Abstract
High mutation rates select for the evolution of mutational robustness where populations inhabit flat fitness peaks with little epistasis, protecting them from lethal mutagenesis. Recent evidence suggests that a different effect protects small populations from extinction via the accumulation of deleterious mutations. In drift robustness, populations tend to occupy peaks with steep flanks and positive epistasis between mutations. However, it is not known what happens when mutation rates are high and population sizes are small at the same time. Using a simple fitness model with variable epistasis, we show that the equilibrium fitness has a minimum as a function of the parameter that tunes epistasis, implying that this critical point is an unstable fixed point for evolutionary trajectories. In agent-based simulations of evolution at finite mutation rate, we demonstrate that when mutations can change epistasis, trajectories with a subcritical value of epistasis evolve to decrease epistasis, while those with supercritical initial points evolve towards higher epistasis. These two fixed points can be identified with mutational and drift robustness, respectively.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
When a population is in mutation–selection balance, it is able to maintain its mean fitness while still generating genetic variation that may increase its fit to the environment via adaptive mutations (Goyal et al. 2012). However, this balance between the evolutionary forces of selection and mutation can sometimes be precarious. When mutation rates become too high, for example, mutations can overpower selection leading to the extinction of a population via lethal mutagenesis (Bull et al. 2007). Similarly, when population size dwindles, selection can become so weak that deleterious mutations cannot be eliminated, leading to fitness decline via Muller’s ratchet (Haigh 1978) or population extinction through a mutational meltdown (Lynch et al. 1993). Populations can adapt to high mutation rates and/or small population sizes by evolving “mutational robustness” (Wilke and Adami 2003) or “drift robustness” (Kondrashov 1994; LaBar and Adami 2017; Lan et al. 2017). Populations evolve mutational robustness by moving onto flat fitness peaks, where they experience a reduction in maximum fitness counterbalanced by an increased fraction of new mutations that are either neutral or have a small fitness effect (Wilke et al. 2001; Franklin et al. 2019); this phenomenon is often referred to as the “survival-of-the-flattest” effect (Wilke et al. 2001). Robustness to drift, on the other hand, appears to involve favoring fitness peaks that have steep flanks, enabled by mutations that are synergistic in their deleterious effect (Kondrashov 1994; LaBar and Adami 2017), while reducing (rather than increasing) the likelihood of mutations with small effect, and increasing the fraction of mutations that are lethal. Interestingly, a recent re-analysis of the survival-of-the-flattest effect has shown that an increase in the fraction of lethal mutations is also seen in the response to high mutation rates (Franklin et al. 2019), suggesting that resistance to drift and resistance to mutations are intertwined (see also Lan et al. 2017).
The threat of high mutation rates and small population sizes to genetic survival is particularly real for populations that periodically undergo bottlenecks during transmission between hosts and cannot rely on sexual recombination to protect against gene loss, such as the mitochondria of the salivarian Trypanosomes T. brucei and T. vivax (Speijer 2006). For those organisms, population size often drops into the single digits (Oberle et al. 2010) while mutation rates are elevated due to oxidative stress (Koffi et al. 2009). High mutation rates and small population sizes are also important for viral populations. Mutational robustness (and possibly drift robustness) has been observed in some strains of the RNA virus vesicular stomatitis virus (VSV) that differ in the rate at which deleterious mutations accumulate at small population size (Sanjuán et al. 2007).
How genomes respond to mutations is determined to a large extent by how mutations interact. In general, the effect of a mutation on host fitness is influenced by the genetic background within which that mutation occurs, a phenomenon known as epistasis (Wolf et al. 2000). Epistasis has a direction: the effect of a pair of mutations can either be larger or smaller than what is expected from a single mutation, so that the deleterious effect of two mutations can be either amplified (synergistic), or buffered (antagonistic). The average direction between pairs (also called directional epistasis, see for example Wilke and Adami 2001) plays an important role in determining linkage equilibria (Charlesworth 1976; Barton 1995), canalization (Scharloo 1991; Gibson and Wagner 2000), as well as theoretical investigations of the origin of sex (Kondrashov 1982, 1988; Westy et al. 1999). Epistasis has been measured quantitatively for a number of model organisms, and both antagonistic and synergistic trends have been observed (de Visser et al. 1997; Elena and Lenski 1997; Bonhoeffer et al. 2004; Burch and Chao 2004; Sanjuán et al. 2005; Beerenwinkel et al. 2007; Jasnos and Korona 2007).
When faced with changed conditions, one of the ways in which populations can adapt is by changing the way information is encoded in the genome, leading to changes in epistasis (Gros et al. 2009). Here we study the impact of epistasis on both drift and mutational robustness in a simple model fitness landscape. We show that whether a population predominantly displays drift or mutational robustness is largely determined by the average value of directional epistasis: populations occupying a peak with synergistic epistasis above a critical value will tend to evolve towards drift-robust peaks (by moving towards peaks with increased positive epistasis), while those inhabiting peaks with subcritical epistasis will respond by lowering epistasis until mutations are mostly neutral, consistent with mutational robustness. Thus, evolutionary trajectories for populations under evolutionary stress will bifurcate towards drift-robust or mutationally robust fixed points.
Model
We study a simple fitness landscape in which the wild-type genotype resides on a fitness peak with a height of 1, and the fitness of a k-mutant is given by
where \(s=-\log f(1)\) is the mean effect of a deleterious mutation to the wild type, q determines the degree of directional epistasis, and genotypes have a finite number of binary loci L (see, e.g., Wilke and Adami 2001)Footnote 1. In such a model \(q=1\) signals absence of epistasis (i.e., the fitness landscape is multiplicative), \(q>1\) describes a peak with synergy between deleterious mutations (synergistic epistasis), and \(q<1\) is indicative of buffering mutations (antagonistic epistasis). When \(q>1\) we sometimes speak of negative epistasis (because the combined two-mutant fitness is lower than the multiplicative expectation), while \(q<1\) indicates positive epistasis (the double-mutant is higher in fitness than expected on the basis of the single-mutation effect). Of course, a model that only treats the mean epistatic effect between mutations using a single parameter q has significant limitations. In particular, such a model cannot capture effects that are due to a distribution of pair-wise epistatic effects (something that can be delivered by an NK model, for example Østman et al. 2012). Furthermore, we ignore here all the subtleties of sexual reproduction, which can also affect how epistasis evolves. The loss of realism is offset by our ability to control the parameters of such an effective model precisely (s and q), which in more sophisticated models depend on each other. Furthermore, an analysis in terms of asexual processes is warranted for those genomic stretches in strong linkage disequilibrium.
We can analytically calculate the evolutionary dynamics of a population on this fitness landscape in the weak mutation limit, \(N\mu \ll 1\), where N is the effective population size and \(\mu \) is the mutation rate per genome per generation. In this limit, the population is monomorphic and individual mutations either rapidly go to fixation or are lost to drift (McCandlish and Stoltzfus 2014). The dynamics of an evolving population in this limit can be mathematically represented as a Markov process, where the state of the Markov process at time t corresponds to the predominant genotype present in the population at that time, and a transition to a new state corresponds to the fixation of a new mutation (Sella and Hirsh 2005; McCandlish and Stoltzfus 2014). For sufficiently large times t, the Markov process reaches stationarity, at which point its probability to reside in any given state is provided by the equilibrium distribution \(p_k\). We can interpret \(p_k\) as the probability to observe the population centered around a genotype carrying k mutations at any point in time.
Assuming that we know the equilibrium distribution \(p_k\), we can calculate the population mean fitness \(f_{\rm eq}\) by averaging over the stationary distribution of k-mutants,
Importantly, \(f_{\rm eq}\) represents an average over time. It is the mean fitness in the population when averaged both over all individuals in the population and over a long period of time.
The distribution \(p_k\) can be calculated using the transition probability \(P(0\rightarrow k)\) in the Markov process, solving the detailed balance equations (Sella and Hirsh 2005). More precisely, detailed balance entails that in a process where the transitions \(i\rightarrow j\) and \(j\rightarrow i\) are both possible, the number of changes \(n_{i\rightarrow j}\) must equal the number \(n_{j\rightarrow i}\). To obtain \(n_{i\rightarrow j}\) and \(n_{j\rightarrow i}\), we calculate the number of k-mutants that go to fixation, starting with the wild-type sequence with fitness f(0), and compare this with the number of k-mutants that are replaced by the wild type. Using the Sella–Hirsh fixation formula (2005) that is appropriate for a haploid Wright–Fisher process, for a k-mutant with fitness f(k), we find
The reverse rate is then
so that the detailed balance condition becomes
In Eq. (5), \(p_0\) is the equilibrium density of the wild type, while \(p_k\) is the (combined) equilibrium density of all individual k-mutants. Equations (3)–(5) then lead to the solution (Sella and Hirsh 2005)
In this expression, Z is the partition function
For \(q=1\) we can obtain a closed-form expression for the equilibrium fitness,
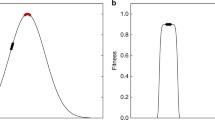
that shows clearly the steep fitness drop with decreasing population size that is due to genetic drift. But different values for q affect the fitness drop differently. In Fig. 1a, we can see the dependence of \(f_{\rm eq}\) on the population size for the multiplicative model (\(q=1\)), a model with positive epistasis (\(q=2.0\)), as well as the case of negative epistasis (\(q=0.5\)), evaluated at \(s = 0.01\) and \(L=100\). The model suggests that while positive epistasis protects from a fitness drop for moderate population sizes (higher mean equilibrium fitness), the drop becomes severe once populations dwindle below 100.
a Equilibrium fitness Eq. (2) as a function of population size N for strong positive epistasis (\(q=2.0\), dark gray), no epistasis (\(q=1.0\), teal), and strong negative epistasis (\(q=0.5\), light gray). b Equilibrium fitness as a function of epistasis q, for three different population sizes (red: \(N=10\), dark red: \(N=50\), black: \(N=100\)). In both panels, we used \(s = 0.01\) and \(L=100\). The equilibrium fitness tends to increase with increasing N but displays a minimum at intermediate q (colour figure online)
In fact, plotting \(f_{\rm eq}\) against q as in Fig. 1b reveals a fitness minimum as a function of q, suggesting that fitness loss via drift can be prevented in two different ways: high positive epistasis or high negative epistasis, while populations with weak or no epistasis appear to be the most vulnerable.
Two Regimes: Selection and Neutral Drift
The minimum in mean equilibrium fitness apparent in Fig. 1b can be seen as interpolating between two regimes: the neutral drift regime and the selection regime. To formalize these two regimes, we define the critical epistasis parameter \(q^\star \) at which mean fitness is minimal. Then, the neutral drift regime corresponds to \(q\ll q^\star \) and the selection regime corresponds to \(q\gg q^\star \). In the selection regime, an organism’s fitness declines rapidly with increasing number of mutations, and this rapid decline effectively limits the maximum number of mutations an organism can carry. By contrast, in the neutral drift regime additional mutations have increasingly smaller effects on organism fitness, and as a consequence selection cannot effectively purge deleterious mutations.
When \(q\ll q^\star \), selection cannot effectively purge deleterious mutations, and consequently the evolutionary dynamics are dominated by neutral drift. We can estimate the mean equilibrium fitness in the neutral regime by using
that is, the distribution given by Eq. (6) but with \(f(k)\equiv 1\). To calculate the mean fitness under this distribution, we insert this expression for \(p_k\) into Eq. (2), but note that we need to keep the original expression for f(k) in Eq. (2). The idea is that in the drift regime fitness differences are sufficiently small that they have no influence on the mutant distribution \(p_k\). This does not mean, however, that all organisms have a fitness of 1. The result of this derivation is the dashed line in Fig. 2, which agrees with the full model for sufficiently small q.
On the other hand, when epistasis between deleterious mutations is synergistic (\(q\gg q^{\star }\)), we are operating in the limit of strong selection: In this limit, fitness declines super-exponentially with increasing k, so that fitness is effectively 0 for sufficiently large k. Consequently, genotypes with k exceeding some number \(k_{\max}\) can be ignored, because they are removed by selection and do not contribute to mean fitness. We can model this effect by truncating the sum over k in the expression for mean fitness,
where \(k_{\max}<L\). We see that the truncated mutation model agrees with the full solution for large q (Fig. 2, dotted line).
Equilibrium fitness as a function of the epistasis parameter q for \(N=10\), \(s=0.01\), and \(L=100\). The solid black line represents the full model, Eq. (2), while the dashed blue line represents the drift-driven approximation, Eq. (9), and the dotted brown line represents the selection-driven approximation, Eq. (10) with a maximum number of mutations \(k_{\max}=20\). For \(q\lesssim 0.8\), the solid black line lies exactly on top of the dashed blue line, and for \(q\gtrsim 1.7\), the solid black line lies exactly on top of the dotted brown line. Thus, the two approximations represent the full model well in the two limits of small and large q, respectively (colour figure online)
One of the most striking features of the interplay between the neutral regime and the selection regime is the appearance of a minimum mean fitness (as a function of epistasis) where the drop of fitness is largest. The location of this minimum \(q^\star \) (reflecting the amount of directional epistasis that leads to the largest fitness loss) depends on the population size, the mean deleterious effect of mutations, and the number of loci (Fig. 3). Another less pronounced fitness minimum appears at significantly higher q (\(q>4\)) because epistasis this high essentially leads to truncation selection, limiting the number of mutations that contribute to the equilibrium fitness to just one (a “selection-driven” model as the one shown in Fig. 2, but with \(k_{\max}=1\)). In principle, this minimum gives rise to another attractive fixed point that is separated from the stronger attractor at lower q by an unstable fixed point. Because epistasis levels this high are rarely (if ever) observed in nature, we do not consider these additional fixed points any further.
To estimate the epistasis coefficient at which the steady-state fitness is at its minimum, we analyze the stationary distribution of fitness, Eq. (6), which apart from the normalization constant Z consists of two factors, \(\left( {\begin{array}{c}L\\ k\end{array}}\right) \) and \(f(k)^{2N-2}=\exp [-sk^{q}(2N-2)]\). As discussed in the derivations to Eqs. 9 and 10, these two factors represent neutral drift and selection, respectively. Importantly, for most values of k the binomial coefficient \(\left( {\begin{array}{c}L\\ k\end{array}}\right) \) is much larger than 1, whereas the selection term \(\exp [-sk^{q}(2N-2)]\) is much smaller than 1. Further, to the left of the minimum the binomial coefficient dominates the product, whereas to the right of the minimum the selection term dominates. Thus, at the minimum we expect the two factors to cancel, i.e., have a combined value of \(\sim 1\). To arrive at an expression that is independent of k, we maximize \(\left( {\begin{array}{c}L\\ k\end{array}}\right) \) by setting \(k=L/2\). Then, the condition for maximal fitness loss (where drift maximally balances selection) becomes
We can solve this equation for \(q^\star \) by using the Stirling approximation (\(\log n! = n \log n - n\)) to expand the binomial coefficient. We obtain for the minimum \(q^{\star }\) that
We test this estimate by comparing it to the numerically inferred minimum obtained via numerically minimizing Eq. (2) and find that Eq. (12) generally performs well, though it has a tendency to overestimate the true value of \(q^{\star }\) by a few percent (Fig. 3). Importantly, Eq. (12) captures the correct functional relationship between \(q^{\star }\) and the model parameters. In particular, the location of \(q^{\star }\) is primarily determined by the product of s and N, and not by their individual values. Further, because the product sN enters the expression for \(q^{\star }\) via a double-log, \(q^{\star }\) changes very slowly even if sN changes by orders of magnitude.
Relationship between the critical epistasis value \(q^{\star }\) and the selection coefficient s, for different population sizes N (\(L=100\) throughout). The lines represent analytically derived \(q^{\star }\), as given by Eq. (12). The dots represent \(q^{\star }\) values obtained by numerically minimizing Eq. (2). Throughout the entire parameter range, Eq. (12) provides a good approximation to the true location of the minimum
Increased Mutation Rate Exacerbates Fitness Loss in the Selection Regime
The theoretical results shown above were derived in the weak mutation limit where every mutation is either lost or goes to fixation before another mutation occurs in the population. In this section we study how finite mutation rates modify those results.
We simulate finite populations on a single-peak fitness landscape at finite mutation rates \(\mu \) using stochastic simulation. The population evolves asexually, and the population size is held constant over time for all simulations. For each combination of mutation rates and epistasis parameters, we simulated populations sizes \(N=10\) and \(N=100\), as well as selection coefficients \(s=0.01\) and \(s=0.001\). We recorded the mean fitness of the population over a period of time after a population reached steady state (see Methods), as a proxy for this equilibrium fitness \(f_{\rm eq}\). The simulations of the evolutionary process on the fitness landscape defined by Eq. (1) recover the theoretical results well for small mutation rates, as expected. As the mutation rate increases, we see notable departures from the weak mutation limit for the selection regime (larger q), while the neutral (drift) regime is largely unaffected by the increased rates (Fig. 4).
In particular, we notice that the minimum of the equilibrium fitness shifts towards higher q (Fig. 4). Furthermore, while for small mutation rates an increased epistasis protects from the loss of fitness due to genetic drift (mean fitness does not drop appreciably), it is clear that higher mutation rates negate this effect, and instead exacerbate the loss of fitness. Indeed, the increased mutation rate mimics the effect of a smaller population size (see Fig. 1b), which is expected as the effective population size decreases with mutation rate.
Comparison between theoretical equilibrium fitness and simulated equilibrium fitness. The black line represents theoretical fitness of the population at steady state, Eq. (2). The dots represent mean fitness of a population at steady state over ten simulations for different mutation rates (see legend). The error bars represent the standard error. For almost all cases, error bars are smaller than the symbol size. a–d Theoretical fitness and simulated fitness for different population sizes (N), epistasis coefficient (q), and selection coefficient (s). Mutation rate seems to have no effect on equilibrium fitness to the left of the fitness minimum, but to the right higher mutation rates lead to systematically lower equilibrium fitness values
While the depressed equilibrium fitness suggests that there are two routes to withstand genetic drift at small population sizes, it is not clear whether evolutionary trajectories could indeed bifurcate.
Bifurcation Analysis of Survival Strategies
The minimum in \(f_{\rm eq}\) at \(q^\star \) suggests that if q were a dynamical variable, then \(q^\star \) represents an unstable fixed point of the evolutionary dynamics. While q is not a dynamical variable in the usual sense, we can simulate it by endowing each genotype with a particular value of q that can be changed via mutation. In such a simulation, the statistics of the mutational process affecting q (the rate of change \(\mu _q\) as well as the mean change per mutation \(\Delta q\)) matter, so we test multiple different values for each.
It is worth pointing out that a genotype-dependent q appears to contradict the idea of fitness optimization in a landscape with a fixed fitness function such as Eq. (1). Such a function suggests that as a population climbs this peak, the parameters q and s are unaffected by this climb. While this is true for such a simple fitness function, it does not hold for more realistic evolutionary landscapes (for example in digital life Adami 1998; Adami et al. 2000; Wilke and Adami 2001; Ofria et al. 2009), where the mean effect of mutations s and the directional epistasis q are not fixed properties of the landscape, but instead emerge as properties of the local neighborhood in genetic space. As a consequence, moving in this space (via mutations) will affect both s and q. We attempt to simulate part of that dynamics by allowing q to adapt (while keeping s fixed). If selection favors a particular value of epistasis, we should see a gradual change in the mean epistasis \(\bar{q}\) of a population.
Mean epistasis q as a function of time t for different combinations of \(\mu _q\) and \(\Delta {q}\) in simulations with evolving q. Each line corresponds to the mean over ten replicates. The labels on top of each panel specify the mutation rate of q, \(\mu _q\), and the mutation step size, \(\Delta {q}\). The different colors correspond to different starting points of q. Each population had a fixed q until \(t=200,000\), for equilibration, and then q was allowed to evolve. Other simulation parameters were \(\mu =0.01\), \(N=100\), \(s=0.01\), and \(L=100\). We observe bistable behavior, such that populations with a mean q below the critical value experience continued decline in mean q, whereas populations with a mean q above the critical value do not (colour figure online)
In Fig. 5, we show how the mean epistasis parameter \(\bar{q}\) (averaged over sequences in the population) changes over time when populations are seeded with different seed organisms with fixed initial q. A bifurcation is indicated when trajectories move to different future fixed points given different initial states. While we can see clear signs of a bifurcation when plotting the mean trajectory in q-space over time (Fig. 5), viewing each trajectory separately reveals significant variation among them. In particular, for trajectories that are initialized with a q above the unstable fixed point \(q^\star \), some trajectories still move towards the low-q fixed point, yet the mean of trajectories may appear constant or nearly so (for example, the third panel in Fig. 6). We also emphasize that for q near or above \(q^\star \), the evolution of q proceeds extremely slowly, over hundreds of millions of generations, so that we cannot guarantee that populations are ever fully equilibrated in our simulations. It is possible that if we ran simulations for billions or trillions of generations, all populations would eventually reach the absorbing boundary at \(q=0.\)
Individual population trajectories for the simulations shown in the middle panel of Fig. 5 (\(\mu _q=0.001, \Delta q =0.001\)). Thin gray lines correspond to the evolution of individual populations, and the thick colored lines trace the mean among the replicate populations, as in Fig. 5. The labels on top of each panel indicate the initial population epistasis \(q(t=0)\) in that panel (colour figure online)
Discussion
The dynamics of evolution in asexual population is well understood in the common population-genetic limits, namely vanishingly small mutation rate and large population (weak mutation and strong selection). When mutation rates are high and selection is weak, the classic theoretical results are undermined by new effects such as mutational robustness (effect of large mutation rate) and drift robustness (effect of small population size), as anticipated by generalized population-genetic models such as “free-fitness” evolution (Iwasa 1988; Aita et al. 2004; Sella and Hirsh 2005; Barton and Coe 2009). Such theoretical models posit that Darwinian evolution does not optimize reproduction rate, but rather a combination of terms (the “free fitness,” in analogy to the free energy concept of statistical physics) that includes the reproduction rate as well as a term proportional to the inverse of population size and one proportional to mutation rate. In such theories, it is possible to increase the free fitness by trading reproduction rate for robustness to mutations, to drift, or both.
In most populations, we expect both mutational and drift robustness to contribute to survival. For example, when the mutation rate is large, the effective population size is diminished, so that both mutational and drift robustness are bound to be intertwined. The mean directional epistasis between mutations plays a role in both effects. While fitness peaks for mutationally robust populations tend to be flatter with little epistasis between mutations, we also observe something akin to truncation selection (Wilke et al. 2001; Franklin et al. 2019). In drift robustness, we observe both an increase in neutral mutations as well as an increase in strongly deleterious and lethal mutations, mediated by strong negative epistasis (\(q>1)\).
Here, we have calculated the mean equilibrium fitness of a population in the limit of small mutation rates using a simple fitness function with variable epistasis and tunable mutation effect-size, and we have found a minimum as a function of the mean directional epistasis parameter q that depends on population size. Stochastic simulations of adaptation on this landscape suggest that the minimum also depends on mutation rate. The model further suggests that there are two attractive fixed points for evolutionary dynamics, namely small q where mutations become nearly neutral, and large q where deleterious mutations interact synergistically. The low-q fixed pointFootnote 2 can be identified with mutational robustness (\(q\approx 0\)). In contrast, the \(q>1\) fixed point is reminiscent of drift robustness.
While the existence of a minimum in equilibrium fitness is suggestive of an unstable fixed point \(q^\star \) at which evolutionary trajectories bifurcate towards a low-q and a high-q fixed point, an agent-based simulation of such trajectories in a bit-string fitness model implementing Eq. (1) but with variable q paints a more complicated picture. It is clear from inspection of Eq. (1) that for every single sequence, a reduction of q while keeping k constant increases fitness (as \(\partial f/\partial q<0\)), no matter what the mean q of the population. This means that the population will sense an evolutionary pressure to reduce q independently of the mean population epistasis. However, if \(q>q^\star \), a secondary selective pressure appears that acts via the fitness distribution of a sequence’s offspring. For sequences with \(q>q^\star \), sequences with higher q have on average offspring with higher fitness than those with lower q, leading to a second-order selective pressure to increase q. However, in any particular fitness trajectory, there is a chance that a sequence with \(q<q^\star \) is among the offspring. Such a sequence may then go to fixation and abrogate the evolutionary trajectories leading towards a \(q>q^\star \), even though the selective pressure towards higher q is still present. We also expect that the likelihood of mutations that create sequences with \(q<q^\star \) in the offspring distribution depends on \(\mu _q\) as well as \(\Delta q\). This is precisely what we observe in Figs. 5 and 6: while for small \(q<q^\star \) the trend towards the mutationally robust fixed point is evident, at \(q>q^\star \) the mean epistasis across ten replicate experiments often shows a decrease (or remains constant) even though theoretically we expect an approach towards the drift-robust fixed point. The distribution of fitness trajectories shown in Fig. 6 shows that while some trajectories indeed move towards higher q, the possibility of mutating towards \(q<q^\star \) leads to trajectories in which the secondary selective pressure towards higher q is muted. Indeed, trajectories towards \(q>q^\star \) are absent among the replicates with initial \(q<q^\star \), reinforcing the conclusion that a critical amount of epistasis separates a population’s response to evolutionary stress either in a mutationally robust, or a drift-robust manner.
Throughout this work, we have used relatively small population sizes around \(N=100\) or less, as any investigation of drift robustness necessitates sufficiently small populations sizes such that drift can play a significant role in the evolutionary dynamics. However, our analytical results provide a more nuanced picture of the conditions under which our results may be relevant. First, we can see from Eq. (12) that the location of \(q^{\star }\) depends only on the product sN (i.e., the scaled selection coefficient) for sufficiently large N. Thus, drift robustness can act even for very large population sizes as long as s is sufficiently small. However, this is only true with one additional caveat: There can only be a fitness minimum separating the drift and selection regimes if the number of sites L is sufficiently large (on the order of 1/s), so that a large number of deleterious mutations can accumulate. By contrast, if both s and L are small, then the mean fitness is always approximately 1 and whether mutations are present or absent in a genotype makes virtually no difference.
While it is difficult to extrapolate results obtained using the abstract fitness function Eq. (1) to more complex landscapes in which many different peaks with different effect sizes and directional epistasis exist at the same time, our results support the notion that mutational robustness and drift robustness are indeed two different effects, which are likely to be intertwined in realistic scenarios. In particular, it would be interesting to study the response of experimental populations exposed to different mutation rates and populations, something that is possible using strains of T. brucei, for example. In those eukaryotic parasites, the directional epistasis between mutations in mitochondrial genes is controlled in part by RNA editing leading to overlapping genes (Kirby and Koslowsky 2017). Because the rate of gene overlap strongly correlates with directional epistasis, the present theory predicts that strains that differ in the number of overlapping genes could take different evolutionary trajectories when subjected to severe bottlenecks. While experimental evolution over prolonged time with parasites through controlled bottlenecks is difficult, such experiments might reveal to us these hidden dimensions of genomic adaptation.
Methods
Evolutionary Model with Fixed Epistasis
For all simulations, we used an individual-based model using Wright–Fisher sampling for reproduction. Individuals in the population are represented by bit strings of length L, where each bit can either be in the wild-type state (0) or in the mutated state (1). The fitness of an individual is given by the number of 1s in the bit string. We refer to this number as k, with \(0\le k \le L\), and write fitness as \(f(k)=e^{-sk^{q}}\), where s is the selection coefficient and q is the epistasis coefficient.
A population of individuals is represented as a vector V of a length \(L+1\). Each component \(V_{k}\) of the vector corresponds to a bin counting the number of individuals with k mutations. Reproduction occurs in discrete generations, and the number of offspring is drawn from a multinomial distribution, such that the probability of reproduction for each mutation number k is given by the number of individuals in bin k, \(V_k\), and their fitness f(k). Population size N is held constant at all times. After reproduction, mutation events move individuals up or down one or more bins. To simplify the mutation process, we limit the total number of mutations that can occur to a single individual during one reproductive event. For most simulations, this limit is set to 3. However, for very high mutation rates, \(\mu = 1\), we set the maximum number of mutations to 4. (The mutation rate \(\mu \) is defined as the expected number of mutations per genome per duplication.)
The probability that an individual carrying k mutations mutates into an individual carrying j mutations is given by
where u is the per-site mutation rate, \(u = \mu /L\).
We simulated population sizes of \(N = 100\) and \(N = 10\). We set selection coefficients to 0.01 and 0.001. We used mutation rates \(\mu \) of 0.1, 0.01, 0.001, and 0.0001. For each combination of population size, selection coefficient, and mutation rate, we simulated ten replicates, each for 2.5 million generations. We used the first 1.5 million generations for equilibration, and we used the subsequent 1 million time steps to measure the mean fitness of the population for the given parameter settings.
Evolutionary Model with Evolving Epistasis
Similarly to simulations with fixed epistasis, we employed an individual-based bit-string model to simulate populations with evolving epistasis. In these simulations, a population is represented by two vectors, each of length N. Individuals are represented by the components of these two vectors. The first vector keeps track of the number of mutations k of each individual, and the second vector keeps track of the epistasis coefficient q of each individual. Fitness is again given by \(f(k)=e^{-sk^{q}}\).
Replication again is implemented as Wright–Fisher sampling. After reproduction, each individual is subjected to a two-step mutation process. First, an individual either gains a mutation (\(k\rightarrow k+1\)), loses a mutation (\(k\rightarrow k-1\)), or does not mutate, with probabilities for these events given by Eq. 13. Second, each individual’s epistasis parameter q may mutate. The probability that the q mutates is given by a second mutation rate, \(\mu _q\). When such a mutation even occurs, q either increases or decreases by a fixed amount (\(\Delta {q}\)), with equal probability. \(\Delta {q}\) remains constant for all individuals and across all generations within each simulation trajectory. However, epistasis is only set to evolve after an initial equilibration phase, which we set to 200,000 generations.
Data Analysis and Code
We wrote our simulations in Python (Python Software Foundation 2019), using the NumPy (Oliphant 2006) and SymPy (Meurer et al. 2017) libraries for numeric and symbolic manipulations of matrices, respectively. Downstream data analysis and visualization was performed in R (R Core Team 2019), making extensive use of the tidyverse family of packages (Wickham et al. 2019). Our simulation and analysis code is available at https://github.com/clauswilke/epistasis_evolution/ and it is archived on Zenodo at https://doi.org/10.5281/zenodo.3558802. Simulation datasets generated with this code are archived in the Texas Data Repository at https://doi.org/10.18738/T8/GUNX76.
Notes
The present model in which fitness declines as a function of genetic distance from the wild type (modulated by epistasis) gives rise to conclusions similar to what Fisher’s geometric model would predict, even though in Fisher’s model the distance from wild type is phenotypic rather than genetic (Tenaillon et al. 2007).
Note that while technically the low-q fixed point is \(q=0\), this value cannot be attained in any realistic population as such a landscape is completely neutral (\(f=1\)) in this limit.
References
Adami C (1998) Introduction to artificial life. Springer, New York
Adami C, Ofria C, Collier TC (2000) Evolution of biological complexity. Proc Natl Acad Sci 97:4463–4468
Aita T, Morinaga S, Husimi Y (2004) Thermodynamical interpretation of evolutionary dynamics on a fitness landscape in a evolution reactor. I Bull Math Biol 66:1371–1403
Barton NH (1995) A general model for the evolution of recombination. Genet Res 65:123–145
Barton NH, Coe JB (2009) On the application of statistical physics to evolutionary biology. J Theor Biol 259:317–324
Beerenwinkel N, Pachter L, Sturmfels B, Elena SF, Lenski RE (2007) Analysis of epistatic interactions and fitness landscapes using a new geometric approach. BMC Evol Biol 7:60
Bonhoeffer S, Chappey C, Parkin NT, Whitcomb JM, Petropoulos CJ (2004) Evidence for positive epistasis in HIV-1. Science 306:1547–1550
Bull JJ, Sanjuan R, Wilke CO (2007) Theory of lethal mutagenesis for viruses. J Virol 81:2930–2939
Burch CL, Chao L (2004) Epistasis and its relationship to canalization in the RNA virus phi 6. Genetics 167:559–567
Charlesworth B (1976) Recombination modification in a fluctuating environment. Genetics 83:181–195
de Visser JA, Hoekstra RF, van den Ende H (1997) An experimental test for synergistic epistasis and its application in chlamydomonas. Genetics 145:815–819
Elena SF, Lenski R (1997) Test of synergistic interactions among deleterious mutations in bacteria. Nature 390:395–397
Franklin J, LaBar T, Adami C (2019) Mapping the peaks: fitness landscapes of the fittest and the flattest. Artif Life 25:250–262
Gibson G, Wagner G (2000) Canalization in evolutionary genetics: a stabilizing theory? BioEssays 22:372–380
Goyal S, Balick DJ, Jerison ER, Neher RA, Shraiman BI, Desai MM (2012) Dynamic mutation-selection balance as an evolutionary attractor. Genetics 191:1309–1319
Gros P-A, Le Nagard H, Tenaillon O (2009) The evolution of epistasis and its links with genetic robustness, complexity and drift in a phenotypic model of adaptation. Genetics 182:277–293
Haigh J (1978) The accumulation of deleterious genes in a population—Muller’s ratchet. Theor Popul Biol 14:251–267
Iwasa Y (1988) Free fitness that always increases in evolution. J Theor Biol 135:265–281
Jasnos L, Korona R (2007) Epistatic buffering of fitness loss in yeast double deletion strains. Nat Genet 39:550–554
Kirby LE, Koslowsky D (2017) Mitochondrial dual-coding genes in Trypanosoma brucei. PLoS Negl Trop Dis 11:e0005989
Koffi M, De Meeûs T, Bucheton B, Solano P, Camara M, Kaba D, Cuny G, Ayala FJ, Jamonneau V (2009) Population genetics of Trypanosoma brucei gambiense, the agent of sleeping sickness in western africa. Proc Natl Acad Sci USA 106:209–214
Kondrashov AS (1982) Selection against harmful mutations in large sexual and asexual populations. Genet Res 40:325–332
Kondrashov AS (1988) Deleterious mutations and the evolution of sexual reproduction. Nature 336:435–440
Kondrashov AS (1994) Muller’s ratchet under epistatic selection. Genetics 136:1469–1473
LaBar T, Adami C (2017) Evolution of drift robustness in small populations of digital organisms. Nat Commun 8:1012
Lan Y, Trout A, Weinreich D M, Wylie C S (2017) Natural selection can favor the evolution of ratchet robustness over evolution of mutational robustness. bioRxiv 122087
Lynch M, Bürger R, Butcher D, Gabriel W (1993) The mutational meltdown in asexual populations. J Hered 84:339–344
McCandlish DM, Stoltzfus A (2014) Modeling evolution using the probability of fixation: history and implications. Q Rev Biol 89:225–252
Meurer A, Smith CP, Paprocki M, Čertík O, Kirpichev SB, Rocklin M, Kumar A, Ivanov S, Moore JK, Singh S, Rathnayake T, Vig S, Granger BE, Muller RP, Bonazzi F, Gupta H, Vats S, Johansson F, Pedregosa F, Curry MJ, Terrel AR, Roučka Š, Saboo A, Fernando I, Kulal S, Cimrman R, Scopatz A (2017) SymPy: symbolic computing in Python. PeerJ Comput Sci 3:e103
Oberle M, Balmer O, Brun R, Roditi I (2010) Bottlenecks and the maintenance of minor genotypes during the life cycle of Trypanosoma brucei. PLoS Pathog 6:e1001023
Ofria C, Bryson DM, Wilke CO (2009) Avida: a software platform for research in computational evolutionary biology. In: Komosinski M, Adamatzky A (eds) Artificial life models in software. Springer, London, pp 3–35
Oliphant TE (2006) A guide to NumPy. Trelgol Publishing, New York
Østman B, Hintze A, Adami C (2012) Impact of epistasis and pleiotropy on evolutionary adaptation. Proc R Soc B 279:247–256
Python Software Foundation (2019) The Python language reference. Python Software Foundation, Wilmington
R Core Team (2019) R: a language and environment for statistical computing. R Foundation for Statistical Computing, Vienna
Sanjuán R, Cuevas JM, Moya A, Elena SF (2005) Epistasis and the adaptability of an RNA virus. Genetics 170:1001–1008
Sanjuán R, Cuevas JM, Furió V, Holmes EC, Moya A (2007) Selection for robustness in mutagenized RNA viruses. PLoS Genet 3:e93
Scharloo W (1991) Canalization: genetic and developmental aspects. Annu Rev Ecol Syst 22:65–93
Sella G, Hirsh AE (2005) The application of statistical physics to evolutionary biology. Proc Natl Acad Sci USA 102:9541–9546
Speijer D (2006) Is kinetoplastid pan-editing the result of an evolutionary balancing act? IUBMB Life 58:91–96
Tenaillon O, Silander OK, Uzan J-P, Chao L (2007) Quantifying organismal complexity using a population genetic approach. PLoS ONE 2:e217
Westy SA, Lively CM, Read AF (1999) A pluralist approach to sex and recombination. J Evol Biol 12:1003–1012
Wickham H, Averick M, Bryan J, Chang W, D’Agostino McGowan L, François R, Grolemund G, Hayes A, Henry L, Hester J, Kuhn M, Lin Pedersen T, Miller E, Milton Bache S, Müller K, Ooms J, Robinson D, Paige Seidel D, Spinu V, Takahashi K, Vaughan D, Wilke C, Woo K, Yutani H (2019) Welcome to the tidyverse. J Open Source Softw 4:1686
Wilke CO, Adami C (2001) Interaction between directional epistasis and average mutational effects. Proc R Soc Lond B 268:1469–1474
Wilke CO, Adami C (2003) Evolution of mutational robustness. Mutat Res 522:3–11
Wilke CO, Wang JL, Ofria C, Lenski RE, Adami C (2001) Evolution of digital organisms at high mutation rates leads to survival of the flattest. Nature 412:331–333
Wolf JB, Brodie ED III, Wade MJ (eds) (2000) Epistasis and the evolutionary process. Oxford University Press, Oxford
Acknowledgements
We are grateful to an anonymous reviewer who drew our attention to the existence of the weaker secondary minimum of Eq. (2) at high epistasis. This work was supported in part by the National Science Foundation’s BEACON Center for the Study of Evolution in Action, under Contract No. DBI-0939454.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: David Liberles.
Rights and permissions
About this article
Cite this article
Sydykova, D.K., LaBar, T., Adami, C. et al. Moderate Amounts of Epistasis are Not Evolutionarily Stable in Small Populations. J Mol Evol 88, 435–444 (2020). https://doi.org/10.1007/s00239-020-09942-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-020-09942-4