Abstract
A fundamental problem in origins of life research is how the first polymers with the properties of nucleic acids were synthesized and incorporated into living systems on the prebiotic Earth. Here, we show that RNA-like polymers can be synthesized non-enzymatically from 5′-phosphate mononucleosides in salty environments. The polymers were identified and analyzed by gel electrophoresis, nanopore analysis, UV spectra, and action of RNases. The synthesis of phosphodiester bonds is driven by the chemical potential made available in the fluctuating hydrated and anhydrous conditions of hydrothermal fields associated with volcanic land masses.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
A substantial weight of evidence supports the conjecture that the earliest forms of life passed through a phase in which RNA served both as a catalyst and as a carrier of genetic information (Woese 1967; Crick 1968; Orgel 1968; Gilbert 1986). However, it is not clear how a non-biological process could have synthesized random polymers of RNA-like molecules as a first step toward living systems. The goal of the research reported here is to discover whether a mixture of nucleotide monomers, when exposed to simulated prebiotic conditions, can be driven toward polymerization.
An earlier study (Rajamani et al. 2008) showed that small amounts of polymers resembling RNA are produced when 5′-phosphate mononucleosides such as AMP or UMP are exposed to multiple cycles of hydration–dehydration (HD) at elevated temperatures. These conditions simulate hydrothermal processes that commonly occur in volcanic hydrothermal fields today and were presumably ubiquitous in the prebiotic environment. The inclusion of an amphiphilic phospholipid as an ordering agent enhanced the reaction and captured the products in lipid vesicles. Single-stranded polymeric products, presumably homopolymers, were detected by nanopore analysis, and could be end-labeled with 32P by the standard enzyme-catalyzed reaction used to label biological RNA. Denaturing gel electrophoresis revealed polymers between 20 and 100 nt to be present. Yields of polymers were quantified by a fluorescent dye method developed for biological RNA, or by absorption at 260 nm, and were determined to represent a few percent of the starting nucleotides.
These results were confirmed by DeGuzman et al. (2014) who used reaction mixtures containing both adenosine 5′-monophosphate and uridine 5′-monophosphate to test whether mixtures of mononucleotides could also form polymers. Instead of radioactive end labeling, precast gels containing ethidium bromide were employed to detect polymers. Because AMP and UMP have the potential to form base pairs, a second aim was to determine whether hydrogen-bonded secondary structures might be present in the products. The products were stained by ethidium bromide, an intercalating dye, which was consistent with the possible presence of polymers containing duplex regions.
The present study extends this simulation to conditions in which mononucleotides are exposed to HD cycles in the presence of monovalent salts. The expectation was that during dehydration, crystallization of the salts can concentrate and organize mononucleotides in such a way that condensation reactions lead to ester bond synthesis.
Materials and Methods
For the model system, two monomers were chosen: adenosine 5′-monophosphate (AMP) and uridine 5′-monophosphate (UMP) in their acid forms rather than as sodium salts (Sigma-Aldrich). When dissolved in water at 10 mM concentration, the pH of the solution is ~2.5. Commercial polyadenylic acid (polyA) and polyuridylic acid (polyU) were used as polynucleotide control standards (Sigma-Aldrich). These were mixed in 1:1 mol ratios with respect to the bases to produce double-stranded RNA (polyA–polyU). The effects of a variety of monovalent salts, including LiCl, NaCl, KCl, and NH4Cl were investigated in terms of their ability to promote oligomerization. During evaporation, the salts formed crystalline films when their solubilities were exceeded. The growing crystals excluded other solutes such as the mononucleotides, producing highly concentrated eutectic phases within the salt matrix in the final stages of drying.
A Laboratory Simulation of HD Cycles
Simulations were carried out using glass slides with two wells on each slide that hold 0.1 mL of the reaction mixture. Four slides were arranged on a laboratory hot plate set at the desired temperature range, and a plastic flow box with 8 small holes (1 mm diameter) was set on the slides. Each hole was placed directly over a well so that carbon dioxide gas flowed onto the mixture at approximately 1 cc/s into each well. The purpose of the gas was to exclude oxygen, but also to carry away water vapor from condensation reactions as ester bonds formed, thereby preventing hydrolytic back reactions.
Reaction Mixtures
Mononucleotides, AMP (10 mM), and UMP (10 mM) were initially mixed in a 1:1 volume ratio. The mononucleotide solutions and 0.1 M monovalent salts were mixed in a 2:1 volume ratio so that the initial concentrations were 3.3 mM AMP and UMP, together with 0.033 M salt. Because water evaporated during dehydration, the dilute solutions became highly concentrated and finally dry, so it is the ratios that are significant rather than the initial concentrations. In a typical experiment, the reactants were exposed to 1–16 cycles of wetting and drying. The temperature (85 °C) and flow of carbon dioxide caused drying within 1–2 min. After each dehydration phase of 30 min, the samples were dispersed in 0.1 mL of 1.0 mM HCl to maintain acidity, followed by the next dehydration cycle. Variable experimental parameters included initial pH, temperature, the time given to each cycle, and the number of cycles. At the end of the cycle series, the samples were dissolved in 0.1 mL of water.
Isolation of Products
The polymer products were isolated by addition of 2.5× volume of 100 % ethanol, 1/10 volume 3.0 M sodium acetate pH 5.2, and 1.6 μL linear acrylamide (5 mg/mL) (Fischer Scientific) in 700 μL of reaction mixtures, followed by overnight incubation at −20 °C. After centrifugation, analysis of the pellet was performed by UV absorbance with a NanoVue instrument calibrated for RNA to estimate the yields of products. Depending on the conditions and number of HD cycles, typical yields ranged from 1 to 40 %, expressed as the fraction of the total weight of mononucleotides present, and over 55 % if additional monomers were added during cycling.
Intercalation of Ethidium Bromide
Products of the reaction were initially monitored by gel electrophoresis on precast 4 % agarose gels with ethidium bromide staining (Invitrogen) with DNA and RNA ladders used as size markers (Thermoscientific). The gel images were inverted for contrast, so that fluorescence is dark against a light background. Relative densities of products in the gel were analyzed with ImageJ software (available as freeware from National Institutes of Health).
Alkaline Hydrolysis
Alkaline hydrolysis was carried out in 2 M NaOH, 1 mM EDTA, 65 °C for 10 min, then stopped with 2 N HCl, 0.5 M HEPES pH 7.5. Hydrolysis was performed on standard polyA–polyU, polyA, polyU, and the products of HD simulations. The hydrolysis products and mononucleotide references AMP and UMP were spotted on a C18 reverse-phase thin-layer chromatography (TLC) plate that contained a fluorescent compound, and then developed with 1.0 M LiCl as solvent. The products were detected as dark spots when the plate was illuminated with 254 nm UV light.
Ribonuclease Effects on Products
If the polymers incorporated phosphodiester bonds between AMP and UMP, it was expected that RNase A should hydrolyze the products. To test this prediction, pellets produced by ethanol precipitation were suspended either in 10 μL of water or 0.1 M NaCl, and RNase A was added to a final concentration of 1 μg/mL. The solutions were incubated 30 min at 37 °C, followed by ethanol precipitation. The pellets were resuspended in 10 μL water and analyzed by gel electrophoresis and NanoVue spectrophotometry.
Hyperchromicity
It is well known that duplex ribonucleic acids exhibit increased absorbance at 260 nm when the chains become single stranded as temperature is increased (Borer et al. 1974). This effect is referred to as hyperchromicity, and represented a useful test for the presence of duplex structures stabilized by hydrogen bonding in the products. Aliquots of products were dissolved in 1.0 mL of 25 mM phosphate buffer, pH 7.0, to give an initial absorbance value near 0.5. The sample was placed in a cuvet and heated by a circulating water bath, while absorbance was monitored over a temperature range of 15–70 °C.
Nanopore Analysis
Samples were analyzed by the nanopore technique described by DeGuzman et al. (2014). Briefly, a single α-hemolysin channel was inserted into a planar lipid bilayer membrane bathed in 70 μL of 1.0 M KCl buffered at pH 7 with 10 mM HEPES. An aliquot of the products was mixed with the same buffer and added to the well and a voltage of 120 mV was applied to the membrane. The voltage establishes an electric field in the nanopore that captures anionic polymers in solution and transports them from the cis to the trans side of the membrane, which is supported on a thin Teflon aperture with a diameter of 30 μm. Only single-stranded nucleic acids can pass through the 2.0 nm limiting aperture of the pore. When a polymer passes through the pore, it causes a transient blockade of the ionic current that can be interpreted in terms of the structure and molecular dynamics of the polymer (Vercoutere et al. 2001).
Microscopy
In order to better understand the mechanism of salt-induced polymerization, thin films of dried ammonium chloride with the AMP and UMP additions were examined by phase and fluorescence microscopy. A small amount of 0.1 mM pyranine dye was added as a marker.
Results
Intercalation of Ethidium Bromide into dsRNA
The fluorescence of certain dyes is markedly enhanced when they intercalate into double-stranded polynucleotides. This effect is illustrated in Fig. 1a, which shows a gel with control standards of polyA, polyU, and the duplex species polyA–polyU (produced by mixing the two homopolymers in a 1:1 mol ratio of A:U). Neither of the homopolymers (2 μg each) bind the dye, but mixing the polyA and polyU to produce 2 μg of the duplex species polyA–polyU gives a strongly fluorescent band. This result is significant, because the products of an HD reaction of monomers AMP and UMP in the presence of salts also bind ethidium bromide, as described next.
Intercalation of ethidium bromide into dsRNA and products of HD reactions. a Samples of single-stranded polyA, single-stranded polyU, and double-stranded polyA–polyU, 2 µg each, were analyzed with a precast 4 % agarose gel containing ethidium bromide. Only the duplex RNA was stained by the dye. b Products of AMP and UMP exposed to 16 HD cycles in the presence of salts were analyzed by gel electrophoresis. c Products of synthesis analyzed by denaturating PAGE and detected by SYBR Gold staining. Lane A DNA ladder, Lane B products of AMP 10 mM + UMP 10 mM + NH4Cl 0.1 M exposed to 16 HD cycles, Lane C RNA ladder. d Control gel illustrating effect of NH4Cl in promoting polymerization during 16 HD cycles. Lane A 10 mM + UMP 10 mM, Lane B NH4Cl alone, Lane C AMP 10 mM + UMP 10 mM + NH4Cl 0.1 M. See text for details
Effect of Monovalent Cations on Polymerization
When the HD cycles were run with monovalent salts in the reaction mixture, yields of polymer were dramatically increased compared to the absence of salts. Furthermore, the products were stained by ethidium bromide, an intercalating dye, suggesting that base pairing was present. Figure 1b compares the effects of four monovalent salts. Sodium, potassium, and ammonium chloride all promoted synthesis of polymers containing AMP and UMP as monomers. A smear of products ranging from 10 to 300 apparent base pairs with a peak around 100 mers is apparent. NH4Cl had the greatest effect, but products from LiCl produced only a weak band in the gel even though the yield measured by ethanol precipitation was in the same range as NH4Cl (see Table 1).
It is well known that the apparent lengths of the smeared product bands in agarose gels (Fig. 1b) could be caused by a variety of factors other than actual chain length. Therefore as a control, we also ran a denaturing polyacrylamide gel stained with the SYBR dye which does not depend on intercalation (Fig. 1c). After 16 HD cycles, the products again produced a smeared fluorescence consistent with a chain length extending from ~10 to >300 mers.
An additional control was run to demonstrate the effect of NH4Cl in promoting the polymerization reaction. Figure 1d shows AMP + UMP alone (lane A) and in the presence of NH4Cl at an initial concentration of 0.1 M (lane C). The effect is dramatic.
The A 260/A 280 ratio in Table 1 provides an estimate of how much of the absorbance is due to polymers and how much to monomers. A ratio of 2 corresponds to RNA, while a ratio of 3.4 is observed for monomers. The high ratio with LiCl indicates that the product has relatively short strands of oligomers lacking base pairing compared with the other salts. This is consistent with the weakly staining LiCl band in the gel.
Cycling Increases Yield of Polymers
The polymerization process was optimized by performing a series of preliminary experiments in which conditions such temperature, pH, and the number and duration of cycles were varied. The synthesis of polymers is optimal at an elevated temperature (~85 °C), and acidic pH range with a gentle CO2 stream flushing away water vapor. This suggests that synthesis of the ester bond is an acid-catalyzed mechanism and that CO2 plays an essential role in the polymerization process by excluding oxygen and removing water vapor. The dehydration phase appeared to be essential for the polymerization process since a minimum of 30 min of drying per cycle was necessary to synthesize longer oligomers species (data not shown).
Multiple shorter HD cycles were more effective than a single long cycle, as shown in Fig. 2. It is significant that longer products accumulate in later cycles, suggesting that ligation of shorter chains may be occurring.
Role of NH4 + Cations in Promoting Polymerization
Because NH4Cl seemed to have the greatest effect on yields of polymers, we performed a series of experiments to better understand the mechanism. Figure 3 shows results with different ammonium salts, including ammonium phosphate, ammonium molybdate, and ammonium formate. Only the ammonium formate yielded polymers ranging from 10 to 300 apparent base pairs in length but in lesser amounts compared to ammonium chloride (Fig. 3a, b). The importance of the chemical effect of the ammonium cation in this polymerization process was also tested by substituting tetramethylammonium chloride for ammonium chloride. Figure 3c, d shows that tetramethylammonium chloride also produced polymers ranging from 10 to 300 nucleotides in length but with lower efficiency than ammonium chloride. This suggests that NH4 + might promote oligomerization by interacting with and ordering the concentrated mononucleotides produced during drying and crystallization (see Fig. 10).
Comparison of polymerization process between NH4Cl and other ammonium salts. a Gel patterns and b polymer synthesis after 8 h of 30 min HD cycles. Yields are normalized for comparison, taking the products in the presence of NH4Cl as 1.0. Salts, lane A NH4Cl, lane B C19H42BrN, lane C HCO2NH4, lane D (NH4)6Mo7O244H2O, lane E NH4H2PO4. c Gel patterns with a mixture of AMP 10 mM + UMP 10 mM + 0.1 M (CH3)4NCl (1:1:1 volume ratio) compared to AMP 10 mM + UMP 10 mM + NH4Cl 0.1 M (1:1:1 volume ratio) after 4 and 8 h. d Polymer synthesis after 4 and 8 h. Yields are normalized for comparison, taking the products in the presence of NH4Cl as 1.0. (M DNA ladder, hrs hours)
Hydrolysis of Polymers
We next tested whether the chemical and biochemical properties of the polymers matched those expected of a polynucleotide. Polymers were exposed to alkaline hydrolysis as described in “Materials and methods” section, followed by separation of products using thin-layer chromatography (fluorescent C18 reverse-phase TLC and 1.0 M LiCl solvent). The products formed a single spot at the origin, and after hydrolysis two new spots appeared with Rf values matching AMP and UMP (Fig. 4a). The fact that the polymer was labile to alkaline hydrolysis is consistent with ester bonds being the primary linkage. Furthermore, the appearance of two spots confirmed that both the mononucleotides were present in the polymer in approximately equal amounts. However, a certain amount of the polymer remained at the origin, suggesting that some of the products resisted hydrolysis and contained bonds other that ester bonds.
Hydrolysis of HD products exposed to 16 HD cycles and standards. a Alkaline hydrolysis products separated by thin-layer chromatography (see “Materials and Methods” section for reaction conditions). (AH after alkaline hydrolysis). b RNase A digestion. Reaction conditions presence (gray) or absence (black) of 0.1 M NaCl, 1 μg/mL RNase A, 20 μg RNA, 30 min at 37 °C. c Lead cleavage. Reaction conditions 1 mM lead acetate, 20 μg RNA, 100 mM HEPES pH 7.3, 50 mM NaCl, 5 mM MgCl2, 24 h at room temperature (Meli et al. 2002). The percentage was calculated from the quantity of ethanol-precipitated polymers measured by a NanoVue instrument using UV absorbance. For instance, if the pellet before hydrolysis had 20 μg of RNA and 2 μg after hydrolysis, 90 % had been hydrolyzed. The negative values were the result of minor experimental variations in measuring the quantities of products in the pellets
Ribonuclease mapping was also performed to test the expectation that a polynucleotide containing both AMP and UMP should be attacked by an endonuclease that cleaved ssRNA. The polymers were exposed to ribonuclease A both in water and in the presence of 0.1 M NaCl. The comparison was made because RNase A specifically cleaves ssRNA at pyrimidine residues, but is unable to cleave dsRNA that assembles from ssRNA in NaCl. We found that the polymers were in fact digested by RNase A in water, whereas in the presence NaCl they were not (Fig. 4b). Because the presence of NaCl favors base pairing, this result is consistent with duplex species present in 0.1 M NaCl that resist RNase A cleavage. The products were also exposed to lead cations which catalyze the cleavage of ssRNA without base specificity. Figure 4c shows that products are not hydrolyzed by lead cleavage in NaCl solutions, again consistent with a significant amount of duplex structure in the polymers.
These results confirm that RNA-like products having AMP and UMP as monomers are products of the reaction, and the polymers appear to have a certain amount of hydrogen-bonded secondary structure.
Kinetics of Oligomerization
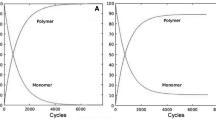
The oligomerization process in the presence of ammonium chloride follows an exponential curve, and reaches a plateau after 30 h of HD cycles with a yield of 40 % (Fig. 5). The plateau can be due to an equilibrium between synthesis and hydrolysis, although degradation of nucleotides over time may also contribute. Figure 6 shows that longer products accumulate in later cycles. Indeed, there is an enhancement of the production of short fragments (10 and 150 apparent bps) after a few cycles, which then decrease over time while longer polymers accumulate in the later cycles. Lengthening may occur either by elongation or by ligation of short fragments.
Kinetics of oligomerization experiments using AMP and UMP as monomers. The total amounts and yields of polymeric products over multiple cycles are shown. Each hour had two 30 min cycles, so 40 h represents 80 cycles. (See “Reaction Mixtures” section in “Materials and Methods” for conditions.)
a Gel pattern of products is shown in Fig. 5. Virtually, no product appeared after a single 30 min HD cycle, but became apparent after 4 cycles (2 h). The shorter strands dominating after 10 cycles (5 h) declined with increasing numbers of cycles, while longer strands emerged. The evolution can be seen in b in which gel densities were quantified by ImageJ analysis at specific strand lengths specified by the ladder. (a DNA ladder, ctrl negative control of the mixture of AMP 10 mM + UMP 10 mM + NH4Cl 0.1 M (1:1:1 volume ratio) without HD cycles, room temperature)
Control of Nucleotide Concentration: Nucleotide Feeding
To determine whether the observed plateau in yield of products was due to exhaustion of monomers, a feeding experiment was performed in which fresh monomers were added every 2 h (4 cycles). Figure 7a shows an enhancement of oligomerization when cycling is accompanied by regular additions of monomers. Figure 7b shows that a yield of 58 % is obtained after 5 feeding steps (final concentration of nucleotides equal to 60 mM), whereas for the same concentration (60 mM) present at the beginning of the experiment, the yield is 37 %. We conclude that controlling nucleotide concentration by stepwise additions enhances the polymerization when compared to the nucleotide pool at an equivalent concentration but without additions.
Feeding experiment. The initial reaction mixture was 0.2 mL of AMP 5 mM + UMP 5 mM + NH4Cl 0.1 M (1:1:1 volume ratio). Every 2 h, an 0.1 mL aliquot of AMP 5 mM + UMP 5 mM (1:1 volume ratio) was added. a, b show the quantity and yield of polymers synthesized. The mean and standard deviation bars were from 4 repeats. Legend: white circle feeding; black circle no feeding. It was important to test whether the increased yields were due to feeding or simply related to the increasing amounts of monomers being added. Therefore, single control experiments were run in which monomer concentration and incubation time were varied to the same extent as the feeding experiment: times 4 h with AMP 10 mM + UMP 10 mM + NH4Cl 0.1 M (1:1:1 volume ratio); plus 6 h with AMP 15 mM + UMP 15 mM + NH4Cl 0.1 M (1:1:1 volume ratio); white up-pointing triangle 8 h with AMP 20 mM + UMP 20 mM + NH4Cl 0.1 M (1:1:1 volume ratio); white diamond 10 h with AMP 25 mM + UMP 25 mM + NH4Cl 0.1 M (1:1:1 volume ratio); white square 12 h with AMP 30 mM + UMP 30 mM + NH4Cl 0.1 M (1:1:1 volume ratio). Although significant amounts of products were synthesized, none of the controls matched the yields achieved with feeding
Hyperchromicity
Temperature-dependent hyperchromicity occurs when double-stranded nucleic acids melt because the bases in single-stranded molecules have a greater intrinsic absorbance than the same bases in a duplex strand. Aliquots of the HD products that were precipitated in ethanol and then dissolved in water were added to 1.0 mL of 25 mM phosphate buffer, pH 7.0, and UV spectra from 200 to 300 nm were obtained while the sample was heated. This experiment was carried out multiple times and a typical result is shown in Fig. 8.
The spectra revealed the expected peaks at 260 and 205 nm. Both peaks increased with temperature between 15 and 50 °C, but the degree of hyperchromicity varied from one experiment to the next, typically ranging from 10 to 50 % of the hyperchromicity exhibited by a sample of duplex polyA–polyU. The observed hyperchromicity between 20 and 40 °C is consistent with the presence of stacked bases in the HD products, and perhaps duplex structures approximately 15 base pairs long. Interesting downward deflections were often observed, perhaps reflecting relaxation of variable strand lengths so that base stacking occurred but then melted as the temperature increased further.
Nanopore Analysis of Products
Ionic current blockades are produced when products interact with the hemolysin nanopore (Fig. 9). The majority of blockades were approximately a third of the open channel current, but others blocked over 90 % of the current. As will be discussed later, the deep blockades indicate that linear anionic polymers are traversing the channel.
Microscopic Appearance of Ammonium Chloride Phase
In order to better understand the mechanism of salt-induced polymerization, we examined thin crystalline films that formed when 100 μL of the standard reaction mixture (ammonium chloride with the AMP and UMP additions) was dried at 85 °C in one well of the microscope slide used for the HD cycling. A small amount (0.1 mM) pyranine dye was added as a marker. Figure 10a shows the film by phase microscopy, and Fig. 10b is a fluorescence image of the same area. We assume that the dye indicates mononucleotides within the crystal matrix, and it is apparent that they are excluded from the main crystals of ammonium chloride during crystallization and accumulate within concentrated regions between the crystals. This is analogous to the eutectic phase concentration that occurs when an aqueous solution undergoes freezing and thereby concentrates solutes between ice crystals (Kanavarioti et al. 2001; Attwater et al. 2013).
Phase (a) and fluorescence (b) micrographs of mononucleotides mixed with NH4Cl and dried in the glass well as described in “Materials and Methods” section. Pyranine dye (0.1 mM) was present to mark the eutectic regions containing the mononucleotides. The mixture was not cycled, but simply dried once at 85 °C. Bar shows 25 μm
Discussion
The goal of our research is to close the gap related to the transition from an entirely sterile prebiotic Earth to the emergence of polymers capable of catalysis and genetic information. All life today exists in a steady state in which energetically uphill synthetic condensation reactions are balanced against spontaneous downhill hydrolysis reactions. A similar balance was likely to have been in place on the prebiotic Earth in which a variety of processes cycled monomers and their polymers. It follows that the origins of life occurred within a steady state in which an uphill synthesis of polymers balanced polymer hydrolysis.
The approach described in this report is guided by consideration of the actual conditions on the early Earth in which life could begin. As a plausible site that is feasible to simulate in the laboratory, we are testing whether polymerization of mononucleotides can be driven by HD cycles that occur in small pools produced by hydrothermal conditions associated with volcanic activity on early Earth. Field studies in analog environments in Kamchatka, Iceland, Hawaii, and northern California have shown that such pools range in temperature from 60 °C to near boiling (Deamer et al. 2006). They are acidic, typically between pH 2 and 3. HD cycles occur around the edges of the pools in time frames of minutes to hours, and the pools themselves evaporate and refill over weeks to months.
We have developed a simple laboratory simulation of HD cycles which we consider to be an essential factor driving polymerization and increasing complexity, analogous to the thermal cycles required for the polymerase chain reaction. If the solutes are mononucleotides, we propose that they are organized under these conditions into highly concentrated mixtures that bring phosphate and ribose groups into close proximity, thereby promoting condensation reactions. In the results reported here, we observed that monovalent salts undergoing crystallization during dehydration could act as an organizing matrix similar to that described earlier for phospholipids. However, the amount of polymer synthesis was more than tenfold greater than that observed with lipid-enhanced reactions, exceeding 50 % yields under some conditions. Yields were highest with LiCl, NH4Cl, KCl, and NaCl, in that order, but the LiCl product was less well stained by ethidium bromide, probably because the oligomers were shorter with relatively little base pairing.
Nature of Products
The following results are consistent with RNA-like polymers being components of the mixture of products:
-
1.
The presence of ester bonds was confirmed by hydrolysis in alkaline conditions expected to hydrolyze RNA, and thin-layer chromatography showed that both AMP and UMP were components of the oligomers.
-
2.
The products could be partially hydrolyzed by ribonuclease A, indicating that a substantial number of the bonds were 3′-5′ phosphodiester linkages involving UMP.
-
3.
The 1:1 mixture of AMP and UMP has the potential to form base pairs if polymers are synthesized. The polymers are stained by ethidium bromide, an intercalating dye, and exhibit hyperchromicity, both properties indicating that a certain amount of secondary structure was present.
-
4.
The products produce blockades when subjected to nanopore analysis. Transient blockades with durations in the range of 10–100 microseconds can only be caused by linear polyanions traversing the pore, so at least some of the products are linear polymers resembling RNA.
Although these results indicate that HD reactions synthesize polymers of mononucleotides linked by phosphodiester bonds, the products appear to be a complex mixture of thousands of different chain lengths having both 2′–5′ and 3′–5′ ester bonds and folded into a variety of secondary structures. There are also degradation products in the mixture produced by browning reactions and depurination. The complexity of the mixture makes it particularly difficult to analyze by standard methods used to study biological RNA. Despite this, it is important to note that the same will be true of any non-biological polymerization process, so the mixtures of products produced by HD cycling are plausible models of similar products that would be synthesized under prebiotic conditions. Our goal now is to discover ways in which such mixtures can evolve into functional systems of catalytic, replicating molecules.
Reaction Mechanism
In regard to the mechanism underlying the condensation reaction, one contributing factor may be the observation by Rode and co-workers that peptide bonds can be synthesized if amino acids are exposed to HD cycles in the presence of sodium chloride (Schwendinger and Rode 1992; Schwendinger et al. 1995). This reaction was referred to as salt-induced peptide formation (SIPF) and it was proposed that as drying occurred, water molecules in the hydration layer were lost from sodium cations. The water of hydration could be replaced by removing it from two neighboring amino acids, so the sodium cations effectively became condensing agents. The fact that LiCl is less effective in promoting polymerization of mononucleotides is consistent with this explanation, because Li cations are more strongly hydrated due to their smaller radius and less able to become dehydrated.
On the other hand, salt ions as condensing agents cannot be the whole story, because polymerization occurs in multilamellar lipid matrices in the complete absence of salt (DeGuzman et al. 2014). We must also account for the fact that ammonium chloride is significantly better at promoting polymerization than the other salts because of its ability to maintain the acidic medium during evaporation. This may be related to the recent observation of Safaee et al. (2013), who found that ammonium cations stabilize the double-stranded structure of polyadenylic acid.
Given that the chemical potential driving the reaction is provided by reduced water activity when the mixture is dried at elevated temperatures, we can extrapolate from the well-known acid-catalyzed ester bond synthesis between carboxylic acids and alcohols to propose a similar mechanism of phosphoester bond synthesis in which water serves as a leaving group (Fig. 11).
An Organizing Matrix Concentrates Reactants and Reduces Entropy Barriers
Orgel and co-workers pioneered the use of imidazole esters of mononucleotides and demonstrated that imidazole esters can assemble on RNA templates to produce longer complementary RNA strands up to 30 nucleotides in length (Inoue and Orgel 1983; Orgel 1987). Ferris and co-workers have shown that montmorillonite clay surfaces also provide an organizing interface that enhances polymerization of activated mononucleotides by adsorbing and concentrating them on the mineral surface (Huang and Ferris 2003, 2006).
The question posed here is whether an alternative organizing medium could impose an additional amount of order that promotes polymerization processes in the absence of activated chemical species. This depends on the thermodynamic prediction that it should be possible to drive phosphodiester bond synthesis under conditions in which water can be removed from the reactants, as illustrated in Fig. 11. Fluctuating environments in the form of wet–dry cycles have long been considered as possible sources of free energy for uphill polymerization reactions. Verlander et al. (1973) showed that anhydrous heating of nucleotides could drive the formation of mixed 2′–5′ and 3′–5′ phosphodiester bonds. Lohrmann and Orgel (1973) also explored this possibility but were only able to achieve dimers and trimers in low yields. Usher and McHale (1976) proposed that cycles of heating and drying, followed by rehydration, could lead to phosphodiester bond formation and promote the accumulation of 3′–5′ bonds in the system due to the relative lability of 2′–5′ bonds to hydrolysis.
Anhydrous conditions do not ordinarily produce long polymers when monomers are dried. The reason is that potential reactants are disorganized and immobilized within the solid matrix of a dry bulk phase, so that reactive groups only rarely come into contact to undergo condensation reactions. However, if a microenvironment not only organizes and concentrates monomers, thereby decreasing entropy compared to the bulk phase solution, but also permits diffusional mobility, it is possible that polymers are synthesized that cannot form under ordinary conditions of polymerization. To account for the results described here, we propose that eutectic phases are produced during crystallization of monovalent salts that organize and concentrate mononucleotides, as shown in the micrograph of dried ammonium chloride (Fig. 10). Under these conditions, it is plausible that the highly concentrated monomers self-assemble into aggregates stabilized by hydrogen bonding and base stacking, then undergo condensation reactions to form polymers linked by phosphodiester bonds.
Conclusions and Implications for the Origin of Life
In summary, we have established a laboratory simulation of hydrothermal pools undergoing cycles of hydration and dehydration. In these conditions, when 1:1 mixtures of AMP and UMP are cycled in the presence of monovalent salts, a polymerization reaction yields a product with properties similar to RNA. The products can be precipitated with ethanol, visualized by gel electrophoresis, and nanopore analysis reveals ionic current blockades expected of linear polyanionic polymers. Furthermore, the products are hydrolyzed when heated in alkaline pH ranges, and are partially hydrolyzed by RNase A. These results confirmed that UMP is incorporated into the polymer, because RNase A attacks pyrimidine bonds.
Remarkably, when both AMP and UMP are present, the products appear to have significant base pairings and perhaps even duplex character as indicated by dye intercalation and hyperchromicity. Previous reports indicated that mRNA and synthetic RNA readily fold into duplex structures stabilized by hydrogen bonding (Doty et al. 1959). Computational modeling suggests that the extent of folding can approach 50 % base pairing, Gralla and Delisi (1974). This fraction should be even greater when only two bases are present, because of the increased likelihood of finding favorable sequences of complementary pairs. The implication is that cycles of hydration and dehydration in hydrothermal fields on the early Earth could drive polymerization reactions, and the resulting polymers could then fold into duplex structures.
Given a process that can synthesize polymers, future research can be directed at establishing the properties of random polymers encapsulated within self-assembled boundary membranes. Given that a single milligram of phospholipid can form a trillion or more lipid vesicles, each different in composition from all the rest, a plausible hypothesis is that life emerged when a few such vesicles happened to contain systems of interacting polymers that could capture energy and nutrients from the environment in order to grow and reproduce. The power of this approach is illustrated by the demonstration that an encapsulated peptide catalyst can be a selective factor in an evolutionary process involving competing protocells (Adamala and Szostak 2013).
References
Adamala K, Szostak J (2013) Competition between model protocells driven by an encapsulated catalyst. Nat Chem 5:495–501
Attwater J, Wochner A, Holliger P (2013) In-ice evolution of RNA polymerase ribozyme activity. Nat Chem 5:1011–1018
Borer PN, Dengler B, Tinoco I, Uhlenbeck OC (1974) Stability of ribonucleic acid double-stranded helices. J Mol Biol 86:843–853
Crick FH (1968) The origin of the genetic code. J Mol Biol 38:367–379
Deamer D, Singaram S, Rajamani S, Kompanichenko V, Guggenheim S (2006) Self-assembly processes in the prebiotic environment. Philos Trans R Soc Lond B 361:1809–1818
DeGuzman V, Vercoutere W, Shenasa H, Deamer D (2014) Generation of oligonucleotides under hydrothermal conditions by non-enzymatic polymerization. J Mol Evol 78:251–262
Doty P, Boedtker H, Fresco JR, Haselkorn R, Litt M (1959) Secondary structure in ribonucleic acids. Proc Natl Acad Sci USA 45:482–499
Gilbert W (1986) The RNA world. Nature 319:618
Gralla J, Delisi C (1974) mRNA is expected to form stable secondary structures. Nature 248:330–332
Huang W, Ferris JP (2003) Synthesis of 35–40 mers of RNA oligomers from unblocked monomers. A simple approach to the RNA world. Chem Commun 21:1458–1461
Huang W, Ferris JP (2006) One-step, regioselective synthesis of up to 50-mers of RNA oligomers by montmorillonite catalysis. J Am Chem Soc 128:8914–8919
Inoue T, Orgel LE (1983) A nonenzymatic RNA polymerase model. Science 219:859–862
Kanavarioti A, Monnard PA, Deamer DW (2001) Eutectic phases in ice facilitate nonenzymatic nucleic acid synthesis. Astrobiology 1:271–281
Lohrmann R, Orgel LE (1973) Prebiotic activation processes. Nature 244:418–420
Meli M, Vergne J, Décout JL, Maurel MC (2002) Adenine–aptamer complexes: a bipartite RNA site that binds the adenine nucleic base. J Biol Chem 277:2104–2111
Orgel LE (1968) Evolution of the genetic apparatus. J Mol Biol 38:381–393
Orgel LE (1987) Evolution of the genetic apparatus: a review. Cold Spring Harb Symp Quant Biol 52:9–16
Rajamani S, Vlassov A, Benner S, Coombs A, Olasagasti F, Deamer D (2008) Lipid-assisted synthesis of RNA-like polymers from mononucleotides. Orig Life Evol Biosph 38:57–74
Safaee N, Noronha AM, Rodionov D, Kozlov G, Wilds C, Sheldrick GM, Gehring K (2013) Structure of the parallel duplex of poly(A) RNA: evaluation of a 50 year-old prediction. Angew Chem 52:1–5
Schwendinger MG, Rode BM (1992) Investigations on the mechanism of the salt-induced peptide formation. Orig Life Evol Biosph 22:349–359
Schwendinger MG, Tattler R, Saetia S, Liedl KR, Kroemer RT, Rode BM (1995) Salt induced peptide formation: on the selectivity of the copper induced peptide formation under possible prebiotic conditions. Inorg Chim Acta 228:207–214
Usher DA, McHale AH (1976) Nonenzymatic joining of oligoadenylates on a polyuridylic acid template. Science 192:53–54
Vercoutere W, Winters-Hilt S, Olsen H, Deamer DW, Haussler D, Akeson M (2001) Rapid discrimination among individual DNA molecules at single nucleotide resolution using a nanopore instrument. Nat Biotechnol 19:248–250
Verlander MS, Lohrmann R, Orgel LE (1973) Catalysts for the self-polymerization of adenosine cyclic 2′,3′-phosphate. J Mol Evol 2:303–316
Woese CR (1967) The genetic code: the molecular basis for genetic expression. Harper & Row, New York, p 186
Acknowledgments
The hydrothermal simulation research is supported by a generous gift from the Harry Lonsdale Research Award. We are grateful to Jacques Vergne and to Jean-Luc Décout for valuable discussions related to this work, and Veronica De Guzman for expert nanopore analysis of the hydrothermal polymers.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
About this article
Cite this article
Da Silva, L., Maurel, MC. & Deamer, D. Salt-Promoted Synthesis of RNA-like Molecules in Simulated Hydrothermal Conditions. J Mol Evol 80, 86–97 (2015). https://doi.org/10.1007/s00239-014-9661-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-014-9661-9