Abstract
Skin pigmentation is a complex, conspicuous, highly variable human trait that exhibits a remarkable correlation with latitude. The evolutionary history and genetic basis of skin color variation has been the subject of intense research in the last years. This article reviews the major hypotheses explaining skin color diversity and explores the implications of recent findings about the genes associated with skin pigmentation for understanding the evolutionary forces that have shaped the current patterns of skin color variation. A major aspect of these findings is that the genetic basis of skin color is less simple than previously thought and that geographic variation in skin pigmentation was influenced by the concerted action of different types of natural selection, rather than just by selective sweeps in a few key genes.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
“The ancients believed that the dark skin of some peoples is explained by the heat of the sun. Indeed, it is observed from experience that with increased proximity to the hot countries of the south, people become darker; on the other hand, if we travel up north, we find that men become increasingly fair-skinned, like the French, the Germans, the English and others. However, we can be certain that along the equator, some men are born almost white, as […] on the island of São Tomé. This island was colonized by the Portuguese, and after one hundred years, their children are still white […]. For these reasons, my lord Duarte thinks that the black color is not due to the action of the sun, but stems from the nature of sperm” (translated from Pigafetta and Lopes 2002).
Such were the views about skin color variation of the Portuguese trader Duarte Lopes, who narrated his adventures in the Kingdom of Congo to the Italian writer Filipo Pigafetta in 1591. Today, more than 400 years after Lopes’s and many other travelers’ accounts, the major questions about pigmentation variation remain basically the same: what is the genetic basis of pigmentation diversity, and how can the striking correlation between skin color and latitude be explained? Fortunately, while the questions have not changed, our means to answer them have substantially improved. In this article, I will review the implications of recent findings about the genes involved in normal skin color variation for understanding the evolutionary history of skin pigmentation. For a more comprehensive overview of the biology and evolution of different pigmentary traits, including also eye and hair color, the reader is referred, among others, to the excellent works of Rees (2003), Jablonski (2004), McEvoy et al. (2006), Parra (2007), Sturm (2009), Sturm and Duffy (2012), Liu et al. (2013), Deng and Xu (2018), and Quillen et al. (2019).
The Distribution of Human Skin Pigmentation: Patterns and Evolutionary Explanations
Skin color is a highly heritable complex trait that is determined by the type and total amount of melanin produced within melanocyte cells located in the basal layer of the epidermis (Byard 1981; Barsh 2003; Martin et al. 2017). Although the number of melanocytes is basically constant, individuals of varying skin color may differ in size and distribution of the melanin-producing melanosome organelles that transport melanin from melanocytes into surrounding keratinocyte cells (Barsh 2003; Parra 2007). These differences are the major cause for variation in the melanin content (Barsh 2003), which can be objectively quantified by measuring the amount of light reflected by the skin for different wavelengths, or by estimating a melanin index from reflectance of narrowband light in the red spectrum (Shriver and Parra 2000; Parra 2007).
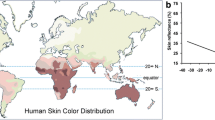
Beyond the impressionistic observations of early travelers, several studies using objective measures of skin pigmentation have shown that skin color varies continuously across different geographic regions, being darker in tropical areas and becoming gradually lighter with increasing distance to the equator (Relethford 1997; Jablonski and Chaplin 2000) (Fig. 1). Due to this striking association, the proportion of skin color variation between geographic regions is unusually high when compared with most genetic markers and other quantitative traits (0.88 vs. 0.15) (Relethford 2002), suggesting that skin color variation was shaped by natural selection in response to environmental factors that are correlated with latitude.
Relationship between distance to the equator (absolute latitude) and skin color measured by the Melanin Index (r = 0.69; p < 0.0001). Data were retrieved from Martin et al. (2017)
Ultraviolet radiation (UVR) has often been considered to be the most important environmental selective pressure for skin color variation, due to its high correlation with latitude and because of observable differences in the risks and benefits of UVR exposure between individuals with lightly and darkly pigmented skin (Jablonski and Chaplin 2010, 2012). Skin pigmentation is more correlated with UVR measured from remote sense technology than with other environmental factors like precipitation, snow, frost-free days, or average temperature (Chaplin 2004). Moreover, UVR levels in autumn were found to be even more correlated with melanin reflectance than latitude, and UVR is strongly predictive of skin reflectance values on a global scale (Jablonski and Chaplin 2000; Chaplin 2004).
The most common harmful effects of UVR that can be counteracted by high concentrations of melanin include sunburn, skin cancer, and photodegradation of micronutrients (Branda and Eaton 1978; Rees and Flanagan 1999; Jablonski and Chaplin 2000; Borradale and Kimlin 2012; Greaves 2014). While it is not clear whether sunburn and skin cancer cause enough differential mortality before reproductive age, photodegradation of folate—a compound of vitamin B—might have a noticeable impact on reproductive ability (Blum 1961; Rees and Flanagan 1999; Jablonski and Chaplin 2000; but see Greaves 2014 for an alternative view on the importance of skin cancer as a selective pressure). This hypothesis seems to be supported by studies showing that folate can be subjected to photolysis at high levels of UVR, and that folate deficiency is associated with neural tube defects and increased incidence of male infertility (Branda and Eaton 1978; Jablonski and Chaplin 2000; Borradale and Kimlin 2012). As UVR declines with increased distance to the equator its damaging effects become less important and the major selective pressure influencing skin color is thought to be the lack of sufficient radiation to stimulate the cutaneous synthesis of vitamin D. In regions far away from the equator, dark skin would be disadvantageous since melanin can block UVR stimulation, increasing the risk for conditions like rickets and osteomalacia that are caused by lack of adequate levels of vitamin D (Loomis 1967; Holick 1995; Jablonski and Chaplin 2000, 2010; Chaplin and Jablonski 2009; but see Robins 2009 and Elias and Williams 2016 for alternative views on the importance of vitamin D for the evolution of light skin color).
According to the framework outlined above, the first UVR-driven shift in skin pigmentation in the human lineage occurred when the white, chimpanzee-like skin of our early hominin ancestors became dark to protect the naked skin from the damaging effects of excess radiation after body hair was lost to enhance thermoregulation in tropical savannas (Wheeler 1985; Jablonski 2004; Jablonski and Chaplin 2017). Consideration of the paleo-ecological and anatomical evidence during the early stages of human evolution has led to the proposal that most thermoregulatory changes, including loss of thick body hair, might have occurred about 2 million years ago, as the earliest members of the genus Homo became mostly savanna dwellers, increasing their levels of activity and exposure to daylight (Klein 1999; Rogers et al. 2004; Jablonski 2006). If this hypothesis is accepted, a 2-million years date can be set as the upper limit for dark skin becoming the ancestral state of later Homo species. However, other estimates based on the divergence times of human and primate lice species suggest that the loss of body hair may be as old as 3.3 million years, raising the possibility of an even earlier change from light to dark pigmentation in the human lineage (Reed et al. 2007; Stoneking 2017). As later Homo groups occupied Eurasia, expanding their range to regions with low UVR, natural selection could have favored genetic variants associated with skin lightening to facilitate vitamin D synthesis (Lalueza-Fox et al. 2007).
In modern humans, the available phenotype information can be combined with current knowledge about major migration routes to derive general models of skin color change (Nielsen et al. 2017). In the simplest scenario, populations migrating out of Africa around 55–65 kya resembled their African ancestors and were dark-skinned. As the ancestors of Europeans and north Asians migrated northwards, they became exposed to the risks of low vitamin D synthesis and light skin was favored by selection. Populations migrating along southern Asia, eventually reaching Australia and Melanesia, remained close to the equator and preserved the ancestral dark-skinned condition or developed novel genetic adaptations as a protection against the damaging effects of UVR (Norton et al. 2007).
Exceptions to these trends can be caused by a number of factors, like insufficient time for selective pressures to act due to recent migration, differences in cultural practices influencing exposure to the sun, availability of vitamin D in the diet, and barriers to gene flow limiting the spread of favorable variants (Quillen et al. 2019). For example, the lighter skin of indigenous populations of the New World compared to peoples from the Old World living at the same latitude is likely to reflect both cultural adaptations and the relatively recent occupation of the Americas by peoples of Asian descent (Jablonski 2006; Jablonski and Chaplin 2017). Cultural practices may also have influenced the evolution of skin pigmentation in the Tibetan plateau, where local populations have lighter pigmentation than predicted by the UVR levels of the region, probably because of the need to wear clothes (Jablonski and Chaplin 2000). Moreover, mate choice based on skin color might have shaped local patterns of skin color variation, altering the expected correlation between melanin content and intensity of UVR (Iliescu et al. 2018).
In spite of these complications, the general framework linking human migrations with natural selection has the advantage of providing broad expectations that can be assessed with the increasingly available information about genes contributing to skin color variation.
The Genetic Basis of Skin Pigmentation
Several candidate genes with a role in human skin color have been identified through studies of model organisms or in humans affected by pigmentation disorders, like albinism. Since not all genes involved in pigmentation pathways determine normal pigmentation variability, some studies additionally used molecular signatures of positive selection as a complementary tool to narrow down the search for genomic regions of interest (McEvoy et al. 2006; Lao et al. 2007; Myles et al. 2007). This approach mostly relies on the assumption that selective events could be modeled as hard sweeps in which new favored alleles rise rapidly in frequency, reducing haplotype diversity, and creating extreme allele frequency differences between populations (Pritchard and Di Rienzo 2010). However, since these methods are phenotype-independent, further analyses are necessary to effectively link candidate genes to phenotypic variation (Jeong and Di Rienzo 2014).
The most common way to show that a gene influences normal skin color diversity is to perform association studies to assess how melanin levels vary between individuals with different genotypes, using other markers to control for the effects of population structure. Association studies can be performed with candidate genes pre-selected on the basis of their known involvement in pigmentation pathways, or use hundreds of thousands of polymorphic markers dispersed across the genome (GWAS—Genome-Wide Association studies) (Quillen et al. 2019). Some of the genes identified by these studies are listed in Table 1. Interestingly, several genes identified by GWAS do not play an obvious role in pigmentation variation and would have remained undetected in candidate gene approaches (e.g., DDB1). These genes raise the possibility that skin color variation may be partially driven by pleiotropic effects. Moreover, many causal variants are located in non-coding regions and are likely to act through regulation of gene expression (e.g., rs12913832 in HERC2 and rs12203592 in IRF4).
The discovery of the role of the SLC24A5 gene in human skin color variation provides a good example of an association approach based on a candidate gene. The gene encodes a cation exchanger with an important function in melanosome biogenesis and melanin synthesis, and was initially found to be involved in pigment variation in zebrafish. Subsequent comparative analyses found that the human SLC24A5 homologue harbored several Single-Nucleotide Polymorphisms (SNPs) with very different allele frequencies between Africans and Europeans, including a non-synonymous polymorphism at rs1426654 (Ala111Thr). Finally, the involvement of SLC24A5 in human skin color variation was validated by association studies showing that rs1426654 genotypes were significantly correlated with melanin levels in admixed Afro-Americans and Afro-Caribbeans (Lamason 2005). Several other association studies, including GWAS, have firmly established the role of SLC24A5 as one of the main loci influencing human skin color variation.
The KITLG gene, which encodes a signaling molecule important for melanocyte migration, illustrates a less straightforward situation. Like SLC24A5, the involvement of KITLG in pigmentation diversity was first suggested by studies in a model organism (the stickleback fish) (Miller et al. 2007). Moreover, allelic variation in potentially regulatory regions of the human homologue gene was found to be significantly associated with skin color in African Americans (Miller et al. 2007). However, this association could not be replicated in a GWAS study of the African European admixed population of Cape Verde (Beleza et al. 2013a). More recently, an association between skin color and KITLG was found to be marginally significant in the southern Africa Nama and ǂKhomani Khoisan, who have lighter skin than their Bantu-speaking neighbors, although no genome-wide significance could be replicated (Martin et al. 2017).
Discrepancies between studies may be due to false-positive associations caused by inefficient correction of population structure, differences in power to detect small effects on the phenotype, or differences in patterns and levels of linkage disequilibrium when the marker SNP is not causal. In addition, epistatic interactions among variants from different loci can modify the effect size of a casual variant mutation in different populations (Quillen et al. 2019). These complications emphasize the need for developing and using standardized methods to validate discovered associations through functional assays (Visser et al. 2012, 2014; Tsetskhladze et al. 2012; Praetorius et al. 2013; Crawford et al. 2017).
Until recently, most genetic variants associated with pigmentation diversity were identified among Europeans or African European admixed populations (Quillen et al. 2019). Studies in admixed populations typically identify genes of high effect displaying elevated allele frequency differences between parental populations, while studies on Europeans focus on the lighter portion of the human skin pigmentation range and detect genes with smaller effects. However, more recent studies have started to correct these biases by focusing on understudied regions like Africa, where the greatest range of pigmentation is observed (Crawford et al. 2017; Martin et al. 2017). These studies are providing an increasingly sharp picture of the genetic basis for skin pigmentation diversity.
In a GWAS study performed in the admixed population of Cape Verde, Beleza et al. (2013a, b) estimated that four loci (SLC24A5, SLC45A2, GMR5-TYR, and APAB2) explain 35% of the skin color variation, suggesting a genetic architecture in which the moderate effects of a few genes are combined with small effects of many genes. A recent study focusing on skin color variation across eastern and southern African populations (Crawford et al. 2017) showed that 29% of pigmentation diversity in these populations can be attributed to variation in four genomic regions (SLC24A5, MFSD12, DDB1/TMEM138, OCA2/HERC2). Still in Africa, Martin et al. (2017) found that only 23% of the phenotypic variance in the Nama and ǂKhomani Khoisan could be explained by the 50 variants most significantly associated with skin pigmentation in the two populations, showing that the complexity of skin color itself may vary between populations. Together, these studies indicate that the genetic architecture of skin color, in spite of being simpler than traits like body height (Wood et al. 2014), is more polygenic than other pigmentary traits like eye color, where a single locus (HERC2/OCA) may explain at least 30% of the observed variation (Sturm et al. 2008; Eiberg et al. 2008; Lloyd-Jones et al. 2017).
The complexity of skin color is also illustrated by the observation that similar phenotypes can be determined by different sets of variants and/or genes in different populations (convergence). For example, the genes SCLC24A5 and SLC45A2 have derived alleles associated with lighter skin that are almost fixed in Europeans but are virtually absent in East Asians, who instead share high frequencies of the ancestral allele with dark-skinned Africans and Australo-Melanesians (McEvoy et al. 2006; Norton et al. 2007). On the other hand, the derived allele of a non-synonymous polymorphism (His615Arg; rs1800414) located in the OCA2 gene is significantly associated with skin lightening in East Asians but is absent in Europeans (Edwards et al. 2010; Yang et al. 2016). More recently, Adhikari et al. (2019) have shown that the derived allele in a non-synonymous SNP (Tyr182His; rs2240751) in the MFSD12 gene is associated with lighter skin pigmentation in a large sample of Latin Americans with high Native American ancestry. Although other variants of MFSD12 had been previously related with skin color variation in Africans (see above; Table 1), the derived allele at rs2240751 was found to be common only in East Asians and Latin Americans, suggesting that it started to rise in frequency in East Asia and was carried into the Americas by migration (Adhikari et al. 2019). These findings suggest that, at least in part, different genes and variants are responsible for decreased melanin content in Europe and East Asia, emphasizing the importance of studying non-European populations to understand the full complexity of skin color variation.
The complexity of skin color also has implications for the accuracy of models predicting phenotypes using a small number of DNA variants in forensic contexts or in ancient DNA studies (Lazaridis et al. 2014; Liu et al. 2015; Walsh et al. 2017; Brace et al. 2019). In fact, while several predictive tests have been developed to assign categorical pigmentation phenotypes with increasing accuracy, SNPs identified in one particular continental region may have no power to predict phenotypes in other regions (Martin et al. 2017; Walsh et al. 2017; Quillen et al. 2019).
Evolution of Skin Pigmentation
The identification of key genes underlying pigmentation diversity made it possible to assess the evolutionary history of skin pigmentation in modern humans by studying the geographical distribution, frequency and age of different variants, and by investigating molecular signatures of natural selection. In this context, it is important to distinguish ancestral and derived skin color phenotypes from ancestral and derived alleles in skin pigmentation loci. As mentioned above, early Homo ancestors are believed to have become more darkly pigmented than the chimpanzee as a protection against the damaging effects of UVR after body hair was lost. According to this hypothesis, dark skin is a derived trait and derived alleles are expected to be associated with dark skin at loci that were involved in increasing melanin content relative to the chimpanzee. However, derived alleles can also be associated with lower melanin content at loci that were not involved in human/chimpanzee skin color differentiation. This is the case of several alleles associated with lighter skin in modern human populations that migrated out of Africa.
One of the first genes used to infer the evolutionary history of skin pigmentation was MC1R (Table 1). Based on the contrast between lack of sequence variation in Africans and abundance of non-synonymous variants in Europeans, Harding et al. (2000) concluded that MC1R was under strong purifying selection for maintaining dark skin in Africa. This observation has often led to the view that most genes involved in skin color diversity would be under functional constraint in dark-skinned populations (Quillen et al. 2019). However, recent studies focusing on skin color variation in African populations provided evidence against this generalization (Crawford et al. 2017; Martin et al. 2017). By analyzing skin color diversity in populations from eastern and southern Africa, Crawford et al. (2017) found that none of the eight SNPs most strongly associated with pigmentation variation displayed allelic fixation. Even among eastern African Nilo-Saharan speakers, who display the darkest pigmentation, the average frequency of alleles associated with light skin is higher than 25% (Fig. 2a). Moreover, melanin content is positively correlated (r = 0.96; p = 0.003) with average heterozygosity at those eight SNPS in a small sample of populations encompassing regions with very different levels of UVR, presented here for illustration purposes (Fig. 2b–d). These findings are consistent with the observation that dark-skinned populations have wider distributions of melanin content than light-skinned populations (Martin et al. 2017), and suggest that selection for dark skin involved small shifts in allele frequency from standing variation as expected in polygenic adaptation (Pritchard and Di Rienzo 2010; Pritchard et al. 2010; Berg and Coop 2014; Field et al. 2016). A clear indication of the role of standing variation is provided by estimated allele ages. According to Crawford et al. (2017), seven out of the eight SNPs most involved in skin color diversity in their sample have derived allele ages ranging from 250,000 to 1,200,000 years, showing that allelic variation in these polymorphisms originated much before the emergence of modern humans.
a Average frequencies of alleles associated with light skin color in regions with different levels of UVR, calculated from eight SNPs most strongly associated with pigmentation variation in African populations: rs1800404, rs4932620, rs7948623, rs10424065, rs6510760, rs6497271, rs1426654, rs11230664. b Average heterozygosities calculated from the same SNPs and geographical regions displayed in a. c Average Melanin Indices for the regions displayed in a and b. d Correlation between average heterozygosity and Melanin Index (r = 0.96; p = 0.003). Data were retrieved from Martin et al. (2017) and Crawford et al. (2017)
The study of African populations has also important implications for a better understanding of global variation in skin color. So far, several genes involved in skin color lightening in Eurasians were found to bear strong signals of recent positive selection acting on new alleles (hard sweeps) (Voight et al. 2005; McEvoy et al. 2006; Lao et al. 2007; Myles et al. 2007; Pickrell et al. 2009). In genes like KITLG (rs642742), the selective sweep probably started around 30,000 ago (Beleza et al. 2013b; Chen et al. 2015; Smith et al. 2018; Yang et al. 2018), before the divergence of Europeans and East Asians and after modern humans left Africa (Nielsen et al. 2017). Consequently, high allele frequency differences between Africans and non-Africans are now observed (Miller et al. 2007; Pickrell et al. 2009). Date estimates for the onset of selection on other genes, like OCA2 (rs1800414), SLC24A5 (rs1426654), SLC45A2 (rs16891982), and MFSD12 (rs2240751) fall within the past 11,000–19,000 years, much after the first migrations into Eurasia (Beleza et al. 2013b; Yang et al. 2016), though older coalescent ages around a 28–31,000 years date for SLC24A5 have been recently calculated (Basu Mallick et al. 2013; Crawford et al. 2017). Moreover, as alleles associated with light pigmentation in these genes have non-overlapping distributions in Eurasia (see above), it is likely that evolution of light skin pigmentation in the two regions was partially independent and still ongoing until recently (McEvoy et al. 2006; Norton et al. 2007). This scenario is further supported by genotype data from ancient genomes showing that light skin variants in SLC24A5 and SLC45A2 genes were not fixed in western Europe as recently as 8000 years ago, and that dark and intermediate skin colors might have been common by that time (Wilde et al. 2014; Mathieson et al. 2015; Brace et al. 2019). Although these findings suggest that skin lightening in Eurasia mostly evolved through strong selection of relatively recent mutations, it is now clear that old variants associated with lighter skin in Africa have also played an important role in non-African populations. These variants reach high frequencies among Europeans and East Asians, indicating that migration out of Africa was accompanied by allele frequency shifts from African standing variation (Fig. 1 from Crawford et al. 2017; Fig. 2a). Furthermore, several old alleles associated with light skin have very similar frequencies in Europeans and southern African Khoisan, suggesting that, as expected in polygenic adaptation (Hancock et al. 2010; Pritchard and Di Rienzo 2010), parallel frequency shifts from standing variation occurred in both populations to respond to selective pressures associated with absolute distance from the equator (Fig. 3a). A similar, though less pronounced, parallelism is found in non-Khoisan Africans and Australo-Melanesians, who share higher frequencies of variants associated with darker skin, probably because these variants were not lost by populations that remained close to the equator after leaving Africa (Crawford et al. 2017).
Pairwise absolute allele frequency differences in eight SNPs most strongly associated with pigmentation variation in African populations (as in Fig. 2). a Differences between Khoisan and Europeans. b Differences between East Africans and Europeans. The dashed lines show the average allele frequency differences. Data were retrieved from Crawford et al. (2017)
Taken together, these results suggest that the shift to light skin in Eurasia involved a combination of polygenic adaptation with hard selective sweeps. Since adaptation through hard sweeps is expected to predominate when genetic variance is reduced and populations are still far from the phenotypic optimum (Chevin and Hospital 2008), it is conceivable that hard selective sweeps occurred because standing variation in skin color genes was lost in Eurasian populations. Moreover, it is possible that this relative simplification of genetic architecture was due to the joint effects of selection favoring “African” alleles associated with light skin and bottlenecks occurring during the out of Africa migration, as expected from the connection between selection and demographic processes that is typically observed in human populations (Coop et al. 2009).
A further example for the combination of migration and selection is provided by the distribution of the SLC24A5 allele associated with light skin pigmentation. Besides being virtually fixed in Europe, this variant is also associated with lighter skin color in Africans, reaching high frequencies (28–50%) in Afro-Asiatic speakers from East Africa, as well as in several Khoisan peoples from southern Africa, especially among the Nama and the ǂKhomani (33–53%) (Pagani et al. 2012; Crawford et al. 2017; Martin et al. 2017; Lin et al. 2018). Taking into account the population history of these regions, it is likely that the SLC24A5 variant entered East Africa from the Middle East around 3000–9000 years ago (Pagani et al. 2012; Crawford et al. 2017) and was then carried by migrating eastern African pastoralists into southern Africa around 2000 years ago, where it was favored by natural selection (Lin et al. 2018). The latter dispersion is remarkably similar to the spread of the − 14,010*C lactase-persistence allele from eastern into southern Africa (Breton et al. 2014; Macholdt et al. 2014; Pinto et al. 2016, Lin et al. 2018).
Conclusion
The remarkable progresses in the understanding of normal skin color variation include not only the identification of an increasing number of genes associated with pigmentation diversity, but also a better perception of the complexity of this trait. It seems now clear that the genetic basis of skin color is less simple than previously thought and that the geographic variation in skin pigmentation is not exclusively driven by hard selective sweeps in a few key genes. The recent increase in the number of populations studied for pigmentation variation, including African groups from a wide range of geographic origins, has revealed that the complexity of skin color can vary across populations and that the evolutionary history of pigmentation involved adaptations achieved by the concerted action of different types of selection (Fig. 4). While this increasing complexity does not challenge the general view that human migration towards regions of lower UVR levels favored lighter skin colors, future studies on fitness differences between individuals with different pigmentation will be needed to investigate the additional impact of epistasis, pleiotropic effects, and cultural practices in shaping the geographical patterning of skin color variation.
Allele frequencies at 5 genes/SNPs associated with skin color variation exemplifying the variety of selection patterns underlying skin pigmentation differences across populations: parallel shifts in allele frequencies from standing variation (HERC2/rs6497271); selective sweeps in proto-Eurasians (KITLG/rs642742); selection on standing variation in proto-Eurasians (DDB1/rs11230664); selective sweeps in Europeans (SLC24A5/rs1426654), and selective sweeps in Asians (OCA2/rs1800414). a = Botswana Khoisan; b = West Africans; c = East Africans; d = Australo-Melanesians; e = East Asians; f = Europeans. Alleles associated with darker skin are shown in black
References
Abe Y, Tamiya G, Nakamura T et al (2013) Association of melanogenesis genes with skin color variation among Japanese females. J Dermatol Sci 69:167–172. https://doi.org/10.1016/j.jdermsci.2012.10.016
Adhikari K, Mendoza-Revilla Sohail A et al (2019) A GWAS in Latin Americans highlights the convergent evolution of lighter skin pigementation in Eurasia. Nat Commun 10:358. https://doi.org/10.1038/s414667-018-08147-0
Barsh GS (2003) What controls variation in human skin color? PLoS Biol 1:e27. https://doi.org/10.1371/journal.pbio.0000091
Basu Mallick C, Iliescu FM, Möls M et al (2013) The light skin allele of SLC24A5 in South Asians and Europeans shares identity by descent. PLoS Genet 9:e1003912. https://doi.org/10.1371/journal.pgen.1003912
Beleza S, Johnson NA, Candille SI et al (2013a) Genetic architecture of skin and eye color in an African-European admixed population. PLoS Genet 9:e1003372. https://doi.org/10.1371/journal.pgen.1003372
Beleza S, Santos AM, McEvoy B et al (2013b) The timing of pigmentation lightening in Europeans. Mol Biol Evol 30:24–35. https://doi.org/10.1093/molbev/mss207
Berg JJ, Coop G (2014) A population genetic signal of polygenic adaptation. PLoS Genet 10:e1004412. https://doi.org/10.1371/journal.pgen.1004412
Blum HF (1961) Does the melanin pigment of human skin have adaptive value? An essay in human ecology and the evolution of race. Q Rev Biol 36:50–63. https://doi.org/10.1086/403275
Bonilla C, Boxill LA, Mc Donald SA et al (2005) The 8818G allele of the agouti signaling protein (ASIP) gene is ancestral and is associated with darker skin color in African Americans. Hum Genet. https://doi.org/10.1007/s00439-004-1251-2
Borradale DC, Kimlin MG (2012) Folate degradation due to ultraviolet radiation: possible implications for human health and nutrition. Nutr Rev. https://doi.org/10.1111/j.1753-4887.2012.00485.x
Brace S, Diekmann Y, Booth TJ et al (2019) Ancient genomes indicate population replacement in Early Neolithic Britain. Nat Ecol Evol 3:765–771. https://doi.org/10.1038/s41559-019-0871-9
Branda R, Eaton J (1978) Skin color and nutrient photolysis: an evolutionary hypothesis. Science 201:625–626. https://doi.org/10.1126/science.675247
Breton G, Schlebusch CM, Lombard M et al (2014) Lactase persistence alleles reveal partial east African ancestry of southern African Khoe pastoralists. Curr Biol 24:852–858. https://doi.org/10.1016/j.cub.2014.02.041
Byard PJ (1981) Quantitative genetics of human skin color. Am J Phys Anthropol 24:123–137. https://doi.org/10.1002/ajpa.1330240506
Chaplin G (2004) Geographic distribution of environmental factors influencing human skin coloration. Am J Phys Anthropol 125:292–302. https://doi.org/10.1002/ajpa.10263
Chaplin G, Jablonski NG (2009) Vitamin D and the evolution of human depigmentation. Am J Phys Anthropol 139:451–461. https://doi.org/10.1002/ajpa.21079
Chen H, Hey J, Slatkin M (2015) A hidden Markov model for investigating recent positive selection through haplotype structure. Theor Popul Biol 99:18–30. https://doi.org/10.1016/j.tpb.2014.11.001
Chevin LM, Hospital F (2008) Selective sweep at a quantitative trait locus in the presence of background genetic variation. Genetics 180:1645–1660. https://doi.org/10.1534/genetics.108.093351
Cook AL, Chen W, Thurber AE et al (2009) Analysis of cultured human melanocytes based on polymorphisms within the SLC45A2/MATP, SLC24A5/NCKX5, and OCA2/P loci. J Invest Dermatol 129:392–405. https://doi.org/10.1038/jid.2008.211
Coop G, Pickrell JK, Novembre J et al (2009) The role of geography in human adaptation. PLoS Genet 5:e1000500. https://doi.org/10.1371/journal.pgen.1000500
Crawford NG, Kelly DE, Hansen MEB et al (2017) Loci associated with skin pigmentation identified in African populations. Science 358:887. https://doi.org/10.1126/science.aan8433
Deng L, Xu S (2018) Adaptation of human skin color in various populations. Hereditas 155:1. https://doi.org/10.1186/s41065-017-0036-2
Edwards M, Bigham A, Tan J et al (2010) Association of the OCA2 polymorphism His615Arg with melanin content in East Asian populations: further evidence of convergent evolution of skin pigmentation. PLoS Genet 6:e1000867. https://doi.org/10.1371/journal.pgen.1000867
Eiberg H, Troelsen J, Nielsen M et al (2008) Blue eye color in humans may be caused by a perfectly associated founder mutation in a regulatory element located within the HERC2 gene inhibiting OCA2 expression. Hum Genet 123:177–187. https://doi.org/10.1007/s00439-007-0460-x
Elias PM, Williams ML (2016) Basis for the gain and subsequent dilution of epidermal pigmentation during human evolution: the barrier and metabolic conservation hypotheses revisited. Am J Phys Anthropol 161:189–207. https://doi.org/10.1002/ajpa.23030
Field Y, Boyle EA, Telis N et al (2016) Detection of human adaptation during the past 2000 years. Science 354:760–764
Graf J, Hodgson R, van Daal A (2005) Single nucleotide polymorphisms in theMATP gene are associated with normal human pigmentation variation. Hum Mutat 25:278–284. https://doi.org/10.1002/humu.20143
Greaves M (2014) Was skin cancer a selective force for black pigmentation in early hominin evolution? Proc R Soc B Biol Sci 281:20132955. https://doi.org/10.1098/rspb.2013.2955
Han J, Kraft P, Nan H et al (2008) A genome-wide association study identifies novel alleles associated with hair color and skin pigmentation. PLoS Genet 4:e1000074. https://doi.org/10.1371/journal.pgen.1000074
Hancock AM, Witonsky DB, Ehler E et al (2010) Human adaptations to diet, subsistence, and ecoregion are due to subtle shifts in allele frequency. Proc Natl Acad Sci 107:8924–8930. https://doi.org/10.1073/pnas.0914625107
Harding RM, Healy E, Ray AJ et al (2000) Evidence for variable selective pressures at MC1R. Am J Hum Genet 66:1351–1361. https://doi.org/10.1086/302863
Hernandez-Pacheco N, Flores C, Alonso S et al (2017) Identification of a novel locus associated with skin colour in African-admixed populations. Sci Rep 7:44548. https://doi.org/10.1038/srep44548
Holick MF (1995) Environmental factors that influence the cutaneous production of vitamin D. Am J Clin Nutr 61:638S–645S. https://doi.org/10.1093/ajcn/61.3.638S
Iliescu FM, Chaplin G, Rai N et al (2018) The influences of genes, the environment, and social factors on the evolution of skin color diversity in India. Am J Hum Biol 30:1–15. https://doi.org/10.1002/ajhb.23170
Jablonski NG (2004) The evolution of human skin and skin color. Annu Rev Anthropol 33:585–623. https://doi.org/10.1146/annurev.anthro.33.070203.143955
Jablonski NG (2006) Skin: a natural history. University of California Press, Berkeley
Jablonski NG, Chaplin G (2000) The evolution of human skin coloration. J Hum Evol 39:57–106. https://doi.org/10.1006/jhev.2000.0403
Jablonski NG, Chaplin G (2010) Human skin pigmentation as an adaptation to UV radiation. Proc Natl Acad Sci 107:8962–8968. https://doi.org/10.1073/pnas.0914628107
Jablonski NG, Chaplin G (2012) Human skin pigmentation, migration and disease susceptibility. Philos Trans R Soc B Biol Sci 367:785–792. https://doi.org/10.1098/rstb.2011.0308
Jablonski NG, Chaplin G (2017) The colours of humanity: the evolution of pigmentation in the human lineage. Philos Trans R Soc B Biol Sci 372:20160349. https://doi.org/10.1098/rstb.2016.0349
Jacobs LC, Wollstein A, Lao O et al (2013) Comprehensive candidate gene study highlights UGT1A and BNC2 as new genes determining continuous skin color variation in Europeans. Hum Genet 132:147–158. https://doi.org/10.1007/s00439-012-1232-9
Jeong C, Di Rienzo A (2014) Adaptations to local environments in modern human populations. Curr Opin Genet Dev 29:1–8. https://doi.org/10.1016/j.gde.2014.06.011
Klein RG (1999) The human career: human biological and cultural origins. University of Chicago Press, Chicago
Lalueza-Fox C, Rompler H, Caramelli D et al (2007) A melanocortin 1 receptor allele suggests varying pigmentation among neanderthals. Science 318:1453–1455. https://doi.org/10.1126/science.1147417
Lamason RL (2005) SLC24A5, a putative cation exchanger, affects pigmentation in zebrafish and humans. Science 310:1782–1786. https://doi.org/10.1126/science.1116238
Lao O, de Gruijter JM, van Duijn K et al (2007) Signatures of positive selection in genes associated with human skin pigmentation as revealed from analyses of single nucleotide polymorphisms. Ann Hum Genet 71:354–369. https://doi.org/10.1111/j.1469-1809.2006.00341.x
Lazaridis I, Patterson N, Mittnik A et al (2014) Ancient human genomes suggest three ancestral populations for present-day Europeans. Nature 513:409–413. https://doi.org/10.1038/nature13673
Lin M, Siford RL, Martin AR et al (2018) Rapid evolution of a skin-lightening allele in southern African KhoeSan. Proc Natl Acad Sci 115:13324–13329. https://doi.org/10.1073/pnas.1801948115
Liu F, Wen B, Kayser M (2013) Colorful DNA polymorphisms in humans. Semin Cell Dev Biol 24:562–575. https://doi.org/10.1016/j.semcdb.2013.03.013
Liu F, Visser M, Duffy DL et al (2015) Genetics of skin color variation in Europeans: genome-wide association studies with functional follow-up. Hum Genet 134:823–835. https://doi.org/10.1007/s00439-015-1559-0
Lloyd-Jones LR, Robinson MR, Moser G et al (2017) Inference on the genetic basis of eye and skin color in an admixed population via bayesian linear mixed models. Genetics 206:1113–1126. https://doi.org/10.1534/genetics.116.193383/-/DC1.1
Loomis WF (1967) Skin-pigment regulation of Vitamin-D biosynthesis in man: variation in solar ultraviolet at different latitudes may have caused racial differentiation in man. Science 157:501–506. https://doi.org/10.1126/science.157.3788.501
Macholdt E, Lede V, Barbieri C et al (2014) Tracing pastoralist migrations to southern Africa with lactase persistence alleles. Curr Biol 24:875–879. https://doi.org/10.1016/j.cub.2014.03.027
Martin AR, Lin M, Granka JM et al (2017) An unexpectedly complex architecture for skin pigmentation in Africans. Cell 171:1340–1353. https://doi.org/10.1016/j.cell.2017.11.015
Mathieson I, Lazaridis I, Rohland N et al (2015) Genome-wide patterns of selection in 230 ancient Eurasians. Nature 528:499–503. https://doi.org/10.1038/nature16152
McEvoy B, Beleza S, Shriver MD (2006) The genetic architecture of normal variation in human pigmentation: an evolutionary perspective and model. Hum Mol Genet 15:R176–R181. https://doi.org/10.1093/hmg/ddl217
Miller CT, Beleza S, Pollen AA et al (2007) cis-Regulatory changes in kit ligand expression and parallel evolution of pigmentation in sticklebacks and humans. Cell 131:1179–1189. https://doi.org/10.1016/j.cell.2007.10.055
Myles S, Somel M, Tang K et al (2007) Identifying genes underlying skin pigmentation differences among human populations. Hum Genet 120:613–621. https://doi.org/10.1007/s00439-006-0256-4
Nielsen R, Akey JM, Jakobsson M et al (2017) Tracing the peopling of the world through genomics. Nature 541:302–310. https://doi.org/10.1038/nature21347
Norton HL, Kittles RA, Parra E et al (2007) Genetic evidence for the convergent evolution of light skin in Europeans and east Asians. Mol Biol Evol 24:710–722. https://doi.org/10.1093/molbev/msl203
Pagani L, Kivisild T, Tarekegn A et al (2012) Ethiopian genetic diversity reveals linguistic stratification and complex influences on the Ethiopian gene pool. Am J Hum Genet 91:83–96. https://doi.org/10.1016/j.ajhg.2012.05.015
Parra EJ (2007) Human pigmentation variation: evolution, genetic basis, and implications for public health. Yearb Phys Anthropol 50:85–105. https://doi.org/10.1002/ajpa
Pickrell JK, Coop G, Novembre J et al (2009) Signals of recent positive selection in a worldwide sample of human populations. Genome Res 19:826–837. https://doi.org/10.1101/gr.087577.108
Pigafetta F, Lopes D (2002) Le Royaume de Congo & les contrées environnantes (1591); traduite & annotée par Willy Bal. Chandeigne/Unesco, Paris
Pinto JC, Oliveira S, Teixeira S et al (2016) Food and pathogen adaptations in the Angolan Namib desert: tracing the spread of lactase persistence and human African trypanosomiasis resistance into southwestern Africa. Am J Phys Anthropol 161:436–447. https://doi.org/10.1002/ajpa.23042
Praetorius C, Grill C, Stacey SN et al (2013) A polymorphism in IRF4 affects human pigmentation through a tyrosinase-dependent MITF/TFAP2A pathway. Cell 155:1022–1033. https://doi.org/10.1016/j.cell.2013.10.022
Pritchard JK, Di Rienzo A (2010) Adaptation—not by sweeps alone. Nat Rev Genet 11:665–667. https://doi.org/10.1038/nrg2880
Pritchard JK, Pickrell JK, Coop G (2010) The genetics of human adaptation: hard sweeps, soft sweeps, and polygenic adaptation. Curr Biol 20:R208–R215. https://doi.org/10.1016/j.cub.2009.11.055
Quillen EE, Bauchet M, Bigham AW et al (2012) OPRM1 and EGFR contribute to skin pigmentation differences between Indigenous Americans and Europeans. Hum Genet 131:1073–1080. https://doi.org/10.1007/s00439-011-1135-1
Quillen EE, Norton HL, Parra EJ et al (2019) Shades of complexity: new perspectives on the evolution and genetic architecture of human skin. Am J Phys Anthropol 168:4–26. https://doi.org/10.1002/ajpa.23737
Reed DL, Light JE, Allen JM, Kirchman JJ (2007) Pair of lice lost or parasites regained: the evolutionary history of anthropoid primate lice. BMC Biol 5:1–11. https://doi.org/10.1186/1741-7007-5-7
Rees JL (2003) Genetics of hair and skin color. Annu Rev Genet. https://doi.org/10.1146/annurev.genet.37.110801.143233
Rees JL, Flanagan N (1999) Pigmentation, melanocortins and red hair. QJM 92:125–131. https://doi.org/10.1093/qjmed/92.3.125
Relethford JH (1997) Hemispheric difference in human skin color. Am J Phys Anthropol 104:449–457
Relethford JH (2002) Apportionment of global human genetic diversity based on craniometrics and skin color. Am J Phys Anthropol 118:393–398. https://doi.org/10.1002/ajpa.10079
Robins AH (2009) The evolution of light skin color: role of vitamin D disputed. Am J Phys Anthropol 139:447–450. https://doi.org/10.1002/ajpa.21077
Rogers AR, Iltis D, Wooding S (2004) Genetic variation at the MCIR locus and the time since loss of human body hair. Curr Anthropol 45:105–108
Shriver MD, Parra EJ (2000) Comparison of narrow-band reflectance spectroscopy and tristimulus colorimetry for measurements of skin and hair color in persons of different biological ancestry. Am J Phys Anthropol 112:17–27
Shriver M, Parra E, Dios S et al (2003) Skin pigmentation, biogeographical ancestry and admixture mapping. Hum Genet 112:387–399. https://doi.org/10.1007/s00439-002-0896-y
Smith J, Coop G, Stephens M, Novembre J (2018) Estimating time to the common ancestor for a beneficial allele. Mol Biol Evol 35:1003–1017. https://doi.org/10.1093/molbev/msy006
Stoneking M (2017) An introduction to molecular anthropology. Wiley, New York
Sturm RA (2009) Molecular genetics of human pigmentation diversity. Hum Mol, Genet
Sturm RA, Duffy DL (2012) Human pigmentation genes under environmental selection. Genome Biol 13:248. https://doi.org/10.1186/gb-2012-13-9-248
Sturm RA, Duffy DL, Zhao ZZ et al (2008) A single SNP in an evolutionary conserved region within intron 86 of the HERC2 gene determines human blue-brown eye color. Am J Hum Genet 82:424–431. https://doi.org/10.1016/j.ajhg.2007.11.005
Tsetskhladze ZR, Canfield VA, Ang KC et al (2012) Functional assessment of human coding mutations affecting skin pigmentation using zebrafish. PLoS ONE 7:e47398. https://doi.org/10.1371/journal.pone.0047398
Visser M, Kayser M, Palstra R-J (2012) HERC2 rs12913832 modulates human pigmentation by attenuating chromatin-loop formation between a long-range enhancer and the OCA2 promoter. Genome Res 22:446–455. https://doi.org/10.1101/gr.128652.111
Visser M, Palstra R-J, Kayser M (2014) Human skin color is influenced by an intergenic DNA polymorphism regulating transcription of the nearby BNC2 pigmentation gene. Hum Mol Genet 23:5750–5762. https://doi.org/10.1093/hmg/ddu289
Voight BF, Adams AM, Frisse LA et al (2005) Interrogating multiple aspects of variation in a full resequencing data set to infer human population size changes. Proc Natl Acad Sci. https://doi.org/10.1073/pnas.0507325102
Walsh S, Chaitanya L, Breslin K et al (2017) Global skin colour prediction from DNA. Hum Genet 136:847–863. https://doi.org/10.1007/s00439-017-1808-5
Wheeler PE (1985) The loss of functional body hair in man: the influence of thermal environment, body form and bipedality. J Hum Evol 14:23–28. https://doi.org/10.1016/S0047-2484(85)80091-9
Wilde S, Timpson A, Kirsanow K et al (2014) Direct evidence for positive selection of skin, hair, and eye pigmentation in Europeans during the last 5,000 y. Proc Natl Acad Sci 111:4832–4837. https://doi.org/10.1073/pnas.1316513111
Wood AR, Esko T, Yang J et al (2014) Defining the role of common variation in the genomic and biological architecture of adult human height. Nat Genet 46:1173–1186
Yang Z, Zhong H, Chen J et al (2016) A genetic mechanism for convergent skin lightening during recent human evolution. Mol Biol Evol 33:1177–1187. https://doi.org/10.1093/molbev/msw003
Yang Z, Shi H, Ma P et al (2018) Darwinian positive selection on the pleiotropic effects of KITLG explain skin pigmentation and winter temperature adaptation in eurasians. Mol Biol Evol 35:2272–2283. https://doi.org/10.1093/molbev/msy136
Acknowledgements
I thank Anne-Maria Fehn for her comments and suggestions, and for her help in compiling the bibliography and preparing the figures. I would like also to thank two anonymous reviewers, whose comments helped to improve the initial version of this article. My own research in skin color was supported by the Portuguese Foundation for Science and Technology (FCT; PTDC/BIA-BDE/64044/2006).
Author information
Authors and Affiliations
Corresponding author
Additional information
Handling Editor: Konstantinos Voskarides.
Rights and permissions
About this article
Cite this article
Rocha, J. The Evolutionary History of Human Skin Pigmentation. J Mol Evol 88, 77–87 (2020). https://doi.org/10.1007/s00239-019-09902-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-019-09902-7