Abstract
Our previous calculations of ionic interactions indicated that the Caenorhabditis elegans intermediate filament (IF) IFA proteins, in addition to IFA/IFB-1 heterodimers, may also form homodimers. In order to prove the significance of these calculations, we analysed the dimerization potential of the IFA chains in blot overlays. Unexpectedly, we found here that the dimerization of the IFA-1 protein was of both homotypic and heterotypic nature, and involved all proteins immobilized on the membrane (IFA-1, IFA-2, IFA-4, IFB-1, IFB-2, IFC-1, IFC-2, IFD-1, IFD-2 and IFP-1). A similar interaction profile, though less complex, was observed for two biotinylated proteins (IFA-2 and IFA-4). These and previous results indicate that the IFA proteins are able to form many different heteropolymeric and homopolymeric complexes in the C. elegans tissue, but that only those triggered by the IFA-specific IFB-1 protein result in mature IFs. Moreover, the calculations of the possible ionic interactions between the individual rod sequences as well as their various deletion variants indicated a special role in this process for the middle part of the C. elegans IF coil 1B segment that is deleted in all vertebrate cytoplasmic IFs. We hypothesized here, therefore, that the striking promiscuity of the C. elegans IFs originally involved a nuclear lamin which, due to a two-heptad-long rod deletion, prevented formation of a functional lamin/cIF dimer. This, in concert with an efficient dimerization and a strict tissue-specific co-expression, may allow expansion and maintenance of the multiple Caenorhabditis IFs. A possible implication for evolution of chordate IFs proteins is also discussed.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Intermediate filament (IF) proteins are important components of the cytoskeleton in many metazoan cells. All IF proteins show a common tripartite structure comprising variable head and tail domains flanking a central alpha-helical rod domain divided into three segments (termed 1A, 1B and 2), each of which forms a parallel, double-stranded coiled coil and two linker regions termed L1 and L12. Five major subfamilies of IF have been identified in vertebrates. The largest of these encode the type I and type II cytoplasmic keratins, which give rise to the obligate heteropolymeric keratin filaments of epithelia and epidermal appendages. The type III IF (desmin, vimentin, glial fibrillary acidic proteins) can form, at least in part, homopolymeric IF. The type IV IF subfamily is represented by neurofilament proteins and α-internexin, while the type V IF proteins are the nuclear lamins, which have a unique tail containing an Ig-like segment, a nuclear localization signal and, in most cases, a CaaX box. Finally, the eye lens filensin and phakinin are classified as type VI IF proteins. It seems reasonable to assume that lamins represent an ancestral sequence of both the long/L-type protostomic (sharing with the lamins the long version of the coil 1B subdomain) and the short/S-type deuterostomic (lacking a middle 42 residue-long part of the coil 1B subdomain) cytoplasmic IFs (cIF; for reviews see Fuchs and Weber 1994; Parry and Steinert 1995; Erber et al. 1998, 1999; Herrmann et al. 2009; Peter and Stick 2015; Kollmar 2015; Suzuki et al. 2017; Ehrlich et al. 2019). Orthologs of the vertebrate cytoplasmic type I to III IF proteins have also been found in early chordates (Karabinos et al. 2004b; Karabinos 2013 and references therein).
The primary (but not sole) function of the IF cytoskeleton is to provide resistance against mechanical stress as revealed by a variety of epidermal fragility syndromes induced by mutated human keratins (Irvine and McLean 1999), by knockouts in mice (i.e. Hesse et al. 2000; Vijayaraj et al. 2009) as well as by genetics in the nematode Caenorhabditis elegans (Karabinos et al. 2001, 2003a, 2004a; Hapiak et al. 2003; Woo et al. 2004; Hüsken et al. 2008; Zhang et al. 2011; Geisler et al. 2019).
The nematode C. elegans has a single nuclear lamin LMN-1, which is an essential protein of the early embryo (Liu et al. 2000; Fraser et al. 2000), and eleven cIF genes, some of which give rise to alternative splice variants (Dodemont et al. 1994; Karabinos et al. 2001, 2003a, 2004a, 2019; Woo et al. 2004; Al-Hashimi et al. 2018; Geisler et al. 2019). Interestingly, the C. elegans IF proteins IFA-1 and IFB-1 represent a basic heteropolymeric IF cytoskeleton in all nematode clades investigated thus far, in contrast to the remaining nine C. elegans IF sequences restricted to the clade III-V (IFA-2, IFA-4) and V (IFB-2) taxa, or even to the Caenorhabditis genus (IFA-3, IFC-1, IFC-2, IFD-1, IFD-2, IFP-1; Karabinos 2019). Previously, we and others have reported that the six C. elegans IF proteins IFB-2, IFC-1, IFC-2, IFD-1, IFD-2 and IFP-1 (originally referred to as IFE-1; Karabinos et al. 2001) are expressed in the intestinal terminal web (referred to as the endotube) during different developmental stages and reveal various differently penetrant RNA interference (RNAi) phenotypes (Karabinos et al. 2001, 2002a, 2004a; Hüsken et al. 2008; Geisler et al. 2016, 2019). Moreover, using the recombinantly expressed IFB-2, IFC-1, IFC-2, IFD-1, IFD-2 and IFP-1 proteins and the overlay assay as a tool, we previously found that the latter proteins form another heteropolymeric IFB-2/IFCDP-1 IF system in C. elegans (Karabinos et al. 2017).
Using RNAi on all eleven genes, we previously identified four genes essential for C. elegans development (IFA-1, IFA-2, IFA-3 and IFB-1; Karabinos et al. 2001). Moreover, using IFB-1 RNAi by feeding on wild-type as well as on transgenic worms expressing the IFA-1-GFP protein, we observed not only a postembryonic phenotype similar to that found earlier for IFA-2 and IFA-3 but additional defects of the IFA-1-GFP-containing tonofilament bundles in marginal cells of the pharynx (Karabinos et al. 2003a). These phenotypes and a strict tissue-specific co-expression pattern for the IF proteins IFB-1, IFA-1, IFA-2 and IFA-3 indicate an essential role in the transmission of muscle force to the cuticle and to the maintenance of the correct hypodermis/muscle relationship in development (Francis and Waterston 1991; Hresko et al. 1994, 1999; Karabinos et al. 2002a, 2003a). In addition, the RNAi experiments using the ifb-1 gene cause morphogenetic defects and defective outgrowth of the excretory cells (Karabinos et al. 2003a; Woo et al. 2004; Kolotuev et al. 2013) and the same holds true for its excretory cells’ expressed dimerization partner IFA-4 (Karabinos et al. 2003a; Al-Hashimi et al. 2018; for a review about intestinal and non-intestinal C. elegans IFs see Carberry et al. 2009; Coch and Leube 2016).
Thus, related phenotypes and a strict co-expression of IFB-1 and IFA proteins, described above, correlate well with the in vitro polymerization properties of these proteins. In gel overlays of all eleven recombinant IF proteins, the IFB-1 chain strongly decorates only the IFA-1 to IFA-4 chains, while filament assembly studies show that IFB-1 gives rise to obligate heteropolymeric IFs when mixed with equal molar amounts of IFA-1, IFA-2 or IFA-3 (Karabinos et al. 2003a). Moreover, using the murine IFA-1 to IFA-3-specific monoclonal Ab MH4 and the immunoprecipitation assay as a tool, we identified the heteropolymeric IFA-1/IFB-1 complexes in the whole nematode protein extract, confirming their existence also in vivo (Karabinos 2019). Interestingly, however, the previous calculation of ionic interactions indicates, in addition to IFB-1/IFA heterodimers, a tendency also for IFA/IFA homodimers (Karabinos et al. 2003a), and similarly the previously analysed IFA-5 and IFB-1 IF proteins from the Ascaris lumbricoides body musculature (Geisler et al. 1998; Karabinos 2019) indicate both homotypic and heterotypic modes of assembly. Thus, these findings as a whole raise an important question regarding the rules which control dimerization and assembly processes of the multiple IF proteins in the C. elegans tissue.
In this study, we completed the characterization of the assembly of the IF proteins in C. elegans by searching for in vitro interaction properties of the A-type IF sequences. Using recombinantly expressed IFA-1, IFA-2 and IFA-4 proteins and the overlay assay as a tool, we were able to identify unexpected broad heterotypic properties of these proteins. However, all three proteins also revealed an ability to form a homodimer in blot overlays. The latter results fit well with the calculation of the number of potential interchain ionic interactions of individual IFA proteins within the two-stranded coiled coil. This emphasized, in addition, a special role in dimerization for the middle part of the C. elegans IF coil 1B rod segment, deleted in all chordate S-type cIFs (see above). These highly promiscuous interaction properties of the individual IFA proteins, as revealed in this study, contrast with a striking IFA-specific IFB-1 recognition profile. The same contrast was previously observed between the IFC/IFD/IFP-1 and IFB-2 proteins from another intestine-specific heteropolymeric IFB-2/IFCDP-1 IF system in C. elegans (see above). Thus, the combined results enable us to speculate about the basis of IF heteropolymerization in C. elegans tissue and heterodimer IF formation, which evolved independently on several occasions in the course of IF evolution (Karabinos 2013 and references therein). In addition, we hypothesize here a supportive role of an efficient heterodimerization, a strict tissue-specific co-expression and/or a fine rod lamin versus cIF deviation in the evolution of the unique Caenorhabditis and chordate IF complex.
Materials and Methods
Cloning, expression and purification of the recombinant IF proteins IFA-1, IFA-2, IFA-4, IFB-1, IFC-1, IFC-2, IFD-1, IFD-2 and IFP-1 were reported previously (Karabinos et al. 2001, 2003a). Cloning, expression and purification of the Branchiostoma recombinant IF proteins A3 and B2 and of Styela clava keratins C and D were also reported previously (Karabinos et al. 2002b; Wang et al. 2000). Biotinylation of recombinant C. elegans (IFB-1, IFA-1, IFA-2, IFA-4), Branchiostoma (A3, B2), Styela (C, D) and human (keratin 8 and 18) IF proteins and the overlay assay were essentially as described (Karabinos et al. 2003a). Briefly, biotinylation of recombinant proteins was achieved with the EZ-Link TM Sulfo-NHS-LC-Biotinylation kit from Pierce (Rockford, IL) in 20 mM sodium borate (pH 8.5) containing 4 M urea and 1 mM 2-mercaptoethanol. The overlay experiments using the biotinylated C. elegans (IFB-1, IFA-1, IFA-2 or IFA-4) and chordate (Branchiostoma, Styela and human keratins 8 and 18) IF proteins were performed on nitrocellulose membranes containing the ten immobilized, non-modified C. elegans IF proteins (IFA-1, IFA-2, IFA-4, IFB-1, IFC-1, IFC-2, IFD-1, IFD-2, IFP-1) and the six immobilized, non-modified chordate (Branchiostoma A3/B2, Styela C/D and human keratin 8/18) IF proteins, respectively. Membranes were blocked by incubation for 1 h at room temperature in 50 (w/v) skim milk in TBS (20 mM Tris–HCl, pH 7.4, 150 mM NaCl), 0.05% (v/v) Tween 20 (TBST) and then incubated for two hours at room temperature with the corresponding biotinylated IF probe in 20 mM Tris–HCl buffer (pH 7.5) containing 4 M urea. After washing in TBST, 1% (v/v) Triton, in TBST, 0.5% (w/v) NaCl and again in TBST, each for 5 min, the membrane was treated with horse-radish peroxidase-conjugated streptavidin (Pierce, Rockford, IL, USA) used at a concentration of 0.2 μg/ml. The blots were finally developed using the ECL chemiluminiscence kit (Amersham Pharmacia, Uppsala, Sweden) according to the manufacturer’s instructions.
Results and Discussion
Interaction Properties of the Biotinylated IFA-1, IFA-2 and IFA-4 IF Proteins in Blot Overlay
In order to prove the significance of the previous calculated interchain ionic interactions indicating, in addition to IFB-1/IFA heterodimers, a tendency for IFA/IFA homodimers (Karabinos et al. 2003a), the recombinant IFA-1, IFA-2 and IFA-4 proteins were individually used as a probe to investigate their binding to ten recombinant C. elegans IF proteins (IFA-1, IFA-2, IFA-4, IFB-1, IFB-2, IFC-1, IFC-2, IFD-1, IFD-2 and IFP-1) in blot overlays (see Materials and Methods). This technique was performed in the standard 4 M urea-containing buffer, which was previously successfully applied for dimerization in different IF proteins, including vertebrate keratins (Hatzfeld and Franke 1985), vimentin (Herrmann et al. 1996) and the cephalochordate-specific IF proteins A3 and B2 (Karabinos et al. 2002b, 2012). In addition, the biotinylated C. elegans IF protein IFB-1, the human keratin pair K8/K18 (expressed in all vertebrates; for reviews see Peter and Stick 2015), the single urochordate Styela clava keratin pair C/D (Wang et al. 2000) as well as the Branchiostoma-specific pair A3/B2 (Karabinos et al. 2002b, 2012) was used in the study as a positive control.
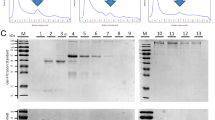
In line with our previous experiments (Karabinos et al. 2003a), the biotinylated IFB-1 strongly recognized proteins IFA-1, IFA-2 and IFA-4 (and also weakly IFC-2 and IFP-1) in blot overlay (IFA-1-biotin overlay panel in Fig. 1a). Figure 1a (IFA-1-biotin overlay panel) also shows that the biotinylated IFA-1, somewhat surprisingly, recognizes all proteins immobilized on the membrane, including the homotypic IFA-1/IFA-1 interactions. This latter result fits the previously described IFA-1 interchain ionic calculations which indicate, in addition to heterodimers, a tendency for IFA-1/IFA-1 homodimers (Karabinos et al. 2003a). A similar binding pattern was also seen with the biotinylated IFA-2 protein which, in addition to IFA-2/IFA-2 homodimers, also revealed a binding to IFA-4, IFC, IFD and IFP-1 proteins (IFA-2-biotin overlay panel in Fig. 1a) as well as with the biotinylated IFA-4 protein. The latter, in addition to IFA-4/IFA-4 homodimers, also revealed a binding to IFA-2, IFB-1, IFC and IFP-1 proteins (IFA-4-biotin overlay panel in Fig. 1a). These results suggest that the IFA-1, IFA-2, IFA-4 and, by analogy, the IFA-2-homologous IFA-3 proteins (Dodemont et al. 1994; Karabinos 2019), may form heterocomplexes with multiple C. elegans IF proteins. Homopolymeric complexes are also clearly possible.
Blot overlay assays of biotin-labelled C. elegans and chordate IF proteins. a Equal amounts of purified recombinant C. elegans proteins IFA-1, IFA-2, IFA-4, IFB-1, IFB-2, IFC-1, IFC-2, IFD-1, IFD-2 and IFP-1 were separated by 10% SDS-PAGE and stained with Coomassie dye (upper panel) or transferred to nitrocellulose membranes. The lower panels show the blots after incubation and staining with the IFB-1-, IFA-1-, IFA-2- and IFA-4-biotin proteins (marked on the left). b Equal amounts of purified recombinant human (huK18, huK8), Branchiostoma floridae (BfB2, BfA3) and Styela clava (ScD, ScC) IF proteins were separated by 10% SDS-PAGE and stained with Coomassie dye (upper panel) or transferred to nitrocellulose membranes. The lower panels show the blots after incubation and staining with the huK18-, huK8-, ScC-, ScD-, BfA3- and BfB2-biotin proteins (marked on the left). The summary of heterotypic and homotypic interactions for individual blot overlay experiments is presented on the right. Bold letters on the right indicate strong detected protein bands on the individual blot overlay membranes. Very weakly detected protein bands on individual blot overlay membranes are marked by asterisks. Note the homotypic interactions seen on presented blot membranes in (a), while no homotypic interactions were seen on presented blot membranes in (b)
In line with our previous blot overlay experiments (Karabinos et al. 2017), we also observed that some biotin-labelled proteins failed to interact with another protein, which exclusively decorated it in the reciprocal blot overlay assay (i.e. inability of IFA-2 to bind IFB-1 or a broad promiscuity of IFA-1/IFA-2 with IFB-2 and/or IFC/D/P-1 proteins). This has not been previously observed (for details see Karabinos et al. 2017). As already mentioned in the latter study, this might arise because a putative protein-binding sequence of the biotinylated protein may have been blocked due to biotinylation of lysines and/or, because partial SDS-denaturation of the proteins fixed on overlay membranes may have compromised interactions of the biotinylated protein and its partner in the blot overlay experiment. However, some signals in blot overlay experiments might reflect interactions of the terminal head and/or tail domains of the investigated proteins and not a true double-stranded coiled coil.
Taken together, the here experimentally assessed polyvalent/homopolymeric IFA interaction properties, documented in Fig. 1a, resemble those of the C. elegans IFC-1, IFC-2, IFD-1 and IFD-2 proteins, which have been previously shown in blot overlay to form many different heteropolymeric and three (IFC and IFD-2) homopolymeric complexes (Karabinos et al. 2017). Similarly, the here (Fig. 1a) and previously presented (Karabinos et al. 2003a) blot overlay-assessed obligate heteropolymeric IFA-profile of the IFB-1 protein resembles a strict IFC/D/P-1 heterodimeric profile of the biotinylated IFB-2 (Karabinos et al. 2017), while none from the two IFB proteins reveal in the latter blot overlay experiments a self-interaction ability. These results enabled us to conclude that both these related heteropolymeric IFB-1/IFA and IFB-2/IFCDP-1 IF systems probably retained similar heterodimer formation principles during the course of Caenorhabditis IF evolution (for details about evolution of nematode IF genes see Karabinos 2019). Moreover, the polyvalent/homopolymeric IFA protein ability presented here also deviates from the control blot overlay experiments in Fig. 1b of the human keratin pair K8/K18 (Hatzfeld and Franke 1985), the urochordate keratin pair C/D (Wang et al. 2000), as well as the cephalochordate-specific IF pair A3/B2 (Karabinos et al. 2002b, 2012), which all strongly decorate their partner chains (or huK18 protein in the case of the Styela clava keratin D), while not one homodimer is formed.
Calculation of Interchain Ionic Interactions of Individual IFA and IFB-1 Rod Homo and Heterodimers
In order to find sequences/segments that might be responsible for different dimerization properties of the individual IFA and IFB-1 members of the heteropolymeric IFB-1/IFA IF system in C. elegans, described above, all possible ionic interactions were calculated between pairs of chains in the two-stranded molecules for different combinations involving the full-length IFA-1, IFA-2, IFA-3, IFA-4 and IFB-1 rod sequences (Table 1; the sequences with the corresponding interchain ionic interactions are provided in Fig. S1A), as well as their various deletion variants lacking either the coil segment 1A, 1B, the first (42 residues), middle (42 residues) and the last (59 residues) part of the coil segment 1B, or the entire coil segment 2 (Table 1). It should be noted, however, that some of the calculated values presented here deviate from those previously published (Karabinos et al. 2003a) because they now represent the maximum number of possible interactions within the whole two-stranded coiled coil segments, in contrast to our previous approach, mentioned above, in which parts of the rod domain with low-coiled coil potential were omitted from the calculations (for details see also Citi et al. 2000).
In line with our previous ionic interaction score calculations (Karabinos et al. 2003a), the IFA-1 and IFA-4 rod would, in order, have a preference for the formation of IFA-1/IFA-1 (26) and IFA-4/IFA-4 (26) homodimers over the formation of IFA-1/IFB-1 (22) and IFA-4/IFB-1 (22) heterodimers (upper panel of Table 1). In contrast, the highly related IFA-2 and IFA-3 rods would favour IFA-2/IFB-1 (24) and IFA-3/IFB-1 (21) heterodimerization over the formation of IFA-2 (22) and IFA-3 (16) homodimers (upper panel of Table 1). The calculated net number of ionic interactions for the IFA-1, IFA-2, IFA-3 and IFA-4 homodimers minus that for the corresponding IFA/IFB-1 heterodimer, would be 4, − 2, − 5 and 4, respectively (see the “T” values in the upper panel of Table 1). When we calculated the net numbers of ionic interactions for the individual parallel and in-register arrangements of the various deletion variants (see above) for IFA-1, IFA-2, IFA-3 and IFA-4 homodimers minus the corresponding IFA/IFB-1 heterodimer, only the deletion of the middle 42 residue-long part of the coil 1B segment would consistently support heteropolymeric over homopolymeric assembly in all of the analysed dimers. All other deletions would have a variable effect on different analysed dimers (i.e. neutral (ne), homodimer (ho) or heterodimer (he) supporting; upper panel of Table 1). Interestingly, when we did a reciprocal calculation of the net number of ionic interactions for the individual parallel and in-register arrangements of the various deletion variants of the IFB-1 homodimer minus the corresponding IFB-1/IFA heterodimer, only deletion of the whole coil 1B or its two 42 residue-long derivatives would constantly support the homodimer over the heterodimer, while all other deletions would have a variable effect on different analysed dimers (i.e. neutral (ne), homodimer (ho) or heterodimer (he) supporting; lower panel of Table 1). Thus, the middle 42 residue-long part of the coil 1B segment consistently supports the homodimerization in all IFA proteins, while in the IFB-1 protein it seems to have an opposite effect by supporting heterodimerization with the IFA binding partners.
Calculation of Interchain Ionic Interactions of Individual IFCDP-1 and IFB-2 Rod Homo- and Heterodimers
In order to directly compare theoretical dimerization principles of the two heteropolymeric IFB-1/IFA and IFB-2/IFCDP-1 IF systems in C. elegans, we also applied the in silico approach, used in the previous section, for the IFB-2, IFC, IFD and IFP-1 proteins (Table 2). In line with our previous calculations (Karabinos et al. 2017), the IFC-2 and IFD-2 rod would, in order of ionic interaction score, prefer IFC-2/IFC-2 (26) and IFD-2/IFD-2 (24) homodimers over the formation of the IFC-2/IFB-2 (18) and IFD-2/IFB-2 (23) heterodimers (upper panel of Table 2). In contrast, the IFC-1, IFD-1 and IFP-1 rod would significantly prefer IFC-1/IFB-2 (15), IFD-1/IFB-2 (21) and IFP-1/IFB-2 (14) heterodimerization, respectively, over the formation of the IFC-1 (8), IFD-1 (16) and IFP-1 (4) homodimers (upper panel of Table 2). The calculated net charge for the IFC-1, IFC-2, IFD-1, IFD-2 and IFP-1 homodimers minus the corresponding IFCDP-1/IFB-2 heterodimer would be − 7, 8, − 5, 1 and − 10, respectively (see the “T” values on upper panel of Table 2). When we calculated the net number of ionic interactions for the individual parallel and in-register arrangements of the various deletion variants (see above) of the IFC-1, IFC-2, IFD-1, IFD-2 and IFP-1 homodimers minus the corresponding IFCDP-1/IFB-2 heterodimer, a deletion of the middle 42 residue-long part of the coil 1B segment would support heterotypic over the homotypic assembly of the majority (IFC-1, IFC-2 and IFD-1) of the analysed proteins (this result resembles that of the IFA/IFB-1 calculation, described above), while for the remaining IFD-2 and IFP-2 proteins it is neutral and homodimer supporting, respectively. Similarly, a deletion of the whole coil 2 segment would support heterotypic over the homotypic assembly of the three (IFC-2, IFD-2 and IFP-1) analysed proteins, while for the two remaining IFC-1 and IFD-1 proteins it is homodimer supporting. All other calculated deletions would have a variable effect on different analysed dimers (i.e. neutral (ne), homodimer (ho) or heterodimer (he) supporting; upper panel of Table 2). When we did a reciprocal calculation of the net number of ionic interactions for the individual parallel and in-register arrangements of the various deletion variants of the IFB-2 homodimer minus the corresponding IFB-2/IFCDP-1 heterodimer, only deletion of the first 42 residue-long part of the coil 1B would constantly support homodimeric over heterodimeric assembly, while all other deletions would have a variable effect on different analysed dimers (i.e. neutral (ne), homodimer (ho) or heterodimer (he) supporting; lower panel of Table 2).
Taken together, using this simple in silico analysis we were able to identify a potentially significant feature of the middle 42 residue-long part of the coil 1B segment, which in all IFA, IFC and IFD-1 proteins constantly supports their homodimerization, while in IFB-1 it might be (at least partly) responsible for the striking IFA-specificity in the blot overlay experiments (Fig. 1a; Karabinos et al. 2003a, 2017).
The Coil 1B Segment Probably Determines the Specificity of the IFB-1 Dimerization
It is known that charged residues often occupy positions a and d of the heptads in trigger sequences in coiled coil proteins and that these are involved in both intra- and inter-helical salt bridges (Burkhard et al. 2000). A detailed inspection of the heptads in the 1B segments of IFA-1, IFA-2, IFA-3 and IFB-1 arranged as homodimers or IFA/IFB-1 heterodimers showed two regions (marked with arrowheads in Fig. 2) with several charged residues (R, D, E) in positions g, a, d and/or e. These have the potential for multiple repulsive interchain salt bridges of the 2e′-1g, 1g′-2e, 2a′-1g and 1g′-2a type (indicated with dashed lines in Fig. 2), as well as for two attractive inter-helical salt bridges of the 2e′-1d and 1d′-2e type (indicated with normal lines in Fig. 2). In addition, the position of several other potential attractive salt bridges of the classical 2e′-1g and 1g′-2e type were identified in all coil 1B sequences (marked by arrows in Fig. 2). Interestingly, when we summarized the ability of the corresponding dimers to produce potential attractive and repulsive salt bridges, the IFA homodimers revealed no repulsive salt bridges (all were attractive), while 60%, 33% and 14% of repulsive salt bridges were found in the IFB-1 homodimer, the IFB-1/IFA-1 to -3 heterodimer and the IFB-1/IFA-4 heterodimer, respectively (right panel of Fig. 2). The evidence suggests that the middle 42 residue region in the 1B sequence of the IFA chains is actively involved in homodimerization of these proteins. This is in contrast to the corresponding IFB-1 coil 1B segment which harbours a sequence motif (i.e. a trigger-like motif) that prevents IFB-1 homodimer formation due to the repulsion of like charged amino acids as a pre-requisite for an efficient IFB-1/IFA heterodimerization and subsequent mature IF assembly. Moreover, the latter IFB-1/IFA heterodimers have the potential to form several repulsive interchain ionic interactions, possibly rendering the heterodimers less stable and thereby controlling a dynamic turnover of the IFB-1/IFA tonofibrils, as previously documented by fluorescence recovery after photobleaching in the marginal cells of the pharynx (Karabinos et al. 2003a). Further assembly and/or X-ray crystallography experiments, including both the wild-type and the coil 1B-deletion mutant proteins, are needed to explore the function of the coil 1B segment and its individual fragments in dimerization of the multiple C. elegans IF proteins in more detail. In this respect, we note our previous work, which identified a putative trigger-like sequence in segment 1B of the Branchiostoma-specific IF protein B2 (Karabinos et al. 2012).
Alignment of the individual homodimers (ho) and heterodimers (he) of the IFA-1, IFA-2, IFA-3, IFA-4 and IFB-1 rod coil 1B fragments. Asterisks between the rod fragments of the individual proteins mark the a and d positions of the heptad repeat pattern. The 42 residue-long middle part of the coil 1B segment is marked by dots above the sequences. The vertical lines between the individual sequences mark the positions of the charged residues in the presented coil segments which are involved in potential 2e′-1g, 1g′-2e, 2a′-1g, 1g′-2a, 2e′-1d, and 1d′-2e attractive (normal lines) or repulsive (dashed lines) salt bridges. The two arrowheads shown above the sequences indicate positions of the two groups of repulsive salt bridges, while the position of five other potential attractive salt bridges of the classical 2e′-1g and 1g′-2e type are marked by arrows (see text for details). The summary of the predicted attractive and repulsive salt bridges for individual dimers is presented on the right. Note 60% of the predicted repulsive salt bridges occur for the IFB-1 homodimer, while none (0%) comprise the individual homotypic IFA dimers
Possible Mechanisms Allowing Expansion and Maintenance of the Caenorhabditis and Chordate IFs
As previously reported, the C. elegans IFA-1 and IFB-1 proteins probably appeared at the base of the nematode branch and represent a basic heteropolymeric IF cytoskeleton of all nematode clades, while the other nematode IFA, IFB and IFC/D/P-1 IF proteins arose later during nematode evolution from an AB-type IF ancestor (see Introduction). This nematode-specific radiation may indicate that the IFA-1 ancestral sequence originally possessed still retains a high promiscuous mode of assembly, as reported here, while its evolutionary descendant IFs, due to a sequence/functional diversification in the nematode tissue (see Introduction), lost a part of it (IFA-2 to IFA-4, IFC/D/P-1) or switched to the heterotypic mode of assembly (IFB; see Karabinos et al. 2003a, 2017 and text above). Moreover, the IFA-1 and IFB-1 proteins, as we previously reported, also have a striking lamin-like character comprising a longer rod domain, a significant length and sequence conservation in their N- and C-terminal extensions and about 23–25% identity over the rod domain (including several conserved charged a and d positions of heptads), which is in the range found among the C. elegans IF proteins (Karabinos 2019). Thus, these and our present findings enable us to propose (though it is not yet experimentally proved), that some of the promiscuous C. elegans cIF may form chimeric heterocomplexes with a nuclear lamin (LMN-1) during a known cytoskeletal reorganization at mitosis. If this proposal is correct, such cIF/lamin interactions would likely be harmful as a consequence of interference with lamin assembly and function in the C. elegans cell, since LMN-1 is a ubiquitously expressed and essential gene in the early embryo (see Introduction; for a review about invertebrate lamins see Melcer et al. 2007), which forms another 10-nm lamin-like IF system based on a parallel/double-stranded dimer (Karabinos et al. 2003b; Ben-Harush et al. 2009). There are several known mechanisms by which animal cells could prevent such proposed collisions. Firstly, an efficient heterodimerization of the fully compatible IF binding partners might preclude their connections to lamins or to other less compatible IF partners. The latter mechanism in C. elegans seems to be represented by the two homologous IFB-1 and IFB-2 proteins which, due to a striking specificity for the IFA-1 to IFA-4 and IFC-1 to IFP-1 chains, respectively (as described above), might readily be able to involve them in the formation of stable heterodimers and subsequently into mature heteropolymeric IFs, precluding thereby a formation of undesired complexes. This suggestion is further directly supported by our immunoprecipitation experiments using the murine IFA-1 to IFA-3-specific monoclonal Ab MH4 identifying only the heteropolymeric IFA-1/IFB-1 complexes in the whole nematode protein extract (Karabinos 2019), as well as by a recent study demonstrating that the single ifb-2 gene knockout prevents endotube formation in the IF-rich C. elegans intestine (Geisler et al. 2019). As indicated by the in silico study described here, the efficient heteropolymerization process may be controlled (at least in part) by a short trigger-like sequence located in the coil 1B segment of the C. elegans IFs (see also below). Moreover, it seems that heterodimer formation occurs even more efficiently in chordates, as documented by the above described blot overlay experiments of the chordate keratins (for details see also Hatzfeld and Franke 1985) and related proteins, all of which strongly decorate their partner chains, while no homodimers at all are formed (Fig. 1b and text above). Secondly, a strict tissue-specific co-expression pattern, well documented from studies of vertebrate type I and type II keratins (for reviews see Fuchs and Weber 1994; Parry and Steinert 1995), is also seen in the C. elegans tissue (see Introduction) and may allow only the appropriate IF partner chains, precluding thereby a formation of undesired IF complexes. Thirdly, a specific structural deviation between the cIF proteins and lamins would also be expected to compromise the formation of undesired cIF/lamin complexes, as we proposed above. Interestingly, the C. elegans LMN-1, like other lamins of the nematode Caenorhabditis genus (Fig. S1B), harbours a unique two-heptad-long deletion in the coil 2 segment of the rod, which would be expected to compromise formation of a functional (i.e. symmetrical/unstaggered) cIF/lamin dimer in the C. elegans cell and a similar six heptad-long (42 residues) rod deletion is also seen in all chordate S-type cIF proteins (for details see Introduction). Curiously, the latter chordate-specific deletion exactly matches the IFA, IFB-1, IFC and IFD-1 coil 1B segments, which are (at least partly) responsible for the different behaviour of these C. elegans proteins in blot overlay (Figs. 1, 2 and text above). Finally, additional but as yet unknown mechanisms must exist (at least) in the mammalian cell which are able to separate different co-expressed IF chains from each other, as demonstrated by studies with the mammalian B1 and A-type lamins, which show mixed filaments in vitro while, a homopolymeric assembly is defined in vivo (Kolb et al. 2011; for review see Osmanagic-Myers et al. 2016).
Taken together, a surprising promiscuity revealed in our present experiments enable us to hypothesize that (a) the metazoan cytoplasmic IFs are subject to evolutionary constraints (most of the metazoans contain only a few genes; for a summary on the number of currently known metazoan IF genes see Peter and Stick 2015) in order to preclude inappropriate interactions between wrong IF chains, including lamin and that (b) the latter complexes might be prevented by an efficient heterodimerization, a strict tissue-specific IF co-expression and/or a fine structural cIF versus lamin rod deviation (i.e. several heptad-long deletions), which together released the selective constraints on the cIFs and promoted, at least in part, their expansion from the few genes existing in a common ancestor of nematodes (Karabinos 2019), cephalochordates (Karabinos 2013) and vertebrates (Vandebergh and Bossuyt 2012). A more detailed picture of the mechanisms which influence dimer/IF formation is predicted to emerge from in vitro and in vivo assembly studies of the full-length or truncated mutant IF proteins (see above), from transgenic worms, as well as from X-ray crystallography. An important experiment would also be to either delete/mutate the coil 1B domain (or its individual fragments) in the IFB-1 protein or to replace it with the homologous IFA sequences. The mutated IFB-1 proteins may then be able to form homotypic interactions in blot overlay. Such an altered recognition ability has already been described with the Branchiostoma-specific IF protein B2 (Karabinos et al. 2012). In conclusion, our study has resulted in a number of interesting hypotheses that are amenable to molecular experimentation. This will be required to better understand the evolution, structure and function of the multiple IF proteins in nematodes and chordates.
Abbreviations
- GFP:
-
Green fluorescent protein
- IF:
-
Intermediate filament
- RNAi:
-
RNA interference
References
Al-Hashimi H, Hall DH, Ackley BD, Lundquist EA, Buechner M (2018) Tubular excretory canal structure depends in intermediate filaments EXC-2 and IFA-4 in Caenorhabditis elegans. Genetics 210:637–652
Ben-Harush K, Wiesel N, Frenkiel-Krispin D, Moeller D, Soreq E, Aebi U, Herrmann H, Gruenbaum Y, Medalia O (2009) The supramolecular organization of the C. elegans nuclear lamin filament. J Mol Biol 386:392–1402
Blaxter M (2011) Nematodes: the worm and its relatives. PLoS Biol 9(4):1–9
Burkhard P, Kammerer RA, Steinmetz MO, Bourenkov GP, Aebi U (2000) The coiled-coil trigger site of the rod domain of cortexillin I unveils a distinct network of interhelical and intrahelical salt bridges. Structure 8:223–230
Carberry K, Wiesenfahrt T, Windoffer R, Bossinger O, Leube RE (2009) Intermediate filaments in Caenorhabditis elegans. Cell Motil Cytoskelet 66:852–864
Citi S, D’Atri F, Parry DAD (2000) Human and Xenopus cingulin a modular organization of the coiled-coil rod domain: predictions for intra- and intermolecular assembly. J Struct Biol 131:135–145
Coch RA, Leube RE (2016) Intermediate filaments and polarization in the intestinal epithelium. Cells 5:32. https://doi.org/10.3390/cells5030032
Dodemont H, Riemer D, Ledger N, Weber K (1994) Eight genes and alternative RNA processing pathways generate an unexpectedly large diversity of cytoplasmic intermediate filament proteins in the nematode Caenorhabditis elegans. EMBO J 13:2625–2638
Ehrlich F, Fischer H, Langbein L, Praetzel-Wunder S, Ebner B, Figlak K, Weissenbacher A, Sipos W, Tschachler E, Eckhart L (2019) Differential evolution of the epidermal keratin cytoskeleton in terrestrial and aquatic mammals. Mol Biol Evol 26(2):328–340
Erber A, Riemer D, Bovenschulte M, Weber K (1998) Molecular phylogeny of metazoan intermediate filament proteins. J Mol Evol 47:751–762
Erber A, Riemer D, Hofemeister H, Bovenschulte M, Stick R, Panopoulou G, Lehrach H, Weber K (1999) Characterisation of the Hydra lamin and its gene; a molecular phylogeny of metazoan lamins. J Mol Evol 49:260–271
Francis R, Waterston RH (1991) Muscle cell attachement in Caenorhabditis elegans. J Cell Biol 114:465–479
Fraser AG, Kamath RS, Zipperlen P, Martinez-Campos M, Sohrmann M, Ahringer J (2000) Functional genomic analysis of C. elegans chromosome I by systematic RNA interference. Nature 408:325–330
Fuchs E, Weber K (1994) Intermediate filaments: structure, dynamics, function and disease. Annu Rev Biochem 63:345–382
Geisler N, Schünemann J, Weber K, Häner M, Aebi U (1998) Assembly and architecture of invertebrate cytoplasmic intermediate filaments reconcile features of vertebrate cytoplasmic and nuclear lamin-type intermediate filaments. J Mol Biol 282:601–617
Geisler F, Gerhardus H, Carberry K, Davis W, Jorgensen E, Richardson C, Bossinger O, Leube RE (2016) A novel function for the MAP kinase SMA-5 in intestinal tube stability. Mol Biol Cell 27(24):3855–3868
Geisler F, Coch RA, Richardson C, Goldberg M, Denecke M, Bossinger O, Leube RE (2019) The intestinal intermediate filament network responds to and protects against microbial insults and toxins. Development 146(2):dev169482. https://doi.org/10.1242/dev.169482
Hapiak V, Hresko MC, Schriefer LA, Saiyasisongkhram K, Bercher M, Plenefisch JD (2003) mua-6, a gene required for tissue integrity in Caenorhabditis elegans, encodes a cytoplasmic intermediate filament. Dev Biol 263:330–342
Hatzfeld M, Franke WW (1985) Pair formation and promiscuity of cytokeratins: formation in vitro of heterotypic complexes and intermediate-sized filaments by homologous and heterologous recombinations of purified polypeptides. J Cell Biol 101:1826–1841
Herrmann H, Häner M, Brettel M, Müller S, Goldie K, Fedtke B, Lustig A, Franke WW, Aebi U (1996) Structure and assembly properties of the intermediate filament protein vimentin: the role of its head, rod and tail domains. J Mol Biol 264:933–953
Herrmann H, Strelkov SV, Burkhard P, Aebi U (2009) Intermediate filament: primary determinants of cell architecture and plasticity. J Clin Investig 119:1772–1783
Hesse M, Franz T, Tamai Y, Taketo MM, Magin TM (2000) Targeted deletion of keratin 18 and 19 leads to trophoblast fragility and early embryonic lethality. EMBO J 19:5060–5070
Hresko MC, Williams BD, Waterston RH (1994) Assembly of body wall muscle and muscle cell attachment structures in Caenorhabditis elegans. J Cell Biol 124:491–506
Hresko MC, Schriefer LA, Shrimankar P, Waterston RH (1999) Myotactin, a novel hypodermal protein involved in muscle-cell adhesion in Caenorhabditis elegans. J Cell Biol 146:659–672
Hüsken K, Wiesenfahrt T, Abraham Ch, Windoffer R, Bossinger O, Leube RE (2008) Maintenance of the intestinal tube in Caenorhabditis elegans: the role of the intermediate filament protein IFC-2. Differentiation 76:881–896
Irvine AD, McLean WH (1999) Human keratin diseases: the increasing spectrum of disease and subtlety of the phenotype-genotype correlation. Br J Dermatol 140:815–828
Karabinos A (2013) The cephalochordate Branchiostoma genome contains 26 intermediate filament (IF) genes: implication for evolution of chordate IF proteins. Eur J Cell Biol 92(8–9):295–302
Karabinos A (2019) Intermediate filament (IF) proteins IFA-1 and IFB-1 represent a basic heteropolymeric IF cytoskeleton of nematodes: a molecular phylogeny of nematode IFs. Gene 692:44–53
Karabinos A, Schmidt H, Harborth J, Schnabel R, Weber K (2001) Essential roles for four cytoplasmic intermediate filament proteins in Caenorhabditis elegans development. Proc Natl Acad Sci USA 98(14):7863–7868
Karabinos A, Schulze E, Klisch T, Wang J, Weber K (2002a) Expression profiles of the essential intermediate filament (IF) protein A2 and the IF protein C2 in the nematode Caenorhabditis elegans. Mech Dev 117:311–314
Karabinos A, Schünemann J, Parry DAD, Weber K (2002b) Tissue-specific coexpression and in vitro heteropolymer formation of the two small Branchiostoma intermediate filament proteins A3 and B2. J Mol Biol 316:127–137
Karabinos A, Schulze E, Schünemann J, Parry DAD, Weber K (2003a) In vivo and in vitro evidence that the four essential intermediate filament (IF) proteins A1, A2, A3 and B1 of the nematode Caenorhabditis elegans form an obligate heteropolymeric IF system. J Mol Biol 333:307–319
Karabinos A, Schünemann J, Meyer M, Aebi U, Weber K (2003b) The single nuclear lamin of Caenorhabditis elegans forms in vitro stable intermediate filaments and paracrystals with a reduced axial periodicity. J Mol Biol 325:241–247
Karabinos A, Schünemann J, Weber K (2004a) Most genes encoding cytoplasmic intermediate filament (IF) proteins of the nematode Caenorhabditis elegans are required in late embryogenesis. Eur J Cell Biol 83(9):457–468
Karabinos A, Zimek A, Weber K (2004b) The genome of the early chordate Ciona intestinalis encodes only five cytoplasmic intermediate filament proteins including a single type I and type II keratin and a unique IF-annexin fusion protein. Gene 326:123–129
Karabinos A, Schünemann J, Parry DAD (2012) A rod domain sequence in segment 1B triggers dimerisation of the two small Branchiostoma IF proteins B2 and A3. Eur J Cell Biol 91:800–808
Karabinos A, Schünemann J, Parry DAD (2017) Assembly studies of six intestinal intermediate filament (IF) proteins B2, C1, C2, D1, D2, and E1 in the nematode C. elegans. Cytoskeleton 74(3):107–113
Karabinos A, Schulze E, Baumeister R (2019) Analysis of the novel excretory cell expressed ECP-1 proteins and its proposed ECP-1/IFC-2 fusion proteins EXC-2 in the nematode Caenorhabditis elegans. Gene Expr Patterns 34:119061
Kolb T, Maaβ K, Hergt M, Aebi U, Herrmann H (2011) Lamin A and lamin C form homodimers and coexist in higher complex forms both in the nucleoplasmic fraction and in the lamina of cultured cells. Nucleus 2(5):425–433
Kollmar M (2015) Polyphyly of nuclear lamin genes indicates an early eukaryotic origin of the metazoan-type intermediate filament proteins. Sci Rep 5:10652. https://doi.org/10.1038/srep10652
Kolotuev I, Hyenne V, Schwab Y, Rodriguez Y, Labouesse M (2013) A pathway for unicellular tube extension depending on the lymphatic vessel determinant Prox1 and on osmoregulation. Nat Cell Biol 15(2):157–168
Liu J, Ben-Shahar TR, Riemer D, Treinin M, Spann P, Weber K, Fire A, Gruenbaum Y (2000) Essential roles for Caenorhabditis elegans lamin gene in nuclear organization, cell cycle progression, and spatial organization of nuclear pore complexes. Mol Biol Cell 11:3937–3947
Melcer S, Gruenbaum Y, Krohne G (2007) Invertebrate lamins. Exp Cell Res 313:2157–2166
Osmanagic-Myers S, Dechat T, Foisner R (2016) Lamins at the crossroads of mechanosignaling. Genes Dev 29:225–237
Parry DAD, Steinert PM (1995) Intermediate filament structure. Springer, New York
Peter A, Stick R (2015) Evolutionary aspects in intermediate filament proteins. Curr Opin Cell Biol 32:48–55
Suzuki KT, Suzuki M, Shigeta M, Fortriede JD, Takahashi S, Mawaribuchi S, Yamamoto T, Taira M, Fukui A (2017) Clustered Xenopus keratin genes: a genomic, transcriptomic, and proteomic analysis. Dev Biol 426:384–392
Vandebergh W, Bossuyt F (2012) Radiation and functional diversification of alpha keratins during early vertebrate evolution. Mol Biol Evol 29:995–1004
Vijayaraj P, Kröger K, Reuter U, Windoffer R, Leube RE, Magin TM (2009) Keratins regulate protein biosynthesis through localization of GLUT1 and -3 upstream of AMP kinase and Raptor. J Cell Biol 187:175–184
Wang J, Karabinos A, Schünemann J, Riemer D, Weber K (2000) The epidermal intermediate filament proteins of tunicates are distant keratins; a polymerisation-competent hetero coiled coil of the Styela D protein and Xenopus keratin 8. Eur J Cell Biol 79:478–487
Woo W-M, Goncharov A, Jin Y, Chisholm AD (2004) Intermediate filaments are required for C. elegans epidermal elongation. Dev Biol 267:216–229
Zhang H, Landmann F, Zahreddine H, Rodriguez D, Koch M, Labouesse M (2011) A tension-induced mechanotransduction pathway promotes epithelial morphogenesis. Nature 471:99–103
Acknowledgements
This work was funded by the European Regional Development OPVaV-2009/2.2/05-SORO (ITMS code:26220220143).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors proclaim no conflict of interests.
Additional information
Handling Editor: Kerry Geiler-Samerotte.
Electronic supplementary material
Below is the link to the electronic supplementary material.
239_2019_9904_MOESM1_ESM.doc
Fig. S1. (A) Alignment of the rod amino acid sequences of the C. elegans lamin (LMN-1) and cytoplasmic proteins IFA-1 to IFA-4, IFB-1, IFB-2, IFC-1, IFC-2, IFD-1, IFD-2 and IFP-1. Arrows pointing downwards and upwards mark the beginning and the end of the rod segments 1A, 1B and 2. The linkers L1 and L12, as well as the stutter in coil 2 are also indicated. Asterisks between the rod segments of the individual proteins mark the a and d positions of the heptad repeat pattern. Dashes are used to optimize the sequence alignment. Bold letters indicate residues that are identical in all twelve proteins. The 42 residue-long region of the C. elegans IF coil 1B segments, which is deleted by all chordate short/S-type IF proteins, is marked by dots above the sequences. The vertical lines between the individual sequences mark the positions of the charged residues in the corresponding coil segments which are involved in potential 2e′-1 g, 1 g′-2e, 2a′-1 g, 1 g′-2a, 2e′-1d, and 1d′-2e attractive (normal lines) or repulsive (dashed lines) salt bridges (see text for details). (B) Alignment of the full-length (fl) C. elegans Lamin (CeLamin_fl) and the seventeen nematode Lamins (the presented alignment was retrieved from the CyMoBase; Kollmar 2015). The arrowheads pointing downwards and upwards mark the beginning and the end of the rod segments 1A, 1B and 2, which are connected by linkers L1 and L12, respectively (for details, see the text). Dashes are used to optimize the sequence alignment. Note that only C. elegans Lamin harbours 14 deleted residues in the coil 2B but the same deletion is also seen in the related Caenorhabditis remanei, Caenorhabditis brenneri and Caenorhabditis briggsae species (data not shown). Also note that Lamins from another two clade V Heterorhabditis and Oscheius species have a coil 2 which is only one heptad shorter. The compared full-length (fl) Lamin sequences are derived from the following nematode species: Clade I: Trichinella spiralis (Trs), Trichuris muris (Trm); Clade III: Ascaris suum (Ass), Brugia malayi (Brm), Onchocerca volvulus (Ov), Wuchereria bancrofti (Wb); Clade IV: Bursaphelenchus xylophilus (Bx), Globodera pallida (Glp), Heterodera glycines (Heg), Meloidogyne incognita (Mi), Meloidogyne hapla (Mh), Panagrellus redivivus (Par); Clade V: Caenorhabditis elegans (Ce), Heterorhabditis baderophora (Hb), Oscheius tipulae (Oct), Pristionchus pacificus (Psp). The clades III (Spirurina), IV (Tylenchina and other) and V (Rhabditina) belong to Chromadoria, which diverged during nematode evolution from the more ancient Dorylaimia (clade I) and Enoplia (clade II; for review see Blaxter, 2011). Supplementary material 1 (DOC 288 kb)
Rights and permissions
About this article
Cite this article
Karabinos, A., Schünemann, J. & Parry, D.A.D. Promiscuous Dimerization Between the Caenorhabditis elegans IF Proteins and a Hypothesis to Explain How Multiple IFs Persist Over Evolutionary Time. J Mol Evol 87, 221–230 (2019). https://doi.org/10.1007/s00239-019-09904-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-019-09904-5