Abstract
Pitviper sensory perception incorporates diverse stimuli through the integration of trichromatic color vision, bifocal heat-sensing, and dual-system chemoperception. Chemoperception, or olfaction, is mediated by chemoreceptors in the olfactory bulb and the vomeronasal organ, but the true genomic complexity of the gene families and their relative contributions is unknown. A full genomic accounting of pitviper chemoperception directly complements our current understanding of their venoms by generating a more complete polyphenic representation of their predatory arsenal. To characterize the genetic repertoire of pitviper chemoperception, we analyzed a full-genome assembly for Crotalus adamanteus, the eastern diamondback rattlesnake. We identified hundreds of genes encoding both olfactory receptors (ORs; 362 full-length genes) and type-2 vomeronasal receptors (V2Rs; 430 full-length genes). Many chemoreceptor genes are organized into large tandem repeat arrays. Comparative analysis of V2R orthologs across squamates demonstrates how gene array expansion and contraction underlies the evolution of the chemoreceptor repertoire, which likely reflects shifts in life history traits. Chromosomal assignments of chemosensory genes identified sex chromosome specific chemoreceptor genes, providing gene candidates underlying observed sex-specific chemosensory-based behaviors. We detected widespread episodic evolution in the extracellular, ligand-binding domains of both ORs and V2Rs, suggesting the diversification of chemoreceptors is driven by transient periods of positive selection. We provide a robust genetic framework for studying pitviper chemosensory ecology and evolution.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The genetic basis of receptor-mediated sensation in vertebrates ranges from deceptively simple to highly complicated (Baldwin and Ko 2020). Infrared heat-sensing in pitvipers largely depends on a single calcium-channel that is activated by heating of the pit organ (Gracheva et al. 2010). In contrast, vertebrate chemoperception (e.g., olfaction) depends on hundreds of receptor genes (Bear et al. 2016) that enable detection of a diverse set of scent molecules. Evolution of the sensory receptor underlies diverse adaptive strategies and natural history traits in vertebrates (Baldwin and Ko 2020). A well characterized example is the vertebrate vision system. The complement of photoreceptor paralogs and evolutionary spectral tuning enable vertebrates to utilize diverse light environments (Davies et al. 2012). A wealth of experimental molecular studies has shown links between specific amino acid substitutions and receptor spectral sensitivity (Bowmaker 2008; Davies et al. 2012; Van Hazel et al. 2013; Hunt et al. 2009; Schweikert et al. 2018; Yokoyama 2008; Yokoyama et al. 2014, 2016). Recently, this experimental work has led to functionally predictive, computational modeling of receptor structures (Patel et al. 2018). Clearly, a full genetic accounting of sensory receptors and characterization of the evolutionary forces that shape their distributions and sequences are critical to elucidating the mechanistic basis of sensory perception.
Chemoperception is the sensory detection of chemicals originating outside the body, including airborne, dissolved, and surface-bound compounds. At the molecular level, chemoperception depends on the physical interaction of these compounds with membrane-bound receptor proteins in the sensory tissue (Mombaerts 2004). Vertebrate chemoreceptors involved in chemoperception belong to the large family of G-protein coupled receptors (GPCRs) (Buck and Axel 1991). GPCRs share a highly conserved structure and functional mechanism (Gether and Kobilka 1998; Rosenbaum et al. 2009). Chemical ligands bind to the variable extracellular domain triggering a conformational change that is propagated through the highly conserved seven-transmembrane domain. The conformational change leads to a change in the interaction state of the GPCR intracellular domain with the G-protein. G-protein activation then leads to a signaling cascade and a change in cell state, which, in the case of vertebrate chemoreceptors, is the depolarization of afferent sensory neurons (Rosenbaum et al. 2009).
Vertebrates utilize several types of GPCRs for chemoperception. Among these are the large groups of olfactory receptors (ORs), vomeronasal type-1 receptors (V1Rs), vomeronasal type-2 receptors (V2Rs), and trace amine-associated receptors (TAARs). Copies of these types of chemoreceptors are typically abundant, with large numbers of pseudogenes indicating evolution through a birth–death process (Dong et al. 2009; Hughes et al. 2018; Nei and Rooney 2005; Niimura and Nei 2007). Lineage specific presence/absence of these groups and the relative abundance is evolutionarily responsive to ecology (Grus et al. 2005; Hayden et al. 2010; Khan et al. 2015; Silva and Antunes 2017). For example, several marine mammals (Kishida et al. 2007; Liu et al. 2019) and sea snakes (Kishida and Hikida 2010) show a convergent loss of OR genes upon adaptation to aquatic environments. The abundance of chemoreceptors can also be responsive to the evolution of other sensory systems, as in the decline of functional ORs in primates upon greater reliance on the visual system (Niimura et al. 2018). Lineage specific expansions are associated with behavior as in the expansion of V2R genes in ostariophysian fish with pheromone-stimulated fright reaction (Yang et al. 2019).
A shift in abundance from V2Rs to V1Rs has been postulated to correlate with the shift from aquatic to terrestrial lifestyle during early tetrapod evolution (Shi and Zhang 2007). However, this trend is apparently contradicted in squamate reptiles, including snakes and lizards, that have few V1Rs and abundant V2Rs (Brykczynska et al. 2013). Squamates have distinct chemosensory morphologies (Baeckens et al. 2017), and the olfactory bulb and vomeronasal organ are anatomically separate (Graves 1993; Halpern 1987; Halpern and Martínez-Marcos 2003; Keverne 1999). Airborne molecules enter the nares and pass over the main olfactory bulb (MOB) (Schwenk 1993, 1995) for detection by ORs. Snakes and some lizards utilize elongated tongues that bifurcate into microscopic points, optimized in form and kinematic function for bi-directional scent collection and delivery to the vomeronasal organ (VNO) through distinctive oscillatory tongue-flicking behaviors (Cooper 1995; Daghfous et al. 2012; Filoramo and Schwenk 2009) and subsequent detection by V2Rs (Cooper, 1997a). This dual-system arrangement (Fig. 1a) provides squamates with a unique chemoperception system that can detect volatile, airborne chemicals via the olfactory bulb and ORs as well as water-soluble chemicals via the VNO and V2Rs (Cooper 1997b). Previous work assessing the genomic and chromosomal context of genes in the five-pace viper (Yin et al. 2016) reported signatures of adaptive evolution in ORs. This work further identified significantly more OR genes located on the Z chromosome than any other chromosome. These findings set the stage for precise genomic accounting of snake chemoperception by also considering V2Rs in a similar way.
Graphical summary of ORs and V2Rs found in Crotalus adamanteus. a Chemical cues (shown as red circles) are collected from the environment and processed by chemoreceptors localized in the MOB (above) and VNO (below). b 2D representations of membrane-bound ORs and V2Rs with extracellular regions colored dark green and the intracellular (cytoplasmic and transmembrane) regions colored purple; V2R dimerization is represented by a duplicated, faded and mirrored structure with the dimer-interface highlighted by a red shaded oval; red circles with red arrows conceptualize ligand-binding potential in the extracellular regions of ORs and V2Rs. c Chromosomal distributions of 362 OR paralogs and 430 V2R paralogs; genes occurring on contigs which failed to map to the reference Crotalus viridis assembly are labeled “Unplaced”; genes occurring on contigs that mapped to sub-chromosome resolution regions of the reference assembly are labeled “Unplaced contiguous”
Rattlesnakes display chemosensory-based behaviors, such as the ability to distinguish envenomated verses non-envenomated prey (Saviola et al. 2013), the detection and tracking of potential mates by male rattlesnakes over harsh terrain from distances of up to a kilometer (Weldon et al. 1992; Duvall et al. 1992), and maternal care behaviors between female rattlesnakes and her offspring (as reviewed in Weldon et al. 1992). These distinct behaviors present rattlesnakes as pertinent study systems for understanding the mechanistic basis and biological relevance for chemoperception. Additionally, their reliance on thermal-sensing and venom for predation provides an opportunity for understanding the evolutionary response of phenotypic integration and identifying genetic signatures of prey specificity (Holding et al. 2015).
As a first step to understanding the rattlesnake chemosensory system, we used genomic and transcriptomic approaches to generate a genetic framework of the complete chemoreceptor repertoire of the eastern diamondback rattlesnake (Crotalus adamanteus). We characterized chromosomal localizations and gene family distributions within Crotalus adamanteus and among other squamates, as well as analyzed protein structure and selection signatures across chemoreceptor phylogenies. Our combined bioinformatic approach suggests genomic mechanisms underlying sex-biased chemoperception, evolutionary mechanisms underlying chemoreceptor diversification, and creates a framework for elucidating the roles of chemoreceptors in rattlesnake ecology and evolution.
Results and Discussion
Highly Diverse Complement of Rattlesnake ORs and V2Rs
To characterize rattlesnake chemosensory gene repertoires, we sequenced and assembled a high-coverage reference genome for Crotalus adamanteus using an adult female rattlesnake. The final assembly represents the first full-length genome for this species. Our combined Illumina, Nanopore, and Pacbio sequencing resulted in 51.6 × coverage based on the final assembly size (see methods for sequencing details). The final assembly is 1.6 Gbp in size and consists of 11,369 scaffolded contigs with a N50 of 338 Kb. Gaps comprise < 0.0% and contigs > 50 Kbp comprise 93.07% of assembled bases. Gene annotations were hand-curated for individual chemoreceptor genes guided by full-genome splice aware mapping of mRNA transcripts extracted from olfactory bulb and VNO tissues. Altogether, we annotated 796 full-length chemosensory genes, 164 pseudogenes, and 51 truncated genes (chromosomal distributions summarized in Table 1). Our gene annotations outline exonic structures, splice sites, protein coding sequences (CDSs), mRNAs, UTRs, poly-A tail signals, and assign unique IDs to individual genes. Pseudogenes and truncated genes occurring on contigs containing full-length genes were annotated to provide context for future synteny analyses.
Rattlesnake chemoperception integrates olfactory and vomeronasal acuity through comparable OR and V2R genetic contributions. C. adamanteus possess hundreds of OR and V2R genes (n = 362 and 430, respectively) in addition to two V1R genes, one TAAR2-like gene, and one TAAR5-like gene. High numbers of V2R paralogs were previously described in the corn snake using joint transcriptomics and mRNA in situ hybridization (Brykczynska et al. 2013). More recently, Kishida et al. (2019) reported high numbers of paralogs for both ORs and V2Rs in sea snakes. RNA-seq alone limits the ability to assemble large, multi-exon genes such as V2Rs, as evidenced by the recovery of many partial coding sequences from the corn snake (Brykczynska et al. 2013). The approach used here, RNA-seq read mapping to a reference genome, provides a more complete assembly of full-length OR and V2R genes as demonstrated by Kishida et al. (2019) with sea snakes. The hand-curated full-length gene annotations generated in the present study further distinguish pseudogenes from putatively functional chemoreceptors, which would otherwise obscure the ability to assign chemoreceptor orthology downstream. Our identification of chemoreceptor genes enables future research expanding on multi-species chemoreceptor orthology and recovering clade-specific gene expansions.
Our annotations show that rattlesnake OR and V2R genes generally occur as large tandem-repeat arrays spread across the genome. The largest V2R array of 64 putatively functional genes spans ∼4 Mbp on chromosome 2 (Fig. 2). This particular array includes V2R genes in both the 3′ and 5′ direction, pseudogenes mid-array, and truncated genes (i.e., potentially functional yet only partially recovered in the genome assembly). Conservative estimates suggest that ∼25% of all gene duplication events in vertebrates are tied to tandemly arrayed genes (Pan and Zhang 2008). Array duplications can instantaneously and dramatically increase gene copy numbers and influence gene expression phenotypes (Stranger et al. 2007), suggesting an efficient mechanism for OR and V2R gene expansion. High OR diversity and genomic architecture appear consistent across terrestrial vertebrates (Niimura and Nei 2006; Niimura 2009, 2012) and has previously been extended to non-viperid snakes. For example, Perry et al. (2018) reported high numbers of OR paralogs from the garter snake genome and further confirmed an early expansion in snake ORs when compared to non-varanid lizards. In a comparison of V2Rs from the Komodo dragon and other squamates, Lind et al. (2019) demonstrated lineage-specific V2R expansions. Additionally, Komodo dragon V2Rs occur in tandem repeat clusters, similar to mammalian VNO receptors (Yang et al. 2005) and ORs (Niimura and Nei 2006), suggesting that gene duplication via tandem arrays is a common mechanism for OR and V2R gene family expansion across vertebrates.
Chemosensory genomic architecture for Crotalus adamanteus. Gene annotations are overlaid on the chromosome-level RaGOO scaffolding to visualize high-density regions (*chromosome W is included for size only). OR genes are colored blue and V2Rs are colored green. The yellow and orange zoomed-in regions illustrate the breakdown of a large V2R gene array present on chromosome 2; pseudogenes and truncated genes (dark green) are also shown in the orange ∼600 Kb close-up
Using a comparative syntenic and phylogenetic analysis, we traced the evolution of a distinguishable orthologous V2R array between five squamate species with available genomes (Fig. 3). Based on our established V2R exonic structure template (Fig. 3b), we identified 47 full-length V2R genes for C. adamanteus, 59 for Crotalus tigris, 82 for Thamnophis elegans, 18 for Varanus komodoensis, and 6 for Anolis carolinensis. V2R pseudogenes were also identified for C. tigris and T. elegans in addition to C. adamanteus. There are clear V2R array expansions and contractions potentially corresponding to life history changes during squamate evolution. For example, there is a large expansion of V2R genes among snake lineages but a contraction among pitvipers, which may correlate with the evolution of the heat-sensing organ. Additionally, lineage specific changes in V2R distributions are evident among the two rattlesnake species, which may correlate with differences in diet and habitat. The disparity in genome completeness and annotation among these species prohibits a genome-wide analysis of V2R arrays. However, our approach of combining long and short-read genomic data with transcriptomics provides a guide for annotation and analysis of large copy-number gene families.
Combined synteny and phylogenetic assessment of a shared squamate V2R gene array. a Genomic schematics illustrating V2R gene numbers, order, and orientations identified from the genomes of five species. Missing sequences are shown as red “Ns” with the number of bases underneath in red, and manual scaffold locations are shown as double grey dashes. This array is distinguished by several conserved immunoglobulin type-V subunit containing regions (light-blue arrows), and collectively highlights a substantial V2R gene expansion in snakes with a potential reduction following the evolution of the pit organ. Red and orange shading connects snake V2Rs which grouped with lizard genes on the gene phylogeny. b Generalized exonic structure shared across all V2R genes shown in a. Violations to this structure such as missing exons or premature stops were identified as pseudogenes, and these were excluded for clarity. c Gene phylogeny of all V2R genes shown in a. The tree represents a maximum likelihood reconstruction of full-length gene sequences spanning non-coding regions (i.e., introns and UTRs) for accurate recovery of recent duplication events. The red clade includes V2Rs from all five species, and the orange clade includes V2Rs from the three snakes and Varanus komodoensis, suggesting a potential gene expansion linked to a forked-tongue common ancestor. Image of V. komodoensis courtesy of Daniel Dashevsky
To further confirm our CDS annotations for C. adamanteus and characterize receptor structural features, we iteratively assessed translated chemosensory genes for conserved elements during the genome annotation process. Extracellular ligand binding domains were identified with the Conserved Domains Database (NCBI; Supp_file_01). We used SignalP (Supp_file_02) to identify signal sequences in our set of receptors. Most of the V2R genes contained signal sequences (327/430). We found only one OR (1/362) gene with a signal sequence, which conforms with the enigmatic trafficking of ORs in mature olfactory neurons (McClintock and Sammeta 2003). Additionally, structural homology searches confirm that C. adamanteus ORs are monomeric (Fig. 1b), which follows the one allele-one neuron rule found across vertebrate ORs (Chess et al. 1994). In contrast, V2Rs are likely dimeric (Fig. 1b) based on homology with the metabotropic glutamate receptors (Wu et al. 2014). These structural predictions lack the precision needed to resolve if V2Rs form homo- or heterodimers, although VNO neurons have been shown to contain transcripts from at least two V2R subfamilies (Martini et al. 2001; Francia et al. 2014; Akiyoshi et al. 2018). The combinatorial potential of V2Rs would greatly increase the spectrum of ligands potentially detected by the VNO. Regardless, the large number of OR (362) and V2R (430) genes in C. adamanteus suggests a complex basis for the chemosensory phenotype.
Chromosomal Distribution of ORs and V2Rs Suggests Sex-Biased Chemoperception
Male rattlesnakes are the homogametic sex with two copies of the Z chromosome, and females are heterogametic with Z and W chromosomes. To assign genomic contigs to sex chromosomes for our C. adamanteus assembly, we first mapped short genomic reads from a male and a female to the assembly. Based on male:female scaled mean read depth ratios, we assigned 977 contigs to chromosome W (42 Mbp; 2.63% genomic contribution), 1484 contigs to chromosome Z (91 Mbp; 5.68% genomic contribution), and the remaining 8908 contigs as autosomal (1.46 Gbp; 91.68% genomic contribution). The Z chromosome genomic contribution of 5.68% for our female rattlesnake is consistent with the male Crotalus viridis Z contribution of 8.5% (Schield et al. 2019). We further assigned 7387 of the 8908 autosomal contigs to specific chromosomes via whole genome scaffolding using RaGOO (Alonge et al. 2019) and the chromosome-level C. viridis genome assembly as a reference (Schield et al. 2019). Of the remaining 1521 contigs, 1041 were considered contiguous but unplaced and the remaining 480 were considered unplaced. Inferred chromosome placements of chemoreceptor genes, pseudogenes, and truncated genes are shown in Table 1. Unplaced contiguous chemosensory gene groupings can be found in the Supplementary Materials online (Supp_file_03).
The presence of ORs and V2Rs on the Z and W sex chromosomes suggests a potential route for sex-biased olfaction. The top three chromosomes ranked by descending OR copy-number were chromosomes Z, 1, and 4, and no ORs were detected on the W chromosome (Fig. 1c). Viperids, including rattlesnakes, lack a mechanism to compensate for the reduced gene expression in the heterogametic sex, i.e. no dosage compensation (Vicoso et al. 2013). The chromosomal distribution of ORs suggests that differences in expression levels of Z chromosome ORs between sexes could be a potential mechanism of observed sex-biased olfaction behaviors (Duvall et al. 1992). Similarly, 17 V2R genes mapped to the Z chromosome (Fig. 1C), suggesting a similar mechanism could also modulate V2R based chemoperception. Additionally, five V2R genes present on the W chromosome represents a potential genetic basis for female-biased chemoperception such as detection of offspring (Weldon et al. 1992; Duvall et al. 1992).
Diversification of Extracellular, Ligand Binding Domains Driven by Transient Positive Selection
We evaluated selection signatures across ORs and V2Rs found in the C. adamanteus genome. We generated maximum likelihood gene phylogenies with IQ-TREE based on full-length CDS alignments of putatively functional paralogs. Consensus trees and codon alignments were tested for directional selection within the HyPhy suite v2.5.8 (Pond et al. 2019). Generally, the statistical approaches implemented via HyPhy estimate nonsynonymous substitution rates and synonymous substitution rates to generate a ratio (i.e., dN/dS) which provides information about the evolution of each codon. Estimates of pervasive selection assumes a constant dN/dS across the entire phylogeny for a single codon site. We tested for pervasive selection using FEL v2.1 (Pond and Frost 2005) using a selection cutoff of 0.05 for significance. With FEL, we detected positive selection at 3 sites for ORs and 22 sites for V2Rs. We detected pervasive purifying selection at 295 sites for ORs and 629 sites for V2Rs.
We tested for the signature of episodic evolution in chemoreceptor coding sequences using the MEME program (v2.1.2) in the HyPhy suite using a selection cutoff of 0.05 for significance. This method allows the estimate of the dN/dS ratio for a codon site to vary across the phylogeny revealing periods of positive and negative selection along a lineage (Murrell et al. 2012). The analysis also reports the number of branches under selection corresponding to each codon site, which we called "branch-codons" (total branch-codons = # codon sites in alignment × # branches in phylogeny). We detected episodic selection at 41 sites and 197 branch-codons in ORs and 445 sites and 3442 branch-codons in V2Rs suggesting transient periods of positive selection drove chemoreceptor diversification across the coding sequences.
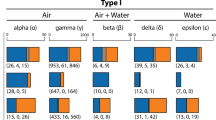
The ligand binding domains of chemoreceptors that provide scent specificity are generally located in the extracellular region, while the highly conserved 7-transmembrane domains and intracellular region provide the machinery for canonical GPCR signaling. We hypothesized that codon sites undergoing periods of positive selection would be biased toward the extracellular domains (Fig. 1b). For rattlesnake ORs and V2Rs, we confirmed all ligand binding and related features detected via NCBI’s CDD mapped to extracellular regions predicted by TMHMM (Lu et al. 2019; Krogh et al. 2001). We ran proportion z-tests evaluating whether the proportion of sites experiencing selection were significantly higher or lower for extracellular or intracellular regions. The proportions used for z-tests were based on the total number of sites per region detected by FEL and MEME using a selection cutoff of 0.05 for significance. For FEL, we tested if the proportion of positively selected sites was significantly higher or lower for extracellular or intracellular regions, and this test was repeated for the proportion of negatively selected sites (p < 0.05; Table 2). Signals for negative selection were significantly higher in the intracellular regions for both ORs and V2Rs, consistent with these regions performing conserved GPCR functions. For our measurements of episodic selection, we tested if the proportion of sites identified by MEME was significantly higher or lower for either extracellular or intracellular regions, and this test was repeated for branch-codons (p < 0.05; Table 2). Both ORs and V2Rs had proportionally more sites that had experienced episodic selection in their respective extracellular regions, however only the V2Rs were significant in the site-only analysis. For the branch-codon model, both ORs and V2Rs had significantly higher proportions of positive sites in extracellular regions, and this result is apparent in both gene phylogenies (Figs. 4 and 5). We colored branches shades of green corresponding with ( +) selection detected for extracellular sites (empirical Bayes factor > 100). A yellow star on the OR gene tree (Fig. 4) marks a particular Z-linked clade with widespread extracellular episodic selection, suggesting an evolutionary mechanism underlying potential sex-biased chemoperception. Among the autosomal V2Rs, we find lineages in the phylogeny with long branches extending from sub-clades experiencing positive selection in extracellular regions (Fig. 5). These rapidly evolving branches may be currently undergoing adaptive evolution, potentially for prey detection, suggesting the chemoreceptor repertoire is highly dynamic in both number of paralogs and sequence.
OR gene tree for Crotalus adamanteus colored by chromosome placement and relative signals of extracellular selection. The gene phylogeny represents a maximum likelihood reconstruction of full-length nucleotide coding sequences. Branches are shaded green based on signatures of episodic positive selection detected in extracellular sites (MEME analysis; empirical Bayes factor > 100): dark green distinguishes fewer sites detected per branch and light green distinguishes more sites detected per branch. Branches with no extracellular sites detected are colored black. The outside ring is colored according to chromosomal placements for individual genes (i.e., tips or leaves) on the tree. A yellow star on the tree marks a particular clade with candidate genes potentially evolving sex-biased function. Gene labels and bootstrap values were excluded for clarity; fully labeled consensus trees with gene names and bootstrap values can be found in the Supplementary Materials online (Supp_file_07a and Supp_file_07b). Tree figures were generated using iTOL via their online server (Letunic and Bork, 2019)
V2R gene tree for Crotalus adamanteus colored, labeled, and generated identical to the OR gene tree (Fig. 4). Fully labeled consensus trees with gene names and bootstrap values can be found in the Supplementary Materials online (Supp_file_07a and Supp_file_07b)
Conclusion
Rattlesnake sensory systems encompass phenotypic extremes representing a refined suite of predatory features involved in both the perception of prey (Saviola et al. 2013; Katti et al. 2018; Geng et al. 2011; Schraft et al. 2019) and prey capture (Margres et al. 2013, 2015; Rokyta et al. 2012, 2013). The majority of previous genomic studies on rattlesnakes focus largely on venom (Schield et al. 2019; Vonk et al. 2013; Yin et al. 2016). The eastern diamondback (Crotalus adamanteus) possesses one of the most thoroughly characterized venom systems (Rokyta et al. 2012; Margres et al. 2013) with reports of intraspecific venom variation tied to regional adaption to sympatric prey (Margres et al. 2016, 2017), phenotypic integration with fang and head morphologies (Margres et al. 2015), and ontogenetic shifts in venom composition as individual rattlesnakes mature (Rokyta et al. 2017). Although venom represents a crucial aspect of the rattlesnake predatory arsenal, the complete ecological context in which venom is deployed includes the chemosensory-based perception of prey-derived stimuli. Hence, characterizing the sensory repertoire of rattlesnakes directly complements our current understanding of their venoms by generating a more complete polyphenic representation of their predatory arsenal. Additionally, other behaviors directly relevant to fitness, including mate and offspring detection, depend on chemoperception.
We generated a comprehensive characterization of the chemoreceptor repertoire for a representative species, the eastern diamondback rattlesnake (Crotalus adamanteus) using genomic and transcriptomic approaches. We found that OR and V2R genes occur in large tandem-repeat arrays, suggesting a genomic mechanism where array duplications provide the extensive gene duplications needed to generate the large chemoreceptor complement. Additionally, we show for V2Rs that changes in this complement correspond with major changes in life-history traits and exhibits lineage-specific expansions and contractions demonstrating the potential ecological and evolutionary responsiveness of the chemoreceptor repertoire. We determined the chromosomal distribution of chemoreceptors, which suggests sex-biased chemoperception as a potential mechanism for observed male and female specific behaviors. Our selection analyses revealed the signature of episodic selection in the extracellular domains of chemoreceptors, an evolutionary mechanism for the diversification of ORs and V2Rs. Our combined results provide a basis for understanding the origins of a complex phenotype, chemoperception, and a stepping-off point for future comparative studies on the role of chemoperception in rattlesnake ecology and evolution.
Materials and Methods
Sampling, Sequencing, and Genome Assembly
Two Crotalus adamanteus specimens were utilized for genomic sequencing, and two additional specimens were utilized for chemosensory tissue RNA-seq. Animal ID#s, capture locality, and basic morphometric information is shown in Table 3. All specimens were collected from the wild under Florida Fish and Wildlife Conservation Commission permit #LSSC-13-00004A-C, and all procedures were approved by the Florida State University Animal Care and Use Committee (IACUC) under protocols #1529 and #1836.
For generating the genome assembly, we utilized an adult female eastern diamondback rattlesnake (ID = KW1264) collected from the Apalachicola National Forest in northern Florida (Leon County). Although not utilized for the genome assembly, genomic reads were also generated for later chromosomal assignments from an adult male rattlesnake (ID = KW0944) collected from southern Florida (Miami-Dade county). We extracted genomic DNA using a standard Phenol–Chloroform-Isoamyl alcohol extraction from blood that was frozen in 95% EtOH upon collection. High molecular weight DNA aliquots were set aside for long read sequencing, and the remaining genomic DNA was sheared using a Covaris S220 Focused-ultrasonicator with a target average fragment size of 500 bp. DNA libraries were prepared from the sheared DNA using the Illumina TruSeq DNA PCR-Free library prep kit and quality checked using a 1% agarose gel. KAPA PCR was performed to determine the amplifiable concentration prior to performing 250 bp paired-end sequencing. The complete genome was assembled de novo using a hybrid approach, utilizing Oxford Nanopore (MinION; 195,747 total reads; read N50 value of 12,913) and Pacific Biosciences (performed by GENEWIZ in South Plainfield, NJ using a Sequel II; 6,111,768 total reads; read N50 value of 1,468,549) long-reads at 25 × coverage combined and Illumina 250 bp paired-end data (Illumina HiSeq 2500; 71,531,898 total read pairs) at 60 × coverage, employing MaSuRCA version 3.2.8 (Zimin et al. 2013). The final assembly was checked for duplicate sequences resulting from haplotype variants or assembly artifacts using purge_dups v1.2.5 (https://github.com/dfguan/purge_dups.git). No duplicate or chimeric sequences were detected, and the genome was considered a haploid assembly moving forward. The male snake Illumina short reads were generated using the same protocol as the female (Illumina HiSeq 2500; 35,572,858 total read pairs).
Transcriptomics were implemented to identify chemosensory genes and outline exonic structures for genome annotations. We utilized two Crotalus adamanteus specimens (IDs = DRR0044 and MM0114; additional information on Table 3) for chemosensory tissue transcriptomics. Tissue samples were dissected from chemosensory regions postmortem via the roof of the buccal cavity in the mouth. We targeted olfactory epithelial tissues representing both the vomeronasal organ and the main olfactory bulb for DRR0044 and only the main olfactory bulb for MM0114. Tissue samples were buffered and stored in RNA-later at − 80 °C until RNA extraction. Total RNA was extracted using a TRIzol-chloroform extraction as previously described (Rokyta et al. 2012). Buffered chemosensory tissue was added to 500 µL of TRIzol (Invitrogen) and homogenized with a 20-gauge syringe. The TRIzol-tissue mixture was then thoroughly mixed with 20% chloroform followed by an additional 500 µL of TRIzol. Next, the mixture was transferred to a 5Prime phase lock heavy gel tube and centrifuged at 20 °C. The supernatant was extracted and mixed with glycogen before precipitating with isopropyl alcohol and centrifugation. The resulting totalRNA pellets were subject to a 75% ethanol wash and reconstitution in rnase-free water. Extracted RNA was quantified with a Qubit RNA Broad-range kit (Thermo Fisher Scientific) and quality checked on an Agilent Bioanalyzer using an RNA 6000 Pico chip, per the manufacturer’s instructions. Next, an NEBNext Poly(A) mRNA Magnetic Isolation Module (New England Biolabs) was used to isolate the mRNA. The isolated mRNA was fragmented for exactly 13 min to generate an average fragment size of 370 base pairs. Transcript cDNA libraries were prepped from isolated mRNA using the NEBNext Ultra RNA Library Prep kit with the High-Fidelity 2X Hot Start PCR Master Mix, Agencourt AMPure XP PCR Purification Beads, and Multiplex Oligos for Illumina sequencing (New England Biolabs). The resulting libraries were quantified and quality controlled on an Agilent Bioanalyzer using a High Sensitivity DNA chip. KAPA PCR was performed on the purified cDNA libraries by the Molecular Cloning Facility at Florida State University to determine amplifiable concentrations for sequencing. cDNA libraries were diluted to concentrations of ∼5 nM and pooled with other libraries to optimize sequencing capacity. The pooled sample of cDNA libraries was then quality checked with another High Sensitivity DNA Bioanalyzer chip, and the amplifiable concentrations were determined with KAPA PCR. The pooled cDNA libraries were sequenced on an Illumina HiSeq 2500 to generate 150 bp paired-end reads at the Florida State University College of Medicine Translational Laboratory. The resulting chemosensory tissue RNA-sequencing yields were 14,021,239 read pairs for MM0114 (olfactory bulb only) and 22,067,312 read pairs for DRR0044 (olfactory bulb & VNO). The % contamination and % trimmed was 0.014 and 0.5 respectively for MM0114, and 0.008 and 0.5 for DRR0044.
Chemosensory Gene Identification and Annotation
We implemented a conservative approach for chemosensory gene identification and annotation. Gene annotations and downstream analyses were visualized using Geneious v9.1.5 (Kearse et al. 2012). Genomic contigs potentially containing chemosensory genes were identified using BLAST and a custom reference gene database (Camacho et al. 2009). This database comprised full-length nucleotide coding sequences (e.g., start and stop codons confirmed; intact and complete 7-trans membrane domains) compiled from three snake species and two lizard species (including genera Anolis, Gekko, Protobothrops, Thamnophis, and Python; sequences downloaded from the NR database October, 2018). The finalized database included coding sequences spanning all genes identified or predicted to be GPCRs. Any genomic contigs with one or more BLAST hits from this database were isolated from the full-genome assembly, and temporary BLAST-hit annotations were overlaid to highlight target regions for downstream assessment and gene annotations.
Chemoreceptor gene annotations were hand-curated and guided by the RNA-seq transcriptome data aligned to the genome (transcriptome sequencing described in previous section). To identify and reduce sample cross contamination, raw reads were filtered and quality checked using custom scripts and FastQC v0.11.5. Adapters and low quality reads were trimmed from raw reads using Trim Galore! v0.4.5. (Krueger 2015; accessed October 2018). Cleaned read pairs were mapped to the genome using the splice/intronaware aligner HISAT2 v2.1.0 (Kim et al. 2015), and indexed BAM alignments were generated using SAMtools v1.6 (Li et al. 2009), effectively outlining exonic structures for expressed genes from mRNA transcripts. Annotations for putative chemoreceptors were first generated for highly expressed genes with > 100 × coverage overlapping the BLAST-hit regions on the isolated contigs. These initial high-confidence annotations later served as references for predicting structures (e.g., splice sites, starts, stops, UTRs) for genes with low or partial coverage. Stop codons occurring within the highly conserved regions of GPCRs (e.g., the 7-trans membrane domain) signaled pseudogenization (Hilger et al. 2018). Unexpressed genes (0 × coverage) similar to expressed chemoreceptors with intact protein domains were also annotated based on the expressed genes using BLAST and gene prediction tools available in Geneious (Kearse et al. 2012). Annotated genes were back-searched against the genome using BLAST as a final check to confirm all recognizable paralogs had been annotated. The finalized gene annotations include gene IDs, exons, coding sequences, mRNAs, UTRs, and poly-A tail signals. Pseudogenes and truncated genes were not explicitly targeted for this study; however, any occurring on contigs containing full-length genes were annotated to provide context for downstream synteny analyses. Due to the extreme diversity of ORs and V2Rs and high potential for novelty in snakes, these genes were named as OR_### and V2R_###, respectively, where increased numbering corresponds with increased CDS length (e.g., we named the shortest olfactory receptor “OR_001”). Finalized gene coding sequences were isolated for downstream phylogenetic analyses. Top BLAST-p hits and corresponding conventional names for translated coding sequences of the final consensus chemosensory genes can be found in the Supplementary Materials online (Supp_file_04; final BLASTp search completed July, 2020).
Qualitative protein assessments using translated chemosensory gene coding sequences were iteratively performed throughout the genome annotation process. We utilized NCBI’s Conserved Domain Database via their online server to check for completeness of conserved GPCR protein features from translated coding sequences (Marchler-Bauer et al. 2014; Lu et al. 2019; searches completed between October, 2017 and June, 2020). We also checked all annotated coding sequences for the presence of signal peptides using SignalP v5.0 via their online server (Armenteros et al. 2019; Emanuelsson et al. 2007; searches completed August 2019). Structure homology and oligomerization predictions based on highly similar amino acid sequences (i.e., above 30% identity) were also considered using SWISSMODEL Expasy via their online server (Waterhouse et al. 2018; Guex et al. 2009; Bertoni et al. 2017).
Chromosomal Inferences
To identify candidate sex chromosome linked scaffolds, genomic reads from an adult male (ID = KW0944) and an adult female (ID = KW1264) were mapped to the genome assembly and evaluated for read depth and coverage. Male and female short read data (Illumina sequencing described above) were aligned separately using BWA v0.7.12 and read depth and coverage was calculated using SAMtools v1.10 (Li et al. 2009). Sex chromosomes were assigned mirroring methods described by Yin et al. (2016), with several adjustments. Specifically, we adjusted depth criteria and coverage filtering to improve predictions by accounting for noise of potential chimeric contigs with sex chromosome identity for more than half their length. Males are the homogametic sex in snakes with two copies of the Z chromosome, so the expected W chromosome read mapping ratio of male to female (M:F) should be close to zero. We defined W chromosome contigs as those with M:F scaled mean read depth ratios less than 0.3, at least 50% female coverage, and less than 50% male coverage. The expected M:F scaled ratio of a Z chromosome contig is 2:1. We defined Z chromosome contigs as those with M:F scaled mean read depth ratios greater than 1.6 and at least 50% coverage for both male and female. All contigs not identified as Z or W were considered autosomal. We assigned autosomal contigs using RaGOO v1.1 with the chromosome-level Crotalus viridis genome assembly as the reference (Alonge et al. 2019; Schield et al. 2019). The reference Z-chromosome and previously assigned sex-linked contigs were excluded from RaGOO scaffolding to prevent errors from misplacing previously assigned sex-linked contigs. Any genomic contigs which failed to map to the reference were considered “Unplaced”, and contigs that mapped to sub-chromosome resolution regions of the reference assembly were considered “Unplaced contiguous”. Chimeric contigs identified and split by RaGOO were assigned to whatever chromosome comprised the larger percentage of the contig. Genes were assigned to chromosomes based on the final placements of their corresponding genomic contigs, and these assignments were included in the phylogenetic gene trees for Crotalus adamanteus ORs and V2Rs (Figs. 4 and 5), which were each overlaid with color rings representing chromosomal placement of tips. The final gene-chromosome assignments were also utilized for generating the proportional donut plots in Fig. 1c.
Phylogenetics and Selection Analyses
We limited phylogenetic inferences and selection analyses to coding sequences extracted from the C. adamanteus genome, and thus only interpreted gene family diversity and relationships across paralogs in a single species. To generate the most functionally informative alignments for membrane-bound chemosensory genes, we aligned translated OR and V2R coding sequences using PRALINE (Pirovano and Feenstra, 2008) via their online server with default parameters plus the TMHMM v2.0 transmembrane prediction option (Krogh et al. 2001; alignments completed June 2020). Detailed reports for these alignments and the extracted coding sequences can be found in the Supplementary Materials online (Supp_file_05 and Supp_file_06). Resulting amino acid alignments were back-translated using PAL2NAL v14.0 via their online server (Suyama et al. 2006). Back-translated alignments were utilized for downstream phylogenetic reconstructions and selection analyses.
Maximum likelihood trees were inferred for each alignment using IQ-TREE v2.0.5 (Nguyen et al. 2014; Chernomor et al. 2016; Kalyaanamoorthy et al. 2017; Hoang et al. 2017). Input alignments were specified as codon data using the -st CODON option, and the best substitution models were chosen automatically using ModelFinder (Kalyaanamoorthy et al. 2017). Branch support was assessed using ultrafast bootstrap approximations and single branch SH-like approximate likelihood ratio tests using the -B and -alrt commands with 1000 replicates each (Hoang et al. 2017; Guindon et al. 2010). Tree figures were generated using iTOL via their online server (Letunic and Bork 2019; the full IQ-TREE outputs with gene names and bootstrap values can be found in the Supplementary Materials online as Supp_file_07a and Supp_file_07b). Consensus trees and back-translated alignments were consolidated into nexus files and tested for directional selection within the HyPhy suite v2.5.8 (Pond et al. 2019). Specifically, we tested sites for pervasive selection using FEL v2.1 (Pond and Frost, 2005) and episodic selection using MEME v2.1.2 (Murrell et al., 2012). The compiled raw outputs of both FEL and MEME are available in the Supplementary Materials online (Supp_file_08). To investigate if selected sites were correlated with structural regions, we ran proportion z-tests comparing selected sites from extracellular and intracellular regions. Proportions were based on the number of selected extracellular and intracellular sites out of all extracellular and intracellular designated sites. We first tested this using the FEL and MEME default selection cutoff of 0.1 for significance, and this was repeated using 0.05 for significance. Since significant findings were identical between the repeated analysis, only the results from the latter are shown in Table 2. For FEL, we tested if the proportion of positively selected sites was significantly higher or lower for extracellular or intracellular regions, and this was repeated for the proportion of negatively selected sites (p < 0.05). For MEME, we tested if the proportion of episodically diversifying sites was significantly higher or lower for extracellular or intracellular regions, and this was repeated for branch-codons (p < 0.05). Test result summaries can be found in Table 2, and full outputs from the selection analyses and statistical tests can be found in the Supplementary Materials online (Supp_file_08).
Comparative Analysis of V2R Arrays
To demonstrate the utility of our thorough chemosensory characterization for C. adamanteus, we traced the evolution of a distinguishable V2R array between five squamate species with available genomes (Fig. 3). In addition to our C. adamanteus assembly, we downloaded the following genomes from Genbank (accessed January 18th, 2021) and assessed them in Geneious Prime v2021.0.3: Crotalus tigris (Genbank accession GCA_016545835.1), Thamnophis elegans (GCA_009769535.1), Varanus komodoensis (GCA_004798865.1), and Anolis carolinensis (GCA_000090745.2). Full length V2R gene sequences were extracted from the C. adamanteus genome and used as queries for BLAST searches of the other genomes. We selected a V2R array with tractable homology between the five genomes for further analysis. This array is distinguished by several conserved immunoglobulin type-V (IG-V) subunit containing regions flanking and interspersing the tandemly repeated V2R genes (Fig. 3a). The full array was recovered from a single contiguous genomic region from Crotalus tigris (contig ID = VORL01000046.1), Thamnophis elegans (NC_045558.1 (reversed)), and Anolis carolinensis (NW_003339038.1 (reversed)), while manual scaffolding was necessary to recover the complete array in the correct orientation from C. adamanteus (contig IDs = Cadam-autosomal_5021_of_8908, Cadam-autosomal_949_of_8908, Cadam-autosomal_239_of_8908 (reversed), Cadam-autosomal_7936_of_8908 (reversed), and Cadam-autosomal_8446_of_8908) and V. komodoensis (SJPD01000131.1 (reversed), SJPD01000180.1 (reversed), SJPD01000374.1 (reversed), and SJPD01000152.1). The manually scaffolded arrays were simply full-length contig concatenations with no trimming.
To identify potential homology between the lizard and snake V2Rs, we assessed synteny and generated a reduced V2R gene phylogeny using only the genes identified from this array across the five genomes. Most V2R gene annotations among the downloaded genomes were either incomplete or missing from our BLAST-matched regions. For consistency, we manually annotated these regions as described above for C. adamanteus. Full-length gene sequences spanning the predicted exons, introns, and UTRs (Fig. 3b) were extracted, combined, and aligned using MUSCLE v3.8.425 with default parameters and two iterations as is recommended for larger sequences (Edgar, 2004). The nucleotide alignment was manually trimmed to the shortest sequences on each end to minimize end-gap biases. The final alignment we used to generate the phylogeny is available in the supplementary materials online (Supp_file_9). We generated a maximum likelihood gene phylogeny representing this array using IQ-TREE v2.0.5 (Nguyen et al. 2014; Chernomor et al. 2016; Kalyaanamoorthy et al. 2017; Hoang et al. 2017), with identical input commands described earlier minus the -st CODON option. The resulting IQ-TREE output with gene names and bootstrap values can be found in the Supplementary Materials online (Supp_file_9). A handful of snake V2R genes grouped with lizard V2R genes on the phylogeny, and these were colored red and orange in Fig. 3c. To visualize this relationship overlaid onto the syntenic representation of the gene array, we included red and orange color shading connecting these "lizard-like" V2R regions in Fig. 3a. General information corresponding to the additional genomes can be found in the supplemental materials online (Supp_file_08).
Data Availability
All gene annotations and sequence analyses correspond with the final Crotalus adamanteus genome assembly (MaSuRCA) available at NCBI: assembly version Cadam-genome_11369, GenBank accession JADPQB000000000. Raw sequencing data from PacBio (SRR12802468), Nanopore (SRR12802467), Illumina WGS for female (SRR12802470) and male (SRR12802469), and RNA-seq (SRR12802472 and SRR12802471) are available under BioProject PRJNA667573. The extracted chemosensory gene coding sequences utilized for downstream phylogenetics and selection analyses are included in the supplemental materials online (Supp_file_06).
References
Akiyoshi S, Ishii T, Bai Z, Mombaerts P (2018) Subpopulations of vomeronasal sensory neurons with coordinated coexpression of type 2 vomeronasal receptor genes are differentially dependent on Vmn2r1. Eur J Neurosci 47:887–900
Alonge M, Soyk S, Ramakrishnan S, Wang X, Goodwin S, Sedlazeck F, Lippman Z, Schatz M (2019) RaGOO: fast and accurate reference-guided scaffolding of draft genomes. Genome Biol 20:224
Armenteros J, Tsirigos K, Sønderby C, Nordahl Petersen T, Winther O, Brunak S, Heijne G, Nielsen H (2019) SignalP 5.0 improves signal peptide predictions using deep neural networks. Nat Biotechnol 37:420–425
Baeckens S, Herrel A, Broeckhoven C, Vasilopoulou-Kampitsi M, Huyghe K, Goyens J (2017) Evolutionary morphology of the lizard chemosensory system. Sci Rep 7:1–13
Baldwin M, Ko MC (2020) Functional evolution of vertebrate sensory receptors. Horm Behav 124:104771
Bear DM, Lassance JM, Hoekstra H, Datta SR (2016) The evolving neural and genetic architecture of vertebrate olfaction. Curr Biol 26:R1039–R1049
Bertoni M, Kiefer F, Biasini M, Bordoli L, Schwede T (2017) Modeling protein quaternary structure of homo- and hetero-oligomers beyond binary interactions by homology. Sci Rep 7:10480
Bowmaker J (2008) Evolution of vertebrate visual pigments. Vision Res 48:2022–2041
Brykczynska U, Tzika AC, Rodriguez I, Milinkovitch MC (2013) Contrasted evolution of the vomeronasal receptor repertoires in mammals and squamate reptiles. Genome Biol Evol 5:389–401
Buck L, Axel R (1991) A novel multigene family may encode odorant receptors: a molecular basis for odor recognition. Cell 65:175–187
Camacho C, Coulouris G, Avagyan V, Ma N, Papadopoulos J, Bealer K, Madden TL (2009) Blast+: architecture and applications. BMC Bioinform 10:421
Chernomor O, von Haeseler A, Minh B (2016) Terrace aware data structure for phylogenomic inference from supermatrices. Syst Biol 65:997–1008
Chess A, Simon I, Cedar H, Axel R (1994) Allelic inactivation regulates olfactory receptor gene expression. Cell 78:823–834
Cooper W (1995) Evolution and function of lingual shape in lizards, with emphasis on elongation, extensibility, and chemical sampling. J Chem Ecol 21:477–505
Cooper W (1997a) Correlated evolution of prey chemical discrimination with foraging, lingual morphology and vomeronasal chemoreceptor abundance in lizards. Behav Ecol Sociobiol 41:257–265
Cooper W (1997b) Independent evolution of squamate olfaction and vomerolfaction and correlated evolution of vomerolfaction and lingual structure. Amphibia-Reptilia 18:85–105
Daghfous G, Smargiassi M, Libourel PA, Ruddy W, Bels V (2012) The function of oscillatory tongue-flicks in snakes: Insights from kinematics of tongue-flicking in the banded water snake (Nerodia fasciata). Chem Senses 37:883–896
Davies W, Collin S, Hunt D (2012) Molecular ecology and adaptation of visual photopigments in craniates. Mol Ecol 21:3121–3158
Dong D, He G, Zhang S, Zhang Z (2009) Evolution of olfactory receptor genes in primates dominated by birth-and-death process. Genome Biol Evol 1:258–264
Duvall D, Arnold SJ, Schuett GW (1992) Pitviper mating systems: ecological potential, sexual selection, and microevolution. In: Campbell JA, Brodie ED (eds) Biology of the pitvipers. Selva Publishing, Tyler, TX, pp 321–336
Edgar R (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Emanuelsson O, Brunak S, von Heijne G, Nielsen H (2007) Locating proteins in the cell using TargetP, SignalP and related tools. Nat Protoc 2:953–971
Filoramo N, Schwenk K (2009) The mechanism of chemical delivery to the vomeronasal organs in squamate reptiles: a comparative morphological approach. J Exp Zool 311:20–34
Francia S, Silvotti L, Ghirardi F, Catzeflis F, Percudani R, Tirindelli R (2014) Evolution of spatially coexpressed families of type-2 vomeronasal receptors in rodents. Genome Biol Evol 7:272–285
Geng J, Liang D, Jiang K, Zhang P (2011) Molecular evolution of the infrared sensory gene TRPA1 in snakes and implications for functional studies. PLoS ONE 6:e28644
Gether U, Kobilka B (1998) G protein-coupled receptors. J Biol Chem 273:17979–17982
Gracheva E, Ingolia N, Kelly Y, Cordero-Morales J, Hollopeter G, Chesler A, Sánchez E, Pérez J, Weissman J, Julius D (2010) Molecular basis of infrared detection by snakes. Nature 464:1006–1011
Graves B (1993) Chemical delivery to the vomeronasal organs and functional domain of squamate chemoreception. Brain Behav Evol 41:198–202
Grus W, Shi P, Zhang Y, Zhang J (2005) Dramatic variation of the vomeronasal pheromone receptor gene repertoire among five orders of placental and marsupial mammals. Proc Natl Acad Sci USA 102:5767–5772
Guex N, Peitsch M, Schwede T (2009) Automatedcomparativeproteinstructuremodeling with SWISS-MODEL and Swiss-PdbViewer:a historical perspective. Electrophoresis 30(suppl):S162–S173
Guindon S, Dufayard JF, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321
Halpern M (1987) The organization and function of the vomeronasal system. Annu Rev Neurosci 10:325–362
Halpern M, Martínez-Marcos A (2003) Structure and function of the vomeronasal system: an update. Prog Neurobiol 70:245–318
Hayden S, Bekaert M, Crider T, Mariani S, Murphy W, Teeling E (2010) Ecological adaptation determines functional mammalian olfactory subgenomes. Genome Res 20:1–9
Hilger D, Masureel M, Kobilka B (2018) Structure and dynamics of GPCR signaling complexes. Nat Struct Mol Biol 25:4–12
Hoang D, Chernomor O, von Haeseler A, Minh B, Vinh L (2017) UFBoot2: Improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522
Holding ML, Kern EH, Denton RD, Gibbs HL (2015) Fixed prey cue preferences among Dusky Pigmy Rattlesnakes (Sistrurus miliarius barbouri ) raised on different long-term diets. Evol Ecol 30:1–7
Hughes G, Boston E, Finarelli J, Murphy W, Higgins D, Teeling E (2018) The birth and death of olfactory receptor gene families in Mammalian niche adaptation. Mol Biol Evol 35:1390–1406
Hunt D, Carvalho L, Cowing J, Davies W (2009) Evolution and spectral tuning of visual pigments in birds and mammals. Philos Trans R Soc Lond B 364:2941–2955
Kalyaanamoorthy S, Minh B, Wong T, von Haeseler A, Jermiin L (2017) ModelFinder: Fast model selection for accurate phylogenetic estimates. Nat Methods 14:587–591
Katti C, Stacey-Solis M, Coronel-Rojas NA, Davies W (2018) Opsin-based photopigments expressed in the retina of a South American pit viper, Bothrops atrox (Viperidae). Vis Neurosci 35:e027
Kearse M, Moir R, Wilson A, Stones-havas S, Sturrock S, Buxton S, Cooper A (2012) Geneious Basic: an integrated and extendable desktop software platform for the organization and analysis of sequence data. Bioinformatics 28:1647–1649
Keverne E (1999) The vomeronasal organ. Science 286:716–720
Khan I, Yang Z, Maldonado E, Li C, Zhang G, Gilbert M (2015) Olfactory receptor subgenomes linked with broad ecological adaptations in sauropsida. Mol Biol Evol 32:2832–2843
Kim D, Langmead B, Salzberg SL (2015) HISAT: a fast spliced aligner with low memory requirements. Nat Methods 12:357-U121
Kishida T, Hikida T (2010) Degeneration patterns of the olfactory receptor genes in sea snakes. J Evol Biol 23:302–310
Kishida T, Kubota S, Shirayama Y, Fukami H (2007) The olfactory receptor gene repertoires in secondary-adapted marine vertebrates: evidence for reduction of the functional proportions in cetaceans. Biol Let 3:428–430
Kishida T, Go Y, Tatsumoto S, Tatsumi K, Kuraku S, Toda M (2019) Loss of olfaction in sea snakes provides new perspectives on the aquatic adaptation of amniotes. Philos Trans R Soc Lond B 286:20191828
Krogh A, Larsson B, Heijne G, Sonnhammer E (2001) Predicting transmembrane protein topology with a hidden markov model: application to complete genomes. J Mol Biol 305:567–580
Krueger F (2015) Trim Galore. A wrapper tool around Cutadapt and FastQC to consistently apply quality and adapter trimming to FastQ files, with some extra functionality for MspI-digested RRBS-type (Reduced Representation Buisulfite-Seq) libraries. URL https://github.com/FelixKrueger/TrimGalore. Accessed Oct 2018
Letunic I, Bork P (2019) Interactive Tree Of Life (iTOL) v4: recent updates and new developments. Nucleic Acids Res 47:W256–W259
Li H, Handsaker B, Wysoker A, Fennell T, Ruan J, Homer N, Marth GT, Abecasis GR, Durbin R (2009) The sequence alignment/map format and SAMtools. Bioinformatics 25:2078–2079
Lind A, Lai Y, Mostovoy Y, Holloway KA, Iannucci A, Mak CYA, Fondi M, Orlandini V, Eckalbar W, Milan M, Rovatsos M, Kichigin GI, Makunin A, Trifonov V, Schijlen E, Kratochvil L, Fani R, Jessop ST, Patarnello T, Bruneau B (2019) A high-resolution,chromosome-assignedKomododragongenomerevealsadaptationsin the cardiovascular, muscular, and chemosensory systems of monitor lizards. Nat Ecol Evolut 3:1241–1252
Liu A, He F, Shen L, Liu R, Wang Z, Zhou J (2019) Convergent degeneration of olfactory receptor gene repertoires in marine mammals. BMC Genomics 20:1–14
Lu S, Wang J, Chitsaz F, Derbyshire M, Geer R, Gonzales N, Gwadz M, Hurwitz D, Marchler G, Song J, Thanki N, Yamashita R, Yang M, Zhang D, Zheng C, Lanczycki C, Marchler-Bauer A (2019) CDD/SPARCLE: the conserved domain database in 2020. Nucleic Acids Res 48:1–4
Marchler-Bauer A, Derbyshire M, Gonzales N, Lu S, Chitsaz F, Geer L, Geer R, He J, Gwadz M, Hurwitz D, Lanczycki C, Lu F, Marchler G, Song J, Thanki N, Wang Z, Yamashita R, Zhang D, Zheng C, Bryant S (2014) CDD: NCBI’s conserved domain database. Nucleic Acids Res 43:D222–D226
Margres MJ, Mcgivern JJ, Wray KP, Seavy M, Calvin K, Rokyta DR (2013) Linking the transcriptome and proteome to characterize the venom of the eastern diamondback rattlesnake (Crotalus adamanteus). J Proteomics 96:145–158
Margres MJ, Wray KP, Seavy M, Mcgivern JJ, Sanader D, Rokyta DR (2015) Phenotypic integration in the feeding system of the eastern diamondback rattlesnake (Crotalus adamanteus). Mol Ecol 24:3405–3420
Margres MJ, Walls R, Suntravat M, Lucena S, Sánchez EE, Rokyta DR (2016) Functional characterizations of venom phenotypes in the eastern diamondback rattlesnake (Crotalus adamanteus) and evidence for expression-driven divergence in toxic activities among populations. Toxicon 119:28–38
Margres MJ, Wray KP, Hassinger A, Mcgivern JJ, Lemmon EM, Rokyta DR (2017) Quantity, not quality: rapid adaptation in a polygenic trait proceeded exclusively through expression differentiation. Mol Biol Evol 34:3099–3110
Martini S, Silvotti L, Shirazi A, Ryba N, Tirindelli R (2001) Co-expression of putative pheromone receptors in the sensory neurons of the vomeronasal organ. J Neurosci 21:843–848
McClintock T, Sammeta N (2003) Trafficking prerogatives of olfactory receptors. Neuro Rep 14:1547–1552
Mombaerts P (2004) Genes and ligands for odorant, vomeronasal and taste receptors. Nat Rev Neurosci 5:263–278
Murrell B, Wertheim J, Moola S, Weighill T, Scheffler K, Pond S (2012) Detecting individual sites subject to episodic diversifying selection. PLoS Genet 8:e1002764
Nei M, Rooney AP (2005) Concerted and birth-and-death evolution of multigene families. Annu Rev Genet 39:121–152
Nguyen LT, Schmidt H, von Haeseler A, Minh B (2014) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum-likelihood phylogenies. Mol Biol Evol 32:268–274
Niimura Y (2009) On the origin and evolution of vertebrate olfactory receptor genes: comparative genome analysis among 23 chordate species. Genome Biol Evol 1:34–44
Niimura Y (2012) Olfactory receptor multigene family in vertebrates: from the viewpoint of evolutionary genomics. Curr Genomics 13:103–114
Niimura Y, Nei M (2006) Evolutionary dynamics of olfactory and other chemosensory receptor genes in vertebrates. J Hum Genet 51:505–517
Niimura Y, Nei M (2007) Extensive gains and losses of olfactory receptor genes in mammalian evolution. PLoS ONE 2:e708
Niimura Y, Matsui A, Touhara K (2018) Acceleration of olfactory receptor gene loss in primate evolution: Possible link to anatomical change in sensory systems and dietary 782 transition. Mol Biol Evol 35:1437–1450
Pan D, Zhang L (2008) Tandemly arrayed genes in vertebrate genomes. Comp Funct Genomics 2008:545269
Patel J, Brown C, Ytreberg F, Stenkamp D (2018) Predicting peak spectral sensitivities of vertebrate cone visual pigments using atomistic molecular simulations. PLoS Comput Biol 14:e1005974
Perry B, Card D, McGlothlin J, Pasquesi G, Adams R, Schield D, Hales N, Corbin A, Demuth PJ, Hoffmann F, Vandewege M, Schott R, Bhattacharya N, Chang B, Casewell N, Whiteley G, Velasco RJ, Mackessy S, Gamble T, Castoe T (2018) Molecular adaptations for sensing and securing prey and insight into amniote genome diversity from the gartersnake genome. Genome Biol Evolut 10:2110–2129
Pirovano W, Feenstra KA (2008) PralineTM: a strategy for improved multiple alignment of transmembrane proteins. Bioinformatics (Oxford, England) 24:492–497
Pond S, Frost S (2005) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22:1208–1222
Pond S, Poon A, Velazquez R, Weaver S, Hepler N, Murrell B, Shank S, Rife Magalis B, Bouvier D, Nekrutenko A, Wisotsky S, Spielman S, Frost S, Muse S (2019) Hyphy 2.5-a customizable platform for evolutionary hypothesis testing using phylogenies. Mol Biol Evol 37:295–299
Rokyta DR, Lemmon AR, Margres MJ, Aronow K (2012) The venom-gland transcriptome of the eastern diamondback rattlesnake (Crotalus adamanteus). BMC Genomics 13:312
Rokyta DR, Wray KP, Margres MJ (2013) The genesis of an exceptionally lethal venom in the timber rattlesnake (Crotalus horridus) revealed through comparative venomgland transcriptomics. BMC Genomics 14:e394
Rokyta DR, Margres MJ, Ward M, Sánchez EE (2017) The genetics of venom ontogeny in the eastern diamondback rattlesnake (Crotalus adamanteus). PeerJ 5:e3249
Rosenbaum D, Rasmussen S, Kobilka B (2009) The structure and function of G-protein-coupled receptors. Nature 459:356–363
Saviola AJ, Chiszar D, Busch C, Mackessy SP (2013) Molecular basis for prey relocation in viperid snakes. BMC Biol 11:20
Schield D, Card D, Hales N, Perry B, Pasquesi G, Blackmon H, Adams R, Corbin A, Smith C, Ramesh B, Demuth PJ, Betrán E, Tollis M, Meik J, Mackessy S, Castoe T (2019) Theoriginsandevolutionofchromosomes,dosagecompensation,and mechanismsunderlyingvenomregulationinsnakes. Genome Res 29:590–601
Schraft HA, Bakken G, Clark RW (2019) Infrared-sensing snakes select ambush orientation based on thermal backgrounds. Sci Rep 9:e3950
Schweikert L, Fitak R, Caves E, Sutton T, Johnsen S (2018) Spectral sensitivity in rayfinnedfishes:diversity,ecology,andshareddescent. J Exp Biol 221:jeb.189761
Schwenk K (1993) The evolution of chemoreception in squamate reptiles: a phylogenetic approach. Brain Behav Evol 41:124–137
Schwenk K (1995) Of tongues and noses: chemoreception in lizards and snakes. Trends Ecol Evol 10:7–12
Shi P, Zhang J (2007) Comparative genomic analysis identifies an evolutionary shift of vomeronasal receptor gene repertoires in the vertebrate transition from water to land. Genome Res 17:166–174
Silva L, Antunes A (2017) Vomeronasal receptors in vertebrates and the evolution of pheromone detection. Annu Rev Anim Biosci 5:353–370
Stranger BE, Forrest MS, Dunning M, Ingle CE, Beazley C, Thorne N, Redon R, Bird CP, de Grassi A, Lee C, Tyler-Smith C, Carter N, Scherer SW, Tavaré S, Deloukas P, Hurles ME, Dermitzakis ET (2007) Relative impact of nucleotide and copy number variation on gene expression phenotypes. Science 315:848–853
Suyama M, Torrents D, Bork P (2006) PAL2NAL: robust conversion of protein sequence alignments into the corresponding codon alignments. Nucleic Acids Res 34:W609–W612
Van Hazel I, Sabouhanian A, Day L, Endler J, Chang B (2013) Functional characterization of spectral tuning mechanisms in the great bowerbird short-wavelength sensitive visual pigment (SWS1), and the origins of UV/violet vision in passerines and parrots. BMC Mol Biol 13:250
Vicoso B, Emerson J, Zektser Y, Mahajan S, Bachtrog D (2013) Comparative sex chromosome genomics in snakes: differentiation, evolutionary strata, and lack of global dosage compensation. PLoS Biol 11:1001643
Vonk FJ, Casewell NR, Henkel CV, Heimberg AM, Jansen HJ, McCleary RJR, Kerkkamp HM, Vos RA, Guerreiro IM, Calvete JJ, Williams DJ, Woods AE, Logan JM, Harrison RA, Castoe TA, de Koning APJ, Pollock DD, Yandell M, Calderón DRV, Renjifo C, Currier RB, Salgado DC, Pla D, Sanz LC, Hyder AS, Ribeiro JMC, Arntzen JW, van den Thillart G, Boetzer M, Pirovano W, Dirks RP, Spaink HP, Duboule D, McGlinn E, Kini RM, Richardson MK (2013) The king cobra genome reveals dynamic gene evolution and adaptation in the snake venom system. Proc Natl Acad Sci USA 110:20651–20656
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer TF, de Beer APT, Rempfer C, Bordoli L, Lepore A, Schwede T (2018) SWISS-MODEL: homologymodellingofproteinstructuresandcomplexes. Nucleic Acids Res 46:W296–W303
Weldon PJ, Ortiz R, Sharp TR (1992) Thechemicalecologyincrotalinesnakes. In: Campbell JA, Brodie ED (eds) Biology of the Pitvipers. Selva Publishing, Tyler, TX, pp 309–319
Wu H, Wang C, Gregory K, Han G, Cho H, Xia Y, Niswender C, Katritch V, Meiler J, Cherezov V, Conn P, Stevens R (2014) Structure of a class C GPCR metabotropic glutamate receptor 1 bound to an allosteric modulator. Science 344:58–64
Yang H, Shi P, Zhang PY, Zhang J (2005) CompositionandevolutionoftheV2R vomeronasalreceptorgenerepertoireinmiceandrats. Genomics 86:306–331
Yang L, Jiang H, Wang Y, Lei Y, Chen J, Sun N, Lv W, Wang C, Near JT, He S (2019) Expansion of vomeronasal receptor genes (OlfC) in the evolution of fright reaction inOstariophysanfishes. Commun Biol 2:235
Yin W, Zj W, Qy Li, Jm L, Zhou Y, Bz Lu, Lj J, Px Q, Zhang P, Wb Z, Wen B, Yj H, Zl L, Qiu Bt Su, Xw YH, Gj Z, Gm Y, Zhou Q (2016) Evolutionary trajectories of snake genes and genomes revealed by comparative analyses of five-pacer viper. Nat Commun 7:13107
Yokoyama S (2008) Evolution of dim-light and color vision pigments. Annu Rev Genomics Hum Genet 9:259–282
Yokoyama S, Xing J, Liu Y, Faggionato D, Altun A, Starmer W (2014) Epistatic adaptive evolution of human color vision. PLoS Genet 10:e1004884
Yokoyama S, Tada T, Liu Y, Faggionato D, Altun A (2016) A simple method for studying the molecular mechanisms of ultraviolet and violet reception in vertebrates. BMC Evol Biol 16:64
Zimin AV, Marçais G, Puiu D, Roberts M, Salzberg SL, Yorke JA (2013) The MaSuRCA genome assembler. Bioinformatics 29:2669–2677
Acknowledgements
The authors thank Kenneth Wray, Mark J. Margres, Alyssa Hassinger, Schyler A. Ellsworth, Gunnar S. Nystrom, Kylie Lawrence, Laura E. Koffinas, Rachel Saul, Alex Oliver, and Simone Gable for assistance with animal collection and maintenance. Special thanks to Jack Facente for providing his invaluable expertise regarding venomous snake husbandry.
Funding
This work was supported by the National Science Foundation (DEB 1145978 and DEB 1638902 to DRR and DEB 1638872 to HLG).
Author information
Authors and Affiliations
Contributions
MPH conceived the study. MPH, ACW, MBB, HLG, and DRR are responsible for the intellectual and technical development of the project. MPH, MJW, MBB, and DRR generated and assembled the sequencing data. MPH generated genome annotations, performed analyses, and made the figures. MPH and ACW wrote the manuscript. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Additional information
Handling editor: David Liberles.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Hogan, M.P., Whittington, A.C., Broe, M.B. et al. The Chemosensory Repertoire of the Eastern Diamondback Rattlesnake (Crotalus adamanteus) Reveals Complementary Genetics of Olfactory and Vomeronasal-Type Receptors. J Mol Evol 89, 313–328 (2021). https://doi.org/10.1007/s00239-021-10007-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-021-10007-3