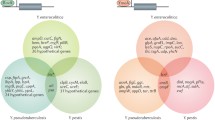
Abstract
A fundamental tenet of evolution is that alleles that are under negative selection are often deleterious and confer no evolutionary advantage. Negatively selected alleles are removed from the gene pool and are eventually extinguished from the population. Conversely, alleles under positive selection do confer an evolutionary advantage and lead to an increase in the overall fitness of the organism. These alleles increase in frequency until they eventually become fixed in the population. Francisella tularensis is a zoonotic pathogen and a potential biothreat agent. The most virulent type of F. tularensis, Type A, is distributed across North America with Type A.I occurring mainly in the east and Type A.II appearing mainly in the west. F. tularensis is thought to be a genome in decay (losing genes) because of the relatively large number of pseudogenes present in its genome. We hypothesized that the observed frequency of gene loss/pseudogenes may be an artifact of evolution in response to a changing environment, and that genes involved in virulence should be under strong positive selection. To test this hypothesis, we sequenced and compared whole genomes of Type A.I and A.II isolates. We analyzed a subset of virulence and housekeeping genes from several F. tularensis subspecies genomes to ascertain the presence and extent of positive selection. Eleven previously identified virulence genes were screened for positive selection along with 10 housekeeping genes. Analyses of selection yielded one housekeeping gene and 7 virulence genes which showed significant evidence of positive selection at loci implicated in cell surface structures and membrane proteins, metabolism and biosynthesis, transcription, translation and cell separation, and substrate binding and transport. Our results suggest that while the loss of functional genes through disuse could be accelerated by negative selection, the genome decay in Francisella could also be the byproduct of adaptive evolution driven by complex interactions between host, pathogen, and thier environment, as evidenced by several of its virulence genes which are undergoing strong, positive selection.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The evolution of virulence is driven by complex interactions between host, pathogen, and their environment (Read 1994; Brown et al. 2006; Masri et al. 2015). Alleles that are under negative selection are usually thought to be either deleterious or neutral and are eventually eliminated from the gene pool. Conversely, alleles under positive selection can confer an evolutionary advantage leading to an increase in the overall fitness of the organism, eventually becoming fixed in the population. Both positive and negative selection can lead to changes in genome architecture. For example, in combination with gene duplication, positive selection can facilitate the expansion of gene families and functional diversity (Demuth and Hahn 2009; Lan et al. 2009). Similarly, negative selection can lead to increased numbers of pseudogenes and is implicated in genome decay (Rohmer et al. 2007). Both positive and negative selection are involved in the evolution of virulence, as pathogens respond to changing host and physical environments (Ward et al. 2009; Nandi et al. 2010; Feng et al. 2011).
Francisella tularensis is a Gram-negative facultative intracellular pathogen and is the causative agent of the zoonotic disease, tularemia (rabbit fever). While many mammals are susceptible to the disease, they are not known to act as environmental reservoirs (Jones et al. 2012). Arthropods such as ticks and deer flies are likely the primary vectors in the spread of tularemia, with human infection usually resulting from the bite of an insect or inhalation of aerosolized bacteria (Sjostedt 2007; Oyston 2008). While there are no confirmed reservoirs for F. tularensis, it has been demonstrated that some strains cause the rapid encystment of Acanthamoeba castellanii and are able to survive within these ameba cysts for up to 3 weeks, thus increasing the environmental persistence of the bacteria (El-Etr et al. 2009; Jones et al. 2012).
There are currently three recognized species within the Francisella genus: F. tularensis, F. philomiragia, and F. novicida. It has been suggested that F. novicida be reclassified as a subspecies of F. tularensis, and much of the scientific literature, including Bergey’s manual of Systemic Bacteriology, already reflects this change (George 2005; Svensson et al. 2005; Keim et al. 2007; Sjostedt 2007; Gunnell et al. 2012). Excluding F. novicida, there are three recognized subspecies of F. tularensis: tularensis, holarctica, and mediasiatica. Two of the subspecies of Francisella tularensis are often abbreviated simply as Type A (tularensis) and Type B (holarctica). Furthermore, Type A tularensis has been further divided into A.I and A.II types based on geographic distribution and genome architecture (Johansson et al. 2004). Type A tularensis is generally found in North America with Type A.I dominating the central and eastern portions of the continent, while Type A.II is typically found in the western United States (Staples et al. 2006). Type B tularensis (or holarctica) exhibits a worldwide prevalence, while the mediasiatica subspecies appears to be confined to the central Asian republics of the former Soviet Union (Broekhuijsen et al. 2003; Johansson et al. 2004) (Table 1). The phylogenetic relationships of these subspecies are shown in Fig. 1. Type A tularensis is the most virulent subspecies of F. tularensis (Owen et al. 1964; Weiss et al. 2007).
Maximum likelihood tree inferring the phylogenetic relationships of the F. tularensis subspecies described in this work. Tree was constructed by concatenating 10 housekeeping genes (recA, gyrB, groEL, dnaK, rpoA1, rpoB, rpoD, rpoH, fopA, and sdhA) followed by alignment with ClustalW and generation of the tree with MEGA 5.2. Taxa names use the shorthand described in Table 2
In general, virulence genes are those whose products contribute to an organism’s ability to infect and colonize a host. Virulence genes are often located within mobile genetic elements such as plasmids, transposons, bacteriophages, and pathogenicity islands, which enhance genetic diversity through the exchange with other bacteria (Kaper 2005). Despite these varied mechanisms for sharing genes among bacteria, molecular evidence suggests that the four subspecies of F. tularensis have evolved and acquired virulence genes by vertical descent rather than by horizontal gene transfer (Svensson et al. 2005). A whole-genome analysis of horizontal gene transfer (HGT) in F. tularensis identified 30 candidate genes as having been acquired through HGT (Sridhar et al. 2012), but none of them were putative virulence genes. Genetic variation in the subspecies of F. tularensis seems to have arisen by mutation rather than HGT (Gestin et al. 2010; Siddaramappa et al. 2012; Sutera et al. 2014). Furthermore, the subspecies novicida has been shown to possess a CRISPR/Cas system to defend against invading genetic elements, further supporting the notion that mutation and selection are the driving factors of evolution in F. tularensis virulence rather than the acquisition of mobile genetic elements by HGT (Gallagher et al. 2008; Schunder et al. 2013).
Whole-genome comparisons between pathogenic and non-pathogenic subspecies of F. tularensis reveal that pathogenic subspecies evolved from the non-pathogenic subspecies by mechanisms of genomic rearrangements, point mutations, and small indels (Rohmer et al. 2007). The many pseudogenes present in the F. tularensis genome suggest that it is a genome in decay (Santic et al. 2010). While these pseudogenes may have arisen via negative selection through disuse, it is also possible that that this evidence of decay is a byproduct of an adaptive response to a changing environment (Ward et al. 2009; Feng et al. 2011) or even the evolution of avirulence (Hain et al. 2007; Harris et al. 2015). Alternatively, in addition to genome decay through the creation of pseudogenes, we hypothesize that positive natural selection could also be a driving force in the continued evolution of virulence of this important pathogen.
Analyses of selection are often conducted by calculating the ratio of nonsynonymous (dN) to synonymous (d S) substitutions within a gene. It has been suggested that this method of identifying selection is somewhat conservative and a more sensitive approach is needed (Crandall et al. 1999; Woolley et al. 2003; Fay 2011; Messer and Petrov 2013). To detect the presence of selection in virulence genes, and gain insights into the molecular evolution of F. tularensis, we tested for selection based on an expected random distribution of possible amino acid substitutions based on 31 physiochemical properties of amino acids.
Materials and Methods
Bacterial Strains and Culture Conditions
The isolates used in this study (Table 2) are a part of a select agent archive housed at the special pathogens laboratory at Brigham Young University (Provo, UT). The collection largely consists of isolates obtained from the State Health Departments of Utah and New Mexico over the past two decades. All F. tularensis isolates were grown on modified Mueller–Hinton agar (MMHA) (Becton–Dickinson and Company, Franklin Lakes, New Jersey, USA) for 3–4 days with 5 % CO2 at 35 °C. MMHA was prepared by autoclaving the Mueller–Hinton base, which was chocolatized by adding 5 % sheep blood, while the medium was approximately 80 °C. After the medium cooled to 50 °C, 10 mL of 10 % glucose and 20 mL of IsoVitaleX (Becton–Dickinson and Company) were added to 1 L.
Genome Sequencing and Annotation
Total genomic DNA was extracted from each isolate using the MagNA Pure System (Roche) and the MagNA Pure LC DNA Isolation Kit III (Roche) according to the manufacturer’s directions. Briefly, cells grown on MMHA agar were suspended in 250 μL of TE buffer containing 1.8 μg/μL lysozyme and incubated for 1 h at 37 °C. To this tube, 270 μL of bacterial lysis buffer and 100 μL of proteinase K were added and the tube was incubated for 10 min at 65 °C. Samples were then incubated in boiling water for 10 min to inactivate pathogens. DNA was eluted in a total volume of 100 µL. DNA concentration was measured using a PicoGreen assay (Invitrogen) and TBS-380 fluorometer (Turner Biosystems).
Prior to genome sequencing, isolates were typed using the multiplex PCR assay as described previously (Gunnell et al. 2012). Four Type A.I and six Type A.II strains were selected for sequencing in order to maximize the available geographic and genetic diversity. Whole-genome shotgun sequencing was accomplished using the Roche/454 GS FLX Pyrosequencer (Roche) and the Titanium® chemistry according to the manufacturer’s recommendations. Raw sequencing reads were aligned to reference sequences using Newbler version 2.6. The reference sequence for A.I strains was SCHU S4 (GenBank accession number AJ749949.2), while WY96-3418 (NC_009257.1) was used for A.II strains. Aligned shotgun sequences were annotated using the Prokaryotic Genome Automatic Annotation Pipeline (PGAAP) (Angiuoli et al. 2008).
Analysis of Selection
A total of 22 taxa were included in the analysis of selection: the 10 new F. tularensis subspecies tularensis genomes described in this work and 12 other F. tularensis genomes of the tularensis, holarctica, and mediasiatica subspecies retrieved from GenBank (Table 2).
In addition to 5 Type IV pilus genes which are known virulence factors of F. tularensis, 89 other previously identified virulence genes (Su et al. 2007) were analyzed for positive selection. In total, 94 virulence genes were extracted from the genomes of the 22 taxa in Table 2 for analysis. While the taxa used in this work were isolated from various geographical locations (Table 2), the genes selected for analysis are common to all the isolates. Individual genes were aligned with ClustalW (Larkin et al. 2007) using the default parameters. A hierarchical likelihood ratio test was performed on the aligned sequences to determine which model of evolution best fit the data under the Akaike Information Criterion (AIC) (Nei and Kumar 2000) using MEGA v. 5.2 (Tamura et al. 2011). MEGA v. 5.2 was also used to generate phylogenetic trees via maximum likelihood assuming the previously identified best-fit substitution model (Tamura et al. 2011). Maximum likelihood computations of d N and d S were conducted using HyPhy (Hasegawa et al. 1985; Muse and Gaut 1994; Suzuki and Gojobori 1999; Pond and Frost 2005; Pond et al. 2005).
In order to perform a more sensitive test of selection, we measured the strength of selection on 31 physiochemical properties of homologous amino acids among the different genomes using TreeSAAP version 3.2 (Woolley et al. 2003). TreeSAAP compares aligned sequences in the context of the specified phylogenetic topology, codon by codon, to infer amino acid replacement events. Because TreeSAAP is computationally intensive, following d N/d S analysis using HyPhy, we selected 11 genes for further analysis. Results were divided into one of eight categories based on the magnitude of the change: categories 1–3 indicate a conservative change, categories 4–6 represent moderate change, while categories 7–8 represent more drastic changes and indicate positive selection. Only genes that returned changes in categories 7–8 were examined further. A z-score was calculated by TreeSAAP for each of the 31 physiochemical properties. A z-score > 3.09 indicated a dramatic change (categories 7–8). Changes where p < 0.001 were determined to indicate statistically significant positive selection. To summarize, we explored 94 virulence-associated genes. Of those, 64 were indicated by HyPhy analysis to be under positive selection, though the results were not statistically significant. From the 64 genes that showed some evidence of positive selection, we chose a subset of 11 genes for subsequent TreeSAAP analysis. As a control measure, 10 housekeeping genes were selected and subjected to these same analyses of selection. To better visualize the evolutionary changes in the proteins, secondary structures of the proteins were predicted using the PSIPRED secondary structure prediction (http://bioinf.cs.ucl.ac.uk/psipred/) (Jones 1999) (Figs. S1, S2, S3, S4, S5, S6, S7).
Results and Discussion
Genome Sequencing and Annotation
Of the 10 novel genomes sequenced, 4 were F. tularensis Type A.I and 6 were F. tularensis Type A.II. A summary of the sequencing statistics of these newly sequenced genomes can be found in Table 3.
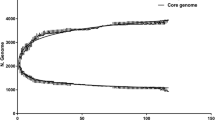
The lengths of the reference sequences were 1,892,775 and 1,898,476 for the SCHU S4 and WY96-3418 strains, respectively. The average length of the sequenced Type A.I genomes was 1,814,938 and the average length of the Type A.II genomes was 1,814,544. Thus, the sequenced genomes are about 80 kb shorter than the reference sequences. This is not surprising since the shotgun method of sequencing cannot adequately sequence through large repetitive regions of DNA resulting in gaps in the final alignments.
After the sequences were aligned to their respective references, they were annotated using the Prokaryotic Genome Automatic Annotation Pipeline (PGAAP) (Angiuoli et al. 2008) and submitted to GenBank under the following accessing numbers: AMPP00000000, AMPY00000000, AMPX00000000, AMPV00000000, AMPU00000000, AOUD00000000, APKY00000000, APKX00000000, APKW00000000, and APKV00000000. A summary of the annotation results can be found in Table 4.
The number of predicted protein-coding sequences for the references SCHU S4 and WY96-3418 was 1604 and 1634, respectively. Because of the shortened size of the newly sequenced genomes, it was expected that the total number of protein sequences would also be less. This, however, was not the case. The new genomes were processed through an automated annotation pipeline, and were not manually curated. This resulted in many sequences that the automated pipeline mistook for genes that the curators of the reference genomes had already removed. Further study and curation of these newly sequenced genomes will be necessary for detailed comparative analyses, but our analyses provided sufficient resolution to facilitate analysis of selection on virulence-associated genes.
Analysis of Selection
The analysis of d N/d S ratios is a well-known benchmark for identifying selection and was used as a first pass to identify genes that are most likely to be under positive selection. Results revealed that of the 94 previously identified virulence genes (Su et al. 2007), 64 gave some indication of positive selection (although with p values ranging from 0.40 to 0.90, none of them were statistically significant). It has been suggested that some of the assumptions of the d N/d S ratio test, as well as the McDonald–Kreitman test of adaptive evolution (McDonald and Kreitman 1991) can be too conservative for accurately detecting positive selection and adaptation (Crandall et al. 1999; Woolley et al. 2003; Fay 2011; Messer and Petrov 2013). For example, d N/d S ratios alone were unable to detect positive selection in rapidly mutating genes of HIV in patients showing signs of drug resistance (Crandall et al. 1999). Furthermore, Sharp (1997) asserts that another problem with d N/d S ratio tests is that their significance falls as selection continues to “weed out” the effects of detrimental amino acid changes. Thus, more nuanced methods may be needed to accurately predict positive selection in biological systems (Messer and Petrov 2013). Finally, while d N/d S ratios and the McDonald–Kreitman test may indicate the presence of selection on a gene, they do not give any indication of how selection affects the structure and function of the protein (Taylor et al. 2005).
To perform a more sensitive analysis, we used TreeSAAP (Selection on Amino Acid Properties using phylogenetic Trees) which tests for selection of expected random distributions of possible amino acid changes based on 31 physiochemical properties of amino acids and associated phylogenetic trees (Woolley et al. 2003). The methodological approach that TreeSAAP uses to detect selection has been shown to be better suited at detecting selection where more traditional methods such as d N/d S ratios and McDonald–Kreitman analyses cannot (Sharp 1997; Woolley et al. 2003). We note that TreeSAAP does not have a built-in correction for multiple-hypothesis testing when calculating p values. Although a Bonferroni correction of TreeSAAP significance tests is indicated for comparative purposes, previous work has cast doubt on whether this will yield an appropriate level of confidence (McClellan and Ellison 2010). Thus, we employed TreeSAAP as the best-suited approach currently available for detecting selection.
Since the HyPhy results were not statistically significant, 11 virulence genes were selected for further analyses of selection using TreeSAAP (Table 5). In genes FTT_1611, FTT_1156c, FTT_0935c, and FTT_1525c, no significant positive selection was detected. However, significant positive selection (categories 7–8) was detected in one or more physiochemical properties of the remaining seven genes: FTL_1134, FTT_0683, FTT_0881c, FTT_0504c, FTT_0936c, FTT_0766, and FTT_1125. The physiochemical properties for which positive selection was detected include the following: isoelectric point, power to be at the N-terminal, composition, power to be at the middle of alpha helix, solvent-accessible reduction ratio, equilibrium constant, power to be at the C-terminal, coil tendencies, compressibility, bulkiness, turn tendencies, and average number of surrounding residues.
For analytical purposes, these seven genes were placed into four functional categories (Su et al. 2007): FTL_1134 and FTT_0683 fall into the cell surface structures and membrane proteins category, FTT_0881c, FTT_0504c, and FTT_0936c are metabolism and biosynthesis proteins, FTT_0766 is categorized as a protein involved in transcription, translation, and cell separation, and FTT_1125 is classified in the substrate binding and transport functional group.
Cell Surface Structures and Membrane Proteins
In the functional group of cell surface structures and membrane proteins, the gene FTL_1134 showed signatures of selection for three of the 31 physiochemical properties tested: isoelectric point in codons 183–203 and 383–394, power to be at the N-terminal in codons 271–285 and 323–337, and composition in codons 183–197 (Fig. 2). Codons 183–203 begin in an α-helix and end in a loop, codons 271–285 include portions of two α-helices and a loop in between, codons 323–337 begin in an α-helix and end in a loop, and codons 383–394 occur near the C-terminal of an α-helix (Fig. S1). The product of FTL_1134 is a hypothetical membrane protein of unknown function. This gene (as were all other genes cited in this work) was identified as a virulence gene by a whole-genome transposon screen for virulence genes of F. tularensis (Su et al. 2007). Thus, in this case, a protein of unknown function has been identified as a virulence gene.
Selection on FTL_1134. Areas of the gene under selection are indicated where the test statistic is greater than 3.09 (horizontal line) (p < 0.001). Selection is present in the gene for the following amino acid properties where the lines cross the threshold: isoelectric point (orange line), power to be at the N-terminal (gray line), and composition (yellow line)
Francisella tularensis is an intracellular pathogen which is routinely phagocytized by host cells and taken into a phagosome where the pH is lowered in an effort to rid the cell of the bacterium. Selection of FTL_1134 for the isoelectric point, the pH at which a molecule carries no net electrical charge, hints that this membrane protein may be exposed to changes in pH and is adapting to withstand these changes.
Also in FTL_1134, TreeSAAP identified signatures of selection for the amino acid property of power to be at the N-terminal. This amino acid property is defined as the intrinsic property of an amino acid residue to be located at the N-terminus of an α-helix (Prahhakaran and Ponnuswamy 1979). The codons under selection occur in varied domains of the secondary structure of this protein including the N-terminus of an α-helix, the C-terminus of an α-helix, and the loop between two α-helices (Fig. S1). While this property and its relation to positive selection are not well understood, we speculate that the regions under selection are important to the ability of this virulence gene to function intercellularly since in an infection model with this gene knocked out, the host is able to clear the infection (Su et al. 2007).
The third and final amino acid property under selection for FTL_1134 is composition. Composition refers to the individual amino acid make-up of a protein. Since FTL_1134 is a hypothetical membrane protein of an unknown function, it is difficult to know how positive selection for the composition property may affect it. Further study is needed to determine the function of this protein and its role in the virulence mechanisms of F. tularensis before we can gain an understanding of how selection is influencing this protein.
The other protein showing signatures of positive selection in the cell surface structures and membrane proteins functional group is FTT_0683, which codes for the type IV pilus protein PilD. Type IV pili are common in Gram-negative pathogens and mediate the attachment of the pathogen to various host receptors (Strom and Lory 1993). Specifically, pilD is a bifunctional, cytoplasmic membrane protein responsible for the cleavage of the N-terminal leader sequences of prepilin and also catalyzes the N-methylation of the N-terminal phenylalanine of mature pilins (Strom et al. 1993). Without a functional PilD, the type IV pilus apparatus cannot be assembled, thus restricting the pathogenicity of F. tularensis. TreeSAAP detected selection in PilD for the amino acid property of power to be at the middle of α-helix, which is described as the intrinsic property of a specific amino acid to be located in the middle of an α-helix (Prahhakaran and Ponnuswamy 1979). The catalytic domains of PilD are predicted to be specific cytoplasmic cysteine motifs found toward the N-terminus of the protein (Lory and Strom 1997), well distanced from the selection occurring toward the C-terminus of the protein in amino acids 190–204 (Fig. 3). Selection in these membrane bound α-helices does not affect the catalytic activity of PilD, but may alter membrane positioning of the protein (Fig. S2).
Selection on FTT_0683 (pilD). Areas of the gene under selection are indicated where the test statistic is greater than 3.09 (horizontal line) (p < 0.001). Selection is present in the gene for the amino acid property of power to be at the middle of alpha helix where the orange line crosses the threshold
Metabolism and Biosynthesis
Three genes in the metabolism and biosynthesis functional group showed signatures of selection, FTT_0881c (rocE), FTT_0504c (sucC), and FTT_0936c (bioF). The study which identified these genes as virulence factors used a signature-tagged mutagenesis approach (Su et al. 2007). Briefly, this approach introduces random transposons to disrupt genes in the genome before infection using a mouse model. If an organism with a particular disrupted gene is not recovered from the infection model, that gene is then classified as necessary for virulence. This approach has been used to identify virulence factors in many other organisms (Hensel et al. 1995; Chiang et al. 1999). In the strictest sense, genes categorized as metabolism and biosynthesis should not be considered virulence factors, since impairing them would naturally restrict growth regardless of the status of infection (Casadevall and Pirofski 1999; Su et al. 2007). However, it is important not to discount the role these genes may play in infection altogether, since they may be valuable targets for the future development of therapeutics or vaccines.
The first gene in the metabolism and biosynthesis category for which TreeSAAP identified selection signatures was FTT_0881c. This locus codes for the protein RocE which is an amino acid permease, specifically, a transmembrane arginine transporter protein. Francisella tularensis lacks the ability to synthesize the amino acid arginine. Furthermore, recent studies have shown that the cytosol of the host cell does not contain sufficient free nutrients to support the rapid growth of the pathogen (Steele et al. 2013). Without access to arginine, the pathogen would not be able to reproduce and infection with F. tularensis would fail. To compensate, F. tularensis induces the host cell into an autophagy state in which proteins are broken down into their constituent amino acids and made available to the pathogen (Steele et al. 2013). F. tularensis with the transmembrane amino acid permease, RocE, may then scavenge for the newly available arginine. Furthermore, macrophages use arginine to produce nitric oxide (NO) as an antimicrobial defense (Kone et al. 2003). In addition to scavenging arginine for growth, the ability of F. tularensis to remove arginine would also deprive the cell of this potent antimicrobial agent, thus allowing the infection to progress.
The rocE gene showed signatures of positive selection for the solvent-accessible reduction ratio property of amino acids in codons 109–116, 438, and 442 (Fig. 4). In the secondary structure of RocE, codons 109–116 encompass a β-sheet and part of a loop, while codons 438 and 442 occur within the same α-helix (Fig. S3). The solvent-accessible reduction ratio is defined as the decrease in solvent accessibility for an amino acid residue when the protein molecule moves from a hypothetically extended state to the native folded state (Ponnuswamy et al. 1980). Consequently, as the solvent accessibility decreases, the hydrophobicity of the protein increases. RocE is a transmembrane protein and hydrophobic domains are necessary for its placement and function in the membrane. Host cells have evolved a mechanism of nutrient restriction as a protection from intracellular pathogens such as F. tularensis (Steele et al. 2013). These results suggest that F. tularensis is also evolving mechanisms to scavenge for, and acquire, the necessary nutrients for intracellular survival.
The second protein in the metabolism and biosynthesis group demonstrated to be under positive selection is SucC, coded for by the FTT_0504c locus. SucC is the β-chain of the succinyl-CoA synthetase which, when completed with another β-chain and 2 α-chains (SucD), catalyzes the reaction of succinyl-CoA to succinate in the citric acid cycle. Positive selection was identified for the amino acid property of equilibrium constant (ionization of COOH) in codons 31–45 and 84–89 (Fig. 5). These codons encompass all three major secondary structures: an α-helix, a β-sheet, and a loop, all near the N-terminus of the protein (Fig. S4). The areas identified to be under selection do not appear to be involved in any of the catalytic regions of the protein (Wolodko et al. 1994; Fraser et al. 1999), thus amino acid substitutions in these non-catalytic regions should still allow the completed enzyme to function normally. It has also been demonstrated that the activity of the succinyl-CoA synthetase increases when Escherichia coli is cultured in an acidic environment. If succinyl-CoA synthetase behaves similarly in F. tularensis, this may be part of a stress response initiated when the bacterium is inside a phagolysosome of a host cell (Walshaw et al. 1997; Stancik et al. 2002).
Selection on FTT_0504 (sucC). Areas of the gene under selection are indicated where the test statistic is greater than 3.09 (horizontal line) (p < 0.001). Selection is present in the gene for the amino acid property of equilibrium constant (ionization of COOH) where the orange line crosses the threshold
The third and final gene identified to be under selection in the metabolism and biosynthesis group is FTT_0936c which codes for the protein BioF. BioF is a 8-amino-7-oxononanoate synthase which is responsible for the first committed step in the biosynthesis of biotin (Alexeev et al. 1998). TreeSAAP identified selection for 4 different amino acid properties: power to be at the N-terminal in codons 8–22 and 198–210, power to be at the C-terminal in codons 8–22 and 198–202, coil tendencies in codons 8–22 and 200–203, and compressibility at codons 8–22 and 198–209 (Fig. 6). The secondary structures involved in the areas of selection include α-helices, part of a β-sheet, and loops (Fig. S5). The active site for BioF is at residues 47 and 361 (Kerbarh et al. 2006), well away from the areas of selection in this gene. It is worth reiterating that genes involved in metabolism and biosynthesis are not considered virulence genes in the strictest sense, but they should not be discounted either as they may prove to be effective drug targets. Such is the case with plumbagin (5-hydroxy-2-methyl-1,4-naphthoquinone), a natural compound found to be an effective herbicide through the inhibition of 8-amino-7-oxononanoate synthase (BioF) (Choi et al. 2012). Because plants, microbes, and some fungi synthesize their own biotin, while animals require trace amounts of biotin in their diets, plumbagin may be a safe and effective herbicide by disrupting the biotin biosynthesis pathway (Nudelman et al. 2004; Kerbarh et al. 2006; Choi et al. 2012).
Selection on FTT_0936c (bioF). Areas of the gene under selection are indicated where the test statistic is greater than 3.09 (horizontal line) (p < 0.001). Selection is present in the gene for the following amino acid properties where the lines cross the threshold: power to be at the N-terminal (orange line), power to be at the C-terminal (gray line), coil tendencies (yellow line), and compressibility (dark blue line)
Transcription, Translation, and Cell Separation
One gene in the transcription, translation, and cell separation functional group determined to be under selection, DeoD, is coded for by the locus FTT_0766. Of the 31 physiochemical properties of amino acids tested, selection was noted for bulkiness in codons 169–183 and turn tendencies in codons 127 and 176–183 (Fig. 7). Bulkiness refers to the relative size of the side chain of a particular amino acid. For example, leucine is considered to be “bulkier” than alanine (Dhanasekaran et al. 2008; Sengupta and Kundu 2012). Substituting amino acids with different bulkiness properties can affect the overall hydrophobicity of a protein or even protein folding and substrate binding where specific steric interactions are important (Dhanasekaran et al. 2008). Turn tendencies refer to the propensity of a particular amino acid to be in a β-turn (Chou and Fasman 1978). In terms of secondary structure, the areas of the gene under selection include an α-helix, a β-sheet, and loops (Fig. S7).
The product of the deoD gene is a purine nucleoside phosphorylase (PNP) which catalyzes the phosphorylation of purine ribonucleosides and 2′-deoxyribonucleosides as part of the purine salvage pathway (Luo et al. 2011). The active site for DeoD is near the N-terminus of the protein, while selection was detected further downstream toward the C-terminus, indicating that selection is not changing the specific function of this enzyme. However, enzymes that are part of the nucleotide biosynthesis pathway, such as PNP, are an optimal target for antimicrobial therapy since they are significantly different from the eukaryotic enzymes that perform the same function (Grenha et al. 2005). Understanding how selection operates on this gene could lead to better, more effective antimicrobial therapies.
Selection on FTT_0766 (deoD). Areas of the gene under selection are indicated where the test statistic is greater than 3.09 (horizontal line) (p < 0.001). Selection is present in the gene for the following amino acid properties where the lines cross the threshold: bulkiness (orange line) and turn tendencies (gray line)
Substrate Binding and Transport
The final functional group to have a gene implicated in selection is substrate binding and transport. Selection was detected for two amino acid properties in the gene metQ (FTT_1125): composition and average number of surrounding residues, both in codons 105–119 (Fig. 8). These regions occur within α-helices (Fig. S7). Composition refers to the individual amino acid make-up of a protein. Most amino acid residues in a protein cannot be replaced without a change in function (Hormoz 2013). The average number of surrounding residues refers to the number of residues surrounding a particular amino acid within the effective distance of influence, and is an important factor in changing the hydrophobicity index of the region (Ponnuswamy et al. 1980).
Selection on FTT_1125 (metQ). Areas of the gene under selection are indicated where the test statistic is greater than 3.09 (horizontal line) (p < 0.001). Selection is present in the gene for the following amino acid properties where the lines cross the threshold: composition (orange line) and average number of surrounding residues (gray line)
MetQ is a d-Methionine-binding protein, which together with MetN (an ATPase) and MetI (a d-methionine permease) make up the d-methionine ABC transporter (Merlin et al. 2002). Similar to RocE discussed earlier in the metabolism and biosynthesis section, which is an arginine scavenger, this d-methionine transporter scavenges for d-methionine from its environment, since F. tularensis cannot synthesize methionine. Once transported inside the cell, d-methionine can be converted to l-methionine and used in protein synthesis (Santic and Abu Kwaik 2013). This is an important strategy used by F. tularensis to survive intracellularly. The fact that MetQ is experiencing selection implies that F. tularensis is continuing to evolve strategies to acquire nutrients from its environment to survive within a host cell.
Housekeeping Genes
As a control to determine if the virulence genes of F. tularensis are experiencing more selection than other genes in the genome, 10 housekeeping genes (recA, gyrB, groEL, dnaK, rpoA1, rpoB, rpoD, rpoH, fopA, and sdhA) (Qu et al. 2013) were subjected to the same tests of selection (HyPhy and TreeSAAP) as the virulence genes described earlier. Since housekeeping genes are responsible for the fundamental functions of the cell, these are typically highly conserved and are not expected to be evolving at the same rate, or under similar selective pressures as the genes responsible for virulence. Of the 10 housekeeping genes surveyed, only one gene, the molecular chaperone GroEL, was shown to experience significant positive selection (category 7–8), confirming the hypothesis that the virulence genes of F. tularensis are more likely to be undergoing concerted selective pressures. The other nine housekeeping genes analyzed showed no significant positive selection by either the HyPhy or TreeSAAP methods.
We initially expected that all of the housekeeping genes would not show significant selection. However, while GroEL is classified as a housekeeping gene, its function as a molecular chaperone includes responding to stress. In this way, GroEL may be functioning like a virulence gene, experiencing positive selection in much the same way as the other virulence genes.
Conclusions
Francisella tularensis virulence genes failed to recover statistically significant evidence of positive selection using d N/d S ratios. However, subsequent analyses of 11 virulence genes via TreeSAAP (Woolley et al. 2003) identified 7 genes undergoing statistically significant positive selection. The biological functions of the identified mutations are cautiously inferred. However, since these genes appear to be under positive selection, they likely confer some evolutionary advantage leading to an increase in the overall fitness of the organism. The genes undergoing selection participate in a variety of functions, such as membrane transport, host defense evasion, stress response, intracellular survival, and even certain metabolic and biosynthetic pathways. Since the genes we identified as showing evidence of selection are virulence genes, it can be inferred that they confer an adaptive benefit and increase the ability of the organism to infect the host and/or evade host defenses. Although a number of pseudogenes present in F. tularensis genomes indicate that they are in decay (Santic et al. 2010), our findings suggest that they are also undergoing an adaptive response to changes in their intracellular environment by way of positive selection on virulence genes.
References
Alexeev D, Alexeeva M, Baxter RL, Campopiano DJ, Webster SP, Sawyer L (1998) The crystal structure of 8-amino-7-oxononanoate synthase: a bacterial PLP-dependent, acyl-CoA-condensing enzyme. J Mol Biol 284(2):401–419. doi:10.1006/jmbi.1998.2086
Angiuoli SV, Gussman A, Klimke W, Cochrane G, Field D, Garrity G, Kodira CD, Kyrpides N, Madupu R, Markowitz V, Tatusova T, Thomson N, White O (2008) Toward an online repository of standard operating procedures (SOPs) for (meta) genomic annotation. Omics-a Journal of Integrative Biology 12(2):137–141. doi:10.1089/omi.2008.0017
Antwerpen M, Schacht E, Kaysser P, Splettstoesser W (2013) Complete genome sequence of a Francisella tularensis subsp. holarctica strain from Germany causing lethal infection in common marmosets. Genome Announc 1(1):e00135-12
Barabote RD, Xie G, Brettin TS, Hinrichs SH, Fey PD, Jay JJ, Engle JL, Godbole SD, Noronha JM, Scheuermann RH, Zhou LW, Lion C, Dempsey MP (2009) Complete genome sequence of Francisella tularensis subspecies holarctica FTNF002-00. Plos One. doi:10.1371/journal.pone.0007041
Beckstrom-Sternberg SM, Auerbach RK, Godbole S, Pearson JV, Beckstrom-Sternberg JS, Deng ZM, Munk C, Kubota K, Zhou Y, Bruce D, Noronha J, Scheuermann RH, Wang AH, Hao J, Wang JJ, Wagner DM, Brettin TS, Brown N, Gilna P, Keim PS (2007) Complete genomic characterization of a pathogenic A.II strain of Francisella tularensis subspecies tularensis. Plos One. doi:10.1371/journal.pone.0000947
Broekhuijsen M, Larsson P, Johansson A, Bystrom M, Eriksson U, Larsson E, Prior R, Sjostedt A, Titball RW, Forsman M (2003) Genome-wide DNA microarray analysis of Francisella tularensis strains demonstrates extensive genetic conservation within the species but identifies regions that are unique to the highly virulent F. tularensis subsp. tularensis. J Clin Microbiol 41(7):2924–2931
Brown NF, Wickham ME, Coombes BK, Finlay BB (2006) Crossing the line: selection and evolution of virulence traits. PLoS Pathog 2(5):e42. doi:10.1371/journal.ppat.0020042
Casadevall A, Pirofski LA (1999) Host-pathogen interactions: redefining the basic concepts of virulence and pathogenicity. Infect Immun 67(8):3703–3713
Champion MD, Zeng QD, Nix EB, Nano FE, Keim P, Kodira CD, Borowsky M, Young S, Koehrsen M, Engels R, Pearson M, Howarth C, Larson L, White J, Alvarado L, Forsman M, Bearden SW, Sjostedt A, Titball R, Michell SL, Birren B, Galagan J (2009) Comparative genomic characterization of Francisella tularensis strains belonging to low and high virulence subspecies. Plos Pathogens. doi:10.1371/journal.ppat.1000459
Chaudhuri RR, Ren CP, Desmond L, Vincent GA, Silman NJ, Brehm JK, Elmore MJ, Hudson MJ, Forsman M, Isherwood KE, Gurycova D, Minton NP, Titball RW, Pallen MJ, Vipond R (2007) Genome sequencing shows that European isolates of Francisella tularensis subspecies tularensis are almost identical to US laboratory strain Schu S4. Plos One. doi:10.1371/journal.pone.0000352
Chiang SL, Mekalanos JJ, Holden DW (1999) In vivo genetic analysis of bacterial virulence. Annu Rev Microbiol 53:129–154. doi:10.1146/annurev.micro.53.1.129
Choi JS, Park NJ, Lim HK, Ko YK, Kim YS, Ryu SY, Hwang IT (2012) Plumbagin as a new natural herbicide candidate for Sicyon angulatus control agent with the target 8-amino-7-oxononanoate synthase. Pestic Biochem Physiol 103(3):166–172. doi:10.1016/j.pestbp.2012.04.007
Chou PY, Fasman GD (1978) Empirical predictions of protein conformation. Annu Rev Biochem 47:251–276. doi:10.1146/annurev.bi.47.070178.001343
Crandall KA, Kelsey CR, Imamichi H, Lane HC, Salzman NP (1999) Parallel evolution of drug resistance in HIV: failure of nonsynonymous/synonymous substitution rate ratio to detect selection. Mol Biol Evol 16(3):372–382
Demuth JP, Hahn MW (2009) The life and death of gene families. BioEssays 31(1):29–39. doi:10.1002/bies.080085
Dhanasekaran M, Negi S, Imanishi M, Suzuki M, Sugiura Y (2008) Effects of bulkiness and hydrophobicity of an aliphatic amino acid in the recognition helix of the GAGA zinc finger on the stability of the hydrophobic core and DNA binding affinity. Biochemistry 47(45):11717–11724. doi:10.1021/bi801306d
El-Etr SH, Margolis JJ, Monack D, Robison RA, Cohen M, Moore E, Rasley A (2009) Francisella tularensis Type A strains cause the rapid encystment of Acanthamoeba castellanii and survive in amoebal cysts for three weeks postinfection. Appl Environ Microbiol 75(23):7488–7500. doi:10.1128/aem.01829-09
Fay JC (2011) Weighing the evidence for adaptation at the molecular level. Trends Genet 27(9):343–349. doi:10.1016/j.tig.2011.06.003
Feng Y, Chen Z, Liu SL (2011) Gene decay in Shigella as an incipient stage of host-adaptation. Plos One. doi:10.1371/journal.pone.0027754
Fraser ME, James MNG, Bridger WA, Wolodko WT (1999) A detailed structural description of Escherichia coli succinyl-CoA synthetase. J Mol Biol 285(4):1633–1653. doi:10.1006/jmbi.1998.2324
Gallagher LA, McKevitt M, Ramage ER, Manoil C (2008) Genetic dissection of the Francisella novicida restriction barrier. J Bacteriol 190(23):7830–7837. doi:10.1128/jb.01188-08
George MG (2005) In: David RB, Richard WC, George MG, Don JB, Noel RK, James TS (eds) Bergey’s Manual® of Systematic Bacteriology. Springer, Germany.
Gestin B, Valade E, Thibault F, Schneider D, Maurin M (2010) Phenotypic and genetic characterization of macrolide resistance in Francisella tularensis subsp. holarctica biovar I. J Antimicrob Chemother 65(11):2359–2367. doi:10.1093/jac/dkq315
Grenha R, Levdikov VM, Fogg MJ, Blagova EV, Brannigan JA, Wilkinson AJ, Wilson KS (2005) Structure of purine nucleoside phosphorylase (DeoD) from Bacillus anthracis. Acta Crystallogr Sect F 61:459–462. doi:10.1107/s174430910501095x
Gunnell MK, Lovelace CD, Satterfield BA, Moore EA, O’Neill KL, Robison RA (2012) A multiplex real-time PCR assay for the detection and differentiation of Francisella tularensis subspecies. J Med Microbiol 61(11):1525–1531. doi:10.1099/jmm.0.046631-0
Hain T, Chatterjee SS, Ghaia R, Kuenne CT, Billion A, Steinweg C, Domann E, Karst U, Jansch L, Wehland J, Eisenreich W, Bacherc A, Joseph B, Schar J, Kreft J, Klumpp J, Loessner MJ, Dorscht J, Neuhaus K, Fuchs TM, Scherer S, Doumith M, Jacquet C, Martin P, Cossart P, Rusniock C, Glaser P, Buchrieser C, Goebel W, Chakraborty T (2007) Pathogenomics of Listeria spp. Int J Med Microbiol 297(7–8):541–557. doi:10.1016/j.ijmm.2007.03.016
Harris SR, Robinson C, Steward KF, Webb KS, Paillot R, Parkhill J, Holden MTG, Waller AS (2015) Genome specialization and decay of the strangles pathogen, Streptococcus equi, is driven by persistent infection. Genome Res 25(9):1360–1371. doi:10.1101/gr.189803.115
Hasegawa M, Kishino H, Yano TA (1985) Dating of the human ape splitting by a molecular clock of mitochondrial-DNA. J Mol Evol 22(2):160–174. doi:10.1007/bf02101694
Hensel M, Shea JE, Gleeson C, Jones MD, Dalton E, Holden DW (1995) Simultaneous identification of bacterial virulence genes by negative selection. Science 269(5222):400–403. doi:10.1126/science.7618105
Hormoz S (2013) Amino acid composition of proteins reduces deleterious impact of mutations. Scientific Reports. doi:10.1038/srep02919
Johansson A, Farlow J, Larsson P, Dukerich M, Chambers E, Bystrom M, Fox J, Chu M, Forsman M, Sjostedt A, Keim P (2004) Worldwide genetic relationships among Francisella tularensis isolates determined by multiple-locus variable-number tandem repeat analysis. J Bacteriol 186(17):5808–5818
Jones DT (1999) Protein secondary structure prediction based on position-specific scoring matrices. J Mol Biol 292(2):195–202. doi:10.1006/jmbi.1999.3091
Jones CL, Napier BA, Sampson TR, Llewellyn AC, Schroeder MR, Weiss DS (2012) Subversion of host recognition and defense systems by Francisella spp. Microbiol Mol Biol Rev 76(2):383–404. doi:10.1128/mmbr.05027-11
Kaper JB (2005) Pathogenic Escherichia coli. Int J Med Microbiol 295(6–7):355–356. doi:10.1016/j.ijmm.2005.06.008
Keim P, Johansson A, Wagner DM (2007) Molecular epidemiology, evolution, and ecology of Francisella. Francisella Tularensis: Biology, Pathogenicity, Epidemiology, and Biodefense 1105:30–66. doi:10.1196/annals.1409.011
Kerbarh O, Campopiano DJ, Baxter RL (2006) Mechanism of alpha-oxoamine synthases: identification of the intermediate Claisen product in the 8-amino-7-oxononanoate synthase reaction. Chem Commun 1:60–62. doi:10.1039/b511837a
Kone BC, Kuncewicz T, Zhang WZ, Yu ZY (2003) Protein interactions with nitric oxide synthases: controlling the right time, the right place, and the right amount of nitric oxide. American Journal of Physiology-Renal Physiology 285(2):F178–F190. doi:10.1152/ajprenal.00048.2003
Lan T, Yang ZL, Yang X, Liu YJ, Wang XR, Zeng QY (2009) Extensive functional diversification of the Populus glutathione S-transferase supergene family. Plant Cell 21(12):3749–3766. doi:10.1105/tpc.109.070219
Larkin MA, Blackshields G, Brown NP, Chenna R, McGettigan PA, McWilliam H, Valentin F, Wallace IM, Wilm A, Lopez R, Thompson JD, Gibson TJ, Higgins DG (2007) Clustal W and clustal X version 2.0. Bioinformatics 23(21):2947–2948. doi:10.1093/bioinformatics/btm404
Larsson P, Oyston PCF, Chain P, Chu MC, Duffield M, Fuxelius HH, Garcia E, Halltorp G, Johansson D, Isherwood KE, Karp PD, Larsson E, Liu Y, Michell S, Prior J, Prior R, Malfatti S, Sjostedt A, Svensson K, Thompson N, Vergez L, Wagg JK, Wren BW, Lindler LE, Andersson SGE, Forsman M, Titball RW (2005) The complete genome sequence of Francisella tularensis, the causative agent of tularemia. Nat Genet 37(2):153–159. doi:10.1038/ng1499
Lory S, Strom MS (1997) Structure-function relationship of type-IV prepilin peptidase of Pseudomonas aeruginosa - A review. Gene 192(1):117–121. doi:10.1016/s0378-1119(96)00830-x
Luo W, Liu YP, Zhu XC, Zhao WJ, Huang L, Cai J, Xu ZN, Cen PL (2011) Cloning and characterization of purine nucleoside phosphorylase in Escherichia coli and subsequent ribavirin biosynthesis using immobilized recombinant cells. Enzyme and Microbial Technology 48(6–7):438–444. doi:10.1016/j.enzmictec.2011.03.008
Masri L, Branca A, Sheppard AE, Papkou A, Laehnemann D, Guenther PS, Prahl S, Saebelfeld M, Hollensteiner J, Liesegang H, Brzuszkiewicz E, Daniel R, Michiels NK, Schulte RD, Kurtz J, Rosenstiel P, Telschow A, Bornberg-Bauer E, Schulenburg H (2015) Host-pathogen coevolution: the selective advantage of Bacillus thuringiensis virulence and its cry toxin genes. PLoS Biol 13(6):e1002169. doi:10.1371/journal.pbio.1002169
McClellan D, Ellison D (2010) Assessing and improving the accuracy of detecting protein adaptation with the TreeSAAP analytical software. Int J Bioinform Res Appl 6(2):120–133
McDonald JH, Kreitman M (1991) Adaptive protein evolution at the Adh locus in Drosophila. Nature 351(6328):652–654. doi:10.1038/351652a0
Merlin C, Gardiner G, Durand S, Masters M (2002) The Escherichia coli metD locus encodes an ABC transporter which includes Abc (MetN), YaeE (MetI), and YaeC (MetQ). J Bacteriol 184(19):5513–5517. doi:10.1128/jb.184.19.5513-5517.2002
Messer PW, Petrov DA (2013) Frequent adaptation and the McDonald–Kreitman test. Proc Natl Acad Sci USA 110(21):8615–8620. doi:10.1073/pnas.1220835110
Modise T, Ryder C, Mane SP, Bandara AB, Jensen RV, Inzana TJ (2012) Genomic comparison between a virulent type A1 strain of Francisella tularensis and its attenuated O-antigen mutant. J Bacteriol 194(10):2775-2776. doi:10.1128/JB.00152-12
Muse SV, Gaut BS (1994) A likelihood approach for comparing synonymous and nonsynonymous nucleotide substitution rates, with application to the chloroplast genome. Mol Biol Evol 11(5):715–724
Nalbantoglu U, Sayood K, Dempsey MP, Iwen PC, Francesconi SC, Barbote RD, Xie G, Brettin TS, Hinrichs SH, Fey PD (2010) Large direct repeats flank genomic rearrangements between a new clinical isolate of Francisella tularensis subsp. tularensis A1 and Schu S4. PLoS One 5(2):e9007. doi:10.1371/journal.pone.0009007
Nandi T, Ong C, Singh AP, Boddey J, Atkins T, Sarkar-Tyson M, Essex-Lopresti AE, Chua HH, Pearson T, Kreisberg JF, Nilsson C, Ariyaratne P, Ronning C, Losada L, Ruan Y, Sung WK, Woods D, Titball RW, Beacham I, Peak I, Keim P, Nierman WC, Tan P (2010) A genomic survey of positive selection in Burkholderia pseudomallei provides insights into the evolution of accidental virulence. PLoS Pathog 6(4):e1000845. doi:10.1371/journal.ppat.1000845
Nei M, Kumar S (2000) Molecular evolution and phylogenetics. Oxford University Press, New York
Nudelman A, Marcovici-Mizrahi D, Nudelman A, Flint D, Wittenbach V (2004) Inhibitors of biotin biosynthesis as potential herbicides. Tetrahedron 60(8):1731–1748. doi:10.1016/j.tet.2003.12.047
Owen CR, Lackman DB, Jellison WL, Buker EO, Bell JF (1964) Comparative studies of Francisella tularensis + Francisella novicida. J Bacteriol 87(3):000–676
Oyston PCF (2008) Francisella tularensis: unravelling the secrets of an intracellular pathogen. J Med Microbiol 57(8):921–930. doi:10.1099/jmm.0.2008/000653-0
Petrosino JF, Xiang Q, Karpathy SE, Jiang HY, Yerrapragada S, Liu YM, Gioia J, Hemphill L, Gonzalez A, Raghavan TM, Uzman A, Fox GE, Highlander S, Reichard M, Morton RJ, Clinkenbeard KD, Weinstock GM (2006) Chromosome rearrangement and diversification of Francisella tularensis revealed by the type B (OSU18) genome sequence. J Bacteriol 188(19):6977–6985. doi:10.1128/jb.00506-06
Pond SLK, Frost SDW (2005) Not so different after all: a comparison of methods for detecting amino acid sites under selection. Mol Biol Evol 22(5):1208–1222. doi:10.1093/molbev/msi105
Pond SLK, Frost SDW, Muse SV (2005) HyPhy: hypothesis testing using phylogenies. Bioinformatics 21(5):676–679. doi:10.1093/bioinformatics/bti079
Ponnuswamy PK, Prabhakaran M, Manavalan P (1980) Hydrophobic packing and spatial arrangement of amino-acid-residues in globular-proteins. Biochim Biophys Acta 623(2):301–316. doi:10.1016/0005-2795(80)90258-5
Prahhakaran M, Ponnuswamy PK (1979) The spatial distribution of physical, chemical, energetic and conformational properties of amino acid residues in globular proteins. J Theor Biol 80(4):485–504. doi:10.1016/0022-5193(79)90090-0
Qu P-H, Chen S-Y, Scholz HC, Busse H-J, Gu Q, Kaempfer P, Foster JT, Glaeser SP, Chen C, Yang Z-C (2013) Francisella guangzhouensis sp nov., isolated from air-conditioning systems. Int J Syst Evol Microbiol 63:3628–3635. doi:10.1099/ijs.0.049916-0
Read AF (1994) The evolution of virulence. Trends Microbiol 2(3):73–76
Rohmer L, Fong C, Abmayr S, Wasnick M, Freeman TJL, Radey M, Guina T, Svensson K, Hayden HS, Jacobs M, Gallagher LA, Manoil C, Ernst RK, Drees B, Buckley D, Haugen E, Bovee D, Zhou Y, Chang J, Levy R, Lim R, Gillett W, Guenthener D, Kang A, Shaffer SA, Taylor G, Chen JZ, Gallis B, D’Argenio DA, Forsman M, Olson MV, Goodlett DR, Kaul R, Miller SI, Brittnacher MJ (2007) Comparison of Francisella tularensis genomes reveals evolutionary events associated with the emergence of human pathogenic strains. Genome Biology. doi:10.1186/gb-2007-8-6-r102
Santic M, Abu Kwaik Y (2013) Nutritional virulence of Francisella tularensis. Frontiers in Cellular and Infection Microbiology. doi:10.3389/fcimb.2013.00112
Santic M, Al-Khodor S, Abu Kwaik Y (2010) Cell biology and molecular ecology of Francisella tularensis. Cell Microbiol 12(2):129–139. doi:10.1111/j.1462-5822.2009.01400.x
Schunder E, Rydzewski K, Grunow R, Heuner K (2013) First indication for a functional CRISPR/Cas system in Francisella tularensis. Int J Med Microbiol 303(2):51–60. doi:10.1016/j.ijmm.2012.11.004
Sengupta D, Kundu S (2012) Do topological parameters of amino acids within protein contact networks depend on their physico-chemical properties? Physica A 391(17):4266–4278. doi:10.1016/j.physa.2012.03.034
Sharp PM (1997) In search of molecular darwinism. Nature 385(6612):111–112. doi:10.1038/385111a0
Siddaramappa S, Challacombe JF, Petersen JM, Pillai S, Kuske CR (2012) Genetic diversity within the genus Francisella as revealed by comparative analyses of the genomes of two North American isolates from environmental sources. Bmc Genomics. doi:10.1186/1471-2164-13-422
Sjostedt A (2007) Tularemia: history, epidemiology, pathogen physiology, and clinical manifestations. Francisella Tularensis: Biology, Pathogenicity, Epidemiology, and Biodefense 1105:1–29. doi:10.1196/annals.1409.009
Sridhar S, Sharma A, Kongshaug H, Nilsen F, Jonassen I (2012) Whole genome sequencing of the fish pathogen Francisella noatunensis subsp orientalis Toba04 gives novel insights into Francisella evolution and pathogenecity. Bmc Genomics. doi:10.1186/1471-2164-13-598
Stancik LM, Stancik DM, Schmidt B, Barnhart DM, Yoncheva YN, Slonczewski JL (2002) pH-dependent expression of periplasmic proteins and amino acid catabolism in Escherichia coli. J Bacteriol 184(15):4246–4258. doi:10.1128/jb.184.15.4246-4258.2002
Staples JE, Kubota KA, Chalcraft LG, Mead PS, Petersen JM (2006) Epidemiologic and molecular analysis of human tularemia, United States, 1964–2004. Emerg Infect Dis 12(7):1113–1118
Steele S, Brunton J, Ziehr B, Taft-Benz S, Moorman N, Kawula T (2013) Francisella tularensis harvests nutrients derived via ATG5-independent autophagy to support intracellular growth. Plos Pathogens. doi:10.1371/journal.ppat.1003562
Strom MS, Lory S (1993) Structure-function and biogenesis of the type-IV pili. Annu Rev Microbiol 47:565–596. doi:10.1146/annurev.micro.47.1.565
Strom MS, Nunn DN, Lory S (1993) A single bifunctional enzyme, PilD, catalyzes cleavage and N-methylation of proteins belonging to the type-IV pilin family. Proc Natl Acad Sci USA 90(6):2404–2408. doi:10.1073/pnas.90.6.2404
Su J, Yang J, Zhao D, Kawula TH, Banas JA, Zhang J-R (2007) Genome-wide identification of Francisella tularensis virulence determinants. Infect Immun 75(6):3089–3101. doi:10.1128/iai.01865-06
Sutera V, Levert M, Burmeister WP, Schneider D, Maurin M (2014) Evolution toward high-level fluoroquinolone resistance in Francisella species. J Antimicrob Chemother 69(1):101–110. doi:10.1093/jac/dkt321
Suzuki Y, Gojobori T (1999) A method for detecting positive selection at single amino acid sites. Mol Biol Evol 16(10):1315–1328
Svensson K, Larsson P, Johansson D, Bystrom M, Forsman M, Johansson A (2005) Evolution of subspecies of Francisella tularensis. J Bacteriol 187(11):3903–3908
Svensson K, Sjodin A, Bystrom M, Granberg M, Brittnacher MJ, Rohmer L, Jacobs MA, Sims-Day EH, Levy R, Zhou Y, Hayden HS, Lim R, Chang J, Guenthener D, Kang A, Haugen E, Gillett W, Kaul R, Forsman M, Larsson P, Johansson A (2012) Genome sequence of Francisella tularensis subspecies holarctica strain FSC200, isolated from a child with tularemia. J Bacteriol 194(24):6965–6966. doi:10.1128/jb.01040-12
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28(10):2731–2739. doi:10.1093/molbev/msr121
Taylor SD, de la Cruz KD, Porter ML, Whiting MF (2005) Characterization of the long-wavelength opsin from Mecoptera and Siphonaptera: does a flea see? Mol Biol Evol 22(5):1165–1174. doi:10.1093/molbev/msi110
Walshaw DL, Wilkinson A, Mundy M, Smith M, Poole PS (1997) Regulation of the TCA cycle and the general amino acid permease by overflow metabolism in Rhizobium leguminosarum. Microbiology-Uk 143:2209–2221
Ward PN, Holden MTG, Leigh JA, Lennard N, Bignell A, Barron A, Clark L, Quail MA, Woodward J, Barrell BG, Egan SA, Field TR, Maskell D, Kehoe M, Dowson CG, Chanter N, Whatmore AM, Bentley SD, Parkhill J (2009) Evidence for niche adaptation in the genome of the bovine pathogen Streptococcus uberis. Bmc Genomics. doi:10.1186/1471-2164-10-54
Weiss DS, Brotcke A, Henry T, Margolis JJ, Chan K, Monack DM (2007) In vivo negative selection screen identifies genes required for Francisella virulence. Proc Natl Acad Sci USA 104(14):6037–6042. doi:10.1073/pnas.0609675104
Wolodko WT, Fraser ME, James MNG, Bridger WA (1994) The crystal-structure of cussinyl-CoA synthetase from Escherichia coli at 2.5 angstrom resolution. J Biol Chem 269(14):10883–10890
Woolley S, Johnson J, Smith MJ, Crandall KA, McClellan DA (2003) TreeSAAP: selection on amino acid properties using phylogenetic trees. Bioinformatics 19(5):671–672. doi:10.1093/bioinformatics/btg043
Acknowledgments
We thank the Utah Department of Health and the New Mexico Department of Health for help in obtaining the isolates used in this study, and for the thoughtful and constructive criticisms of the anonymous reviewers. Dr. Angelo Madonna of Dugway Proving Ground provided expert guidance and assistance in completing this work.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of Interest
The authors declare that they have no conflicts of interest.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Gunnell, M.K., Robison, R.A. & Adams, B.J. Natural Selection in Virulence Genes of Francisella tularensis . J Mol Evol 82, 264–278 (2016). https://doi.org/10.1007/s00239-016-9743-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-016-9743-y