Abstract
Temperature plays a dominating role in protein structure and function, and life has evolved myriad strategies to adapt proteins to environmental thermal stress. Cellular systems can utilize kosmotropic osmolytes, the products of complex biochemical pathways, to act as chemical chaperones. These extrinsic molecules, e.g., trehalose, alter local water structure to modulate the strength of the hydrophobic effect and increase protein stability. In contrast, simpler genetic systems must rely on intrinsic mutation to affect protein stability. In naturally occurring microvirid bacteriophages of the subfamily Bullavirinae, capsid stability is randomly distributed across the phylogeny, suggesting it is not phylogenetically linked and could be altered through adaptive mutation. We hypothesized that these phages could utilize an adaptive mechanism that mimics the stabilizing effects of the kosmotrope trehalose through mutation. Kinetic stability of wild-type ID8, a relative of ΦX174, displays a saturable response to trehalose. Thermal adaptation mutations in ID8 improve capsid stability and reduce responsiveness to trehalose suggesting the mutations move stability closer to the kosmotropic saturation point, mimicking the kosmotropic effect of trehalose. These mutations localize to and modulate the hydrophobicity of a cavern formation at the interface of phage coat and spike proteins—an evolutionary spandrel. Across a series of genetically distinct phages, responsiveness to trehalose correlates positively with cavern hydrophobicity suggesting that the level of hydrophobicity of the cavern may provide a biophysical gating mechanism constraining or permitting adaptation in a lineage-specific manner. Our results demonstrate that a single mutation can exploit pre-existing, non-adaptive structural features to mimic the adaptive effects of complex biochemical pathways.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Organisms must maintain biochemical function in the face of a variety of environmental stressors such as temperature, pressure, and osmotic stress. Protein stability can be modulated intrinsically via mutation and/or by extrinsic factors termed chemical chaperones (Somero 2003). All domains of cellular life utilize these organic osmolyte chaperone systems as counteracting solutes to preserve protein stability (Yancey et al. 1982; Yancey 2005; Bolen and Baskakov 2001). For example, the kosmotrope trehalose is a multifunctional molecule with wide distribution across cellular life (Elbein et al. 2003). In the yeast Saccharomyces cerevisiae, where trehalose is a glucose storage molecule and acts as an in vivo kosmotropic protein stabilizer (Singer and Lindquist 1998; Elbein et al. 2003), endogenous production of trehalose is accomplished by the trehalose synthase protein complex—four genes encoding a trehalose synthase, a trehalose phosphatase, and two regulatory proteins (Francois and Parrou 2001). Trehalose has been shown to stabilize a variety of protein systems in vitro (Xie and Timasheff 1997; Kaushik and Bhat 2003).
Kosmotropic (from the Greek kosmos meaning order) agents increase the ordering of water molecules (Bruździak et al. 2013; Miyawaki et al. 2014; Ferreira et al. 2017) strengthening the hydrophobic effect (Timasheff 1998; Kaushik and Bhat 1998). Solvent water structure (Cheng and Rossky 1998) and dynamics (Mattos 2002) govern protein structure and function (Levy and Onuchic 2006). Water structure—i.e., localized density or surface tension surrounding a solute molecule (Hummer et al. 2000; Dill and Truskett 2005)—dictates the strength of the hydrophobic effect (Sharp and Madan 1997) which mediates protein folding (Dill 1990) and stability (Pace et al. 2011). Chemical chaperone systems like trehalose are the product of complex biosynthetic pathways (Bell et al. 1998) and are restricted to relatively complex organisms. It is not clear how, or if, simpler genetic systems, like microvirid bacteriophages that cannot produce osmolytes nor utilize host osmolytes for survival after lysis, could have access to a kosmotropic mechanism of adaptation as these systems must rely on intrinsic stabilization through mutation.
Laboratory adaptation of the microvirid bacteriophage ID8, a close relative of the model system ΦX174 (subfamily: Bullavirinae), to high heat stress produced stability mutations that localized to a cavern-like structure at the interface of coat (F protein) and spike (G protein) pentamers (Fig. 1a, b; McGee et al. 2014, 2016). Heat stress was applied via “heat-shock”—incubation of isolated phage at 80 °C for a set time interval. The response to heat-shock, death rate, was measured as the time-weighted, logarithmic decrease in phage titer. The heat-shock was used to test the adaptive capacity of a protein system and elucidate the underlying biophysical mechanisms of thermal adaptation. The overall capsid structure, including the interfacial caverns, is highly conserved across the Bullavirinae. Microvirid survival outside of host depends on the stability of the protein–protein interactions between coat and spike proteins (Lee et al. 2011; McGee et al. 2014). Using molecular dynamics simulation, McGee et al (2014) quantified the contribution of two ancestral residues (R38 and R168 of the spike protein) to the intrinsic binding constant between capsid subunits. The ancestral residues contribute very little to capsid stability in the wild-type phage, but mutation of these residues from polar, charged arginine to the more hydrophobic cysteine, improves capsid stability through a large, favorable contribution to the interaction-free energy between spike and coat proteins. The fact that multiple mutations arose in parallel (including a Thr to Ala substitution at position 65 of the spike protein), are localized to the interfacial cavern (Fig. 1c), and show similar changes in physicochemical properties suggests that this cavern location might be primed for adaptive mutations.
ID8 heat-stability mutations localize to a cavern-like structure between coat and spike proteins on the mature phage capsid. a The icosahedral ID8 capsid contains 60 copies each of the major coat protein F (blue), and the spike protein G (red). b Coat (tan, bottom) and spike (cyan, top) monomers are arranged in pentamers forming the 9S and 6S particles, respectively. The 9S and 6S particles combine to form the 12S particle. Heat-shock mutations at R38 and R168 (green) were shown to increase capsid stability through increased affinity between coat and spike proteins (McGee et al. 2014). The box indicates the area detailed in c. c The interface of two coat and two spike proteins forms a cavern with a high frequency of heat-shock mutations that alter the hydrophobicity of the cavern environment. d This change in hydrophobicity (GRAVY score) shows a strong correlation with heat-induced death rates [death rates from (McGee et al. 2016)]. Correlation was determined by Pearson Product Moment. (Color figure online)
The adaptive effect of the ID8 cavern mutations may be mediated by alterations in the hydrophobic effect, similar to kosmotropic agents. We hypothesized that these mutations modulate capsid thermal stability in a manner similar to the kosmotrope trehalose. We utilized structural modeling and phylogenetics of the Bullavirinae to put cavern hydrophobicity and death rates of naturally occurring phages in a phylogenetic context to determine the link between cavern genetic variation and capsid stability. We determined the kinetics and energetics of capsid stability of wild-type ID8 and the two heat-shock mutations in the presence of varying concentrations of trehalose to establish a mechanism of the intrinsic kosmotropic response. We measured capsid stability in other naturally occurring phages to determine if the kosmotropic response is available to other genotypes besides ID8 and what role sequence variation across the Bullavirinae plays in adaptive potentials. Our results suggest a thermal adaptation mechanism where benefits of an extrinsic kosmotropic response to environmental thermal stress, typically restricted to cellular organisms capable of complex biochemical synthesis, are available to simple genetic systems through a single mutation.
Results and Discussion
Capsid Stability in the Bullavirinae
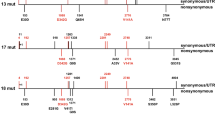
Enterobacteriophages of the subfamily Bullavirinae, typified by the model strain ΦX174, consist of a capsid (Fig. 1a) surrounding a single-stranded genome encoding 11 genes. We estimated a Bayesian phylogeny of the Bullavirinae subfamily based on sequences of the F gene, which codes for the major coat protein, from 70 phages (Fig. 2; Supplementary Dataset 1). The phages were mainly composed of wild-type strains isolated from wastewater treatment plants in Idaho, Washington, and North Carolina (Rokyta et al. 2006). Strain name only indicates the source of the phage and does not indicate phylogenetic relatedness. Additionally, several laboratory strains (ΦX174, G4, α3, S13, St-1, and ΦK) were included. The tree resolved the major clades identified in previous phylogenetic analyses (Rokyta et al. 2006). We also estimated a phylogenetic tree based on the corresponding sequences of the major spike protein G. The spike phylogeny also resolved the major clades of Bullavirinae and there was good agreement between the two trees for taxon placement (Robinson–Foulds ratio = 0.54). Ultimately, we wanted to determine if increases in capsid stability due to mutation are adaptive or driven by phylogenetic effects. Our Bullavirinae phylogeny, based on the capsid protein sequences, allows us to put biophysical and biochemical traits into a phylogenetic context and understand the phylogenetic correlates of capsid stability.
Cavern hydrophobicity and capsid stability are not correlated across the Bullavirinae. Mapping of cavern hydrophobicity, as estimated by GRAVY score, onto a phylogeny estimated in MrBayes from the coat protein (F) gene sequence of 70 phages shows that relative cavern hydrophobicity is phylogenetically linked. However, capsid stability (measured as death rates) of non-heat adapted genotypes does not correlate with phylogeny. This suggests that cavern hydrophobicity and capsid stability are not correlated in naturally occurring phages. Adaptation via modulation of cavern hydrophobicity by mutation is a specific adaptation to heat-shock and is likely available across the Bullavirinae. For clarity, hydrophobicity is shown as both a color map corresponding to GRAVY score and as bar height corresponding to normalized GRAVY score. Death rates were determined by standard fitness assay. Points represent mean death rates (n = 3–5). Error bars represent ± standard error. (Colour figure online)
To determine the role phylogeny plays in cavern hydrophobicity across the Bullavirinae, we measured hydrophobicity of the cavern by estimating the grand average of hydropathy (GRAVY) scores (Kyte and Doolittle 1982) of the constituent residues. We built a structural model of the ID8 12S pentamer (Fig. 1b) using the G4 crystal structure (pdb:1gff) as a template. The 12S pentamer is composed of five coat monomers (F protein; 9S pentamer) and five spike monomers (G protein; 6S pentamer) forming five cavern structures. The 12S pentamer is an intermediate during pro-phage assembly with 12 12S pentamers forming the full capsid in the presence of scaffolding proteins (Doore and Fane 2016). The 12S pentamer provides a tractable structural model (McGee et al. 2014). Cavern residues were identified in the ID8 model as the solvent exposed amino acids forming the walls of the cavern-like structure where two coat and two spike proteins meet (Fig. 1c; Supplementary Dataset 3). Then, we utilized alignments of coat and spike proteins (Supplementary Datasets 1 and 2) to identify the cavern residues in the other phages represented in our phylogeny (Supplementary Dataset 3) and determined their GRAVY scores (Supplementary Dataset 4). Mapping of cavern hydrophobicity measurements onto the Bullavirinae phylogeny shows that cavern hydrophobicity is tightly linked to phylogeny (Fig. 2). We found evidence of phylogenetic signal in GRAVY scores using Blomberg’s K test (capsid tree: K = 0.79, P < 0.001; spike tree: 0.55, P < 0.001; Blomberg et al. 2003) and Pagel’s continuous λ (capsid tree λ = 0.95, P ≪ 0.001; spike tree: λ = 0.92, P ≪ 0.001: Pagel 1999). This follows as GRAVY score is a sequence-based metric.
If resistance to heat-shock in wild-type phages is driven by cavern hydrophobicity, we would expect to also see phylogenetic signal when stability is mapped onto the Bullavirinae phylogeny. In contrast to cavern hydrophobicity, capsid stability in response to heat-shock appears to be independent of phylogeny in naturally occurring, wild-type phages. In addition to ID8, we randomly selected 17 genotypes from among the 70 used for our phylogeny to assay capsid stability in response to heat-shock by measuring phage death rates in response to a 5-min heat-shock at 80 °C. Death rate measures the time-weighted decrease in phage titer on a logarithmic scale. The distribution of death rates across the phylogeny (Fig. 2) indicates no link between resistance to extreme temperature and phylogeny. Tests for phylogenetic signal in death rates confirm this (Blomberg’s K-capsid tree: K = 0.05, P = 0.77; spike tree: K = 0.04, P = 0.40; Pagel’s-capsid tree: λ ≪ 0.001, P = 1.00; spike tree-λ ≪ 0.001, P = 1.00). In addition, there is no correlation between cavern hydrophobicity and death rates in these naturally occurring phages in a phylogenetically naive comparison (Pearson, r = − 0.402, P-value = 0.0985). When phylogeny is included using Phylogenetic Independent Contrasts, there is no correlation on either the capsid tree (linear regression: R2 = 0.078, n = 16, P = 0.298) or spike tree (linear regression: R2 = 0.0134, n = 15, P = 0.682). This contrasts with the correlation seen for ID8 and the high stability mutants derived under heat-shock conditions (Fig. 1d). Naturally occurring phages are likely never under selection for capsid stability at high temperatures as they never experience such extreme conditions during their life-cycle. These results suggest that the heat-shock mutations that modulate cavern hydrophobicity, previously identified by McGee et al. (2014, 2016), are not simply the result of a phylogenetic effect. Because cavern hydrophobicity is not a determinate of capsid stability in response to heat-shock in un-adapted phages and capsid stability is not lineage specific, modulation of cavern hydrophobicity may be an adaptive mechanism generally available to other phages and not an ID8-specific mechanism.
Single Mutations Mimic the Effect of a Complex Biochemical Pathway
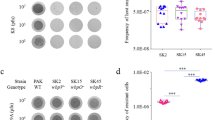
Mutations in ID8 that improve capsid stability (McGee et al. 2014, 2016) appear to modulate the hydrophobicity of a cavern-like structure at the interface of major spike and coat proteins. This hydrophobicity modulation mechanism is similar to the biophysical effects of kosmotropic agents such as trehalose. Trehalose has been shown to stabilize a number of model proteins in vitro including ribonuclease A, lysozyme, and cytochrome C (Xie and Timasheff 1997; Kaushik and Bhat 2003). To determine if ID8 capsid stability responds to such a kosmotropic effect, we measured ID8 death rates in the presence of increasing concentrations of trehalose. Trehalose improves the thermal stability of ID8, an effect that saturates above 0.75 M trehalose (Fig. 3a). We tested for a dilution effect by measuring ID8 death rates in the presence of trehalose and an equal volume of added water instead of trehalose (Fig. S1). Within the range of our measurements (0–1 M trehalose; Fig. 3a), ID8 capsid stability is unaffected by dilution. The water control corresponding to 1.5 M trehalose, well above the range of our measurements, however, shows an improvement in capsid stability that is likely due to changes in ionic strength due to dilution of NaCl and CaCl2. Additionally, the measurement of ID8 capsid stability in the presence of 1.5 M trehalose shows the saturation of the kosmotropic effect. ID8 capsid stability shows a saturable response to the kosmotropic effect.
The kosmotropic effect in capsid stability. a Wild-type ID8 capsid kinetic stability responds to trehalose in a saturable manner. Kosmotropic mutations, R38C and R168C, cause a left-shift in the trehalose curve, moving capsid stability closer to the kosmotropic saturation point. b Other naturally occurring phages show similar responses to trehalose concentration. c The energetic response of kinetic stability (∆∆G) to trehalose correlates with cavern hydrophobicity. GRAVY scores closer to zero represent higher hydrophobicities. Here, the response to trehalose is measured as the slope of the ∆∆G values given by the difference in kinetic stability of the phage at 0 M trehalose and an elevated concentration (Eq. 1). ID102 appears to respond in a different manner and is excluded from the correlation analysis
The response to trehalose is reflected in changes to the free energy barrier to unfolding given by Eq. 1 (Table 1). Death rate measurements probe the kinetic stability of the phage capsid. Kinetic stability represents the free energy barrier separating the native state from the denatured/disassembled states, whereas thermodynamic stability represents the equilibrium populations of the two states. Microvirid capsid stability can be described by the Lumry–Eyring model (Lumry and Eyring 1954; Sanchez-Ruiz 2010) where the native state is more thermodynamically favorable than the disassembled state, but the disassembled capsid represents an irreversible final state due to the requirement of scaffolding proteins for capsid re-assembly (Doore et al. 2017) which are not present outside of the host. Estimation of the thermodynamic stability of ID8 by measuring the difference in free energy of folding in FoldX between the 12S pentamer and the constituent 9S and 6S pentamers indicates that the assembled, native state of the capsid is, indeed, thermodynamically favorable (∆∆G = − 11.22 kcal/mol). Modulation of the kinetic barrier to unfolding via mutations would afford capsid stability in the face of extreme conditions that may disrupt the native state. Similarly, extrinsic agents, such as trehalose, that raise the kinetic barrier to unfolding would show similar effects on capsid stability.
ID8 heat-shock mutations provide the same level of stabilization as 0.5 M trehalose (Fig. 3a; Table 1). These mutants show a limited response to trehalose concentration that saturates at 0.5 M trehalose. This could represent a left-shift of the trehalose response curve of wild-type ID8 suggesting R38C and R168C are closer to the kosmotropic saturation point. Trehalose concentrations above this peak in death rates at 0.5 M are actually disruptive to the heat-stability of the mutants causing a decrease in the kinetic barrier to unfolding (Table 1). These results support our hypothesis that single mutations are potentially capable of recapitulating the stabilization mechanism of trehalose, a product of a complex biochemical pathway. As such, we refer to them as kosmotropic mutations. Further evidence of this saturable kosmotropic effect is provided by the observation that the double-mutant R38C/R168C shows no improvement in heat-stability over the wild-type (Sackman and Rokyta 2018). This saturation effect, thus, also provides a biophysical explanation of observed epistasis, the non-additivity of mutational effects (Starr and Thornton 2016).
A competing mechanism could be mediated by pressure. Many phage genomes are packed into capsids at extremely high densities, reaching an almost crystalline state (Roos et al. 2007). The high packing density leads to high capsid pressure (Roos et al. 2007; Nurmemmedov et al. 2007; Bauer et al. 2015). Increasing temperature could cause an increase in pressure that may lead to capsid rupture. Mutation that modulates the pressure differential across the capsid, as opposed to modulating the hydrophobic effect to increase the interaction energy between capsid protein subunits, may lead to temperature adaptation in contrast to our hypothesized kosmotropic mechanism. However, pressure-based inactivation is relatively ineffective against ΦX174 compared to phages from other families (Guan et al. 2006; Sharma et al. 2008; Cheng et al. 2013; Vo et al. 2014), even when exposed to pressures greater than 300 MPa (Guan et al. 2006). This may be due to the lower genome packing density (Purohit et al. 2005) of ΦX174 as evidenced by the low electron density in the crystal structure (McKenna et al. 1992). This is also seen in the crystal structures of G4 (McKenna et al. 1996) and α (Bernal et al. 2003), other members of the Bullavirinae. Additionally, in our experimental temperature range, the pressure change would be on the order of 0.5 MPa. Temperature adaptation in ID8 and the other Bullavirinae is most likely not mediated by a capsid pressure-based mechanism.
Another potential confounding mechanism could be mediated by direct association of the trehalose molecules with the capsid, and modulation of this interaction by heat-shock mutations. The phage capsids used for crystal structure determination were purified in sucrose gradients (McKenna et al. 1992) and showed a bound glucose molecule on the outer capsid, a reversible association apparently dependent on the presence of calcium ions (Ilag et al. 1994). Trehalose is composed of two glucose molecules joined by a 1,1 glycosidic bond. Capsid glucose binding is likely an artifact of lipopolysaccharide binding, a key step in host infection (Fane et al. 2006). It is unclear what role transient trehalose binding might play in capsid stability, but this association would not be mutually exclusive with our postulated kosmotropic mutation mechanism. Further experiments with a non-glucose-based kosmotropic agent, such as trimethylamine oxide, could provide further evidence of the kosmotropic mutation mechanism and delineate what contribution, if any, direct trehalose association has on capsid stability.
Kosmotropic Response Across the Bullavirinae
We sought to determine if the kosmotropic adaptive response to heat-shock seen in ID8 could be generally available to phages across the Bullavirinae and what role pre-existing cavern hydrophobicity, a phylogenetically linked trait, plays in that response. In the same experiment in which we assayed ID8, R38C, and R168C death rates in response to trehalose, we also assayed five randomly chosen, naturally occurring phages from across the Bullavirinae phylogeny. These naturally occurring phages showed a range of responses to trehalose concentration (Fig. 3b). While heat-stability of naturally occurring phages is not phylogenetically linked, it does appear to dictate how close the phage may move to the phenotypic optimum, a death rate of 0, through the kosmotropic effect, suggesting a potential biophysical constraint for phage thermal adaptation.
Additionally, the results show variation in saturation of the kosmotropic effect across the phages sampled. For example, WA6 shows a trend similar to R38C and R168C saturating at 0.5 M trehalose, while ID102 and ID62 do not saturate up to 1 M trehalose. To quantify the kosmotropic response, we plotted the free energy change in kinetic stability vs trehalose concentration and measured the slope of the resulting curves via linear regression. We then plotted these responses vs cavern hydrophobicity (GRAVY Score) and measured the correlation of the two parameters (Fig. 3c). We found a general, qualitative trend in which the response to trehalose is correlated with cavern hydrophobicity. However, ID102 shows a striking deviation from the trend. When the ID102 response is removed, there is a significant correlation between cavern hydrophobicity and kosmotropic response. Intriguingly, ID102 and ID34 are sister taxa in the ΦX174 clade (Fig. 2) with similar levels of cavern hydrophobicity, yet have different starting death rates and show different kosmotropic responses. Although there is a clear link between cavern hydrophobicity, a phylogenetically linked trait, and the potential for adaptation through the kosmotropic effect, the response of ID102 suggests that even phages that deviate from this trend have access to the same adaptive mechanism. This observation underscores the importance of standing genetic variation in the gating of adaptive paths.
Prior evolutionary events, including the accumulation of neutral mutations in the absence of selection, can shape future adaptive trajectories (Hochberg and Thornton 2017). In naturally occurring phages, it appears that the ability to utilize the kosmotropic adaptive response to extreme temperatures depends on sequence variation in the cavern structure at the interface of coat and spike proteins. The coat and spike proteins play important roles in host cell recognition and attachment, viral DNA delivery, and capsid stability through their interaction with each other Doore and Fane (2016). However, mutational analyses and experimental evolution studies suggest that the cavern-like structure between coat and spike proteins appears to have minimal impact on the phage life-cycle and capsid stability (Rokyta et al. 2009; Fane et al. 2006; Pepin et al. 2008; Wichman and Brown 2010; Sackman and Rokyta 2013; Sackman et al. 2015; Doore and Fane 2016; Doore et al. 2017). Further, in ID8, the ancestral residues at the sites of heat-shock mutations contribute little to coat and spike protein interaction energy (McGee et al. 2014). The cavern-like structure appears to form a literal and metaphorical evolutionary spandrel—“a predictable form that arises as a side consequence rather than a direct adaptation” (Fig. 1; Gould and Lewontin 1979; Gould 1997). The cavern spandrels provide a malleable structural region for adaptive response to extreme environments through kosmotropic mutations that modulate local water structure, the hydrophobic effect, and capsid stability.
Conclusions
Temperature governs the rates and stabilities of biochemical processes and understanding of the molecular adaptive responses of organisms to thermal variation continues to be an area of focus (Viña 2002; Somero 2010; Závodszky and Hajdú 2013; Fields et al. 2015; Nguyen et al. 2017). Our results suggest that sub-structures formed as a by-product of the three-dimensional arrangement of proteins, spandrels, can be co-opted for adaptive response to environmental pressures. Modulation of hydrophobicity via kosmotropic mutation provides a powerful strategy for adaptation to high temperatures. Single mutations can replicate the stability effects of high concentrations of chemical chaperones that are the result of complex biosynthetic pathways in cellular organisms. Additionally, the hydrophobic effect is strengthened with increased temperature (Schellman 1997), suggesting that the increase in stability will be even more pronounced at high temperatures. The modulation of the hydrophobic effect via kosmotropic mutations provides a biophysical mechanism underlying evolutionary phenomena including lineage-specific adaptive constraints and epistasis.
Several reports identify proteins from complex, cellular organisms where surface exposed residues are replaced by hydrophobic amino acids leading to an increase in thermal stability (Van den Burg et al. 1994; Tisi and Evans 1995; Schwehm et al. 1998; Kumar et al. 2000; Machius et al. 2003; Bharadwaj et al. 2008). In the model protein lysozyme, for example, mutation studies suggest that local hydration structures play a role in conformational stability (Funahashi et al. 2000). In combination with the present results, these studies suggest that the kosmotropic mechanism reported here may be applicable to proteins in general and not constrained to microvirid bacteriophages. The kosmotropic mutation mechanism may even find application in protein engineering efforts for design of environmentally robust proteins for industrial applications and viral capsid-based nano-particles for targeted drug delivery (Yildiz et al. 2011; Ma et al. 2012; Rohovie et al. 2017).
Materials and Methods
Phylogenetics
F gene coding sequences from 70 microvirid bacteriophages (SI Dataset 1) were translated to amino acid sequence and aligned with ClustalW (Thompson et al. 1994). The corresponding nucleotide alignment was fitted with nucleotide substitution models in Model-Test v2.1.6 (Darriba et al. 2012). The best fit model, GTR + I + G, was used to estimate a phylogenetic tree by Bayesian inference in MrBayes v3.2.6 (Ronquist et al. 2012). Two independent runs with one cold and three heated chains run for 10 million generations sampling every 10,000 generations with the first 25% of generations discarded as burn-in. The two runs converged on similar trees, and the consensus tree was used for trait mapping. The same procedure was used to estimate a phylogeny based on G gene sequences with the GTR + G substitution model. The Robinson–Foulds estimate of similarity between phylogenies (Robinson and Foulds 1981) was measured in Phylo.io (Robinson et al. 2016). Comparative phylogenetic methods were implemented in aRbor (Harmon et al. 2013). Polytomies were resolved with the modify2di function. Phylogenetic signal in GRAVY score was estimated using the full F and G phylogenies using Pagel’s continuous λ (Pagel 1999) and Blomberg’s K tests (Blomberg et al. 2003). Death rates were only measured for 18 taxa. We manually pruned the trees to match this set of taxa followed by midpoint rooting and tests for phylogenetic signal in death rates. The pruned trees and matching trait sets were used for estimates of trait correlation. We measured a phylogenetically naive correlation using Pearson’s correlation in SigmaPlot (Systat Software, San Jose, CA). We determined phylogenetic independent contrasts (PIC) with GRAVY score as predictor and death rate as response in aRbor. We identified one outlier in PIC values for the F gene tree and two outliers in the G gene tree using Iglewicz and Hoaglins outlier test (Crosby 1994). Phylogenetic generalized least squares method was performed by fitting the remaining PIC values using linear regression in SigmaPlot (Systat Software, San Jose, CA).
Structural Modeling
A homology model of the ID8 12S pentamer consisting of five coat and five spike proteins was built in Modeller v9.18 using the G4 crystal structure as template (pdb:1gff; Šali and Blundell 1993). Ten independent structures were built using the automodel method with slow VFTM optimization for 500 iterations, and slow MD refinement. Refinement was repeated four times. Models were evaluated using the DOPE-HR assessment. The model with the lowest DOPE-HR score of the ten models was chosen for structural analysis. Each cavern structure is formed by residues from two G monomers and two F monomers (Fig. 1). G protein residues were 11–18, 32–41, 61–66, 98, 100, and 167–172 from monomer 1, and 67–70, 94, 107, and 112 from the adjacent monomer in the counter-clockwise direction. F protein residues were 101–104 and 115–117 from monomer 1, and 153–154, 161, 296, 331, 335–336, 338–341, 363–366, 371–372, 377–378, 381, 386, 388–389, and 391–392 from the adjacent monomer in the counter-clockwise direction. These residues were identified in all 70 phages based on the amino acid alignments of F and G proteins (SI Datasets 1–3). The hydrophobicity of cavern residues was estimated by calculating the grand average of hydropathy (SI Dataset 4; http://www.gravy-calculator.de/). In FoldX (Schymkowitz et al. 2005), PDB structures of the 12S pentamer and its constituent 9S and 6S pentamers were repaired using the Repair function followed by measurement of folding stability with the Stability function.
Death Rates
Phage capsid stability was measured as resistance to exposure to extreme temperature (heat-shock) (McGee et al. 2014, 2016). Death rates (doublings/hour) during heat-shock were determined as the population change on a log2 scale. A culture of host cells, Escherichia coli strain C, was grown to ∼ 108 cells/ml in 10 ml of Lysogeny Broth (10 g Tryptone, 10 g NaCl, 5 g yeast extract per liter supplemented with 2 mM CaCl2) within a 125-ml Erlenmeyer flask at 37 °C in an orbital shaking water bath at 200 rpm. Phage isolates were from an individual plaque of sequence confirmed genotypes. Host cell cultures were inoculated with ∼ 105 phage and allowed to propagate for 40 min. Growth was halted and phage isolated by addition of chloroform followed by centrifugation at 14,000 rpm for 3 min in a table top centrifuge. For the negative control, i.e., no trehalose, a 500 µl aliquot of each phage was transferred to a 0.65-ml microfuge tube. To test the effect of trehalose, variable amounts of trehalose were mixed with phage aliquots up to 500 µl. To test for the effect of dilution, the same volume of water was added instead of trehalose. Each sample was vortexed to ensure complete mixing. Tubes were incubated on ice for 5 min to normalize starting temperature, at 80 °C in a hot block with aluminum beads for 5 min for heat-shock, and, finally, on ice for 5 min to halt the heat-shock. The aluminum beads and thin-walled microfuge tubes ensure efficient and uniform heating. Additionally, in this time range for incubation (5 min) relative variations in the rate of heating and cooling will be minor compared to the total incubation time and have minimal influence on the total phage death. Phage populations before and after heat-shock were quantified by plating on host cells on agar plates. The free energy change associated with the change in death rate due to addition of trehalose or to mutation was estimated with the equation
where R is the gas constant and T is temperature.
∆∆G values were estimated for 25 °C. We assumed a two-state model for phage unfolding with a single kinetic barrier that is described by a single exponential decay function. Our death rate measurement is equivalent to the kobs of the decay function.
References
Bauer DW, Li D, Huffman J, Homa FL, Wilson K, Leavitt JC, Casjens SR, Baines J, Evilevitch A (2015) Exploring the balance between DNA pressure and capsid stability in herpesviruses and phages. J Virol 89(18):9288–9298. https://doi.org/10.1128/JVI.01172-15
Bell W, Sun W, Hohmann S, Wera S, Reinders A, De Virgilio C, Wiemken A, Thevelein JM (1998) Composition and functional analysis of the Saccharomyces cerevisiae trehalose synthase complex. J Biol Chem 273(50):33311–33319. https://doi.org/10.1074/jbc.273.50.33311
Bernal RA, Hafenstein S, Olson NH, Bowman VD, Chipman PR, Baker TS, Fane BA, Rossmann MG (2003) Structural studies of bacteriophage α3 assembly. J Mol Biol 325(1):11–24. https://doi.org/10.1016/S0022-2836(02)01201-9
Bharadwaj A, Leelavathi S, Mazumdar-Leighton S, Ghosh A, Ramakumar S, Reddy VS (2008) The critical role of partially exposed N-terminal valine residue in stabilizing GH10 xylanase from Bacillus sp. NG-27 under poly-extreme conditions. PLoS ONE 3(8):e3063. https://doi.org/10.1371/journal.pone.0003063
Blomberg SP, Garland T, Ives AR (2003) Testing for phylogenetic signal in comparative data: behavioral traits are more labile. Evolution 57(4):717–745. https://doi.org/10.1111/j.0014-3820.2003.tb00285.x
Bolen DW, Baskakov IV (2001) The osmophobic effect: natural selection of a thermodynamic force in protein folding. J Mol Biol 310(5):955–963. https://doi.org/10.1006/jmbi.2001.4819
Bruździak P, Panuszko A, Stangret J (2013) Influence of osmolytes on protein and water structure: a step to understanding the mechanism of protein stabilization. J Phys Chem B 117(39):11502–11508. https://doi.org/10.1021/jp404780c
Cheng YK, Rossky PJ (1998) Surface topography dependence of biomolecular hydrophobic hydration. Nature 392(6677):696–699. https://doi.org/10.1038/33653
Cheng X, Imai T, Teeka J, Hirose M, Higuchi T, Sekine M (2013) Inactivation of bacteriophages by high levels of dissolved CO2. Environ Technol 34(1–4):539–544. https://doi.org/10.1080/09593330.2012.704403
Crosby T (1994) How to detect and handle outliers, vol 36. ASQC Quality Press, https://doi.org/10.1080/00401706.1994.10485810
Darriba D, Taboada GL, Doallo R, Posada D (2012) JModelTest 2: more models, new heuristics and parallel computing. Nat Methods 9(8):772. https://doi.org/10.1038/nmeth.2109, arXiv:1011.1669v3
Dill KA (1990) Dominant forces in protein folding. Biochemistry 29(31):7133–7155. https://doi.org/10.1021/bi00483a001, arXiv:1011.1669v3
Dill KA, Truskett TM (2005) Modeling water, the hydrophobic effect, and ion solvation. Annu Rev Biophys Biomol Struct 34(1):173–199. https://doi.org/10.1146/annurev.biophys.34.0402
Doore SM, Fane BA (2016) The microviridae: diversity, assembly, and experimental evolution. Virology 491:45–55. https://doi.org/10.1016/j.virol.2016.01.020
Doore SM, Schweers NJ, Fane BA (2017) Elevating fitness after a horizontal gene exchange in bacteriophage φX174. Virology 501:25–34, https://doi.org/10.1016/j.virol.2016.10.029
Elbein AD, Pan YT, Pastuszak I, Carroll D (2003) New insights on trehalose: a multifunctional molecule. Glycobiology 13(4):17R–27. https://doi.org/10.1093/glycob/cwg047
Fane B, Brentlinger K, Burch A, Chen M, Hafenstein S, Moore E, Novak C, Uchiyama A (2006) ΦwX174 et al., the Microviridae. In: Calendar R (ed) The Bacteriophages, Oxford University Press, Oxford, pp 129–145
Ferreira LA, Breydo L, Reichardt C, Uversky VN, Zaslavsky BY (2017) Effects of osmolytes on solvent features of water in aqueous solutions. J Biomol Struct Dyn 35(5):1055–1068. https://doi.org/10.1080/07391102.2016.1170633
Fields PA, Dong Y, Meng X, Somero GN (2015) Adaptations of protein structure and function to temperature: there is more than one way to ‘skin a cat’. J Exp Biol 218(12):1801–1811. https://doi.org/10.1242/jeb.114298
Francois J, Parrou JL (2001) Reserve carbohydrates metabolism in the yeast Saccharomyces cerevisiae. FEMS Microbiol Rev 25(1):125–145. https://doi.org/10.1016/S0168-6445(00)00059-0
Funahashi J, Takano K, Yamagata Y, Yutani K (2000) Role of surface hydrophobic residues in the conformational stability of human lysozyme at three different positions. Biochemistry 39(47):14448–14456. https://doi.org/10.1021/bi0015717 456
Gould SJ (1997) The exaptive excellence of spandrels as a term and prototype. Proc Natl Acad Sci USA 94(20):10750–10755, https://doi.org/10.1073/pnas.94.20.10750
Gould SJ, Lewontin RC (1979) The spandrels of San Marco and the Panglossian paradigm: a critique of the adaptationist programme. Proc R Soc London Biol Sci 205(1161):581–598. https://doi.org/10.1098/rspb.1979.0086, 9.23
Guan D, Kniel K, Calci K, Hicks D, Pivarnik L, Hoover D (2006) Response of four types of coliphages to high hydrostatic pressure. Food Microbiol 23(6):546–551. https://doi.org/10.1016/J.FM.2005.09.003
Harmon LJ, Baumes J, Hughes C, Soberon J, Specht CD, Turner W, Lisle C, Thacker RW (2013) Arbor: comparative analysis workflows for the tree of life. PLoS Curr 5, https://doi.org/10.1371/currents.tol.099161de5eabdee073fd3d21a44518dc
Hochberg GKA, Thornton JW (2017) Reconstructing ancient proteins to understand the causes of structure and function. Annu Rev Biophys 46(1):247–269. https://doi.org/10.1146/annurev-biophys-070816-033631
Hummer G, Garde S, Garcıa AE, Pratt LR (2000) New perspectives on hydrophobic effects. Chem Phys 258(2–3):349–370. https://doi.org/10.1016/S0301-0104(00)00115-4
Ilag LL, McKenna R, Yadav MP, BeMiller JN, Incardona NL, Rossmann MG (1994) Calcium ion-induced structural changes in bacteriophage φX174. J Mol Biol 244(3):291–300. https://doi.org/10.1006/jmbi.1994.1730
Kaushik JK, Bhat R (1998) Thermal stability of proteins in aqueous polyol solutions: role of the surface tension of water in the stabilizing effect of polyols. J Phys Chem B 102(98):7058–7066. https://doi.org/10.1021/jp981119l
Kaushik JK, Bhat R (2003) Why is trehalose an exceptional protein stabilizer? An analysis of the thermal stability of proteins in the presence of the compatible osmolyte trehalose. J Biol Chem 278(29):26458–26465. https://doi.org/10.1074/jbc.M300815200
Kumar S, Tsai CJ, Nussinov R (2000) Factors enhancing protein thermostability. Protein Eng Des Sel 13(3):179–191. https://doi.org/10.1093/protein/13.3.179
Kyte J, Doolittle RF (1982) A simple method for displaying the hydropathic character of a protein. J Mol Biol 157(1):105–132. https://doi.org/10.1016/0022-2836(82)90515-0, arXiv:1407.5140v1
Lee KH, Miller CR, Nagel AC, Wichman HA, Joyce P, Ytreberg FM (2011) First-step mutations for adaptation at elevated temperature increase capsid stability in a virus. PLoS ONE 6(9):e25640, https://doi.org/10.1371/journal.pone.0025640
Levy Y, Onuchic JN (2006) Water mediation in protein folding and molecular recognition. Annu Rev Biophys Biomol Struct 35(1):389–415. https://doi.org/10.1146/annurev.biophys.35.040405.102134
Lumry R, Eyring H (1954) Conformation changes of proteins. J Phys Chem 58(2):110–120. https://doi.org/10.1021/j150512a005
Ma Y, Nolte RJ, Cornelissen JJ (2012) Virus-based nanocarriers for drug delivery. Adv Drug Deliv Rev 64(9):811–825. https://doi.org/10.1016/j.addr.2012.01.005
Machius M, Declerck N, Huber R, Wiegand G (2003) Kinetic stabilization of Bacillus licheniformis α-amylase through introduction of hydrophobic residues at the surface. J Biol Chem 278(13):11,546– 11,553. https://doi.org/10.1074/jbc.M212618200
Mattos C (2002) Protein–water interactions in a dynamic world. Trends Biochem Sci 27(4):203–208. https://doi.org/10.1016/S0968-0004(02)02067-4
McGee LW, Aitchison EW, Caudle SB, Morrison AJ, Zheng L, Yang W, Rokyta DR (2014) Payoffs, not tradeoffs, in the adaptation of a virus to ostensibly conflicting selective pressures. PLoS Genet 10(10):e1004611, https://doi.org/10.1371/journal.pgen.1004611
McGee LW, Sackman AM, Morrison AJ, Pierce J, Anisman J, Rokyta DR (2016) Synergistic pleiotropy overrides the costs of complexity in viral adaptation. Genetics 202(1):285–295. https://doi.org/10.1534/genetics.115.181628
McKenna R, Xia D, Willingmann P, Ilag LL, Krishnaswamy S, Rossmann MG, Olson NH, Baker TS, Incardona NL (1992) Atomic structure of single-stranded DNA bacteriophage phi X174 and its functional implications. Nature 355(6356):137–143. https://doi.org/10.1038/355137a0
McKenna R, Bowman BR, Ilag LL, Rossmann MG, Fane BA (1996) Atomic structure of the degraded procapsid particle of the bacteriophage G4: induced structural changes in the presence of calcium ions and functional implications. J Mol Biol 256(4):736–750. https://doi.org/10.1006/jmbi.1996.0121
Miyawaki O, Dozen M, Nomura K (2014) Thermodynamic analysis of osmolyte effect on thermal stability of ribonuclease A in terms of water activity. Biophys Chem 185:19–24. https://doi.org/10.1016/j.bpc.2013.10.004
Nguyen V, Wilson C, Hoemberger M, Stiller JB, Agafonov RV, Kutter S, English J, Theobald DL, Kern D (2017) Evolutionary drivers of thermoadaptation in enzyme catalysis. Science 355(6322):289–294, https://doi.org/10.1126/science.aah3717
Nurmemmedov E, Castelnovo M, Catalano CE, Evilevitch A (2007) Biophysics of viral infectivity: matching genome length with capsid size. Q Rev Biophys 40(04):327–356. https://doi.org/10.1017/S0033583508004666
Pace CN, Fu H, Fryar KL, Landua J, Trevino SR, Shirley BA, Hendricks MMN, Iimura S, Gajiwala K, Scholtz JM, Grimsley GR (2011) Contribution of hydrophobic interactions to protein stability. J Mol Biol 408(3):514–528. https://doi.org/10.1016/j.jmb.2011.02.053
Pagel M (1999) Inferring the historical patterns of biological evolution. Nature 401(6756):877–884. https://doi.org/10.1038/44766
Pepin KM, Domsic J, McKenna R (2008) Genomic evolution in a virus under specific selection for host recognition. Infect Genet Evol 8(6):825–834. https://doi.org/10.1016/j.meegid.2008.08.008
Purohit PK, Inamdar MM, Grayson PD, Squires TM, Kondev J, Phillips R (2005) Forces during bacteriophage DNA packaging and ejection. Biophys J 88(2):851–866. https://doi.org/10.1529/biophysj.104.047134
Robinson DF, Foulds LR (1981) Comparison of phylogenetic trees. Math Biosci 53(1–2):131–147. https://doi.org/10.1016/0025-5564(81)90043-2, arXiv:0708.3499v1
Robinson O, Dylus D, Dessimoz C (2016) Phylo.io: interactive viewing and comparison of large phylogenetic trees on the web. Mol Biol Evol 33(8):2163–2166. https://doi.org/10.1093/molbev/msw080, 1602.04258
Rohovie MJ, Nagasawa M, Swartz JR (2017) Virus-like particles: next-generation nanoparticles for targeted therapeutic delivery. Bioeng Transl Med 2(1):43–57. https://doi.org/10.1002/btm2.10049
Rokyta DR, Burch CL, Caudle SB, Wichman HA (2006) Horizontal gene transfer and the evolution of microvirid coliphage genomes horizontal gene transfer and the evolution of microvirid coliphage genomes. J Bacteriol 188(3):1134–1142. https://doi.org/10.1128/JB.188.3.1134
Rokyta DR, Abdo Z, Wichman HA (2009) The genetics of adaptation for eight microvirid bacteriophages. J Mol Evol 69(3):229–239. https://doi.org/10.1007/s00239-009-9267-9
Ronquist F, Teslenko M, Van Der Mark P, Ayres DL, Darling A, H¨ohna S, Larget B, Liu L, Suchard MA, Huelsenbeck JP (2012) Mrbayes 3.2: efficient bayesian phylogenetic inference and model choice across a large model space. Syst Biol 61(3):539–542. https://doi.org/10.1093/sysbio/sys029
Roos WH, Ivanovska IL, Evilevitch A, Wuite GJL (2007) Viral capsids: mechanical characteristics, genome packaging and delivery mechanisms. Cell Mol Life Sci 64(12):1484–1497. https://doi.org/10.1007/s00018-007-6451-1
Sackman AM, Rokyta DR (2013) The adaptive potential of hybridization demonstrated with bacteriophages. J Mol Evol 77(5–6):221–230. https://doi.org/10.1007/s00239-013-9586-8, (NIHMS150003)
Sackman AM, Rokyta DR (2018) Additive phenotypes underlie epistasis of fitness effects. Genetics 208(1):339–348. https://doi.org/10.1534/genetics.117.300451
Sackman AM, Reed D, Rokyta DR (2015) Intergenic incompatibilities reduce fitness in hybrids of extremely closely related bacteriophages. PeerJ 3:e1320. https://doi.org/10.7717/peerj.1320
Šali A, Blundell TL (1993) Comparative protein modelling by satisfaction of spatial restraints. J Mol Biol 234(3):779–815. https://doi.org/10.1006/jmbi.1993.1626
Sanchez-Ruiz JM (2010) Protein kinetic stability. Biophys Chem 148(1–3):1–15. https://doi.org/10.1016/j.bpc.2010.02.004
Schellman JA (1997) Temperature, stability, and the hydrophobic interaction. Biophys J 73(6):2960–2964. https://doi.org/10.1016/S0006-3495(97)78324-3
Schwehm JM, Kristyanne ES, Biggers CC, Stites WE (1998) Stability effects of increasing the hydrophobicity of solvent-exposed side chains in staphylococcal nuclease. Biochemistry 37(19):6939–6948. https://doi.org/10.1021/bi9725069
Schymkowitz J, Borg J, Stricher F, Nys R, Rousseau F, Serrano L (2005) The FoldX web server: an online force field. Nucleic Acids Res 33(SUPPL. 2):382–388. https://doi.org/10.1093/nar/gki387
Sharma M, Shearer AE, Hoover DG, Liu MN, Solomon MB, Kniel KE (2008) Comparison of hydrostatic and hydrodynamic pressure to inactivate foodborne viruses. Innov Food Sci Emerg Technol 9(4):418–422. https://doi.org/10.1016/J.IFSET.2008.05.001
Sharp KA, Madan B (1997) Hydrophobic effect, water structure, and heat capacity changes. J Phys Chem B 101(21):4343–4348. https://doi.org/10.1021/jp9702457
Singer MA, Lindquist S (1998) Thermotolerance in Saccharomyces cerevisiae: the Yin and Yang of trehalose. Trends Biotechnol 16(11):460–468. https://doi.org/10.1016/S0167-7799(98)01251-7
Somero GN (2003) Protein adaptations to temperature and pressure: complementary roles of adaptive changes in amino acid sequence and internal milieu. Comp Biochem Physiol B Biochem Mol Biol 136(4):577–591. https://doi.org/10.1016/S1096-4959(03)00215-X
Somero GN (2010) The physiology of climate change: how potentials for acclimatization and genetic adaptation will determine ‘winners’ and ‘losers’. J Exp Biol 213(6):912–920. https://doi.org/10.1242/jeb.037473
Starr TN, Thornton JW (2016) Epistasis in protein evolution. Protein Sci 25(7):1204–1218. https://doi.org/10.1002/pro.2897
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22(22):4673–4680. https://doi.org/10.1093/nar/22.22.4673, arXiv:1011.1669v3
Timasheff SN (1998) Control of protein stability and reactions by weakly interacting cosolvents: the simplicity of the complicated. Adv Protein Chem 51:355–432
Tisi LC, Evans PA (1995) Conserved structural features on protein surfaces: Small exterior hydrophobic clusters. J Mol Biol 249(2):251–258. https://doi.org/10.1006/jmbi.1995.0294
Van den Burg B, Dijkstra BW, Vriend G, Van der Vinne B, Venema G, Eijsink VG (1994) Protein stabilization by hydrophobic interactions at the surface. Eur J Biochem 220(3):981–985
Viña J (2002) Biochemical adaptation: mechanism and process in physiological evolution, vol 30. Oxford University Press, Oxford. https://doi.org/10.1002/bmb.2002.494030030071
Vo HT, Imai T, Ho TT, Sekine M, Kanno A, Higuchi T, Yamamoto K, Yamamoto H (2014) Inactivation effect of pressurized carbon dioxide on bacteriophage Qβ and ΦX174 as a novel disinfectant for water treatment. J Environ Sci 26(6):1301–1306. https://doi.org/10.1016/S1001-0742(13)60603-8
Wichman HA, Brown CJ (2010) Experimental evolution of viruses: microviridae as a model system. Philos Trans R Soc B Biol Sci 365(1552):2495–2501. https://doi.org/10.1098/rstb.2010.0053
Xie G, Timasheff SN (1997) The thermodynamic mechanism of protein stabilization by trehalose. Biophys Chem 64(1–3):25–43. https://doi.org/10.1016/S0301-4622(96)02222-3
Yancey PH (2005) Organic osmolytes as compatible, metabolic and counteracting cytoprotectants in high osmolarity and other stresses. J Exp Biol 208(15):2819–2830. https://doi.org/10.1242/jeb.01730
Yancey PH, Clark ME, Hand SC, Bowlus RD, Somero GN (1982) Living with water stress: evolution of osmolyte systems. Science 217(4566):1214–1222. https://doi.org/10.1126/science.7112124
Yildiz I, Shukla S, Steinmetz NF (2011) Applications of viral nanoparticles in medicine. Curr Opin Biotechnol 22(6):901–908. https://doi.org/10.1016/j.copbio.2011.04.020
Závodszky P, Hajdú I (2013) Evolution of the concept of conformational dynamics of enzyme functions over half of a century: a personal view. Biopolymers 99(4):263–269. https://doi.org/10.1002/bip.22159
Acknowledgements
Funding for this work was provided by the U.S. National Institutes of Health to D.R.R. (R01 GM-099723).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Whittington, A.C., Rokyta, D.R. Biophysical Spandrels form a Hot-Spot for Kosmotropic Mutations in Bacteriophage Thermal Adaptation. J Mol Evol 87, 27–36 (2019). https://doi.org/10.1007/s00239-018-9882-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-018-9882-4