Abstract
Most members of Siboglinidae (Annelida) harbor endosymbiotic bacteria that allow them to thrive in extreme environments such as hydrothermal vents, methane seeps, and whale bones. These symbioses are enabled by specialized hemoglobins (Hbs) that are able to bind hydrogen sulfide for transportation to their chemosynthetic endosymbionts. Sulfur-binding capabilities are hypothesized to be due to cysteine residues at key positions in both vascular and coelomic Hbs, especially in the A2 and B2 chains. Members of the genus Osedax, which live on whale bones, do not have chemosynthetic endosymbionts, but instead harbor heterotrophic bacteria capable of breaking down complex organic compounds. Although sulfur-binding capabilities are important in other siboglinids, we questioned whether Osedax retained these cysteine residues and the potential ability to bind hydrogen sulfide. To answer these questions, we used high-throughput DNA sequencing to isolate and analyze Hb sequences from 8 siboglinid lineages. For Osedax mucofloris, we recovered three (A1, A2, and B1) Hb chains, but the B2 chain was not identified. Hb sequences from gene subfamilies A2 and B2 were translated and aligned to determine conservation of cysteine residues at previously identified key positions. Hb linker sequences were also compared to determine similarity between Osedax and siboglinids/sulfur-tolerant annelids. For O. mucofloris, our results found conserved cysteines within the Hb A2 chain. This finding suggests that Hb in O. mucofloris has retained some capacity to bind hydrogen sulfide, likely due to the need to detoxify this chemical compound that is abundantly produced within whale bones.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Siboglinid annelids occur throughout the world’s oceans but are best known from hydrothermal vents, cold seeps, and whale bones (Schulze and Halanych 2003; Rouse et al. 2004; Southward et al. 2005). Chemotrophic endosymbiotic bacteria enable these worms to thrive in these extreme environments (Cavanaugh and Gardiner 1981; Southward and Southward 1981; Halanych 2005; Goffredi et al. 2005; Thornhill et al. 2008). Siboglinidae is comprised of four lineages: frenulates, vestimentiferans, monoliferans, and Osedax (Hilário et al. 2011). Frenulates, comprising the majority of known siboglinid species, are often thread-like and found within sediments of reducing habitats (Southward 1978; Southward et al. 2005; Thornhill et al. 2008; Hilário et al. 2010). Vestimentiferans, on the other hand, are large tubeworms that are typically found in hydrothermal vents and cool seeps (McMullin et al. 2003). Monilifera is represented by a single genus (i.e., Sclerolinum) that shares similarities to frenulates in terms of size and preferred habitat, but can also be found on decaying organic material (Halanych et al. 2001). Finally, Osedax, first described in 2004, are worms that colonize whale bones (Rouse et al. 2004; Glover et al. 2005).
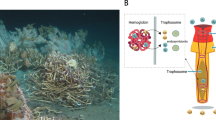
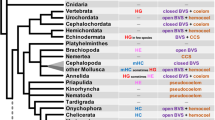
Adult siboglinids lack a functional gut and instead rely on chemosynthetic endosymbionts to supply some or all of their energetic needs (Cavanaugh and Gardiner 1981; Hilário et al. 2011). In this context, hydrogen sulfide (H2S) is absorbed and transported via the blood vascular system to symbiotic bacteria within a specialized organ called the trophosome (Southward 1988; Goffredi et al. 2005; Katz et al. 2011; Bright et al. 2012). Most siboglinid endosymbionts are chemoautotrophic and generally belong to the gammaproteobacteria (Thornhill et al. 2008; Verna et al. 2010). Osedax, whose morphology is more arborescent in appearance, harbor heterotrophic endosymbionts (Oceanospirillales, Gammaproteobacteria) in a root-like system that extends into the whale bone matrix (Goffredi et al. 2005) where endosymbionts utilize the complex compounds released from the bones (Rouse et al. 2004). Approximately 31 lineages of Osedax have been discovered (Smith et al. 2015) and phylogenetic analyses based on ribosomal genes and mitochondrial cytochrome oxidase I usually place Osedax as sister to a moniliferan–vestimentiferan clade (Rouse et al. 2004; Glover et al. 2005), but Glover et al. (2013) and Rouse et al. (2015) suggest a position sister to frenulate siboglinids. Despite this suggestion, recent analyses of whole mitochondrial genome data strongly favor allying Osedax with vestimentiferans and monoliferans (Li et al. 2015; Fig. 1).
Current hypothesized phylogeny of Siboglinidae based on Li et al. (2015, Fig. 3a). Majority rule (50 %) consensus topology of a Bayesian analysis of mitochondrial genome data is shown. Values are shown next to nodes with posterior probabilities left and ML bootstrap support values right. Filled circles indicate fully supported nodes (bs = 100, pp = 1.00). Additional analytical details found in Li et al. (2015)
For some chemoautotroph-bearing siboglinids, H2S uptake and transport is mediated by specialized hemoglobins (Hbs) (Numoto et al. 2005; Meunier et al. 2010). Reversible binding of H2S to Hbs has been best studied in the vestimentiferans Riftia pachyptila and Lamellibrachia luymesi, as well as the frenulate Oligobrachia mashikoi (e.g., Suzuki et al. 1990; Yuasa et al. 1996; Zal et al. 1996a, b, 1997). Hbs are complex structures with individual globin chains assembling into hetero-dimer subunits. Those subunits, in turn, assemble into a tetrameric functional protein, with each heme directly interacting with adjoining subunits whose size varies (Numoto et al. 2008). Vestimentiferans have one large extracellular Hb (V1 ~ 3500 kDa) and one small extracellular Hb (V2: ~400 kDa) in their vascular blood. Additionally, they possess one Hb (C1) in coelomic fluid that is reported to be 400 kDa (Arp and Childress 1981; Zal et al. 1996a). Whereas V1 contains 4 heme-containing globin chains (b–e) and 4 linker chains (L1–L4), V2 is composed of 6 globin chains (a–f), and C1 contains 5 globin chains (a–e). In contrast, the frenulate O. mashikoi possesses a single ~400 kDA Hb composed of 24 globin chains with no linkers, comparable to the small extracellular Hbs of vestimentiferans (Yuasa et al. 1996; Numoto et al. 2005). Binding of H2S has been hypothesized to be mediated, in part, by cysteine residues in the V1 chains and by disulfide bridges formed from cysteine-rich linker chains (R. pachyptila’s V1 chain b—B2 and L. luymesi’s V1 chain AIII—A2; Zal et al. 1996b, 1997). However, this only accounts for part of the binding affinity, and zinc moieties bound to amino acid residues at the interface between pairs of A2 chains may also be involved (Flores et al. 2005). With reference to R. pachyptila’s A2 chain, cysteines at positions 4 and 134 are common to all annelid globin chains studied and form a disulfide bridge while a free cysteine at position 75 is unique to sulfur oxidizing siboglinids (Zal et al. 1997).
Given our understanding of siboglinid phylogeny (Li et al. 2015), the bone-eating Osedax has likely evolved from ancestors dependent upon chemoautotrophic bacteria (Schulze and Halanych 2003; Hilário et al. 2011) at least 100 million years ago (based on fossil and molecular data; Danise and Higgs 2015). Due to its heterotrophic symbiosis, Osedax is apparently no longer dependent on H2S transport or the modified blood physiology to nourish endosymbionts (Rouse et al. 2004; Goffredi et al. 2005). We assume the ability to bind H2S carries a cost to the organism, as most Hbs lack such affinity and may be selected against in sulfide-free habitats (Bailly et al. 2003). Based on this, we hypothesized that the Osedax Hb system would exhibit differences relative to other siboglinids; specifically, amino acid substitutions for carrying H2S should be lacking in Osedax. To avoid a PCR-based approach that would require multiple primers and attempts to isolate single genes, and because Hbs are ubiquitously expressed in the blood vascular system of siboglinids, we employed high-throughput DNA sequencing to generate transcriptomic data. This methodology allowed examination of amino acid sequences of Hbs and linker proteins from O. mucofloris, three frenulates, a moniliferan, and three vestimentiferans, in addition to publically available data. Specific targets were the level of conservation among Cyt residues (especially at positions 4, 75, and 134) in Hb chains across siboglinids as well as conceptually examining how amino acid differences may influence protein-folding characteristics.
Materials and Methods
Siboglinid Sampling
Siboglinid samples were procured for transcriptome sequencing from a variety of sources (Table 1). Specifically, Christoffer Schander kindly provided O. mucofloris from whale bones near Bergen, Norway, and Sclerolinum contortum from the Håkons-Mosby mud volcano off Norway. Samples of Lamellibrachia luymesi, Escarpia spicata, Seepiophila jonesi, and Galathealinum brachiosum were collected in the Gulf of Mexico using the Johnson Sea Link submersible aboard the R/V Seward Johnson. Samples of Siboglinum fiordicum were obtained using a small hand grab on the R/V Aurelia (University of Bergen) and Siboglinum ekmani were obtained by dredge on the R/V Håkons-Mosby from near Bergen, Norway. At the time of collection, all samples were morphologically identified and stored in RNALater.
Extraction and Sequencing
RNA extraction and cDNA preparation for high-throughput sequencing followed Kocot et al. (2011) and Li et al. (2015). Briefly, RNA was extracted using a TRIzol (Invitrogen) protocol, and then purified with the RNeasy kit (Qiagen) using an on-column digestion. Next, single-strand cDNA libraries were reverse transcribed using the SMART cDNA Library Construction kit (Clontech) followed by double-stranded cDNA synthesis using the Advantage 2 PCR system (Clontech). The double-stranded cDNA from O. mucofloris was sequenced on an Illumina MiSeq sequencer at Auburn University using a Nextera (Illumina) protocol, as well as an Illumina HiSeq 2000 sequencer at the Genomics Services Laboratory at the Hudson Alpha Institute for Biotechnology (Huntsville, AL, USA) using the TruSeq v3 (Illumina) protocol. cDNA for Escarpia spicata, G. brachiosum, L. luymesi, and S. jonesi were sent to the University of South Carolina Environmental Genomics Core Facility (Columbia, SC, USA) for Roche 454 GS-FLX sequencing. Additionally, cDNAs for L. luymesi, S. contortum, S. ekmani, and S. fiordicum were sequenced on an Illumina HiSeq 2000 sequencer at Hudson Alpha Institute for Biotechnology.
Sequence Assembly
Sequencing reads were digitally normalized using the normalize-by-median script in the khmer package (https://github.com/ctb/khmer/blob/master/scripts/normalize-by-median.py) to facilitate assembly and decrease the likelihood that overrepresentation of reads would cause assembly artifacts (McDonald and Brown 2013). Transcriptome assemblies from MiSeq and 454 data were done de novo with the October 2012 release of Trinity (Grabherr et al. 2011), while HiSeq 2000 data were assembled with the February 2013 release of the same software. For O. mucofloris and L. luymesi, cDNA was run on two different platforms. In these cases, data were assembled separately and each searched for genes of interest.
BLAST and Sequence Alignment
Hb and linker sequences of interest were obtained from assembled transcriptomes via BLAST (Altschul et al. 1990) by utilizing Hb and linker sequences acquired from GenBank of siboglinids as well as outgroup organisms as queries (Table 2). Specifically, an e value cutoff of 10−5 was utilized in tblastn searches of nucleotide assemblies with the query protein sequences. Arenicola marina, a sulfur-tolerant polychaete, was used as outgroup based on the availability of these sequences. Resulting BLAST hits were filtered using blast2table.pl (available from http://www.genome.ou.edu/informatics.html) with the “top” option, which reports only the best, high-scoring segment pair for each query sequence. Linker sequence hits were manually evaluated based on e value and percent identity to determine similarity. The resulting Hb hits were translated using ESTScan version 3.0.3 (Iseli et al. 1999) and sequences aligned using MUSCLE (Edgar 2004) within MEGA 5.2 (Tamura et al. 2011). The alignment was visually inspected and spuriously aligned data removed based on sequence similarity to the alignment as a whole.
Gene Tree and 3D Structure Prediction of Data
Following alignments, we focused on the A2 Hb because enough sequences were recovered for Osedax and other siboglinids to allow meaningful comparisons. A2 Hb sequences were manually trimmed of missing leading, and trailing positions and Gblocks version 0.91b (Castresana 2000; Talavera and Castresana 2007) was used to trim poorly aligned positions and divergent regions with the following parameters: minimum number of sequences for a conserved position = 7, minimum number of sequences for a flank position = 7, maximum number of contiguous non-conserved positions = 8, minimum length of a block = 2, and gap positions allowed in all blocks. An appropriate amino acid substitution model for phylogenetic reconstruction was selected using Prottest version 3.4 (Darriba et al. 2011). RAxML version 7.3.8 (Stamatakis 2014) was used to infer a maximum-likelihood gene tree with 100 bootstrap replicates using the PROTGAMMAWAG model, with A. marina serving as the outgroup. Osedax mucofloris, Lamellibrachia luymesi, Siboglinum ekmani, Arenicola marina, and Sabella spallanzanii Hb chain A2 structures were predicted as 3D models using the I-TASSER structure prediction server (Yang et al. 2015).
Results
Sequencing Results
High-throughput DNA sequencing produced 283,594–750,876 reads for 454, 3,027,776 reads for MiSeq, and 21,397,136–56,067,578 reads for HiSeq 2000 (Table 1). Contigs per assemblies were 7209–12,080 for 454 data, and 17,617–270,658 for MiSeq and HiSeq 2000 data (Table 1).
BLAST Results
Across the eight transcriptomes, tblastn searches returned 12 top hits (e value cutoff of 10−5) for chain A1, 17 for chain A2, 22 for chain B1, and 12 for chain B2. Upon closer inspection, the singular hit to Osedax mucofloris for chain B2 was a contig that also was returned in searches for chain A2 homologs, and the B2 hit was discarded based on the higher strength match to the A2 hit. These top hits were combined with data acquired from NCBI’s GenBank (Table 2) to generate alignments for each of the four Hb chains. After manual removal of redundant and incorrect sequences, a single contig for each chain was retained per taxon. However, after inspection of the alignment, A1 sequences were not recovered for Escarpia spicata and Galathealinum brachiosum. Additionally, the B2 sequence of Siboglinum ekmani had a single stop codon within the protein-coding region. This sequence was further verified via read mapping with Bowtie 2 (Langmead et al. 2009). Furthermore, the sequence aligned well, but since it was not full length, it was not further considered. All contigs recovered contained complete genes except for E. spicata A2 and B2, Seepiophila jonesi B2, S. ekmani A1, B1, and B2, and all Siboglinum fiordicum contigs.
As linkers aid formation of Hb hexagonal bilayer structure, we also examined Osedax linkers to determine if they are similar to those from vestimentiferans. The tblastn searches for linker sequences resulted in multiple hits for each species. The 454 assemblies of E. spicata, G. brachiosum, S. jonesi, and L. luymesi had relatively few hits at 5, 6, 9, and 18 hits, respectively. Illumina assemblies had higher numbers of hits, with 23 for O. mucofloris, 44 for S. ekmani, 47 for S. fiordicum, 75 for L. luymesi, and 118 for S. brattstromi. Upon manual inspection of each taxon’s BLAST scores, all 8 transcriptomes were found to have an on-average higher score, e value, and percent identity for hits to vestimentiferan linkers than to non-siboglinid linkers (Table 3). Linker sequences showed considerable variation limiting alignment and the ability to produce a meaningful gene tree.
Cysteine Presence/Absence
For chains A1 and B1, no free cysteine occurred at conserved amino acid positions for any taxon. For chain A2, a conserved-free cysteine at position 75, correlating to that found by Zal et al. (1997), was present in all taxa except G. brachiosum (Fig. 2). This species lacked a free cysteine between the two cysteines involved in the formation of disulfide bridges. For chain B2, one incorrect BLAST hit was recovered for O. mucofloris (i.e., an A2 hit returned for the B2 search); however, a conserved-free cysteine was found for all other taxa excluding E. spicata, G. brachiosum, and A. marina.
Amino acid alignment of chain A2 for siboglinids. Alignment was generated in MEGA 5.2 using MUSCLE and visualized using UniPro UGENE (Okonechnikov et al. 2012). Bars at the top of the alignment show percentage of conserved identical amino acid for that position. Conserved cysteines at positions 23, 94, and 153 shown in purple (Color figure online)
Gene tree and 3D Structure Prediction
Final alignment of the 12 A2 chain sequences had 116 amino acid positions. Maximum-likelihood analysis of this alignment placed the O. mucofloris A2 sequence between the A2 sequences of frenulates and a moniliferan/vestimentiferan clade; however, frenulate sequences were recovered as paraphyletic with weak support (Fig. 3). The O. mucofloris chain A2 sequence was recovered as sister to the monilferan/vestimentiferan chain A2 clade with moderate support (bootstrap = 73).
Reconstruction of 3D models resulted in similar predictions for the three siboglinid species examined (Figs. 4, 5). Specifically, I-TASSER predicted a heme ligand binding site for each siboglinid with high confidence (0.99–1.00 C score), and no other ligand binding factor produced a C score >0.03. Ligand binding site residues predicted by I-TASSER (Supplementary Table 1) were identical between L. luymesi and O. mucofloris, while S. ekmani had only one difference and a single codon insertion at the 5′ end of the sequence before the binding pocket region. The predicted A. marina structure was identical to S. ekmani with the exception of an additional binding site residue located seven amino acid positions before the first position in S. ekmani’s binding site. S. spallanzanii started in the same relative position as A. marina, but had differences in predicted ligand binding site residues from all other taxa examined. The 3′ residues were more conserved across all five taxa compared to 5′ residues.
3D structure prediction of (a) Osedax mucofloris, (b) Lamellibrachia luymesi, (c) Siboglinum ekmani, (d) Arenicola marina, and (e) Sabella spallanzanii A2 chain proteins using the I-TASSER protein structure prediction server (Yang et al. 2015). Predicted heme binding site shown in green (Color figure online)
Stereoscopic overlay of the 3D structure predictions of Osedax mucofloris (red), Lamellibrachia luymesi (yellow), Siboglinum ekmani (green), Arenicola marina (orange), and Sabella spallanzanii (purple) A2 chain proteins using the SuperPose webserver version 1.0 (Maiti et al. 2004) (Color figure online)
Discussion
Contrary to our hypothesis, analyses presented here suggest Osedax has the biochemical capability of producing sulfur-binding Hbs. Specifically, Osedax mucofloris possesses a free cysteine at position 76 of the chain A2 of its Hbs. Additionally, the predicted 3D structure of this chain (Fig. 4) is nearly identical among siboglinids and the sulfur-tolerant A. marina, implying identical function. Involvement of Hbs in sulfide detoxification as part of Osedax life history at whale fall habitats may account for selection and retention of residues involved in hydrogen sulfide binding. Assuming that free cysteines in Hbs are subject to negative selection in polychaetes from sulfide-free habitats (Bailly et al. 2003), the presence of free cysteines in Hbs in Osedax is consistent with the idea that Osedax not only copes with hydrogen sulfide, but may use Hbs to interact with hydrogen sulfide in biologically important ways (e.g., Hbs have higher binding affinity than cytochrome-c oxidase, which is inhibited by small amounts of hydrogen sulfide; National Research Council 1979). The ability to bind sulfur for detoxification could even be under positive selection (Eichinger et al. 2014).
O. mucofloris possesses Hb linkers with greater similarity to vestimentiferan siboglinids than to sulfide-tolerant polychaetes; a result consistent with a recent phylogeny for the group (Li et al. 2015). This could indicate that Osedax produces hexagonal bilayer Hbs capable of sulfur binding. In the context of siboglinid phylogeny (Fig. 1), the presence of Hb linkers could indicate that the last common ancestor of vestimentiferan/moniliferan and Osedax possessed Hb that bound sulfur as well as oxygen. However, comparisons between reference sequences of vestimentiferan linkers and our novel transcriptomes recovered frenulate and O. mucofloris hits with similar blast scores (Table 3). Currently, only vestimentiferan and moniliferan siboglinids have been shown to possess the hexagonal bilayer Hbs that self-assemble with linkers. As other annelids have large hexagonal bilayered Hbs, frenulates, possessing ring-shaped Hbs, seem to have lost the ability to produce linkers capable of creating more complex structures. Both ring and hexagonal bilayer Hbs use the same types of globins (Meunier et al. 2010), and similarities across these globin types likely confound the analyses of linker sequences presented here. Quantification of the molecular mass of Osedax Hb would help determine whether Osedax Hbs are a hexagonal bilayer or a ring structure in nature.
Here, we analyzed O. mucofloris Hb as a first step toward determining how these proteins might function in the biology of these siboglinids bearing heterotrophic endosymbionts. The hemoglobin complex shows variation in size and complexity across siboglinid lineages. However, residues of the A2 heme ligand binding site have apparently remained nearly identical over 60 MY. Thus, aspects of the annelid hemoglobin mechanism have evolved at different rates, presumably due to variation in selective pressures. Such pressures may be tied to endosymbiont biology or the need to detoxify H2S in different host environments. Unlike most siboglinids, Osedax should not require sulfur-binding Hb to support its endosymbionts. Yet sulfur-binding Hb has apparently persisted in this group of bone-eating worms. Osedax experience high levels of hydrogen sulfide during their life. They possess a high surface area to volume ratio in its root system, similar to the less branched root of Lamellibrachia where hydrogen sulfide uptake occurs (Julian et al. 1999; Huusgaard et al. 2012). Although the root epidermis of Osedax was suggested as an important site for nutrient uptake (Katz et al. 2010), how the mucus sheath that envelops the trunk and root structures of O. mucofloris (Higgs et al. 2011) effects chemical uptake from bones, including hydrogen sulfide, is unclear. Moreover, the exterior surface of whale bones experiences microbial sulfide production, with the potential for bone interiors to have reducing microbial activity due to degradation of hydrophobic lipids, a process that can be facilitated by Osedax (Treude et al. 2009). The presence of hydrogen sulfide within bones is further supported by observations of iron sulfide staining and white filamentous bacterial mats around Osedax boreholes (Higgs et al. 2011). These factors would indicate that Osedax roots are in an environment with relatively high hydrogen sulfide levels, where the ability to detoxify it may be biologically advantageous.
References
Altschul SF, Gish W, Miller W et al (1990) Basic local alignment search tool. J Mol Biol 215:403–410. doi:10.1016/S0022-2836(05)80360-2
Arp AJ, Childress JJ (1981) Blood function in the hydrothermal vent vestimentiferan tube worm. Science 213:342–344. doi:10.1126/science.213.4505.342
Bailly X, Leroy R, Carney S et al (2003) The loss of the hemoglobin H2S-binding function in annelids from sulfide-free habitats reveals molecular adaptation driven by Darwinian positive selection. PNAS 100:5885–5890. doi:10.1073/pnas.1037686100
Bright M, Eichinger I, Salvini-Plawen L (2012) The metatrochophore of a deep-sea hydrothermal vent vestimentiferan (Polychaeta:Siboglinidae). Org Divers Evol 13:163–188. doi:10.1007/s13127-012-0117-z
Castresana J (2000) Selection of conserved blocks from multiple alignments for their use in phylogenetic analysis. Mol Biol Evol 17:540–552
Cavanaugh CM, Gardiner SL (1981) Prokaryotic cells in the hydrothermal vent tube worm Riftia pachyptila Jones: possible chemoautotrophic symbionts. Science (New York) 213:340–342. doi:10.1126/science.213.4505.340
Danise S, Higgs ND (2015) Bone-eating Osedax worms lived on Mesozoic marine reptile deadfalls. Biol Lett 11:20150072. doi:10.1098/rsbl.2015.0072
Darriba D, Taboada GL, Doallo R, Posada D (2011) ProtTest 3: fast selection of best-fit models of protein evolution. Bioinformatics 27:1164–1165
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797. doi:10.1093/nar/gkh340
Eichinger I, Schmitz-Esser S, Schmid M et al (2014) Symbiont-driven sulfur crystal formation in a thiotrophic symbiosis from deep-sea hydrocarbon seeps. Environ Microbiol Rep 6:364–372. doi:10.1111/1758-2229.12149
Flores JF, Fisher CR, Carney SL et al (2005) Sulfide binding is mediated by zinc ions discovered in the crystal structure of a hydrothermal vent tubeworm hemoglobin. Proc Natl Acad Sci USA 102:2713–2718
Glover AG, Källström B, Smith CR, Dahlgren TG (2005) World-wide whale worms? A new species of Osedax from the shallow north Atlantic. Proc R Soc Lond B 272:2587–2592. doi:10.1098/rspb.2005.3275
Glover AG, Wiklund H, Taboada S et al (2013) Bone-eating worms from the Antarctic: the contrasting fate of whale and wood remains on the Southern Ocean seafloor. Proc R Soc B 280:20131390. doi:10.1098/rspb.2013.1390
Goffredi SK, Orphan VJ, Rouse GW et al (2005) Evolutionary innovation: a bone-eating marine symbiosis. Environ Microbiol 7:1369–1378. doi:10.1111/j.1462-2920.2005.00824.x
Grabherr MG, Haas BJ, Yassour M et al (2011) Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotech 29:644–652. doi:10.1038/nbt.1883
Halanych KM (2005) Molecular phylogeny of siboglinid annelids (a.k.a. pogonophorans): a review. Hydrobiologia 535–536:297–307. doi:10.1007/s10750-004-1437-6
Halanych KM, Feldman RA, Vrijenhoek RC (2001) Molecular evidence that Sclerolinum brattstromi is closely related to vestimentiferans, not to frenulate pogonophorans (Siboglinidae, Annelida). Biol Bull 201:65–75
Higgs ND, Glover AG, Dahlgren TG, Little CTS (2011) Bone-boring worms: Characterizing the morphology, rate, and method of bioerosion by Osedax mucofloris (Annelida, Siboglinidae). Biol Bull 221:307–316
Hilário A, Johnson SB, Cunha MR, Vrijenhoek RC (2010) High diversity of frenulates (Polychaeta: Siboglinidae) in the Gulf of Cadiz mud volcanoes: a DNA taxonomy analysis. Deep Sea Res Part I 57:143–150. doi:10.1016/j.dsr.2009.10.004
Hilário A, Capa M, Dahlgren TG et al (2011) New perspectives on the ecology and evolution of siboglinid tubeworms. PLoS ONE 6:e16309. doi:10.1371/journal.pone.0016309
Huusgaard RS, Vismann B, Kühl M et al (2012) The potent respiratory system of Osedax mucofloris (Siboglinidae, Annelida)—a prerequisite for the origin of bone-eating Osedax? PLoS ONE 7:e35975. doi:10.1371/journal.pone.0035975
Iseli C, Jongeneel CV, Bucher P (1999) ESTScan: a program for detecting, evaluating, and reconstructing potential coding regions in EST sequences. ISMB. pp 138–148
Julian D, Gaill F, Wood E et al (1999) Roots as a site of hydrogen sulfide uptake in the hydrocarbon seep vestimentiferan Lamellibrachia sp. J Exp Biol 202:2245–2257
Katz S, Klepal W, Bright M (2010) The skin of Osedax (Siboglinidae, Annelida): an ultrastructural investigation of its epidermis. J Morphol 271:1272–1280. doi:10.1002/jmor.10873
Katz S, Klepal W, Bright M (2011) The Osedax trophosome: Organization and ultrastructure. Biol Bull 220:128–139
Kocot KM, Cannon JT, Todt C et al (2011) Phylogenomics reveals deep molluscan relationships. Nature 477:452–456. doi:10.1038/nature10382
Langmead B, Trapnell C, Pop M, Salzberg SL (2009) Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol 10:R25. doi:10.1186/gb-2009-10-3-r25
Li Y, Kocot KM, Schander C et al (2015) Mitogenomics reveals phylogeny and repeated motifs in control regions of the deep-sea family Siboglinidae (Annelida). Mol Phylogenet Evol 85:221–229. doi:10.1016/j.ympev.2015.02.008
Maiti R, Van Domselaar GH, Zhang H, Wishart DS (2004) SuperPose: a simple server for sophisticated structural superposition. Nucleic Acids Res 32:W590–W594
McDonald E, Brown CT (2013) Khmer: working with big data in bioinformatics. CoRR, abs/1303.2223, 2013
McMullin ER, Hourdez S, Schaeffer SW, Fisher CR (2003) Phylogeny and biogeography of deep sea vestimentiferan tubeworms and their bacterial symbionts. Symbiosis 34:1–41
Meunier C, Andersen AC, Bruneaux M et al (2010) Structural characterization of hemoglobins from Monilifera and Frenulata tubeworms (Siboglinids): first discovery of giant hexagonal-bilayer hemoglobin in the former “Pogonophora” group. Comp Biochem Physiol A 155:41–48. doi:10.1016/j.cbpa.2009.09.010
National Research Council, Division of Medical Science, subcommittee on Hydrogen Sulfide (1979) Hydrogen sulfide. University Park Press, Baltimore
Numoto N, Nakagawa T, Kita A et al (2005) Structure of an extracellular giant hemoglobin of the gutless beard worm Oligobrachia mashikoi. Proc Natl Acad Sci USA 102:14521–14526
Numoto N, Nakagawa T, Kita A et al (2008) Structural basis for the heterotropic and homotropic interactions of invertebrate giant hemoglobin. Biochemistry 47:11231–11238. doi:10.1021/bi8012609
Okonechnikov K, Golosova O, Fursov M, Team the U (2012) Unipro UGENE: a unified bioinformatics toolkit. Bioinformatics 28:1166–1167. doi:10.1093/bioinformatics/bts091
Rouse GW, Goffredi SK, Vrijenhoek RC (2004) Osedax: bone-eating marine worms with dwarf males. Science 305:668–671. doi:10.1126/science.1098650
Rouse GW, Wilson NG, Worsaae K, Vrijenhoek RC (2015) A dwarf male reversal in bone-eating worms. Curr Biol 25:236–241. doi:10.1016/j.cub.2014.11.032
Schulze A, Halanych KM (2003) Siboglinid evolution shaped by habitat preference and sulfide tolerance. Hydrobiologia 496:199–205. doi:10.1023/A:1026192715095
Smith CR, Glover AG, Treude T et al (2015) Whale-fall ecosystems: recent insights into ecology, paleoecology, and evolution. Annu Rev Mar Sci 7:571–596. doi:10.1146/annurev-marine-010213-135144
Southward EC (1978) A new species of Lamellisabella (Pogonophora) from the north Atlantic. J Mar Biol Assoc UK 58:713–718. doi:10.1017/S0025315400041357
Southward EC (1988) Development of the gut and segmentation of newly settled stages of Ridgeia (Vestimentifera): implications for relationship between Vestimentifera and Pogonophora. J Mar Biol Assoc UK 68:465–487. doi:10.1017/S0025315400043344
Southward AJ, Southward EC (1981) Dissolved organic matter and the nutrition of the Pogonophora: a reassessment based on recent studies of their morphology and biology. Kiel Meeresf 5:445–453
Southward EC, Schulze A, Gardiner SL (2005) Pogonophora (Annelida): form and function. In: Bartolomaeus T, Purschke G (eds) Morphology, molecules, evolution and phylogeny in Polychaeta and related taxa. Springer, Netherlands, pp 227–251
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313. doi:10.1093/bioinformatics/btu033
Suzuki T, Takagi T, Ohta S (1990) Primary structure of a constituent polypeptide chain (AIII) of the giant haemoglobin from the deep-sea tube worm Lamellibrachia. A possible H2S-binding site. http://www.biochemj.org/bj/266/bj2660221.htm
Talavera G, Castresana J (2007) Improvement of phylogenies after removing divergent and ambiguously aligned blocks from protein sequence alignments. Syst Biol 56:564–577
Tamura K, Peterson D, Peterson N et al (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739. doi:10.1093/molbev/msr121
Thornhill DJ, Wiley AA, Campbell AL et al (2008) Endosymbionts of Siboglinum fiordicum and the phylogeny of bacterial endosymbionts in Siboglinidae (Annelida). Biol Bull 214:135–144
Treude T, Smith C, Wenzhöfer F et al (2009) Biogeochemistry of a deep-sea whale fall: sulfate reduction, sulfide efflux and methanogenesis. Mar Ecol Prog Ser 382:1–21. doi:10.3354/meps07972
Verna C, Ramette A, Wiklund H, Dahlgren TG, Glover AG, Gaill F, Dubilier N (2010) High symbiont diversity in the bone-eating worm Osedax mucofloris from shallow whale-falls in the North Atlantic. Environ Microbiol 12:2355–2370
Yang J, Yan R, Roy A, Xu D, Poisson J, Zhang Y (2015) The I-TASSER Suite: protein structure and function prediction. Nat Methods 12:7–8
Yuasa HJ, Green BN, Takagi T et al (1996) Electrospray ionization mass spectrometric composition of the 400 kDa hemoglobin from the pogonophoran Oligobrachia mashikoi and the primary structures of three major globin chains. Biochimica et Biophysica Acta (BBA) 1296:235–244. doi:10.1016/0167-4838(96)00081-7
Zal F, Lallier FH, Green BN et al (1996a) The multi-hemoglobin system of the hydrothermal vent tube worm Riftia pachyptila II. Complete polypeptide chain composition investigated by maximum entropy analysis of mass spectra. J Biol Chem 271:8875–8881. doi:10.1074/jbc.271.15.8875
Zal F, Lallier FH, Wall JS et al (1996b) The multi-hemoglobin system of the hydrothermal vent tube worm Riftia pachyptila I. Reexamination of the number and masses of its constituents. J Biol Chem 271:8869–8874. doi:10.1074/jbc.271.15.8869
Zal F, Suzuki T, Kawasaki Y et al (1997) Primary structure of the common polypeptide chain b from the multi-hemoglobin system of the hydrothermal vent tube worm Riftia pachyptila: an insight on the sulfide binding-site. Proteins: Structure. Funct Bioinform 29:562–574
Acknowledgments
We are indebted to the late Christopher Schander for his contribution of Osedax mucofloris tissue for sequencing. We would like to thank the crews of the R/V Seward Johnson, R/V Aurelia, and R/V Håkons-Mosby for their assistance in procuring samples. We would also like to thank Amanda Shaver and Franziska Franke for preparation of cDNA for samples used in this study. Pamela M. Brannock, Kevin M. Kocot, and Nathan V. Whelan provided guidance and input on procedures used in this study and the writing of versions of this manuscript. This work was funded by National Science Foundation grants IOS-0843473 to K. M. H, S. R. S., and D. J. T., OCE-1155188 to K. M. H., DEB-1036537 to K. M. H. and S. R. S. It represents contributions #141 and #49 to the Auburn University (AU) Marine Biology Program and Molette Biology Laboratory for Environmental and Climate Change Studies, respectively.
Author information
Authors and Affiliations
Corresponding authors
Additional information
Genbank Accession Numbers KT166952–KT166980.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Waits, D.S., Santos, S.R., Thornhill, D.J. et al. Evolution of Sulfur Binding by Hemoglobin in Siboglinidae (Annelida) with Special Reference to Bone-Eating Worms, Osedax . J Mol Evol 82, 219–229 (2016). https://doi.org/10.1007/s00239-016-9739-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-016-9739-7