Abstract
The tRNA nucleotidyltransferases and poly(A) polymerases belong to a superfamily of nucleotidyltransferases. The amino acid sequences of a number of bacterial tRNA nucleotidyltransferases and poly(A) polymerases have been used to construct a rooted, neighbor-joining phylogenetic tree. Using information gleaned from that analysis, along with data from the rRNA-based phylogenetic tree, structural data available on a number of members of the superfamily and other biochemical information on the superfamily, it is possible to suggest a scheme for the evolution of the bacterial tRNA nucleotidyltransferases and poly(A) polymerases from ancestral species. Elements of that scheme are discussed along with questions arising from the scheme which can be explored experimentally.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
RNA nucleotidyltransferases play an important role in the maturation, processing, and degradation of RNAs. The tRNA nucleotidyltransferases (TNTs) and poly(A) polymerases (PAPs) are members of a nucleotidyltransferase superfamily (NTSF) that also includes DNA polymerase β and kanamycin nucleotidyltransferase (Holm and Sander 1995). TNTs add CCA (cytidine–cytidine–adenine) to the 3′-ends of immature and damaged tRNAs and are found in archaea, bacteria, and eukaryotes (Deutscher 1990; Rammelt and Rossmanith 2016). As many bacterial tRNA genes do not encode the 3′-CCA, and as a mature 3′-end is required for the participation of tRNAs in protein synthesis, TNTs are essential enzymes in many species (Hartmann et al. 2009). In bacteria, an intact CCA end is also required for the efficient maturation of the 5′-ends of tRNA by RNase P (Guerrier-Takada et al. 1984; Oh and Pace 1994).
Polyadenylation of RNA 3′-ends, once thought to occur exclusively in eukaryotes, has been shown definitively to occur in bacteria as well (reviewed in (Mohanty and Kushner 2011)). The addition of poly(A) tails to the 3′-ends of bacterial RNAs facilitates their degradation by exoribonucleases with 3′–5′-degradative activities (Cao et al. 1997; Donovan and Kushner 1986).
The structure and function of TNTs and PAPs have been studied in some detail and a wealth of information exists on the activities of these enzymes (Tomita and Yamashita 2014). Less is known about their evolution although a number of researchers have speculated about the evolutionary relationships between them and other members of the nucleotidyltransferase superfamily (e.g., Martin and Keller 2004; Tomita and Weiner 2002; Yue et al. 1996).
In what follows, available phylogenetic, structural, and biochemical data are utilized to explore the evolution of bacterial TNTs and PAPs. This analysis leads to a scheme for the evolution of these important enzymes and also reveals a number of interesting and heretofore unrecognized features of that evolution.
Methods
Phylogenetic Analysis
Sequences of bacterial TNTs and PAPs were retrieved from the microbial genomes database at www.ncbi.nlm.nih.gov. Sequences were selected essentially at random from among the following bacterial phyla: Thermotogae, Thermodesulfobacteria, Synergistetes, Aquificae, Deinococcus, Cyanobacteria, Actinobacteria, Firmicutes, and the α-, β-, γ-, δ-, and ε-Proteobacteria. Sequences from one representative species were utilized for each of the first five phyla listed above, viz. Thermotoga maritima, Caldimicrobium thiodismutans, Synergistetes sp. 53_16, Aquifex aeolicus, and Deinococcus radiodurans, respectively. Sequences were examined from at least nine species from each of the remaining phyla specified above. A list of all the species utilized in this study along with the database annotations of their NTSFs and their probable enzymatic function is presented in Table S1 (Supplementary Material). It should be noted that the identity and activity of the proteins suggested by the phylogenetic tree depicted in Fig. 3 does not always correspond to the database annotation. In many cases, it was possible to assign a probable enzymatic function to each protein based on its similarity to its relatives with known functions. Enzymes whose functions have been verified biochemically are indicated in color in Table S1.
The NTSF sequences were aligned using M-COFFEE (Moretti et al. 2007; Wallace et al. 2006) with the default protein alignment parameters. The multiple sequence alignment (MSA) is provided in CLUSTAL format as Figure S1 in Supplementary Material. A phylogenetic tree was constructed from the MSA using PHYLIP 3.695 (Felsenstein 1989). The sequences were bootstrapped 1000 times and jumbled once. The bootstrapped sequences were used to generate 1000 distance matrices which were then entered into the program NEIGHBOR. The resulting trees were then entered into CONSENSE to produce an unrooted consensus tree using M1 as the consensus type with 0.7 as the input fraction. Thus, the consensus tree shows only those nodes with bootstrap scores of at least 700. The tree was displayed using TreeView 1.6.6 and was rooted using T. maritima as the outgroup.
In addition to the neighbor-joining tree, a maximum likelihood tree was constructed using the PROML program in PHYLIP. This program uses the Jones–Taylor–Thornton model of probabilities of amino acid change (Felsenstein 1989). The consensus tree was constructed using M1 as the consensus type and 0.7 as the input fraction and is depicted in Figure S2 of Supplementary Material.
Molecular Modeling
Structure files for the TNTs from T. maritima, A. aeolicus and for Escherichia coli poly(A) polymerase I were downloaded from the Uniprot database and were manipulated using Jmol 14.29.12 or Deep-View SwissPdb Viewer version 4.1.0. For the construction of Fig. 5, the structure file for the TNT from Geobacillus stearothermophilus (formerly Bacillus stearothermophilus) was also utilized. The relevant region of the structure was then displayed using Deep-View SwissPdb Viewer and hydrogen bonds were identified and inserted using the Compute H-bonds tool in that program. Structures of the TNTs from D. radiodurans and C. thiodismutans were modeled using SWISS-MODEL (Waterhouse et al. 2018). The PDB accession numbers for the structures used in this study are: G. stearothermophilus CCA-adding TNT, 1MIW; T. maritima CCA-adding TNT, 3H38; A. aeolicus A-adding TNT, 1VFG; A. aeolicus CC-adding TNT, 3WFO; and Escherichia coli, poly(A) polymerase I, 3AQK.
Results and Discussion
Features of the Bacterial tRNA Nucleotidyltransferases
As mentioned above, the TNTs are members of a nucleotidyltransferase superfamily (Holm and Sander 1995). Some years ago, Yue et al. demonstrated that the superfamily could be divided into two classes (Yue et al. 1996). Their Class I contains archaeal TNTs, eukaryotic poly(A) polymerases, eukaryotic terminal transferases, DNA polymerase β and several other enzymes. Class II contains the eukaryotic and bacterial TNTs and the bacterial poly(A) polymerases (Yue et al. 1996).
More recent biochemical analyses have revealed that there are two subclasses within the Class II bacterial TNTs. Many bacterial species contain a single enzyme that adds both the two Cs and the terminal A to an immature or damaged tRNA molecule (Rammelt and Rossmanith 2016; Xiong and Steitz 2006), and these enzymes constitute the first Class II subclass. The three-dimensional structures of the TNTs from several such species, including G. stearothermophilus and T. maritima, have been solved and studied in detail (Li et al. 2002; Toh et al. 2009). In their analysis of the G. stearothermophilus TNT, Li et al. compared its structure to that of a seahorse and identified four domains within that structure: the head, neck, body, and tail (Li et al. 2002). The same conformation is adopted by the CCA-adding TNT from T. maritima (Toh et al. 2009, Fig. 1a). Subsequent study has revealed that the head region is the catalytic domain of the enzyme, the neck domain regulates the number of nucleotides incorporated at the tRNA 3′-end and the specificity of that incorporation, the body binds the T and acceptor stems of the tRNA molecule, while the tail interacts with the TψC loop, preventing the tRNA from dissociating from the enzyme (Li et al. 2002; Toh et al. 2009; Tomita et al. 2004).
a Structure of the Thermotoga maritima tRNA nucleotidyltransferase, showing the seahorse-like shape of the enzyme. The head, neck, body, and tail regions are labeled as specified by Toh et al. (2009) and are colored as in that reference. b Structure of E. coli poly(A) polymerase I, showing the sea otter-like shape of the enzyme. Regions are labeled as indicated in Toh et al. (2011)
The second subclass within the bacterial TNTs contains those species that possess two TNTs, one of which adds the two Cs and a second which adds the terminal A residue (Tomita and Weiner 2001, 2002). This division of TNT labor has been observed in the Aquificae (Tomita and Weiner 2001), the Thermodesulfobacteria (based on the phylogenetic analysis presented herein, see further below), Deinococcus (Tomita and Weiner 2002), the Cyanobacteria (Tomita and Weiner 2002), in a few Firmicutes (Bralley et al. 2005; Neuenfeldt et al. 2008), and of particular interest, in some of the δ-Proteobacteria (Bralley et al. 2009). The crystal structure of the A. aeolicus A-adding enzyme has been solved (Tomita et al. 2004) and compared with that of the T. maritima CCA-adding TNT (Toh et al. 2009; Yamashita et al. 2015) and despite the difference in specificity, the seahorse structure is retained by the A. aeolicus enzyme (Tomita et al. 2004). The A. aeolicus CC-adding enzyme also adopts a seahorse-shaped structure (Yamashita et al. 2014) (see further below).
There are significant differences in the sizes of the CCA- and the CC- and A-adding enzymes in various species. The major contributor to these size differences is the presence in the CCA-adding TNT from the Thermotogae and of the A-adding enzymes from the Aquificae, the Cyanobacteria, and the δ-Proteobacteria of domains, which are absent from other bacterial NTSFs. Figure 2 shows a schematic representation of the structure of a putative ancestral CCA-adding enzyme, based on the structure of the T. maritima CCA-adding TNT. This enzyme is almost 900 amino acids in length (Table S1) and contains two N-terminal domains attached to its C-terminal catalytic region. One of the N-terminal domains is a member of the Nrn family, the family of nanoRNases, which degrade small oligoribonucleotides (Liao et al. 2018). The second N-terminal domain is the cystathionine β-synthase domain (CBS), which is known to regulate the activity of enzymes in various species (Anashkin et al. 2017). The A. aeolicus A-adding enzyme and the A-adding TNTs from the Cyanobacteria and the δ-Proteobacteria are similar in size to the T. maritima CCA-adding TNT and also contain the Nrn and CBS domains.
Schematic diagram of the structure of the ancestral CCA-adding TNT containing the Nrn, CBS and “core” CCA-adding domains. The diagram is drawn approximately to scale based on the T. maritima CCA-adding enzyme. The areas labeled NC1 and NC2 are small regions of amino acid sequence which are not conserved in the various CCA- and A-adding TNTs. The phylum names indicated above and below the diagram represent NTSFs derived from those regions of the ancestral CCA-adding enzyme
The possible role of these domains in facilitating the function of the TNTs, if there is one, is unknown. Indeed, it is possible to delete the N-terminal portion of both the T. maritima and the A. aeolicus TNTs, containing the accessory domains, without affecting the activity of the enzymes. In fact, only the C-terminal portion of these proteins was overexpressed in the studies that led to the solution of their crystal structures (Toh et al. 2009; Tomita et al. 2004 and see Fig. 1a). The areas labeled NC1 and NC2 in Fig. 2 are small regions of amino acid sequence which are not conserved in the various CCA- and A-adding TNTs.
Figure 2 indicates the regions of the schematic structure that are retained in the NTSFs from the various phyla discussed in this study. It is apparent that most modern bacterial species lack the Nrn and CBS domains found in the TNTs from the Thermotogae and Aquificae.
Poly(A) polymerases and the Distribution of NTSFs in Bacteria
As indicated above, a number of bacterial species contain PAPs and thus add extended 3′-tails to the ends of RNAs. These tails serve as substrates for the exoribonucleases that degrade RNAs from the 3′-end (reviewed in (Mohanty and Kushner 2011)). The enzyme responsible for 3′-poly(A) tail synthesis in the β- and γ-Proteobacteria is known as poly(A) polymerase I (PAP I) and is a product of the pcnB gene (Cao and Sarkar 1992; Liu and Parkinson 1989). As shown in Table S1, several δ-Proteobacteria contain proteins that are similar in sequence to PAP I and PAP activity has recently been demonstrated in one member of that phylum, Geobacter sulfurreducens (Bralley et al. 2009). No other bacterial phyla are known to contain PAPs of the type found in the β, γ- and δ-Proteobacteria (but see further below).
The crystal structure of Escherichia coli PAP I has been published (Toh et al. 2011). It is interesting that the PAP I conformation is described by the researchers who solved the structure as a sea otter rather than a seahorse (Fig. 1b).
Given the information summarized above on the nature and activities of the TNTs and PAPs, it is of interest to examine further their distribution in the various bacterial phyla represented in Table S1. Most bacterial species contain only a single NTSF, a CCA-adding TNT (e.g., the Thermotogae, Synergistetes (see below), most Firmicutes, the Actinobacteria, and probably the α- and ε-Proteobacteria). Other species contain separate CC- and A-adding TNTs (e.g., the Aquificae, the Thermodesulfobacteria, Deinococcus, the Cyanobacteria, and a few Firmicutes). The δ-Proteobacteria are especially interesting in that some species contain only the CC- and A-adding TNTs while others contain these two TNTS and a PAP (e.g., G. sulfurreducens (Bralley et al. 2009), see above). There are still other variations on the distribution themes. For example, Stigmatella erecta, a δ-proteobacterium, appears to contain a single NTSF, probably a CCA-adding enzyme (Table S1).
A Phylogeny of Bacterial NTSFs
As a first step in understanding the evolution of the bacterial NTSFs, a phylogenetic tree was constructed, using the sequences of the proteins from the species listed in Table S1. The neighbor-joining tree is shown in Fig. 3.
Neighbor-joining phylogenetic tree relating the NTSFs referred to in the text and listed in Table S1. The tree was constructed using PHYLIP 3.695 as described in Methods section and rooted using the T. maritima CCA-adding enzyme as the outgroup. Bootstrap scores are shown at the nodes. In most cases, the enzymes are specified as CCA-, CC-, or A-adding based on their placement in the phylogenetic tree relative to enzymes of known function (Table S1). See text for additional details. M. australis, H. pylori, and B. halodurans are highlighted in color because of the possible acquisition of their NTSF genes by horizontal gene transfer. The “deep branching” species, T. maritima and Synergistetes sp. 53_16, are indicated in dark and lighter red, respectively
Several features of the tree deserve comment. First, it should be noted, as indicated in Methods, that the bacterial species analyzed were chosen essentially at random from the phyla represented. Thus, apart from the choice of phyla, there was no bias in the selection of the species represented in the tree.
Second, the tree presented in Fig. 3 and used as the basis for the evolutionary model presented below is a distance-based neighbor-joining tree. Maximum likelihood (Figure S2) and maximum parsimony trees were also constructed from the multiple sequence alignment (Figure S1). Of the three methods, neighbor-joining produced the tree with a topology closest to that of the phylogenetic trees based on ribosomal RNAs (www.arb-silva.de/projects/living-tree) and conserved signature indels (Gupta 2016). Thus, the neighbor-joining tree was used in the development of the evolutionary scheme presented below.
Third, the tree is rooted with the T. maritima CCA-adding TNT as the outgroup. T. maritima is located at or near the base of the bacterial phylogenetic tree constructed using 16S and 23S ribosomal RNA sequences (www.arb-silva.de/projects/living-tree) and the T. maritima TNT has been well studied (Toh et al. 2009). It might be argued that an outgroup from an archaeal or eukaryotic species would be more suitable to this analysis. There are three nonbacterial species in which the activity of the CCA-adding enzyme has been identified biochemically, two Archaea, Archeoglobus fulgidus (Xiong et al. 2003) and Sulfolobus shibitae (Cho and Weiner 2004) and one Eukaryote, Saccharomyces cerevisiae (Reid et al. 2019). Of the three, only the S. cerevisiae TNT is a Class II enzyme (Yue et al. 1996). To examine the suitability of these species as potential outgroups, the T. maritima CCA-adding TNT sequence was used to perform a BLAST comparison to the CCA-adding enzymes from these three species. In the case of A. fulgidus and S. shibitae, there was no similarity between these two protein sequences and the T. maritima sequence in the region corresponding to the “core” CCA-adding domain (Fig. 2) of T. maritima. The S. cerevisiae CCA-adding enzyme showed only 29% identity to the T. maritima enzyme over 267 amino acids in the “core” CCA-adding region with an E value of 7e−18. Based on these results, and the fact that T. maritima is near the base of the rRNA-derived phylogenetic tree, it was deemed most appropriate to use T. maritima as the outgroup for the phylogenetic analysis rather than an archaeal or eukaryotic species.
Fourth, in addition to T. maritima, several of the other species examined appear near the base of the rRNA-derived phylogenetic tree, and are designated as “deep branching,” viz. Synergistetes, the Thermodesulfobacteria and the Aquificae (Table S1).
A Scheme for the Evolution of Bacterial NTSFs
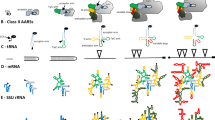
Using the phylogenetic analysis presented in Fig. 3, the rRNA-based phylogenetic tree, the structural data available on a number of NTSFs and summarized above, and other biochemical information for the NTSFs, it is possible to suggest a scheme for the evolution of the NTSFs from an ancestral enzyme form. Such a scheme is shown in Fig. 4. Elements of the scheme are as follows.
Scheme for the evolution of the bacterial RNA nucleotidyltransferases. Details of the scheme are provided in the text. The ancestral CCA-adding enzyme is depicted as in Fig. 2. For convenience, the nonconserved regions are not shown. The numbers indicate the qualitative genetic changes required to produce the various subclasses of tRNA nucleotidyltransferases and poly(A) polymerases that are found in modern bacteria. Note that the order of appearance of the various NTSFs as indicated in the figure may not reflect the actual order in which they appeared in evolutionary time. Nevertheless, the Thermotogae, Synergistetes, Aquificae, Thermodesulfobacteria, Cyanobacteria, and Deinococci are listed in the same vertical order in which they appear along the vertical axis of the rRNA-based bacterial phylogenetic tree (www.arb-silva.de/projects/living-tree). The Actinobacteria, Firmicutes, and Proteobacteria are also placed in the order in which they appear in the rRNA-based tree. HGT denotes horizontal gene transfer. Phyla that are proposed to have acquired NTSF genes by HGT are indicated by asterisks
Ancestral Species (Step 1)
The scheme posits a Most Recent Common Ancestor (MRCA) that contained a gene for a CCA-adding TNT. This ancestor gave rise, certainly through a series of intermediate genetic steps, to the CCA-adding TNT of T. maritima and the other Thermotogae. Because of the location of T. maritima on the rRNA phylogenetic tree (www.arb-silva.de/projects/living-tree), and as has been suggested by others (Neuenfeldt et al. 2008; Tomita and Weiner 2002), it is posited here that the ancestral TNT activity was CCA-addition rather than CC- or A-addition.
Support for this position comes from the studies of Cho et al. on the TNTs from A. aeolicus and G. stearothermophilus (Cho et al. 2007). They observed lower kcat/Km values in the CC- and A-adding reactions for the enzymes from A. aeolicus than were observed for the CCA-adding enzyme from G. stearothermophilus. They argue that these results support the evolution of CC- and A- addition from a CCA-adding ancestor and that the lowered kcat/Km values reflect the kinetic compromises required for the nucleotide binding site in the CC- and A-adding enzymes to exclude one nucleoside triphosphate as a substrate in the reaction (Cho et al. 2007).
Synergistetes, which is also situated near the base of the rRNA-derived bacterial phylogenetic tree (www.arb-silva.de/projects/living-tree), is also postulated to contain a CCA-adding TNT (see further below).
Duplication of the Gene for the CCA-Adding Enzyme (Step 2)
Since the CC- and A-adding enzymes in modern species share significant structural similarities (Toh et al. 2009; Tomita and Weiner 2002) it is likely that both TNTs arose from a common ancestral enzyme. Thus, an early step in the evolution of the two-enzyme system for CCA-addition must have been a gene duplication. Figure 4 posits the duplication of the gene for the ancestral CCA-adding enzyme. In the scheme of Fig. 4, the duplicated genes still encode the Nrn and CBS domains.
Conversion of CCA- to A-Addition (Step 3)
The Aquificae are the oldest extant bacterial phylum known to contain separate CC- and A-adding TNTs (Tomita and Weiner 2001). The question arises, in the evolution of the two-enzyme system, which evolved first, CC-addition or A-addition? There is no experimental evidence available to answer this question but intuitively, it could be argued that the more parsimonious evolutionary scheme would involve the initial appearance of the activity required to add a single A residue to an immature or damaged tRNA rather than the activity required to add the two Cs. Moreover, the capacity to add the Cs would be of little value to a cell without the concomitant ability to add the terminal A. It is noteworthy with regard to this argument, that A. aeolicus contains three tRNA genes that terminate with two C residues at the 3′-end of the gene product, viz. tRNAleu, tRNAsec, and tRNAhis. It is not known whether the two 3′-C residues represent immature forms of the tRNAs that lack only the terminal A, but it is possible that they are vestiges of an ancient system in A. aeolicus in which the organism had the capacity to add A residues but not C residues to immature or damaged tRNAs.
Additional support for the proposal that A-addition evolved before CC-addition comes from inspection of the distance matrix used to generate the phylogenetic tree shown in Fig. 3. Thus, the distance per 100 point accepted mutations (PAMs) between the T. maritima CCA-adding TNT and the A. aeolicus A-adding enzyme is 1.73, while the distance between the T. maritima TNT and the A. aeolicus CC-adding enzyme is 2.97. Similarly, the distances between the T. maritima TNT and the D. radiodurans A- and CC-adding enzymes were 2.23 and 3.00, respectively. Assuming that CCA-addition was the ancestral TNT activity, a larger number of mutations were required to convert the ancestral CCA-adding enzyme to CC-addition than were required to convert that TNT to A-addition. In what follows, therefore, it is postulated that A-addition evolved before CC-addition in bacteria.
The structure of the A. aeolicus A-adding enzyme has been solved by the Tomita research group and compared with the T. maritima CCA-adding TNT (Toh et al. 2009; Tomita et al. 2004; Yamashita et al. 2015). The structures are remarkably similar but Tomita and coworkers argue that a subtle difference between the neck regions of the two proteins is responsible for the switch from CCA- to A-addition. In particular, they identify two pairs of amino acids in a pair of α-helices in the two structures, viz. Glu185Gln186 (using their numbering system) and Arg236Lys232 (Toh et al. 2009). They posit that critical hydrogen bonds form between Glu185 and Arg236 and between Gln186 and Lys232 in the T. maritima enzyme (Fig. 5a) but that due to amino acid changes in the corresponding regions of the A. aeolicus A-adding enzyme, such H-bonds cannot form.
Model showing the amino acids that form hydrogen bonds to create the “springy hinge” in the neck region of the T. maritima (left) and G. stearothermophilus (right) tRNA nucleotidyltransferases. The T. maritima model is based on the structural data from Toh et al. (2009), while the G. stearothermophilus model was derived from the study of Li et al. (2002). The indicated amino acids are components of the α-helices that are shown in the background in light green. Note that the helices were referred to by the letters (in parentheses) in Li et al. (2002)
In the T. maritima TNT, the H-bonds lead to the formation of what Tomita and coworkers refer to as a “springy hinge.” This hinge, in the neck domain of the protein, provides the flexibility required for the catalytic center in the head to reach the residue on the bound tRNA that accepts the two Cs and to add the terminal A to that tRNA as well, to form tRNA-CCA. Because the A. aeolicus A-adding enzyme lacks the “springy hinge,” the head is less flexible and cannot extend to the residue that precedes the first added C and can thus add only the terminal A residue once the Cs are in place. In support of their model, they demonstrate that attaching the head–neck region of the T. maritima enzyme to a fragment of the A. aeolicus A-adding enzyme, lacking the head and neck, confers CCA-adding activity on the chimeric protein (Toh et al. 2009).
One difficulty with the foregoing model is that the Glu185Gln186, Arg236Lys232 pattern is not conserved in the class II CCA-adding enzymes. The researchers argue that other amino acid combinations may still allow the formation of the “springy hinge” in CCA-adding enzymes (Toh et al. 2009). In this regard, it is noteworthy that modeling the relevant region of the G. stearothermophilus CCA-adding enzyme revealed several residues that might form H-bonds to facilitate formation of the “springy hinge” in that enzyme (Fig. 5b). Similar considerations may apply to other CCA-adding enzymes. For example, using an amber suppression assay, Tomita and coworkers identified a number of putative CCA-adding TNT variants in a pool of randomized mutants from the T. maritima neck region (Toh et al. 2009).
It should be noted that Tretbar et al. proposed a different explanation for the specificity of CCA- and A-adding enzymes (Tretbar et al. 2011), and presented data indicating that a deletion of the C-terminal region of the D. radiodurans and B. halodurans A-adding enzymes converted them to CCA-adding enzymes. Tretbar et al. argue that the deleted C-terminal sequences inhibit CC-addition in the A-adding enzymes, and they propose that changes in this region led to the evolution of CCA-adding enzymes from A-adding TNTs. However, Tomita and coworkers demonstrated that deletion of the C-terminal end of the A. aeolicus A-adding enzyme did not confer CCA-adding ability on the resulting mutant enzyme and they argue that the C-terminal end does not dictate the specificity of the A-adding enzyme (Yamashita et al. 2015), at least not absolutely.
As the hypothesis presented here posits that CC- and A-adding enzymes evolved from a CCA-adding ancestor, the data and interpretations advanced by the Tomita group inform the scheme shown in Fig. 4, which proposes that one of the duplicate CCA-adding enzymes in an ancestral species was converted to A-addition by a series of mutations in the duplicate gene which resulted in the loss of the springy hinge (Step 3).
It should be noted here that Fig. 4 posits that the NTSF from Synergistetes 53_16 is a CCA-adding enzyme, rather than an A-adding enzyme. Synergistetes is found near the base of the rRNA-derived phylogenetic tree (www.arb-silva.de/projects/living-tree), in a clade just above the one containing the Thermotogae. The NTSF from Synergistetes 53_16 is approximately of the same size as the CCA-adding enzyme from T. maritima and the A-adding enzyme from A. aeolicus and contains the Nrn and CBS domains. Thus, based on structure and phylogeny, either activity is hypothetically possible for the Synergistetes NTSF. Figure S3 shows though that the “springy hinge” motif, GluGln/ArgLys, is precisely conserved in the Synergistetes enzyme. Thus, the “springy hinge” can presumably form in the Synergistetes NTSF and that would confer CCA-adding activity on that enzyme. The Synergistetes NTSF is thus shown as a CCA-adding TNT derived from the MRCA in Fig. 4.
Generation of the “Core” CCA-Adding TNT (Step 4)
The A-adding enzymes in the Aquificae, the Cyanobacteria, and the δ-Proteobacteria retain the Nrn and CBS domains found in the T. maritima CCA-adding enzyme. However, the CC-adding enzymes (and some A-adding enzymes, see below) have lost those domains and are closer in size to modern bacterial CCA-adding enzymes (Fig. 2). Step 4 of Fig. 4 involves the deletion of the two accessory domains from one of the duplicate genes in an ancestral species, the gene encoding CCA-adding activity. Deletion of those domains in an intermediate ancestor would generate a gene encoding a CCA-adding TNT of the approximate size of modern enzymes, the “core” CCA-adding TNT. As indicated above, the two N-terminal domains are not required for the enzymatic activity of the TNTs that contain them.
Generation of the CC-Adding TNTs (Step 5)
Tomita and coworkers have published the crystal structure of the A. aeolicus CC-adding enzyme (Yamashita et al. 2014). As shown in Fig. 6a, that enzyme adopts a seahorse-like structure similar to the other bacterial TNTs whose structures have been determined.
a Structures of the T. maritima CCA-adding enzyme and the A. aeolicus CC-adding enzyme. The proteins are displayed in a different orientation from that shown in Fig. 1 so that the structural differences between them can be observed. The flexible loop in the T. maritima enzyme is shown in pink. Note that the tail region of the A. aeolicus CC-adding enzyme is not shown in the figure. Data for the tail region were not present in PDB file 3WFO. Additional details are provided in the text. b Sequences of flexible loops implicated in determining the CCA- or CC-adding specificity of bacterial tRNA nucleotidyltransferases (Neuenfeldt et al. 2008). Loops of species from several of the phyla represented in Fig. 2 are shown. Amino acid identities are highlighted in black and similarities in gray. AB Actinobacteria, FC Firmicutes. The basic/acidic amino acid motif (e.g., R–X–[D/E] and the RRD sequence at the C-terminus of the loop are highly conserved (Hoffmeier et al. 2010). Note that the loop sequences of the CC- and A-adding enzymes from S. cerevisiae and S. pombe are included for comparison. Numbers at the left and right of the sequences indicate the N- and C-terminal ends of the loops in the amino acid sequences of the proteins. Note also that the T. maritima sequence is numbered as in (Toh et al. 2009). All the sequences shown derive from CCA-adding enzymes except those from S. pombe. Biochemical function has been verified for the enzymes from T. maritima, E. coli, S. coelicolor, B. subtilis, S. cerevisiae and S. pombe (see Table S1)
However, there are significant differences between the structures of the T. maritima CCA-adding enzyme, the A. aeolicus A-adding enzyme, and the A. aeolicus CC-adding enzyme. In particular, as shown in Fig. 6a, the A. aeolicus CC-adding enzyme possesses two α-helices that are not found in the other two enzymes, viz. Ex-α2 and Ex-α3. Moreover, two of the remaining α-helices, α16 and α17 are longer in the CC-adding enzyme than in the CCA- and A- adding enzymes. These changes result in what Tomita and coworkers (Yamashita et al. 2014) refer to as a “closed structure” with a bulging body domain (compare the two structures shown in Fig. 6a). The upshot of these changes is that the mechanism of CC-addition differs significantly from that of CCA- and A-addition. In particular, the Tomita group (Yamashita et al. 2014) argues that CC-addition involves both translocation and rotation of the tRNA acceptor as the two Cs are added such that after addition of the second C residue, the tRNA dissociates from the enzyme and no further modification of the tRNA is possible by that enzyme. This explains the inability of the CC-adding enzyme to add the terminal A. The dissociated tRNA-CC is then available to serve as a substrate for the A-adding TNT (Yamashita et al. 2014).
It should be noted that Neuenfeldt et al. have proposed an alternative explanation for the generation of CC-adding activity (Neuenfeldt et al. 2008). These workers identified a region of the CCA-adding TNTs that they argue is responsible for the difference in specificity between CCA- and CC-adding enzymes. This region contains a flexible loop of ca. 30 amino acids situated in the head domain of the enzyme and includes conserved basic and acidic residues (Fig. 6b). When this flexible loop was deleted from the E. coli CCA-adding TNT, the resulting mutant retained the ability to add C residues to a model substrate but was unable to catalyze A-addition. Conversely, when the flexible loop from the Bacillus subtilis CCA-adding TNT was inserted into the appropriate position in the CC-adding TNT from Bacillus halodurans, the resulting mutant protein displayed CCA-adding activity (Neuenfeldt et al. 2008).
The flexible loop appears to be present in all bacterial CCA-adding TNTs but absent from all CC-adding TNTs. As seen in Fig. 6b, however, there is very little sequence conservation in the flexible loop between bacterial phyla, except for the N-terminal R-X-[D/E] and the RRD at the C-terminus of the loop. Hoffmeier et al. argue that some sequence conservation exists within phyla although not between phyla but that the sequences in any case provide the determinants required to specify CCA-addition when those sequences are present (Hoffmeier et al. 2010). These researchers argue further that since there is evidence that deletion mutations occur more frequently in nature than insertions (de Jong and Ryden 1981; Petrov 2002), the conversion of an ancestral CCA-adding TNT to a CC-adding TNT resulted from a deletion of the loop in the former protein.
Yamashita et al. (Yamashita et al. 2014) again reported an exception to the general formula advanced by the Mörl group (Hoffmeier et al. 2010; Neuenfeldt et al. 2008). They reported that insertion of the flexible loop from the A. aeolicus A-adding enzyme or the T. maritima CCA-adding enzyme (shown in pink in Figs. 1a and 6a) failed to convert the A. aeolicus CC-adding enzyme to CCA-addition. Thus, the role of the flexible loop in determining CC- and A-adding specificity may not be absolute.
Is it possible to reconcile the mechanisms for the conversion of a CCA-adding to a CC-adding TNT proposed by the Tomita (Yamashita et al. 2014) and Mörl (Hoffmeier et al. 2010; Neuenfeldt et al. 2008) groups? It has been reported recently that the fission yeast, Schizosaccharomyces pombe, utilizes separate CC- and A-adding TNTs to synthesize the CCA-ends of its tRNAs (Reid et al. 2019). This is the first report of the presence of separate CC- and A-adding enzymes in a eukaryote. With regard to the competing mechanisms proposed for CC-addition in bacterial species, it is noteworthy that both the S. pombe CC- and A-adding enzymes contain a loop, quite similar in size and sequence of the conserved regions to those of bacteria (Fig. 6b). The C-terminal sequence of the CC-adding enzyme is RHH, rather than the R–X–[D/E] motif found in bacterial CCA-adding TNTs (Fig. 6b). Reid et al. converted the RHH sequence to RHE but found that this change did not convert the S. pombe CC-adding enzyme to CCA-addition (Reid et al. 2019). They argue that in some circumstances, structural changes other than the presence or absence of the loop are necessary to determine the specificity of CC- and A-adding TNTs. Their modeling of the S. pombe CC- and A-adding TNTs using the T. maritima CCA-adding enzyme as a template revealed several structural differences between the two yeast enzymes. In particular, they argue that the CC-adding enzyme contains a β-sheet that is absent from both the S. pombe A-adding TNT and the S. cerevisiae CCA-adding enzyme. They posit that this β-sheet reduces the flexibility of the loop and that the structural differences revealed by their modeling are responsible for both the loss of A-adding activity by the CC-adding enzyme and the absence of CC-adding activity from the A-adding TNT. Reid et al. also argue that both enzymes evolved from an ancestral CCA-adding TNT, like that found in S. cerevisiae.
Extending their analysis to the bacterial systems, it is possible that, depending on the system, the structural differences identified by the Tomita group (Yamashita et al. 2014) or/and the flexible loop (Hoffmeier et al. 2010; Neuenfeldt et al. 2008) determine the specificity of CCA- and CC-adding TNTs. Similarly, it seems possible that the “springy hinge” (Toh et al. 2009; Yamashita et al. 2015) or/and the C-terminal end of bacterial TNTs (Tretbar et al. 2011) can, depending on the system, determine the specificity of A-adding TNTs. A combination of mutations leading to particular structural changes may then have been responsible for the appearance of the A- and CC-adding enzymes. Figure 4 posits that CC-addition in an ancestral species arose through the effects of those mutations on the gene encoding the “core” CCA-adding TNT. The resulting gene would produce a CC-adding enzyme of the approximate size found in modern bacterial species (Table S1).
Formation of the Two-Enzyme System in the Thermodesulfobacteria (Step 6)
Like the Thermotogae, the Thermodesulfobacteria appear near the base of the rRNA-based bacterial phylogenetic tree (www.arb-silva.de/projects/living-tree). This phylum is represented by C. thiodismutans in Table S1 and Fig. 3. While the presence of the two-enzyme system has not been verified biochemically in C. thiodismutans, the phylogenetic analysis suggests its presence in that species. Moreover, a BLAST search of other Thermodesulfobacteria also suggested the presence in other species of separate A- and CC-adding enzymes.
The putative CC-adding enzyme from C. thiodismutans has lost the Nrn and CBS domains and is approximately of the same size as the corresponding enzyme from the Aquificae. However, the A-adding enzyme from C. thiodismutans has a length of only 416 aa, approximately the length that would be generated by the deletion of the Nrn and CBS domains (ca. 392 aa) from a gene encoding an A-adding enzyme like that from A. aeolicus. These changes are represented by Step 6 of Fig. 4.
Formation of the Two-Enzyme System in the Aquificae and the Cyanobacteria (Steps 7, 8)
Figure 4 posits the formation of an ancestral species containing A- and CC-adding TNTs via the mechanisms described in the preceding sections. The evolution of this ancestor to produce the A-adding TNT found in A. aeolicus and in the Cyanobacteria is indicated as Steps 7 and 8 in Fig. 3. The ancestral A-adding enzyme is posited to be approximately the same size as the A. aeolicus and the cyanobacterial A-adding enzymes and to contain the Nrn and CBS domains found in those enzymes. The δ-proteobacterial A-adding enzymes will be considered separately below. The CC-adding enzymes in the Aquificae are postulated to derive from the “core” CCA-adding enzyme in the relevant ancestor and thus have lost the Nrn and CBS domains, as described above. It is noteworthy that the cyanobacterial CC-adding enzymes appear in two separate clades in Fig. 3. This observation will be discussed further below.
Formation of the Two-Enzyme System in the Deinococci (Step 9)
The CC- and A-adding enzymes from Deinococcus are considerably smaller than the corresponding enzymes from the Aquificae and the Cyanobacteria and have lost not only two the N-terminal domains but some additional sequences as well (Table S1 and Step 9 of Fig. 3). Nevertheless, modeling of the D. radiodurans CC- and A-adding enzymes (and the corresponding enzymes from C. thiodismutans) shows that they can adopt the same structure as the TNTs whose structures have been determined experimentally (Figure S4) and that the head, neck, body, and tail regions are present in the TNTs from the Deinococci and the Thermodesulfobacteria.
Generation of an Ancestral Species Containing the “Core” CCA-Adding Enzyme (Step 10)
Modern CCA-adding enzymes lack the N-terminal extension which is present in the T. maritima CCA-adding TNT and in some of the A-adding TNTs. Figure 4 suggests two possible pathways for the generation of an ancestor containing only the “core” CCA-adding enzyme. Step 10A posits the loss of the gene for the A-adding enzyme from the ancestor that contains that gene and the gene for the “core” CCA-adding enzyme. Alternatively, deletion of the N-terminal extension from the gene encoding the MRCA CCA-adding enzyme (or one of its descendants) would generate the species containing a CCA-adding enzyme similar in size to that found in modern bacterial species (Step 10B). Note that the node indicated by the arrow in Fig. 3 may be the ancestral “core” CCA-adding enzyme, as all the clades radiating from that node contain species with NTSFs that have lost the Nrn and CBS domains.
Generation of Modern CCA-Adding TNTs (Steps 11, 12)
The “core” CCA-adding enzyme is posited to have evolved to produce the CCA-adding TNTs found in the Firmicutes and Actinobacteria (Step 11). As these phyla appear to be ancestral to the Proteobacteria in phylogenetic analyses based on rRNAs (www.arb-silva.de/projects/living-tree) or on conserved signature indels (Gupta 2016), it is postulated that subsequent evolution of the “core” CCA-adding enzyme led to the formation of the ancestor of the proteobacterial TNTs (Step 12), and that this ancestor evolved to yield the modern α- and ε-proteobacterial CCA-adding enzymes and the δ-Proteobacteria that contain a single CCA-adding TNT (Step 12).
Evolution of the Poly(A) polymerases (Step 13)
Step 13 invokes a gene duplication in a species ancestral to modern β, γ-Proteobacteria that contain both a TNT and a PAP. Subsequent mutations in one of those genes would produce, pcnB, the gene for PAP I (Cao and Sarkar 1992; Liu and Parkinson 1989), and subsequent evolution produced the β, γ-Proteobacteria which contain a CCA-adding TNT and PAP I.
The structure of E. coli PAP I has been determined (Toh et al. 2011). Toh et al. invoke a rationale for the restriction of PAP I activity to A-addition similar to that used to explain the formation of A-adding enzymes from CCA-adding ones. Specifically, Toh et al. argue that the head–neck region of PAP I (Fig. 1b) is less flexible than the corresponding region in the CCA-adding enzymes and is constrained in a fashion similar to that posited for the A-adding enzymes, to accommodate only ATP as a substrate (Toh et al. 2011). Thus, the bacterial poly(A) polymerases can only add A residues to RNA 3′-ends.
Supporting this hypothesis, Cho et al. demonstrated that point mutations in three residues situated in the neck region of G. stearothermophilus TNT modified the activity of the enzyme. When R194, M197, and E198 were all replaced by alanines, the resulting mutant protein acquired the ability to add poly(A) tails to a tRNA substrate (Cho et al. 2007).
The Role of Horizontal Gene Transfer in the Evolution of Bacterial NTSFs
It has become increasingly apparent in recent years that the transfer of genetic information between distantly related strains and species (horizontal gene transfer, HGT) plays an important role in cellular evolution (Brown 2003; Koonin 2016; Soucy et al. 2015). The analysis presented here strongly suggests the influence of HGT on the evolution of bacterial NTSFs.
Current literature suggests that the “gold standard” for the identification of HGT is phylogenetic incongruence (Brown 2003; Koonin 2016; Ravenhall et al. 2015; Soucy et al. 2015). Inspection of Fig. 3 reveals several TNTs that are found in clades other than those in which their sister species cluster. For example, the TNT from Helicobacter pylori, an ε-proteobacterium, is found in a clade containing the “deep branching” CC-adding enzymes while the A-adding enzyme from B. halodurans, a Firmicute, appears in the clade containing the “deep branching” A-adding TNTs.
In a similar vein, the TNT from the α-proteobacterium, Magnetofaba australis, appears in the clade containing the β, γ-TNTs. All of these are instances of possible horizontal gene transfer, from a species related to the clade in which the recipient is found to an ancestor of that recipient, and it is noteworthy in this regard that the horizontal transfer of the gene for a CCA-adding TNT from an α-proteobacterium to the eukaryotic Holozoa has been reported (Betat et al. 2015).
To examine further the possibility of HGT of bacterial TNTs, alien indices were calculated for those enzymes genes for which may have been acquired by that mechanism. The alien index was originally defined by Gladyshev et al. (2008), and the principle has been refined by Rancurel et al. (Rancurel et al. 2017). The latter authors define the alien index as
The E values are determined from BLAST searches using appropriate query sequences and donor and recipient sequences. In the case of the bacterial TNTs, the query sequence used was that of the TNT whose gene was suspected of acquisition via HGT. The donor species would be the “alien” species, the potential source of the horizontally transferred gene. The recipient species would be one related to the query species, into which the “alien” gene was transferred during evolution. Results of this analysis are shown in Table 1.
The first example shown in the table is M. australis, classified as an α-proteobacterium (Table S1). However, its TNT appears in the clade containing the β, γ-proteobacterial TNTs in Fig. 3. The best BLAST hit to the M. australis TNT sequence was, indeed, to a γ-proteobacterium (Table 1). When the BLAST search was performed with the exclusion of M. australis and the β, γ-Proteobacteria, the best hit was to an α-proteobacterium from the order Ricksettiae (Table 1). The E values obtained for the potential donor (Legionella worsleiensis) and recipient species (Rickettsiae sp.) yielded an alien index of 40.7. Similar approaches yielded alien indices for the other TNTs shown in Table 1, genes for which may have been acquired by HGT.
Gladyshev et al. proposed an AI value of ≥ 45 as a strong indicator of HGT. Using a different and larger dataset, Rancurel et al. proposed three categories for classifying BLAST results used in the calculation of the AI: (i) very likely HGT (AI > 30 and < 70% identity to candidate donor); (ii) possible HGT (AI > 0 and < 70% identity to candidate donor); and, (iii) likely contamination (AI > 0 and ≥ 70% identity to candidate donor). The stipulation that the query sequence should be < 70% identical to the candidate donor sequence only applies to HGT from prokaryotes to eukaryotes (Ku and Martin 2016; Rancurel et al. 2017).
It is apparent from Table 1 that in all the instances of potential acquisition of bacterial TNTs by HGT suggested by the phylogenetic analysis of Fig. 3, the AI values are > 0. Indeed, with the exception of the B. halodurans CC-adding enzyme, all of the alien indices calculated are > 30. Based on these criteria and on the phylogenetic analysis shown in Fig. 3, there is a high probability that the genes for all of the TNTs shown in Table 1 were acquired by HGT, including the CC- and A-adding enzymes found in modern δ-Proteobacteria. The A-adding and CC-adding enzymes from the δ-Proteobacteria appear near the base of the NTSF tree (Fig. 3) while phylogenies based on rRNA (www.arb-silva.de/projects/living-tree) and on conservative signature indels (Gupta 2016) place the δ-Proteobacteria near the top of the bacterial evolutionary tree. HGT of the relevant genes to the δ-Proteobacteria is shown as Step 14 of Fig. 4.
The B. halodurans CC-adding enzyme appears in the clade containing the Firmicute TNTs in Fig. 3, but is less closely related to the other TNTs in the clade than those enzymes are to each other. The alien index for this enzyme was the lowest of all those calculated for the possible instances of HGT (Table 1). Thus, although HGT remains a possibility for the B. halodurans CC-adding enzyme, it is also possible that this enzyme arose by conversion of a CCA-adding TNT to CC-addition through vertical rather than horizontal transmission from an ancestral species.
The δ-Proteobacterial PAPs appear in the clade containing the corresponding enzymes from the β, γ-Proteobacteria. Table 1 supports the HGT of the pcnB gene from a γ-proteobacterium to an ancestral δ species. This transfer is shown as Step 15 of Fig. 4.
The Cyanobacteria present an intriguing problem. The cyanobacterial A-adding enzymes appear near the base of the tree in Fig. 3, consistent with vertical transmission of the relevant genes and with the position of this phylum in the rRNA-derived phylogenetic tree (www.arb-silva.de/projects/living-tree). However, the cyanobacterial CC-adding enzymes appear more closely related to the α-, ε-proteobacterial TNTs than to the CC-adding enzymes from the “deep branching” species (Fig. 3). It is possible that the Cyanobacteria acquired an ancestral “core” CCA- gene by HGT, different from the one which gave rise to the CC-adding enzymes in the Aquificae, the Deinococci, and the Thermodesulfobacteria, and that a series of mutations converted the gene for that protein to CC-addition. It is noteworthy in this regard that the cyanobacterial CC-adding enzymes are approximately the same size as the CCA-adding enzymes from the α-, ε-, and β, γ-Proteobacteria. Since the CC-adding enzymes of Cyanobacteria do not appear in an incongruent clade for another phylum but in a clade with the enzymes from other cyanobacterial species, it was not possible to calculate an alien index for the Cyanobacteria as there was no straightforward strategy for identifying a viable recipient species.
It does appear that, based on the phylogenetic analysis of Fig. 3, the CC-adding enzymes may have evolved twice in the bacteria. The CC-adding enzymes in the “deep branching” species, unlike those from the Cyanobacteria, appear in clades near the base of the phylogenetic tree of Fig. 3, consistent with the position of the corresponding species in the rRNA-based tree. It has been suggested by others that the CC-adding enzymes arose twice during evolution (Neuenfeldt et al. 2008).
Biochemical Activities of NTSFs that Have Not Been Characterized
Only a few of the enzymatic activities of the enzymes listed in Table S1 have been demonstrated in the laboratory. These are highlighted in color in the table. Is it possible to identify the biochemical function of other NTSFs in the absence of those biochemical analyses? In some cases, putative identification may be possible. NTSFs from particular phyla that are found in the same clade as enzymes of known function from species in those phyla are likely to have the same function. Thus, all of the other cyanobacterial enzymes are found in the same clades as the CC- and A-adding enzymes from Synechocystis (Fig. 3 and Table S1). In a similar vein, all of the Firmicute enzymes (except for B. halodurans, see above) are located in the clade containing the B. subtilis CCA-adding TNT and are likely to be CCA-adding enzymes themselves. This line of reasoning can be applied to many of the other TNTs whose lineages are shown in Fig. 3.
With regard to the poly(A) polymerases, Martin and Keller examined the sequences of a number of bacterial NTSFs and identified signature sequences that distinguish PAPs from TNTs (Martin and Keller 2004). Examples of the PAP-specific signature sequence are shown in Fig. 7, and this sequence is likely to be diagnostic for PAPs. Thus, enzymes that lack this sequence, as do at least some of the NTSFs from Firmicutes, the Actinobacteria, and the α- and ε-Proteobacteria, are likely to be TNTs. Figure 7, for example, shows that the E. coli TNT lacks the PAP signature motif.
The signature sequence motif identified by Martin and Keller (2004) as diagnostic of poly(A) polymerases. Numbers at the left and right of the sequences indicate the N- and C-terminal ends of the motifs in the amino acid sequences of the proteins. The signature consensus is shown at the bottom of the figure. Species names are abbreviated as indicated in Table S1
There are three other species represented in Fig. 7, signature sequences of which differ significantly from the PAP consensus, viz. those for Syntrophobacter fumaroxidans (Sfu), Thermodesulforhabdis norvegica (Tno) and C. thiodismutans (Cth). The putative signature sequences for these species are indicated in red in Fig. 7. The first two of these three species are δ-Proteobacteria containing NTSFs annotated as PAPs in the genome database (Table S1). Figure 3, however, suggests that both of those enzymes are CC-adding TNTs. The C. thiodismutans NTSF is found in the clade containing the β, γ-, and δ-proteobacterial PAPs (Fig. 3). However, that protein, also annotated as a PAP, is significantly smaller than the β, γ-, and δ-proteobacterial PAPs (297 amino acids vs. 450–500 amino acids, Table S1) and given the dissonance in its signature sequence compared with the proteobacterial enzymes, it may not actually possess PAP activity.
While the outcomes of the foregoing analysis are interesting and suggestive, it must be noted that the only definitive method for identifying the activities of these enzymes is biochemical assay.
Other Enzymes Involved in RNA 3′-Tail Synthesis in Bacteria
As indicated above, PAP I is encoded by the pcnB gene in E. coli and related bacteria. Mohanty and Kushner showed some years ago that pcnB null mutants still add tails to the 3′-ends of RNAs (Mohanty and Kushner 2000). However, these tails do not contain A residues exclusively. Rather, they are heteropolymeric, containing G, C, and U as well as A residues. Mohanty and Kushner demonstrated that the enzyme responsible for 3′-tail synthesis in pcnB mutants is polynucleotide phosphorylase (PNPase) (Mohanty and Kushner 2000). PNPase is a 3′–5′-exoribonuclease that catalyzes the following reaction:
As written, the above reaction depicts the phosphorolytic degradation of RNA chains. The reaction is reversible, however, and PNPase will synthesize polyribonucleotide chains using nucleotide diphosphates as substrates. It is this polymerization activity that is responsible for the addition of 3′-tails to E. coli RNAs in the absence of PAP I (Mohanty and Kushner 2000).
PNPases are widely distributed and are found in all bacterial genera examined to date except the Mycoplasmas (Zuo and Deutscher 2001). There is strong evidence for the function of PNPase as the major, if not the sole, RNA 3′-polyribonucleotide polymerase in plant chloroplasts (Yehudai-Resheff et al. 2001), in the Cyanobacteria (Rott et al. 2003) and in Streptomyces (Bralley et al. 2006; Sohlberg et al. 2003). The question is whether it plays the same role in the many bacterial species that do not contain PAP I. It has also been suggested that PNPase and PAP I participate in the synthesis of the CCA-ends of immature and damaged tRNAs in E. coli, at least under certain conditions. Reuven et al. postulated that in situations in which the TNT is absent or present in significantly reduced amounts in E. coli cells, PAP I adds A residues to tRNA-CC and PNPase trims the resulting products as necessary to produce tRNA-CCA (Reuven et al. 1997). These authors also suggest the possibility that PNPase may be able, via its polymerizing activity, to add the two Cs and the terminal A to immature and damaged tRNAs (Reuven et al. 1997). It is noteworthy that Yue et al. have suggested that PNPases may represent a third class of the nucleotidyltransferase superfamily (Yue et al. 1996).
There is some evidence for a third system for synthesizing RNA 3′-tails in addition to PAP I and PNPase. In a study of poly(A) tail synthesis in B. subtilis, Campos-Guillen et al. examined poly(A) tail lengths in a mutant lacking PNPase compared with wild-type cells. The length of the poly(A) tails was essentially the same in both cases (Campos-Guillén et al. 2005). Thus, B. subtilis, which does not contain PAP I (Raynal et al. 1998), is capable of synthesizing RNA 3′-tails in the absence of PNPase. The enzyme responsible for this synthesis has yet to be identified but a potential candidate, if an unlikely one, is the enzyme, RNase PH. Like PNPase, RNase PH is a phosphorolytic 3′–5′-exoribonuclease that plays a role in tRNA maturation (Deutscher et al. 1988). Studies by Bralley et al. demonstrated that, again like PNPase, RNase PH can add A residues to the 3′-ends of an RNA acceptor, at least in vitro (Bralley et al. 2006). Whether this activity has any in vivo relevance remains to be seen.
Given the absence of PAPs from most bacterial species (Fig. 3, Table S1), it is possible that the ancestral enzyme species responsible for the synthesis of RNA 3′-tails was PNPase. There is evidence to suggest that PNPases are ancient enzymes (Leszczyniecka et al. 2004; Sokhi et al. 2014) and as indicated above, there is ample modern precedent for their function as RNA 3′-polyribonucleotide polymerases.
Conclusion
This report has presented a plausible scheme for the evolution of bacterial tRNA nucleotidyltransferases and poly(A) polymerases. Many details of the evolutionary process remain obscure but questions related to some of those unresolved details can be addressed. What are the biochemical functions of the NTSFs whose activities have not yet been characterized? Are PAPs more widely distributed in bacteria than is currently recognized? Are there extant bacterial species containing intermediate forms of the CCA-, CC- or A-adding enzymes? Can modern bacteria with a two-enzyme system survive if the gene for one of those enzymes is inactivated? Are there more than two systems for poly(A) tail synthesis? All of these and other interesting questions related to the evolution of bacterial NTSFs can be addressed experimentally. Of course, perhaps the most intriguing question that derives from this analysis is that of the evolutionary advantage, if there is one, of having two TNTs rather than one, or one, rather than two.
References
Anashkin VA, Baykov AA, Lahti R (2017) Enzymes regulated via cystathionine beta-synthase domains. Biochemistry (Mosc) 82:1079
Betat H, Mede T, Tretbar S, Steiner L, Stadler PF, Morl M, Prohaska SJ (2015) The ancestor of modern Holozoa acquired the CCA-adding enzyme from Alphaproteobacteria by horizontal gene transfer. Nucleic Acids Res 43:6739
Bralley P, Chang SA, Jones GH (2005) A phylogeny of bacterial RNA nucleotidyltransferases: bacillus halodurans contains two tRNA nucleotidyltransferases. J Bacteriol 187:5927
Bralley P, Gust B, Chang S, Chater KF, Jones GH (2006) RNA 3′-tail synthesis in Streptomyces: in vitro and in vivo activities of RNase PH, the SCO3896 gene product and polynucleotide phosphorylase. Microbiology 152:627
Bralley P, Cozad M, Jones GH (2009) Geobacter sulfurreducens contains separate C- and A-adding tRNA nucleotidyltransferases and a poly(A) polymerase. J Bacteriol 191:109
Brown JR (2003) Ancient horizontal gene transfer. Nat Rev Genet 4:121
Campos-Guillén J, Bralley P, Jones GH, Bechhofer DH, Almeda-Alvarez G (2005) Addition of poly(A) and heteropolymeric 3′-ends in Bacillus subtilis wild-type and PNPase-deficient strains. J Bacteriol 187:4698
Cao GJ, Sarkar N (1992) Identification of the gene for an Escherichia coli poly(A) polymerase. Proc Natl Acad Sci USA 89:10380
Cao GJ, Kalapos MP, Sarkar N (1997) Polyadenylated mRNA in Escherichia coli: modulation of poly(A) RNA levels by polynucleotide phosphorylase and ribonuclease II. Biochimie 79:211
Cho HD, Weiner AM (2004) A single catalytically active subunit in the multimeric Sulfolobus shibatae CCA-adding enzyme can carry out all three steps of CCA addition. J Biol Chem 279:40130
Cho HD, Verlinde CL, Weiner AM (2007) Reengineering CCA-adding enzymes to function as (U, G)- or dCdCdA-adding enzymes or poly(C, A) and poly(U, G) polymerases. Proc Natl Acad Sci U S A 104:54
de Jong WW, Ryden L (1981) Causes of more frequent deletions than insertions in mutations and protein evolution. Nature 290:157
Deutscher MP (1990) Ribonucleases, tRNA nucleotidyltransferase, and the 3′ processing of tRNA. Prog Nucleic Acid Res Mol Biol 39:209
Deutscher M, Marshall GT, Cudny H (1988) RNase PH: an Escherichia coli phosphate-dependent nuclease distinct from polynucleotide phosphorylase. Proc Natl Acad Sci USA 85:4710
Donovan WP, Kushner SR (1986) Polynucleotide phosphorylase and ribonuclease II are required for cell viability and mRNA turnover in Escherichia coli K-12. Proc Natl Acad Sci USA 83:120
Felsenstein J (1989) PHYLIP - phylogeny inference package (version 3.2). Cladistics 5:164
Gladyshev EA, Meselson M, Arkhipova IR (2008) Massive horizontal gene transfer in bdelloid rotifers. Science 320:1210
Guerrier-Takada C, McClain WH, Altman S (1984) Cleavage of tRNA precursors by the RNA subunit of E. coli ribonuclease P (M1 RNA) is influenced by 3′-proximal CCA in the substrates. Cell 38:219
Gupta RS (2016) Impact of genomics on the understanding of microbial evolution and classification: the importance of Darwin’s views on classification. FEMS Microbiol Rev 40:520
Hartmann RK, Gossringer M, Spath B, Fischer S, Marchfelder A (2009) The making of tRNAs and more - RNase P and tRNase Z. Prog Mol Biol Transl Sci 85:319
Hoffmeier A, Betat H, Bluschke A, Gunther R, Junghanns S, Hofmann HJ, Mörl M (2010) Unusual evolution of a catalytic core element in CCA-adding enzymes. Nucleic Acids Res 38:4436
Holm L, Sander C (1995) DNA polymerase beta belongs to an ancient nucleotidyltransferase superfamily. Trends Biochem Sci 20:345
Koonin EV (2016) Horizontal gene transfer: essentiality and evolvability in prokaryotes, and roles in evolutionary transitions. F1000Res 5:1805
Ku C, Martin WF (2016) A natural barrier to lateral gene transfer from prokaryotes to eukaryotes revealed from genomes: the 70% rule. BMC Biol 14:89
Leszczyniecka M, DeSalle R, Kang DC, Fisher PB (2004) The origin of polynucleotide phosphorylase domains. Mol Phylogenet Evol 31:123
Li F, Xiong Y, Wang J, Cho HD, Tomita K, Weiner AM, Steitz TA (2002) Crystal structures of the Bacillus stearothermophilus CCA-adding enzyme and its complexes with ATP or CTP. Cell 111:815
Liao H, Liu M, Guo X (2018) The special existences: nanoRNA and nanoRNase. Microbiol Res 207:134
Liu J, Parkinson JS (1989) Genetics and sequence analysis of the pcnB locus, an Escherichia coli gene involved in plasmid copy number control. J Bacteriol 171:1254
Martin G, Keller W (2004) Sequence motifs that distinguish ATP(CTP):tRNA nucleotidyl transferases from eubacterial poly(A) polymerases. RNA 10:899
Mohanty BK, Kushner SR (2000) Polynucleotide phosphorylase functions both as a 3′–5′ exonuclease and a poly(A) polymerase in Escherichia coli. Proc Natl Acad Sci USA 97:11966
Mohanty BK, Kushner SR (2011) Bacterial/archaeal/organellar polyadenylation. Wiley Interdiscip Rev RNA 2:256
Moretti S, Armougom F, Wallace IM, Higgins DG, Jongeneel CV, Notredame C (2007) The M-Coffee web server: a meta-method for computing multiple sequence alignments by combining alternative alignment methods. Nucleic Acids Res 35:W645
Neuenfeldt A, Just A, Betat H, Mörl M (2008) Evolution of tRNA nucleotidyltransferases: a small deletion generated CC-adding enzymes. Proc Natl Acad Sci U S A 105:7953
Oh BK, Pace NR (1994) Interaction of the 3′-end of tRNA with ribonuclease P RNA. Nucleic Acids Res 22:4087
Petrov DA (2002) Mutational equilibrium model of genome size evolution. Theor Popul Biol 61:531
Rammelt C, Rossmanith W (2016) Repairing tRNA termini: news from the 3′ end. RNA Biol 13:1182
Rancurel C, Legrand L, Danchin EGJ (2017) Alienness: rapid detection of candidate horizontal gene transfers across the tree of life. Genes (Basel) 8:248
Ravenhall M, Skunca N, Lassalle F, Dessimoz C (2015) Inferring horizontal gene transfer. PLoS Comput Biol 11:e1004095
Raynal LC, Krisch HM, Carpousis AJ (1998) The Bacillus subtilis nucleotidyltransferase is a tRNA CCA-adding enzyme. J Bacteriol 180:6276
Reid NE, Ngou JS, Joyce PBM (2019) Schizosaccharomyces pombe contains separate CC- and A-adding tRNA nucleotidyltransferases. Biochem Biophys Res Commun 508:785
Reuven NB, Zhou Z, Deutscher MP (1997) Functional overlap of tRNA nucleotidyltransferase, poly(A) polymerase I, and polynucleotide phosphorylase. J Biol Chem 272:33255
Rott R, Zipor G, Portnoy V, Liveanu V, Schuster G (2003) RNA polyadenylation and degradation in cyanobacteria are similar to the chloroplast but different from E. coli. J Biol Chem 278:15771
Sohlberg B, Huang J, Cohen SN (2003) The Streptomyces coelicolor polynucleotide phosphorylase homologue, and not the putative poly(A) polymerase can polyadenylate RNA. J Bacteriol 185:7273
Sokhi UK, DeSalle R, Bacolod MD, Das SK, Dasgupta S, Sarkar D, Fisher PB (2014) Evolutionary dynamics of Polynucelotide phosphorylases. Mol Phylogenet Evol 73:77
Soucy SM, Huang J, Gogarten JP (2015) Horizontal gene transfer: building the web of life. Nat Rev Genet 16:472
Toh Y, Takeshita D, Numata T, Fukai S, Nureki O, Tomita K (2009) Mechanism for the definition of elongation and termination by the class II CCA-adding enzyme. EMBO J 28:3353
Toh Y, Takeshita D, Nagaike T, Numata T, Tomita K (2011) Mechanism for the alteration of the substrate specificities of template-independent RNA polymerases. Structure 19:232
Tomita K, Weiner AM (2001) Collaboration between CC- and A-adding enzymes to build and repair the 3′-terminal CCA of tRNA in Aquifex aeolicus. Science 294:1334
Tomita K, Weiner AM (2002) Closely related CC- and A-adding enzymes collaborate to construct and repair the 3′-terminal CCA of tRNA in Synechocystis sp. and Deinococcus radiodurans. J Biol Chem 277:48192
Tomita K, Yamashita S (2014) Molecular mechanisms of template-independent RNA polymerization by tRNA nucleotidyltransferases. Front Genet 5:36
Tomita K, Fukai S, Ishitani R, Ueda T, Takeuchi N, Vassylyev DG, Nureki O (2004) Structural basis for template-independent RNA polymerization. Nature 430:700
Tretbar S, Neuenfeldt A, Betat H, Mörl M (2011) An inhibitory C-terminal region dictates the specificity of A-adding enzymes. Proc Natl Acad Sci U S A 108:21040
Wallace IM, O’Sullivan O, Higgins DG, Notredame C (2006) M-Coffee: combining multiple sequence alignment methods with T-Coffee. Nucleic Acids Res 34:1692
Waterhouse A, Bertoni M, Bienert S, Studer G, Tauriello G, Gumienny R, Heer FT, de Beer TAP, Rempfer C, Bordoli L, Lepore R, Schwede T (2018) SWISS-MODEL: homology modelling of protein structures and complexes. Nucleic Acids Res 46:W296
Xiong Y, Steitz TA (2006) A story with a good ending: tRNA 3′-end maturation by CCA-adding enzymes. Curr Opin Struct Biol 16:12
Xiong Y, Li F, Wang J, Weiner AM, Steitz TA (2003) Crystal structures of an archaeal class I CCA-adding enzyme and its nucleotide complexes. Mol Cell 12:1165
Yamashita S, Takeshita D, Tomita K (2014) Translocation and rotation of tRNA during template-independent RNA polymerization by tRNA nucleotidyltransferase. Structure 22:315
Yamashita S, Martinez A, Tomita K (2015) Measurement of acceptor-TPsiC helix length of tRNA for terminal A76-addition by A-adding enzyme. Structure 23:830
Yehudai-Resheff S, Hirsh M, Schuster G (2001) Polynucleotide phosphorylase functions as both an exonuclease and a poly(A) polymerase in spinach chloroplasts. Mol Cell Biol 21:5408
Yue D, Maizels N, Weiner AM (1996) CCA-adding enzymes and poly(A) polymerases are all members of the same nucleotidyltransferase superfamily: characterization of the CCA-adding enzyme from the archaeal hyperthermophile Sulfolobus shibatae. RNA 2:895
Zuo Y, Deutscher M (2001) Exoribonuclease superfamilies: structural analysis and phylogenetic distribution. Nucleic Acids Res 29:1017
Acknowledgements
The author thanks Dr. Shozo Yokoyama for his advice on the phylogenetic analysis and Drs. Yokoyama and Paul Joyce for critical readings of this manuscript. Some of the research described herein was supported by Grant Number MCB-0133520 from the National Science Foundation.
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling editor: Ulrich Muller.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Figure S1
Word file containing the multiple sequence alignment in CLUSTAL format for the nucleotidyltransferase superfamily members examined in this study (DOCX 149 kb)
Figure S2
Maximum likelihood phylogenetic tree constructed as described in Methods. Species names are indicated in color as in Fig. 3 (TIFF 16574 kb)
Figure S3
Alignment of the sequence regions containing the Glu185Gln186/Arg236/Lys232 motif from T. maritima and Synergistetes 53_16. The four amino acids which are involved in the formation of the “springy hinge” are highlighted in red. (TIFF 10235 kb)
Figure S4
Models of the CC- and A-adding enzymes from Deinococcus radiodurans and Caldimicrobium thiodismutans. The models were constructed using SWISS-MODEL, from the relevant amino acid sequences and utilized the Thermotoga maritima CCA-adding enzyme as the template (TIFF 19535 kb)
Rights and permissions
About this article
Cite this article
Jones, G.H. Phylogeny and Evolution of RNA 3′-Nucleotidyltransferases in Bacteria. J Mol Evol 87, 254–270 (2019). https://doi.org/10.1007/s00239-019-09907-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-019-09907-2