF7, ORF1ab, Spike, ORF8, ORF6, and Envelope have moderate codon usage bias with mean ENC values of 46.28 ± 0.30, 44.16 ± 1.35, 44.15 ± 0.62, 43.89 ± 0.34, 42.92 ± 0.29, and 42.01 ± 0.49, respectively. High ENC values (> 40), little variation in ENC among different isolates and substantial variations in the ENC of different genes within each genome suggest a highly conserved gene-specific codon usage bias in the SARS-CoV-2 genome.
The Differential Effect of Mutational Biases and Natural Selection on Codon Usage Bias
To determine the factors that influence the SARS-CoV-2 codon usage bias, the ENC plot was constructed using the ENC and GC3 values of 10,114 SARS-CoV-2 isolates. The dotted line in Fig. 5a represents the curve if codon usage is only determined by the GC content at the third-codon position. As shown in Fig. 5a, data points representing SARS-CoV-2 isolates formed a tight cluster below the expected curve, indicating composite effect of mutational pressure and other factors (such as natural selection) in shaping the codon usage pattern of the SARS-CoV-2 genome. Furthermore, analysis of individual SARS-CoV-2 genes revealed varying effect of mutational pressure and natural selection on the codon usage pattern of different genes. The data points representing all SARS-CoV-2 genes except ORF10 formed distinct clusters below the expected curve with variable proximities (Fig. 5a inset), confirming a differential effect of directional mutation pressure and natural selection in shaping the codon usage bias in these genes.
The effect of mutational bias and natural selection on SARS-CoV-2 synonymous codon usage. a Relationship between GC3 and the effective number of codons (ENC). The ENC values of SARS-CoV-2 Strains (concatenated coding sequence) and mean ENC values of individual SARS-CoV-2 genes (upper-right inset) were plotted against the corresponding GC3s. The standard curve (dotted line) indicates the expected codon usage if GC compositional constraints alone account for codon usage bias. b The neutrality plot (GC12 vs. GC3). Neutrality plot analysis of the average GC content in the first- and second-codon positions (GC12) and the GC content at third position (GC3)
Although the ENC–GC3 plot demonstrated the contribution of directional mutational pressure and natural selection on the SARS-CoV-2 codon usage bias, it did not estimate the magnitude of each force. Neutrality plot analysis was performed to determine the relative contribution of mutation pressure or natural selection on the SARS-CoV-2 codon usage pattern (Fig. 5b). When GC3 was plotted against GC12 for all isolates, a weak correlation was found between GC3 and GC12 (r = 0.116, p = < 0.0001), suggesting a minor role of directional mutational pressure in shaping SARS-CoV-2 codon usage bias. The slope of regression line was calculated to be 0.067, indicating a minor effect of directional mutation pressure (6.70%) and major effect of natural selection (93.3%) in shaping SARS-CoV-2 codon usage bias. Similar trends were observed when GC3 of individual genes were plotted against the corresponding GC12 (Fig. S2). Directional mutational pressure accounted for ~ 36% and 10% of selection pressure in N gene and ORF10, respectively, but less than 7% in all other genes. These data suggest that different evolutionary pressures are acting disproportionally on SARS-CoV-2 genes. Mutational pressure is a minor factor, affecting the codon usage preferences of some genes, while natural selection played a major role in shaping the codon usage pattern of almost all SARS-CoV-2 genes.
Dinucleotide Frequency
To determine if dinucleotide composition of SARS-CoV-2 genome is biased, ratio of observed to expected frequencies (O/E ratio) of 16 dinucleotides was computed for all SARS-CoV-2 sequences. The mean dinucleotide O/E ratio of SARS-CoV-2 genome is 1.0 ± 0.235. Dinucleotides with odds ratio > 1.235 or < 0.765 were considered as over- and under-represented, respectively. The dotted box in Fig. 6 represents these conventional thresholds for crude determination of over- or under-representation. The dinucleotide bias in SARS-CoV-2 genome is not acute and only three dinucleotides (CpA, UpG and CpG) were found to have biased usage in protein coding genes. The CpA and UpG dinucleotides were over-represented, whereas CpG was markedly under-represented in SARS-CoV-2 genome (Fig. 6, sold line). In non-coding sequence, two dinucleotides (ApU and CpC) were under-represented, and one nucleotide pair (ApC) was over-represented (Fig. 6, dotted line). Similar dinucleotide biases were observed in individual SARS-CoV-2 genes as well. The CpG dinucleotide was under-represented in all SARS-CoV-2 genes except E and ORF10, whereas CpA/UpG were over-represented in most of the SARS-CoV-2 genes (Fig. S3). Analysis of the RSCU value of the eight CpG containing codons (CGT, CGG, CGC, CGA, TCG, GCG, CCG and ACG) indicated a marked suppression of CpG containing codons with seven codons having RSCU value of < 0.6 and accounting for only 2.5% of total amino acids in SARS-CoV-2 proteins. Taken together, these data indicate a marked suppression of CpG dinucleotides and CpG containing codons in SARS-CoV-2 genome.
Relative dinucleotide abundance in SARS-CoV-2 genome. Line graph represents the mean observed/expected (O/E) frequency ratio of 16 dinucleotides. The mean ± standard deviation of dinucleotide O/E ratios for SARS-CoV-2 coding sequence is 1.0 ± 0.235. Dotted box represents the confidence interval of mean ± 1SD (i.e., O/E ratio 0.765–1.235). Dinucleotide outside dotted box was classified as under- or over-represented in SARS-CoV-2 genome
Distinct patterns of dinucleotide representation could be a consequence of the nucleotide composition of the genome (mutational bias) or natural selection of genes/genomes with specific dinucleotide biases. To explore the mechanism of CpG depletion in the SARS-CoV-2 genome, CpG/GpC odds ratio was calculated. High odds ratio (closer to 1.0) would indicate that depletion was due to pressure specifically acting upon constituent mononucleotides, while low odds ratio would suggest implication of evolutionary pressures specifically acting upon CpG dinucleotide. The mean CpG/GpC odds ratio of 0.362 ± 0.001 indicates that CpG depletion SARS-CoV-2 genome is not a mere consequence of compositional constraints. To further understand the mechanism of CpG depletion, the loss of CpG dinucleotides (1 − (O/E)CpG) and average gain in UpG and CpA dinucleotides (((O/E)TpG – 1) + (O/E)CpA – 1))/2) were calculated. The average loss of CpG in SARS-CoV-2 coding region was 0.607 ± 0.001, and average gain in UpG and CpA dinucleotides was 0.330 ± 0.002. A significant positive correlation between CpG depletion and UpG/CpA gain, (r = 0.31; P = < 0.0001) and a weak but statistically significant positive correlation between CpG depletion and CG% (r = 0.07; P = < 0.0001) suggest that observed under-representation of CpG in SARS-CoV-2 genome genes is mainly due to the evolutionary pressures specifically acting on CpG dinucleotide.
Directional Changes in SARS-CoV-2 Genome Over the Course of Pandemic
To assess the evolution of SARS-CoV-2, we analyzed time-series changes in nucleotide composition and codon usage indices of 10,114 strains isolated from the start of the outbreak till May 4, 2020. To compensate for the compositional diversity resulting from random mutations and sequencing errors, averaged values for strains in each week were plotted against the elapsed week and linear regression lines were used to show the changes in nucleotide composition and codon usage indices over time. The time-series analysis of ENC in Fig. 7a indicates a steady decline in the ENC value of SARS-CoV-2 strains isolated during the course of the pandemic. To assess the degree of ENC variation, virus strains were stratified according to the month of isolation. As shown in Fig. 7b, the ENC value of earlier isolates was significantly higher than the strains isolated at the later stages of the epidemic. This suggests that ENC of SARS-CoV-2 strains progressively declined over the course of the pandemic and SARS-CoV-2 strains tended to use more biased codons as the pandemic progressed.
Time-series change is SARS-CoV-2 ENC. a Averaged ENC value for the strains isolated in each week was plotted according to elapsed week from December 21, 2019. Trend lines were generated using linear regression analysis to facilitate visualization of correlations. b Boxplots of the effective number of codons (ENC) vs month of isolation. Asterisk (***) indicates that there was a significant difference (P < 0.001) between the two groups
To further gain insights into the molecular mechanism of SARS-CoV-2 genome evolution, time-series change in mononucleotides was analyzed. Figure 8(a–d) plots averaged mono-nucleotide compositions (%) for strains in each week according to the date of isolation. The linear regression lines and month-wise boxplot showed a time-dependent increase in U% and a corresponding decrease in C% indicating C-to-U mutations over time in SARS-CoV-2 genome (Fig. 8e, f). The C-to-U mutations accounted for ~ 47% of all the mutations in the SARS-CoV-2 genome during the course of the pandemic (Fig. 8g). Further analyses of C% and U% nucleotides at first-, second- and third-codon positions revealed that most C-to-U mutations occurred at 2nd and 3rd codon positions (Fig. S4).
Time-series change in mono-nucleotide compositions (%) for SARS-CoV-2. a–d Averaged mono-nucleotide compositions (%) for the strains isolated in each week were plotted according to elapsed week from December 21, 2019. The trend lines were generated using linear regression analysis to facilitate visualization of correlations. e, f Boxplot of %U and %C vs month of isolation. g Frequency of mono-nucleotide substitutions in SARS-CoV-2 genome
To further assess the impact of C-to-U mutation on codon usage, time-series analysis was performed on the RSCU value of six amino acids that only use U/C-ending codons (asparagine, histidine, aspartic acid, tyrosine, cysteine and phenylalanine) and ten amino acids that use U/C-ending codons as well as other codons (threonine, isoleucine, proline, leucine, alanine, glycine, valine and serine). Four out six amino acids encoded by U/C-ending codons (phenylalanine, asparagine, histidine and tyrosine) and six out of ten amino acids encoded by U/C-ending codons as well as other codons (threonine, arginine, glycine, valine, leucine and serine) showed time-dependent decline in the usage of C-ending codon and an increase in the usage of U-ending codon (Fig. S5). These data clearly suggest that progressive C-to-U mutations resulted in a more biased codon usage in later strains. To further assess the impact of C-to-U mutation on virus adaptation to its new host, we performed a time-series codon adaptation index analysis. Contrary to nucleotide composition and ENC, the CAI values over time failed to reveal a clear trend. The slope of regression line in Fig. 9a registered a small decrease in the CAI over time relative to human house-keeping genes but the observed difference is so small that few outlying CAI value could effectively change the direction of the trend line. On the other hand, the relative codon deoptimization index, which is also a measure of codon adaptation to the host, showed a steady increase indicating a decreasing adaptability relative to human house-keeping genes over time (Fig. 9b). Furthermore, time course analysis of CpG and UpA dinucleotides also registered a steady increase over time (Fig. 9c–d). Taken together, these data clearly suggest that novel coronavirus is mutating at low levels and the codon usage pattern of the SARS-CoV-2 is moving away from the presumed optimal codon usage in human house-keeping genes over time.
Time-series changes in the codon adaptation indices and dinucleotides compositions (O/E ratio) for SARS-CoV-2. a Averaged CAI and b averaged RCDI values for the strains isolated in each week. c, d Averaged O/E ratios of CpG and UpA dinucleotide for the strains isolated in each week were plotted according to elapsed week from December 21, 2019. The trend lines were generated using linear regression analysis to facilitate visualization of correlations
Discussion
The ENC value of different SARS-CoV-2 genes ranged from 42.01 to 57.80, with mean ENC value of 45.37, indicating weak codon usage bias in the SARS-CoV-2 genome. This result is consistent with previously reported data on other coronaviruses (Chen et al. 2017; Castells et al. 2017; Gu et al. 2004). Weak codon bias seems to be an adaptive strategy used by many RNA and DNA viruses, which may enable these viruses to maintain efficient replication without competing for limited tRNA resources (Jenkins and Holmes 2003). The nucleotide composition of SARS-CoV-2 is more biased than SARS-CoV or closely related bat coronaviruses with higher proportion of A/U nucleotides. Coronaviruses have an atypical codon usage bias characterized by low levels of C and high levels of U nucleotides (Woo et al. 2007; Berkhout and Hemert 2015). Due to higher percentage of A/U nucleotides in the genome, the codon usage pattern tended towards more frequent use of A/U-ending codons with approximately 70% of all amino acids in SARS-CoV-2 proteins being encoded by A/U-ending codon. The intracellular availability of nucleotides (A + T versus G + C) depends on the genomic nucleotide composition of the host. The human genome is AT rich; therefore, the cellular milieu is appropriately enriched for the efficient replication of an AT-rich extra chromosomal genetic elements (Dietel et al. 2019).
Codon usage pattern of SARS-CoV-2 is mostly antagonistic to human codon usage with only one common preferred codon for arginine. Coincidence between viral and host codon usage may lead to improved translation efficiency, whereas antagonism may result in slow viral mRNA translation and viral replication. Some viruses may avoid host immune detection by maintaining low level of replications, while others may prefer stability and folding accuracy over speed by maintaining low prevalence codons in the genome. The presence of low prevalence codons in hepatitis A virus genome has been linked to slow protein translation and correct protein folding (Costafreda et al. 2014). The human immunodeficiency virus and herpesviruses maintain codons in their genes, which show a low prevalence in their host cells (Bradel-Tretheway et al. 2003; Ngumbela et al. 2008). The avian influenza virus (H3N2) overcomes the antiviral action of interferon by skewing its codon usage towards the tRNA pools of interferon-altered cells (Smith et al. 2018). The expression of papillomaviruses capsid proteins L1 and L2 is linked to the specific cell differentiation states in which particular aminoacyl-tRNAs are expressed (Zhou et al. 1999; Zhao et al. 2005). Furthermore, viruses that infect one or few related hosts evolve a more consistent codon usage pattern with their host. When such viruses jump species and infect an unrelated host, they may exhibit a relatively lower codon matching with their new host. Low codon adaption of SARS-CoV-2 to human codon usage may reflect a snapshot of the virus transiting from a well-adapted host to a novel host or an evolutionary strategy adapted by the SARS-CoV-2 to avoid host defense.
The pattern of codon usage is governed by genetic drift and mutational pressure in some organisms, while in others, it is governed by the balance between mutational pressure and natural selection (Bulmer 1991). Both selection and random genetic drift are directly affected by the effective population size. When the effective population size is large, natural selection predominates and random drift is less common (Rouzine et al. 2001). Due to high mutation rates and large populations sizes, mutation pressure and natural selection are generally considered as main forces shaping the RNA virus evolution. Significant differences in ENC and GC3s of different SARS-CoV-2 genes and distinct distribution of genes on ENC-GC3s plot indicate that different SARS-CoV-2 genes have taken different codon evolutionary pathways depending on the function of these genes. A weak correlation between GC3 and GC12 (r = 0.11, p < 0.001) and near-zero regression coefficient (0.067) suggest a strong influence of selective pressure on first- and second-codon positions (GC12) with respect to that on the third-codon position (GC3) in the SARS-CoV-2 genome. The nucleocapsid gene with regression coefficient of 0.373 was found to be the only gene that can accept both synonymous and non-synonymous mutations without compromising its functionality. The near-zero or negative regression lines for all other genes (Fig. S2) suggest that first- and second-codon positions in these genes are constrained to preserve the amino acid sequence.
Time-series analysis revealed that SARS-CoV-2 has not achieved a codon usage bias that balances the selective and mutational pressures on the synonymous codon usage. The SARS-CoV-2 is continuously evolving at low levels. The codon usage pattern of the SARS-CoV-2 is moving away from the presumed optimal codon usage in human house-keeping genes over time. Virus isolates from the later stages of the outbreak showed a relatively more biased nucleotide composition. The C-to-U transitions were by far the most frequent nucleotide substitutions in the SARS-CoV-2 genome over time, accounting for almost one-half of all sequence changes. Further analyses of C% and U% nucleotides at first-, second- and third-codon positions revealed that most C-to-U mutations occurred at 2nd and 3rd codon positions (Fig. S5). Among three codon positions, second-codon position is the most functionally constrained codon position. A nucleotide substitution in the second-codon position causes a non-synonymous change in the coding sequence leading to change in amino acid sequence with potential biological effects. Several amino acid changes in the SARS-CoV-2 spike protein with potential impact on virus transmission and diminished sensitivity to vaccine-elicited antibodies have been reported. A SARS-CoV-2 variant B.1.1.7 with multiple mutations in the spike protein (ΔH69/ΔV70, Δ144, N501Y, A570D, D614G, P681H, T716I, S982A, D1118H) has gained higher transmissibility and was found to be less sensitive to vaccine-elicited neutralizing antibodies (Collier et al. 2020). Another variant B.1.351 with slightly different set of mutations in the spike protein (K417N, E484K, N501Y) also seems refractory to some polyclonal and monoclonal antibodies (Weisblum et al. 2020). A third variant with three mutations in the receptor binding domain of the spike protein (K417T, E484K and N501Y) with potential effects on transmissibility and antigenic profile has been reported from Brazil (Sabino et al. 2021). An emerging lineage (B.1.526) of viral isolates, which accounted for as many as ~ 25% of coronavirus genomes sequenced from New York during February 2021, features mutations on the spike protein (L5F, T95I, D253G, E484K or S477N, D614G and A701V) that allow it to penetrate into host cells and cause infections more effectively (West et al. 2021).
From evolutionary perspective, asymmetric increase in C-to-U mutations cannot have arisen through a mutational effect of the coronavirus RNA-dependent RNA polymerase during virus replication or misincorporation errors in the next-generation sequence methods used to produce the data set (Simmonds 2012). A more plausible explanation of the skewed mutation patterns (towards C-to-U) is spontaneous or enzymatic cytidine deamination processes converting Cs into Us within infected cells (Matyasek and Kovarik 2020). Several interferon-inducible RNA-editing enzymes such as ADAR1 and APOBEC 3 family of RNA-editing enzymes have been implicated in base substitutions in RNA virus genomes with varying outcomes on infection (Gelinas et al. 2011). The ADARs (adenosine deaminases that act on RNA) target double-stranded RNA (dsRNA) for deamination of adenines into inosines and have been implicated in the sequence changes (A-to-G and U-to-C transitions) seen in many double-stranded RNA viruses that replicate in the cytoplasm of infected cells (Samuel 2012). The APOBEC family of cytidine deaminases modulate immune responses by mutating specific nucleic acid sequences of hosts and pathogens. The human genome codes at least seven APOBEC3 proteins (namely A3A, A3B, A3C, A3DE, A3F, A3G and A3H) that exert an antiviral effect against a wide variety of viruses including retrovirus and other DNA viruses, such as herpesviruses, parvoviruses and hepatitis B virus (Chiu and Greene 2009). The role of APOBEC3 in coronavirus restriction and genome editing is anecdotal. Three APOBEC3 proteins (A3C, A3F and A3H) have been shown to restrict coronavirus infection without causing a characteristic C-to-U hypermutation in viral genome suggesting a deaminase-independent mechanism of coronavirus replication inhibition (Milewska et al. 2018). A metagenomic sequencing analysis of SARS-CoV-2 transcriptomic and genomic data revealed over-representation of ADAR-mediated A-to-I mutations as well as APOBEC-mediated C-to-U mutations in the transcriptome data but less in the genomic data suggesting a potential role of ADAR and APOBEC in SARS-CoV-2 RNA editing. The presence of fewer A-to-I and C-to-U mutations in the genomic data indicates the selection against deleterious mutation caused by RNA-editing enzymes (Di Giorgio et al. 2020). In line with these findings, analysis of mononucleotides at fourfold degenerate sites in SARS-CoV-2 coding sequence revealed a near equilibrium usage of C and G mononucleotides, higher than predicted usage of A nucleotide and far below the equilibrium usage of U nucleotide (Rice et al. 2020).
The CpG and UpA dinucleotides, which are markedly suppressed in many RNA and small DNA viruses, have appeared to be increasing in the SARS-CoV-2 genome over time. This observation is particularly intriguing because experimental increase in CpG and UpA dinucleotide resulted in virus attenuation and decreased pathogenicity (Gaunt et al. 2016; Fros et al. 2017; Burns et al. 2009). Studies of influenza and picornaviruses indicate that the innate immune response might recognize RNA-specific CpG motifs, such that the suppression of CpG in viruses could assist immune evasion (Atkinson et al. 2014; Greenbaum et al. 2009). The observed increase in CpG in SARS-CoV-2 genome over time may not necessarily reflect a selection towards high CpG, rather a selection against UU to limit the deleterious effects of high U contents on mRNA stability and gene expression (Rice et al. 2020).
Conclusion
Data presented here alone may not be sufficient to draw any conclusions regarding the viral fitness or its pathogenesis and further in vitro and in vivo studies are needed to establish a direct correlation between the observed changes in the viral genome and its effect on viral fitness or pathogenesis. Nevertheless, this study provided considerable insights into the composition and evolution of SARS-CoV-2 genome. Time-series analysis of large data set can be informative in identifying the directional changes in the genome composition, which is important for the development of the effective diagnostic tools and vaccines. Several candidate vaccines have been granted emergency use authorization, but duration of protective acquired immunity is still an open debate. The duration of protection depends on the genetic stability of the virus as well as magnitude and quality of immune response induced by primary infection or vaccine. All leading SARS-CoV-2 vaccines are recombinant viral-vectored vaccines or nucleic acid-based vaccines engineered to express SARS-CoV-2 spike protein. Such vaccines are generally safe but lack the full antigenic complexity of the virus and may become less effective over time with accumulation of mutations in antigenic epitope. The live attenuated vaccine elicits a strong and lasting immune response, but they raise potential side effects and biosafety concerns. A rational design of subunit vaccine harboring antigenic elements from multiple SARS-CoV-2 proteins may augment the antigenic effects of the vaccine and prevent the viral escape or resistance to vaccine-elicited neutralizing antibodies.
References
Athey J, Alexaki A, Osipova E et al (2017) A new and updated resource for codon usage tables. BMC Bioinform 18(1):391
Atkinson NJ, Witteveldt J, Evans DJ, Simmonds P (2014) The influence of CpG and UpA dinucleotide frequencies on RNA virus replication and characterization of the innate cellular pathways underlying virus attenuation and enhanced replication. Nucleic Acids Res 42(7):4527–4545
Bai HX, Hsieh B, Xiong Z et al (2020) Performance of radiologists in differentiating COVID-19 from viral pneumonia on chest CT. Radiology. https://doi.org/10.1148/radiol.2020200823
Berkhout B, van Hemert F (2015) On the biased nucleotide composition of the human coronavirus RNA genome. Virus Res 202:41–47
Berry M, Gamieldien J, Fielding BC (2015) Identification of new respiratory viruses in the new millennium. Viruses 7(3):996–1019
Bofkin L, Goldman N (2007) Variation in evolutionary processes at different codon positions. Mol Biol Evol 24(2):513–521
Bradel-Tretheway BG, Zhen Z, Dewhurst S (2003) Effects of codon-optimization on protein expression by the human herpesvirus 6 and 7 U51 open reading frame. J Virol Methods 111(2):145–156
Bulmer M (1991) The selection-mutation-drift theory of synonymous codon usage. Genetics 129(3):897–907
Burns CC, Campagnoli R, Shaw J, Vincent A, Jorba J, Kew O (2009) Genetic inactivation of poliovirus infectivity by increasing the frequencies of CpG and UpA dinucleotides within and across synonymous capsid region codons. J Virol 83(19):9957–9969
Campos RK, Wong B, Xie X et al (2017) RPLP1 and RPLP2 are essential flavivirus host factors that promote early viral protein accumulation. J Virol. 91(4):e01706-16
Carroll MW, Matthews DA, Hiscox JA et al (2015) Temporal and spatial analysis of the 2014–2015 Ebola virus outbreak in West Africa. Nature 524(7563):97–101
Carver T, Bleasby A (2003) The design of Jemboss: a graphical user interface to EMBOSS. Bioinformatics 19(14):1837–1843
Castells M, Victoria M, Colina R, Musto H, Cristina J (2017) Genome-wide analysis of codon usage bias in Bovine Coronavirus. Virol J 14(1):115
Chen Y, Xu Q, Yuan X et al (2017) Analysis of the codon usage pattern in Middle East Respiratory Syndrome Coronavirus. Oncotarget 8(66):110337–110349
Chiu YL, Greene WC (2009) APOBEC3G: an intracellular centurion. Philos Trans R Soc Lond B 364(1517):689–703
Collier DA, De Marco A, Ferreira IATM, et al. SARS-CoV-2 B.1.1.7 escape from mRNA vaccine-elicited neutralizing antibodies. medRxiv. 2020.
Costafreda MI, Perez-Rodriguez FJ, D’Andrea L et al (2014) Hepatitis A virus adaptation to cellular shutoff is driven by dynamic adjustments of codon usage and results in the selection of populations with altered capsids. J Virol 88(9):5029–5041
de Chassey B, Aublin-Gex A, Ruggieri A et al (2013) The interactomes of influenza virus NS1 and NS2 proteins identify new host factors and provide insights for ADAR1 playing a supportive role in virus replication. PLoS Pathog 9(7):e1003440
Di Giorgio S, Martignano F, Torcia MG, Mattiuz G, Conticello SG (2020) Evidence for host-dependent RNA editing in the transcriptome of SARS-CoV-2. Sci Adv 6(25):eabb5813
Di Paola N, Freire CCM, Zanotto PMA (2018) Does adaptation to vertebrate codon usage relate to flavivirus emergence potential? PLoS ONE 13(1):e0191652
Dietel AK, Merker H, Kaltenpoth M, Kost C (2019) Selective advantages favour high genomic AT-contents in intracellular elements. PLoS Genet 15(4):e1007778
Dilucca M, Forcelloni S, Georgakilas AG, Giansanti A, Pavlopoulou A (2020) Codon usage and phenotypic divergences of SARS-CoV-2 genes. Viruses. 12(5):498
Drosten C, Gunther S, Preiser W et al (2003) Identification of a novel coronavirus in patients with severe acute respiratory syndrome. N Engl J Med 348(20):1967–1976
Elena SF, Sanjuan R (2005) Adaptive value of high mutation rates of RNA viruses: separating causes from consequences. J Virol 79(18):11555–11558
Faria NR, Azevedo R, Kraemer MUG et al (2016) Zika virus in the Americas: early epidemiological and genetic findings. Science 352(6283):345–349
Fehr AR, Perlman S (2015) Coronaviruses: an overview of their replication and pathogenesis. Methods Mol Biol 1282:1–23
Forster P, Forster L, Renfrew C, Forster M (2020) Reply to Sanchez-Pacheco et al., Chookajorn, and Mavian et al.: Explaining phylogenetic network analysis of SARS-CoV-2 genomes. Proc Natl Acad Sci USA 117(23):12524–12525
Fros JJ, Dietrich I, Alshaikhahmed K, Passchier TC, Evans DJ, Simmonds P (2017) CpG and UpA dinucleotides in both coding and non-coding regions of echovirus 7 inhibit replication initiation post-entry. Elife 6:e29112
Gaunt E, Wise HM, Zhang H et al (2016) Elevation of CpG frequencies in influenza A genome attenuates pathogenicity but enhances host response to infection. Elife 5:e12735
Gelinas JF, Clerzius G, Shaw E, Gatignol A (2011) Enhancement of replication of RNA viruses by ADAR1 via RNA editing and inhibition of RNA-activated protein kinase. J Virol 85(17):8460–8466
Greenbaum BD, Rabadan R, Levine AJ (2009) Patterns of oligonucleotide sequences in viral and host cell RNA identify mediators of the host innate immune system. PLoS ONE 4(6):e5969
Gu W, Zhou T, Ma J, Sun X, Lu Z (2004) Analysis of synonymous codon usage in SARS Coronavirus and other viruses in the Nidovirales. Virus Res 101(2):155–161
Gu H, Chu DKW, Peiris M, Poon LLM (2020) Multivariate analyses of codon usage of SARS-CoV-2 and other betacoronaviruses. Virus Evol 6(1):veaa032
Guan Y, Zheng BJ, He YQ et al (2003) Isolation and characterization of viruses related to the SARS coronavirus from animals in southern China. Science 302(5643):276–278
Hui DS, Azhar EI, Madani TA, Ntoumi F, Kock R, Dar O, Ippolito G, Mchugh TD, Memish ZA, Drosten C, Zumla A (2020) The continuing 2019-nCoV epidemic threat of novel coronaviruses to global health—the latest 2019 novel coronavirus outbreak in Wuhan, China. Int J Infect Dis 1(91):264–6
Jenkins GM, Holmes EC (2003) The extent of codon usage bias in human RNA viruses and its evolutionary origin. Virus Res 92(1):1–7
Kahn JS, McIntosh K (2005) History and recent advances in coronavirus discovery. Pediatr Infect Dis J 24(11):223–227 discussion S226
Kandeel M, Ibrahim A, Fayez M, Al-Nazawi M (2020) From SARS and MERS CoVs to SARS-CoV-2: Moving toward more biased codon usage in viral structural and nonstructural genes. J Med Virol 92(6):660–666
Lahouassa H, Daddacha W, Hofmann H et al (2012) SAMHD1 restricts the replication of human immunodeficiency virus type 1 by depleting the intracellular pool of deoxynucleoside triphosphates. Nat Immunol 13(3):223–228
Lau SK, Woo PC, Li KS et al (2005) Severe acute respiratory syndrome coronavirus-like virus in Chinese horseshoe bats. Proc Natl Acad Sci USA 102(39):14040–14045
Li W, Shi Z, Yu M et al (2005) Bats are natural reservoirs of SARS-like coronaviruses. Science 310(5748):676–679
Matyasek R, Kovarik A (2020) Mutation patterns of human SARS-CoV-2 and Bat RaTG13 coronavirus genomes are strongly biased towards C>U transitions, indicating rapid evolution in their hosts. Genes. 11(7):761
Memish ZA, Mishra N, Olival KJ et al (2013) Middle East respiratory syndrome coronavirus in bats, Saudi Arabia. Emerg Infect Dis 19(11):1819–1823
Milewska A, Kindler E, Vkovski P et al (2018) APOBEC3-mediated restriction of RNA virus replication. Sci Rep 8(1):5960
Ngumbela KC, Ryan KP, Sivamurthy R et al (2008) Quantitative effect of suboptimal codon usage on translational efficiency of mRNA encoding HIV-1 gag in intact T cells. PLoS ONE 3(6):e2356
Novembre JA (2002) Accounting for background nucleotide composition when measuring codon usage bias. Mol Biol Evol 19(8):1390–1394
Paules CI, Marston HD, Fauci AS (2020) Coronavirus infections-more than just the common cold. JAMA 323(8):707–708
Peck KM, Lauring AS (2018) Complexities of viral mutation rates. J Virol. 92(14):e01031-17
Peiris JS, Lai ST, Poon LL et al (2003) Coronavirus as a possible cause of severe acute respiratory syndrome. Lancet 361(9366):1319–1325
Poon LL, Chu DK, Chan KH et al (2005) Identification of a novel coronavirus in bats. J Virol 79(4):2001–2009
Reusken CB, Haagmans BL, Muller MA et al (2013) Middle East respiratory syndrome coronavirus neutralising serum antibodies in dromedary camels: a comparative serological study. Lancet Infect Dis 13(10):859–866
Rice AM, Castillo Morales A, Ho AT et al (2021) Evidence for strong mutation bias toward, and selection against, U content in SARS-CoV-2: implications for vaccine design. Mol. Biol. Evol. 38(1):67–83
Rota PA, Oberste MS, Monroe SS et al (2003) Characterization of a novel coronavirus associated with severe acute respiratory syndrome. Science 300(5624):1394–1399
Rouzine IM, Rodrigo A, Coffin JM (2001) Transition between stochastic evolution and deterministic evolution in the presence of selection: general theory and application to virology. Microbiol Mol Biol Rev 65(1):151–185
Sabino EC, Buss LF, Carvalho MPS et al (2021) Resurgence of COVID-19 in Manaus, Brazil, despite high seroprevalence. Lancet 397(10273):452–455
Samuel CE (2012) ADARs: viruses and innate immunity. Curr Top Microbiol Immunol 353:163–195
Sanjuan R, Domingo-Calap P (2016) Mechanisms of viral mutation. Cell Mol Life Sci 73(23):4433–4448
Sau K, Gupta SK, Sau S, Ghosh TC (2005) Synonymous codon usage bias in 16 Staphylococcus aureus phages: implication in phage therapy. Virus Res 113(2):123–131
Sharp PM, Cowe E (1991) Synonymous codon usage in Saccharomyces cerevisiae. Yeast 7(7):657–678
Sharp PM, Li WH (1986) An evolutionary perspective on synonymous codon usage in unicellular organisms. J Mol Evol 24(1–2):28–38
Simmonds P (2012) SSE: a nucleotide and amino acid sequence analysis platform. BMC Res Notes 5:50
Simmonds P (2020) Rampant C-->U Hypermutation in the Genomes of SARS-CoV-2 and other coronaviruses: causes and consequences for their short- and long-term evolutionary trajectories. mSphere 5(3).
Smith I, Wang LF (2013) Bats and their virome: an important source of emerging viruses capable of infecting humans. Curr Opin Virol 3(1):84–91
Smith BL, Chen G, Wilke CO, Krug RM (2018) Avian Influenza virus PB1 gene in H3N2 viruses evolved in humans to reduce interferon inhibition by Skewing Codon usage toward Interferon-Altered tRNA pools. mBio 9(4):e01222-18
Stremlau M, Owens CM, Perron MJ, Kiessling M, Autissier P, Sodroski J (2004) The cytoplasmic body component TRIM5alpha restricts HIV-1 infection in Old World monkeys. Nature 427(6977):848–853
Sueoka N (1988) Directional mutation pressure and neutral molecular evolution. Proc Natl Acad Sci USA 85(8):2653–2657
Tang JW, Tambyah PA, Hui DSC (2020) Emergence of a novel coronavirus causing respiratory illness from Wuhan, China. J Infect 80(3):350–371
Tort FL, Castells M, Cristina J (2020) A comprehensive analysis of genome composition and codon usage patterns of emerging coronaviruses. Virus Res 283:197976
van Dorp L, Acman M, Richard D et al (2020) Emergence of genomic diversity and recurrent mutations in SARS-CoV-2. Infect Genet Evol 83:104351
Wada Y, Wada K, Iwasaki Y, Kanaya S, Ikemura T (2016) Directional and reoccurring sequence change in zoonotic RNA virus genomes visualized by time-series word count. Sci Rep 6:36197
Weisblum Y, Schmidt F, Zhang F et al (2020) Escape from neutralizing antibodies by SARS-CoV-2 spike protein variants. eLife. 9(e61312)
West AP, Barnes CO, Yang Z, Bjorkman PJ (2021) SARS-CoV-2 lineage B.1.526 emerging in the New York region detected by software utility created to query the spike mutational landscape. bioRxiv 2021:2021.2002.2014.431043.
WHO (2020) COVID-19 Weekly Epidemiological Update–18 October, 2020. In: Organization WH (ed) World Health Organization, Geneva
Woo PC, Wong BH, Huang Y, Lau SK, Yuen KY (2007) Cytosine deamination and selection of CpG suppressed clones are the two major independent biological forces that shape codon usage bias in coronaviruses. Virology 369(2):431–442
Wright F (1990) The “effective number of codons” used in a gene. Gene 87(1):23–29
Zaki AM, van Boheemen S, Bestebroer TM, Osterhaus AD, Fouchier RA (2012) Isolation of a novel coronavirus from a man with pneumonia in Saudi Arabia. N Engl J Med 367(19):1814–1820
Zhao KN, Gu W, Fang NX, Saunders NA, Frazer IH (2005) Gene codon composition determines differentiation-dependent expression of a viral capsid gene in keratinocytes in vitro and in vivo. Mol Cell Biol 25(19):8643–8655
Zhou J, Liu WJ, Peng SW, Sun XY, Frazer I (1999) Papillomavirus capsid protein expression level depends on the match between codon usage and tRNA availability. J Virol 73(6):4972–4982
Zhou P, Yang XL, Wang XG et al (2020) A pneumonia outbreak associated with a new coronavirus of probable bat origin. Nature 579(7798):270–273
Zhu N, Zhang D, Wang W et al (2020) A novel Coronavirus from patients with Pneumonia in China, 2019. N Engl J Med 382(8):727–733
Acknowledgment
We thank College of Clinical Pharmacy for providing necessary support to conduct this research.
Funding
The authors acknowledge the Deanship of Scientific Research at King Faisal University for the financial support under Nasher track (grant number 206156).
Author information
Authors and Affiliations
Contributions
SH and STR identified the research topic and designed the study. PS and SH retrieved and analyzed the data. SH, STR and PS prepared the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The author(s) declared no potential conflicts of interest with respect to the research, authorship, and/or publication of this article.
Ethical Approval
This article does not contain studies with animals performed by any of the authors.
Additional information
Handling editor: Keith Crandall.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hussain, S., Rasool, S.T. & Pottathil, S. The Evolution of Severe Acute Respiratory Syndrome Coronavirus-2 during Pandemic and Adaptation to the Host. J Mol Evol 89, 341–356 (2021). https://doi.org/10.1007/s00239-021-10008-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-021-10008-2