Abstract
Patterns and processes of molecular evolution critically influence inferences in phylogeny and phylogeography. Within primates, a shift in evolutionary rates has been identified as the rationale for contrasting findings from mitochondrial and nuclear DNA studies as to the position of Tarsius. While the latter now seems settled, we sequenced complete mitochondrial genomes of three Sulawesi tarsiers (Tarsius dentatus, T. lariang, and T. wallacei) and analyzed substitution rates among tarsiers and other primates to infer driving processes of molecular evolution. We found substantial length polymorphism of the D-loop within tarsier individuals, but little variation of predominant lengths among them, regardless of species. Length variation was due to repetitive elements in the CSB domain—minisatellite motifs of 35 bp length and microsatellite motifs of 6 bp length. Amino acid evolutionary rates were second highest among major primate taxa relative to nucleotide substitution rates. We observed many radical possibly function-altering amino acid changes that were rarely driven by positive selection and thus potentially slightly deleterious or neutral. We hypothesize that the observed pattern of an increased amino acid evolutionary rate in tarsier mitochondrial genomes may be caused by hitchhiking of slightly deleterious mutations with favored D-loop length variants selected for maximizing replication success within the cell or the mitochondrion.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
One of the biggest conundrums in primatology—the phylogenetic position of tarsiers—finally seems solved. For decades, scientists have shed light on the place of Tarsius from many perspectives, but neither morphologists nor geneticists came to unequivocal conclusions on where to place these small Southeast Asian primates. With the advance of molecular methodology, however, compelling evidence accumulated for a sister group relationship between tarsiers and extant anthropoids, i.e., a monophyletic haplorhine clade (Schmitz et al. 2001; Jameson et al. 2011; Hartig et al. 2013). Moreover, there is now a good understanding of what caused many molecular studies to find an apparently well-supported prosimian clade uniting tarsiers with strepsirrhine primates, i.e., lemurs, lorises, and bushbabies (Hayasaka et al. 1988; Hasegawa et al. 1990; Eizirik et al. 2001; Murphy et al. 2001; Chatterjee et al. 2009). The difficulty, so it seems, chiefly lays in the combined effects of a rapid sequence of basal primate divisions and a pronounced rate shift in primate mitochondrial (mt) DNA evolution. While nuclear (n) DNA studies most often favor a haplorhine clade (Goodman et al. 1998; Zietkiewicz et al. 1999; Page and Goodman 2001; Schmitz et al. 2001; Poux and Douzery 2004; Jameson et al. 2011; Hartig et al. 2013), the majority of cytoplasmic DNA analyses support monophyletic prosimians (Hayasaka et al. 1988; Hasegawa et al. 1990; Chatterjee et al. 2009), identify tarsiers as the basal primate group (Arnason et al. 2002), or even question their inclusion in the order primates (Andrews et al. 1998). Nonetheless, mtDNA studies dictate the recent literature on tarsier systematics and phylogeography—partly with substantial consequence for taxonomy (Groves and Shekelle 2010).
Analyses of complete mitochondrial genomes of the Western tarsier Tarsius bancanus (Schmitz et al. 2002) and the Philippine T. syrichta (Matsui et al. 2009) highlighted effects of mt gene characteristics on the reconstruction of primate phylogeny. Particularly, a pronounced change in nucleotide composition (Schmitz et al. 2002; Matsui et al. 2009), accelerated mutation rates in non-synonymous sites in anthropoids (Adkins et al. 1996; Andrews et al. 1998; Andrews and Easteal 2000), and/or generally increased mutation rates after tarsiers diverged from higher primates (Schmitz et al. 2002) were invoked to explain contradictory results as to the phylogenetic position of Tarsius. Interestingly, such mutation rate shifts have occurred several times and independently on different primate lineages (Adkins et al. 1996; Schmitz et al. 2002). To date, a closer inspection of evolutionary processes within Tarsiidae was hampered by the lack of data on the extraordinary radiation of this group on Sulawesi (see below).
The tarsier lineage is thought to have diverged from other extant primates between 55 and 81 million years ago (mya) (Adkins et al. 1996; Goodman et al. 1998; Fabre et al. 2009; Matsui et al. 2009; Jameson et al. 2011; Meredith et al. 2011; Perelman et al. 2011; Springer et al. 2012). While the long branch leading to extant tarsiers and the rapid succession of strepsirrhine and tarsiiform split-offs from other primate lineages disguise deep primate phylogeny, the more recent past (Neogene to Quaternary) has seen an almost explosive diversification of tarsiers on the Indonesian island of Sulawesi, the hotspot of extant tarsier diversity (Merker and Groves 2006; Shekelle et al. 2008a, b, 2010; Merker et al. 2010). Sulawesi tarsiers are thought to have split from the Western/Philippine tarsier lineage between ca. 10 mya (Shekelle et al. 2008b; Merker et al. 2009) and 20 mya (Shekelle et al. 2010) (cf. Matsui et al. 2009 for an even older implicit date) when the uplift and expansion of landmasses allowed for the colonization of Wallacea, a transitional region between Asian and Australian biotas. Over the past two decades, a reasonable understanding has developed of the close relationship between Wallacea’s “changing patterns of land and sea” (Hall 2001) and the radiation of tarsiers (Shekelle and Leksono 2004; Merker et al. 2009) and other Sulawesi endemics (Bridle et al. 2001; Evans et al. 2003a, b; Mercer and Roth 2003; Lohman et al. 2011). At the same time, however, the necessity of multi-locus phylogeographic studies in order to overcome peculiarities of mtDNA evolution or mitochondrial introgression among species has become obvious (Evans et al. 2003c; Merker et al. 2009). This phylogeographic background and, especially, the position of Tarsius at the base of the observed compositional and mutation rate shift along the primate lineage make this genus exceptionally important to understand the molecular and evolutionary processes behind this shift.
Now integrating complete mt genomes of three Sulawesi tarsier species, our study thus focused on three major goals. First, we aimed at better understanding how tarsiers colonized Sulawesi. Second, we wanted to shed light on common primate and tarsier-specific characteristics of mtDNA molecular evolution, especially regarding amino acid and nucleotide substitution rates. Our third major goal originates in a report of mtDNA length polymorphism due of tandem-repetitive elements in the mt control region (D-loop) of Western tarsiers (Schmitz et al. 2002). Owing to the regulatory function of the D-loop’s secondary structure in mt-genome replication (Mignotte et al. 1990; Saccone et al. 1991; Sbisà et al. 1997; Pereira et al. 2008), the total length of the repeats might be subject to selection on the intracellular or even intramitochondrial level (Rand 2001). We thus asked whether mitochondrial heteroplasmy—not uncommon in other mammals (e.g., Buroker et al. 1990; Rand 1993; Sbisà et al. 1997)—is rule rather than exception in tarsiers and whether length mutations in the non-coding control region are under selective constraints and thus potentially evolutionarily relevant.
Materials and Methods
Tarsier Sampling
We obtained samples from wild tarsiers in central Sulawesi, Indonesia (Fig. 1). Animals were mist-netted at around dawn or dusk in the vicinity of their sleeping site (see Merker 2006 for details). Small ear biopsies were taken from the tip of the pinna and stored in tissue buffer solution [6 M urea, 10 mM Tris/HCl (pH 8), 10 mM EDTA, 125 mM NaCl, 1 % SDS]. Upon completion of this procedure, tarsiers were released at the capture site. From among 30 samples of three species, we chose one male Tarsius dentatus (“SM10” from Kebun Kopi, 0°43.7′S, 120°01.0′E), one male T. lariang (“SM14” from Powelua, 0°46.7′S, 119°43.3′E), and one female T. wallacei (“SM27” from Batusuya, 0°24.3′S, 119°46.6′E)—all sampled between March and May 2008—as voucher specimens for mt-genomic sequencing. Cyt b sequence information for the Wallace’s tarsier female had already been made available (GenBank: HM115973) in the course of the species description (Merker et al. 2010). Reference specimens for all three species—collected during previous studies—can be accessed at the Museum Zoologicum Bogoriense (MZB) in Bogor, Indonesia. All work strictly complied with international and national law and guidelines.
Species ranges of three central Sulawesi tarsiers and provenance of samples. Complete mitochondrial genomes were sequenced from pictured specimens (SM27, T. wallacei; SM10, T. dentatus; SM14, T. lariang; photos S. Merker). Locations are shown for all samples subjected to PCR of the CSB domain; superscript letters correspond to Online Resource 5
DNA Processing
We extracted genomic DNA from ear biopsies using a DNeasy Blood and Tissue Kit (Qiagen). In order to comply with Indonesian export regulations and to amplify the amount of template material, we performed whole genome amplifications (WGA) using a REPLI-g Kit (Qiagen) according to the manufacturer’s protocol. We used 500 ng of the T. lariang sample SM14 for library preparation following the Roche GS FLX Titanium General Library Preparation protocol and sequenced it on 1/8 of a titanium plate on a 454 sequencer. The Roche-provided program Newbler v2.0.1 was used for assembly of the reads to contigs, with standard settings. We subjected reads and contigs to a BlastN search against the mitochondrial genome of the Western tarsier T. bancanus (GenBank: AF348159). Alignments and assemblies were produced using the software Geneious v5.5.8 (Drummond et al. 2011), available at http://www.geneious.com/) which was also used to annotate the genome (Online Resource 1).
We designed oligonucleotide primers (18–27 bp) to amplify and sequence the whole mtDNA genome of three tarsier species from these reads and contigs using the online program Primer3 (Rozen and Skaletsky 2000). In this first step, primers (Online Resource 2) were chosen to amplify overlapping fragments of 900–1,500 bp length. To avoid amplification of numts (nuclear mitochondrial inserts) instead of genuine mtDNA, we used a LongRange PCR Kit (Qiagen) and the manufacturer’s protocol (ca. 100 ng template in a 25-µl reaction volume; see Online Resource 2 for primers, T a = 52 °C) to amplify two fragments of approximately 8.5 and 10 kb in length whose ends overlapped on opposite sides of the mtDNA molecule. Complete sequence identity in the overlapping parts of the amplificates suggested the circular nature and thus, the genuine mitochondrial origin of the template. Subsequent PCRs were all performed using authenticated long-range amplificates from the respective individual as template. In a second step—to validate sequences and to support double-stranded sequencing of PCR products—we designed additional primers allowing for amplification of shorter (400–600 bp) fragments (Online Resource 3). Finally, highly stringent species-specific primers (Online Resource 4) were designed when cross-species amplification with the T. lariang-derived oligonucleotides was not successful. We used the Taq PCR Core Kit (Qiagen) to amplify fragments (400–1,500 bp) of the mtDNA genome of all three species. Reaction mixes were prepared in 25 µl volumes according to the manufacturer’s protocol; an identical cycling protocol was followed for all reactions: initial denaturation at 92 °C for 2 min, 32 cycles of denaturation at 94 °C for 40 s, primer annealing at 55 °C for 1 min, elongation at 72 °C for 1:30 min, and final extension at 72 °C for 5 min. PCR was performed in an MJ Research PTC-225 Thermo Cycler. PCR products were checked on 1.4 % (w/v) agarose gels and purified using an ExoSAP protocol (Werle et al. 1994) or the PureLink PCR Purification Kit (Invitrogen). Both strands of PCR products were sequenced with ABI 3730 (Applied Biosystems) and CEQ 2000 (Beckman Coulter) automated sequencers. The quality of sequences was estimated using Sequence-Scanner 1.0 (Applied Biosystems) and checked by eye. We then aligned sequences using Geneious v5.5.8. We repeated PCR and sequencing until identical and unambiguous sequences from both DNA strands were retrieved. In a few instances, when double-stranded sequencing was not successful, sequences were validated by analyzing only one DNA strand of a fragment, but from at least three separate PCRs .
To test for intraindividual polymorphism in the mitochondrial control region, a segment of the hypervariable region II (HVII) including conserved sequence blocks CSB1 and CSB2 was amplified (approximately 600 bp, Primer20for and Primer19rev, Online Resource 2) and subjected to TA cloning using a pGEM-T Easy Vector (Promega). We sequenced 20 clones per individual using M13 universal primers.
To estimate intra- and interspecific variability in D-loop length polymorphism, we amplified the HVII of ten individuals per species (covering a wide geographic range, see Fig. 1 and Online Resource 5) using the HotStarTaq Plus Master Mix Kit (Qiagen), a reaction setup according to the manufacturer’s protocol (with 0.4 µM of each primer in 20 µl volumes), and the following cycling conditions: 5 min at 95 °C, 30 cycles of 1 min at 94 °C, 1 min at 62 °C, and 1 min at 72 °C, and then 10 min at 72 °C. PCR products were size fractioned on 2 % (w/v)-agarose gels for 4:45 h at 70 V, stained with ethidium bromide, and exposed to UV light. To minimize the risk of misinterpreting possible repeat-induced polymerase slippage in PCR as a natural process, we extensively varied PCR conditions (template concentrations, annealing temperatures, and cycle numbers). Further to using polymerases as noted above, enzymes or enzyme mixes optimized for reliable amplification of repeat-rich regions (e.g., as from Qiagen’s Type-it Microsatellite PCR Kit) or with proofreading activity (from Qiagen’s LongRange PCR Kit) were applied.
We derived annotations of the complete mt genome from comparison of the three Sulawesi tarsier genomes with published sequences of the Western tarsier T. bancanus (GenBank: AF348159) and the Philippine tarsier T. syrichta (GenBank: AB371090). Transfer RNA sequences were identified using the tRNAscan-SE Search Server (Schattner et al. 2005). The CGView Server (Grant and Stothard 2008) was used to visualize sequence feature information on the annotated mitochondrial genome of T. lariang (Online Resource 1).
Phylogenetic Inference
Aiming at recovering mitogenomic relationships within Tarsius and at testing the utility of phylogenetic inference from single mt genes, we applied MEGA5 to determine best-fitting substitution models for single-gene and total mitogenomic sequence alignments. We calculated overall p-distances among the three Sulawesi tarsiers and T. bancanus separately for all 13 protein-coding genes (all codon positions considered), for the hypervariable region I (HVI) of the control region, and for the complete mt genome (stripped of the control region). Gene tree topologies were recovered with MEGA5 using the maximum likelihood method (with 1,000 bootstraps) and with BEAST MC3 v1.7.4 (Drummond et al. 2012) using Bayesian inference (relaxed-lognormal clock; birth–death tree prior; Markov chain length 10,000,000 (stationarity reached); logged every 1,000 steps), rooted with T. bancanus as outgroup. We also used BEAST with abovementioned specifications to infer the maximum clade credibility tree for the full set of known tarsier mt genomes, now also including T. syrichta (leaving out the control region).
Patterns of Molecular Evolution in Tarsiers
To infer differences in the rates of nucleotide and amino acid evolution among genes, we extracted all protein-coding genes, aligned them separately by codons, and concatenated them. The resulting alignments were then used to estimate site-specific rates of nucleotide and amino acid evolution in MEGA5 (Tamura et al. 2011). The rates are scaled such that the average evolutionary rate across all sites is 1. This means that sites showing a rate <1 are evolving slower than average, and those with a rate >1 are evolving faster than average. These relative rates were then averaged in sliding windows (window size 10 for amino acids, 30 for nucleotides, increase of 1) under the Jones–Taylor–Thornton (JTT) model (+Γ) and the general time reversible model (+Γ) for amino acids and nucleotides, respectively.
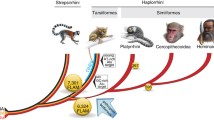
Additionally, we used mitochondrial genomes from GenBank to compare the rate of amino acid evolution relative to nucleotide divergence in tarsiers against other major primate clades (Strepsirrhini, Platyrrhini, Cercopithecidae, and Hominoidea; for GenBank accession numbers see Online Resource 6). First, all protein-coding regions were extracted, per gene codon-wise aligned, and concatenated per taxon. From these alignments, pairwise nucleotide and amino acid distances within clades were calculated under the models described above.
Inference of Selection Patterns
We performed tests for positive selection by applying the codeml algorithm from the PAML program package 4.4 (Yang 2007) to the alignments of all protein-coding genes separately. Calculation of global ω values using the one-ratio model was followed by calculation of branch-specific ω using the free-ratio model. As the free-ratio model is parameter rich and therefore prone to bias, only alignments with significantly better likelihood scores (χ 2, p < 0.02) were used to infer positive selection. We used the inferred maximum clade credibility tree topology (see above) to calculate the branch-specific model. In order to flag potential functional changes, we applied TreeSAAP (Woolley et al. 2003) to compare the differences in 31 physiochemical amino acid properties of observed amino acid changes along the branches of a given phylogenetic tree with the expected random distribution of such differences where every amino acid replacement is equally likely. We used the dataset for all protein-coding mitochondrial genes concatenated with the corresponding maximum clade credibility tree and a sliding window of 20 codons. With the aim of detecting strong positive-destabilizing selection, we only considered most radical amino acid property changes (categories 6, 7, and 8 on a scale of 1–8; at p ≤ 0.001).
Results
Characteristics of Sulawesi Tarsier Mitochondrial Genomes
We sequenced complete mitochondrial genomes of T. dentatus (16,965 bp; GC content 38.8 %; GenBank: KC977310), T. lariang (16,965 bp; GC content 38.6 %; GenBank: KC977309), and T. wallacei (16,957 bp; GC content 38.6 %; GenBank: KC977311). See Fig. 1 for sample provenance and Online Resource 1 for an annotated mt genome of T. lariang. Owing to intraindividual control region length polymorphism (see below), these numbers are not absolute, but correspond to the most frequent D-loop length variant detected in the focal individuals. The gene order and orientation (Online Resources 1 and 7) conform to the ancestral vertebrate state (Inoue et al. 2001). Most protein-coding genes begin with an ATG start codon, but ATA (ND3), ATC (ND2), ATT (ND5), and GTG (ATP6 in T. wallacei) are also employed. Stop codons comprise 10 TAA, three of which are completed by post-transcriptional polyadenylation, two TAG, and one AGA (ND6). As is common in other vertebrate mt genomes, reading frames of some pairs of genes overlap (by up to 43 bases), several genes are separated by non-coding (up to 12-base) spacers (see Online Resource 7).
Phylogenetic Inference
Phylogenetic reconstruction based on complete mtDNA genomes of three Sulawesi tarsiers—rooted with T. bancanus as outgroup—weakly support a sister group relationship between Dian’s and Wallace’s tarsier (Online Resource 8). Inferences based on single genes produced contrasting results with variable—but generally very low—statistical support dependent on the gene and reconstruction method chosen (Online Resource 8). No clear link between substitution rate (measured as DNA distance) and tree topology or support is evident. Including T. syrichta in the analysis resulted in an inferred sister group relationship between Dian’s and Lariang tarsiers (Fig. 2).
Amino acid changes and inferred positive selection along the branches of the tarsier phylogeny. The area of the circles is proportional to the number of amino acid changes along the respective branch. Colors give the proportion of changes in the four mitochondrially encoded OXPHOS complexes. The inner circle is proportional to the number of extreme changes in amino acid properties as inferred by TreeSAAP. Most changes occur in the ND complex. Genes flagged as evolved under positive selection along a branch as inferred by a branch-specific model are given with the respective omega (ω = dN/dS). Very high ω are due to the lack of synonymous substitutions in the respective genes
Patterns of Protein Evolution
The concatenated alignment of all protein-coding mitochondrial genes consists of 11,386 bp corresponding to 3,795 codons (including stop codons). Along the phylogeny, 794 non-synonymous and 2,636 synonymous substitutions were reconstructed. The non-synonymous substitutions affected 619 different codons (16.3 %); some codons were thus substituted more than once. These changes were not equally distributed among genes. Significantly more codons than expected by chance were substituted in the ND complex and significantly less in the COX complex (χ 2 = 126, df = 3, p < 0.00001, Fig. 3, Online Resource 9). Visual inspection of the relative evolutionary rates indicates that (1) these increased rates of amino acid evolution are not equally distributed over the respective genes, but are restricted to certain windows and (2) that increased amino acid evolution is correlated with an increased nucleotide substitution rate (Fig. 3). The rate of amino acid evolution relative to nucleotide divergence differed widely and systematically among different primate lineages. While in all clades, the amino acid divergence scales approximately linear with nucleotide divergence, their slopes were different. It was fastest in the Platyrrhini, followed by tarsiers, the Cercopithecidae, Strepsirrhini, and, finally, Hominoidea (Fig. 4).
Signs of Positive Selection
The omega-based inference yielded signature of positive selection along five branches of the phylogeny (Fig. 2). Positive selection was diagnosed for ATP6, COX2, Cyt b, ND1, ND2, and ND6 on certain branches. Of all 794 observed amino acid changes, 458 (57.6 %) involved a radical change in biochemical properties at the p = 0.001 significance level.
Control Region Heteroplasmy
We found the mitochondrial control region to be length polymorphic in all three species/individuals. Agarose gel electrophoresis of PCR products indicated the section between conserved sequence blocks CSB1 and CSB2 to be the source of this heteroplasmy. Molecular cloning of the region resulted in the identification of 21 different repeat motifs of two length variants: 13 minisatellite motifs of 35 bp length and eight microsatellite motifs of 6 bp length each (Table 1; Fig. 5). The most frequent minisatellite motif was shared between all three species/individuals; others were shared between two species or were found in a single individual only. High sequence similarity among repeat units (Table 1) hints at a common evolutionary origin. The number of tandemly arranged repeats varied between individuals: 0–17 in T. wallacei (16 clones), 2–9 in T. dentatus (13 clones), and 5–20 in five clones of T. lariang containing minisatellite motifs. Microsatellite repeats (3–74 in eight clones containing this motif type) were found in T. lariang only (Fig. 5). Most molecules included several (repeated) motifs, but only of one length variant: either microsatellites or minisatellites. It should be noted, however, that (1) as the analysis was based on one individual of each species only and (2) as the identification of repeat units was disguised by high sequence similarity between repeat motif and flanking regions, this report on the nature and number of motifs and iterations is certainly not exhaustive.
Composition of the tarsier mitochondrial control region between CSB1 (*) and CSB2 (**). Mini- and microsatellite repeats are shown for T. bancanus (GenBank: AF348159; Schmitz et al. 2002), T. wallacei (16 clones [cl]), T. dentatus (13 clones), and T. lariang (13 clones). Colors code for repeat motifs (see Table 1). The orientation is 5′–3′ on the L-strand. Repeat alignment follows their presumed origin by unidirectional replication slippage (5′–3′) on the H-strand (Fumagalli et al. 1996)
To test whether the observed heteroplasmy is common in the three species, we subjected DNA of ten individuals per species to PCR of the hypervariable region II (HVII). All PCR products proved length polymorphic, but showed a concentration of amplified fragments of similar size in all three species (see Online Resource 5).
Discussion
Historic Mitochondrial Introgression
Our phylogenetic reconstructions inferred different gene tree topologies and were dependent on the tree-building method (Online Resource 8) and outgroups included. Moreover, bootstrap support and Bayesian posterior probabilities were generally very low rendering phylogenetic interpretations based solely on tarsier mtDNA little informative. This is even more the case for single-gene analyses. Due to the linked nature of mt genes, conflicting tree topologies—even when receiving reasonable statistical support—are clearly analysis biased rather than reflecting different evolutionary trajectories. If anything, these analyses—with Western tarsiers as outgroup—suggest an association between T. dentatus and T. wallacei. This, however, contradicts Bayesian inference of tree topologies when Philippine tarsiers are included (Fig. 2) and also contrasts findings from nDNA studies providing strong evidence for a closer relationship between T. dentatus and T. lariang (Merker et al. 2010; Driller et al. unpublished data). In view of (1) low statistical support of this and previous studies’ (Shekelle et al. 2008b, 2010) findings, (2) reports of mt introgression in tarsiers (Shekelle et al. 2008b; Merker et al. 2009), and (3) the discordance between mt and nDNA studies, a quick succession of mt lineage divergence events including at least one occasion of mitochondrial capture seems likely. One probable scenario—fitting Online Resource 8—involves asymmetric introgressive hybridization between Dian’s and Wallace’s tarsier soon after the former diverged from (proto-) Lariang tarsiers (or a common ancestor). Another scenario—fitting Fig. 2—implies hybridization between Wallace’s tarsier and an ancestor of Dian’s and Lariang tarsiers shortly before the latter two diverged. Published data suggest reciprocal mt monophyly of these three central Sulawesi tarsier species (Shekelle et al. 2008b; Merker et al. 2010) (but cf. Shekelle et al. 2008b and Merker et al. 2009 for evidence of recent hybridization in species contact zones). This indicates that the implied historic gene flow between widely divergent tarsier lineages resulted in at least one taxon’s complete capture of the mt genome of a congeneric. In view of long speciation intervals, unambiguous tree topologies retrieved from nDNA analyses (Merker et al. 2009, 2010) and low N e of mtDNA, incomplete lineage sorting is an unlikely source of conflicting trees. It is conceivable that an increased accumulation of (slightly) deleterious mutations (see below) may have facilitated the presumed mitochondrial capture, if the introgressing mt genome carried less genetic load.
Origin of Mitochondrial Heteroplasmy
This study documents extensive mitochondrial heteroplasmy in Sulawesi tarsiers. The pattern of variation suggests a high rate of insertion/deletion of repeats in the control region. Additionally, point mutations generate diversity among motifs. The existence of D-loop length polymorphism in Sulawesi tarsiers is hardly surprising: from Western tarsiers (Schmitz et al. 2002) and Philippine tarsiers (Matsui et al. 2009), tandem-repetitive elements between CSB1 and CSB2 were known causing heteroplasmy at least in T. bancanus (Schmitz et al. 2002). Furthermore, mt-genome length polymorphism due to tandemly repeated sequences is widespread across the animal kingdom—in the ETAS and/or CSB domains of the D-loop region (Buroker et al. 1990; Mignotte et al. 1990; Saccone et al. 1991; Ghivizzani et al. 1993; Rand 1993; Hoelzel et al. 1994; Fumagalli et al. 1996; Casane et al. 1997; Sbisà et al. 1997; Wilkinson et al. 1997; Faber and Stepien 1998), but also in other mitochondrially encoded genes (Pfenninger and Bugert 2001). Once such a repeat structure evolved, their length variation seems inevitable: slipped-strand mispairing (Efstratiadis et al. 1980; Levinson and Gutman 1987), intra- and intermolecular recombination and transposition (Rand and Harrison 1989), and replicative misalignment due to competitive displacement between the heavy strand and the D-loop strand (“illegitimate elongation”) (Buroker et al. 1990) have been suggested as responsible for this heteroplasmic variation. Many studies, e.g., on Atlantic cod (Arnason and Rand 1992) thus found the vast majority of mtDNA size variation to lie within rather than between individuals. Our report on repeat motifs and iteration numbers in tarsiers is based on a limited number of mtDNA copies from one individual per species and is thus necessarily incomprehensive. The recognized amount of variability nevertheless exceeds that of most published findings of mitochondrial heteroplasmy.
In accordance with Levinson and Gutman’s (1987) model on the evolution of repeated sequences, the 35-bp repeat motifs can be read—with slight variations—as derivatives of the 6-bp repeats (Table 1). The 3′ end of microsatellite repeat arrays (T. lariang, clone 6-clone 13) is invariably occupied by motif Tl16 (TACACC). The occurrence of this motif in only this position suggests that it is never duplicated or deleted thus corroborating evidence of unidirectional replication of the H-strand in this region (Wilkinson and Chapman 1991). Similar asymmetry in the distribution of variants is known from rabbits (Mignotte et al. 1990), shrews (Fumagalli et al. 1996), elephant seals, and several carnivores (Hoelzel et al. 1994). In minisatellite repeat arrays documented here, however, no difference in sequence divergence between 5′ and 3′ ends is evident.
A Possible Link Between Control Region Heteroplasmy and Protein Evolution
The unevenly increased rates of nucleotide and amino acid evolution in tarsiers (Fig. 4) raise the question after underlying processes. One possibility is that adaptive evolution relating to mitochondrially encoded genes could result in the observed pattern—however, an apparent differential selection pressure on these lowland-rainforest dwellers is lacking, and the number of genes inferred to have evolved under positive selection accounts only for a very small fraction of all observed amino acid changes (Fig. 2). An explanation for a generally increased molecular divergence among tarsier species may be found in their long history of dispersal over the Malay Archipelago. It is conceivable that a series of population bottlenecks and founder events heightened and diversified the relative importance of genetic drift in tarsier evolution. Random drift opposes purifying selection and facilitates accumulation of slightly deleterious alleles (Ohta 1973). Its increased strength owing to low effective population sizes (N e ) of island colonizers thus cannot be ruled out as a source of an increased accumulation of non-synonymous substitutions in Tarsius. We show that large-bodied primates (e.g., hominoids and cercopithecids) have lower average dN/dS rates than small-bodied tarsiers. This contrasts previous findings that in mitochondrial protein-coding genes, larger mammals, generally characterized by lower population sizes (Damuth 1981, 1991, White et al. 2007), have higher dN/dS rates relative to small mammals with high average N e (Popadin et al. 2007, 2013). Thus, the strength of random drift in tarsiers would have been largely shaped by colonization history rather than body size.
Nonetheless, our results on the D-loop length variation allow for suggesting an interesting alternative mechanism which, however, could not be experimentally addressed in the scope of the present study (see Wai et al. 2008 for an example) and remains thus an informed speculation. The mitochondrial genome is subject to selection at various organizational levels, all of which could result in the observed substitution rate variation (Rand 2001). Similar to what is known from evening bats (Wilkinson and Chapman 1991), the existence of intraindividual D-loop length variation with a prevailing length within individuals that is also the major length among individuals suggests selective constraints on the total length of the repeats. Otherwise, if the length variations were neutral, we would expect to see larger differences in main length among unrelated individuals as well as a broader and flatter length distribution within individuals—as shown for rabbits (Mignotte et al. 1990). Given the importance of the loop sequence secondary structure for the regulatory function in mt-genome replication (Mignotte et al. 1990; Saccone et al. 1991; Sbisà et al. 1997; Pereira et al. 2008), it would be, however, not surprising if selection on the intracellular or even intramitochondrial level (Rand 2001) for the most effective replication constantly counteracted the inherent length variation tendency of the repeat structure (Rayko et al. 1988).
We thus hypothesize that constant selection for a certain D-loop length may be invoked to explain the aforementioned pattern of an increased amino acid evolutionary rate in tarsiers (Fig. 4): if selection for replication efficiency is stronger than on gene function or interaction, more and perhaps more than only slightly deleterious mutations could hitchhike to fixation within individuals and eventually populations due to this multilevel selection (Rand 2001). Corroborating evidence for this view arises from the gross disproportion of functionally radical amino acid substitutions to genes flagged as positively selected (Fig. 2). The latter show no consistent pattern but are distributed over all oxidative phosphorylation (OXPHOS) complexes and most branches of the phylogeny. One should perhaps expect such a pattern, if the accumulation of mutations and thus also of (rarer) beneficial mutations was frequent. The accumulation of non-synonymous substitutions is, however, not equally distributed over the protein-coding genes. Regardless of their position in the genome, the genes coding for NADH dehydrogenase subunits have accumulated significantly more amino acid substitutions than the COX subunits (Fig. 3, Online Resource 9). This is in line with other findings in mammals (da Fonseca et al. 2008) and may reflect stronger functional constraints on the mitochondrially encoded COX subunits forming the catalytic core of this enzyme complex.
Conclusions
Our exploratory analysis of tarsier mitochondrial genomes resulted in two major interesting findings: on the one hand, intraindividual D-loop length variation with a major length prevailing within and among individuals in all species, and on the other hand, an increased amino acid evolutionary rate in the protein-coding genes. Bringing tentatively both findings together, we hypothesize that the increased amino acid evolutionary rate may at least partially be caused by hitchhiking of mutations with D-loop length variants selected for maximum replication success within the cell or the mitochondrion. We acknowledge that sophisticated selection experiments or in vitro assays would be necessary to rigorously test our hypothesis which was, however, far beyond the scope of the present study for a multitude of reasons, but may present a way to understand the evolutionary dynamics of the mitochondrial genome. Our results add another twist to recent surprising findings from other parts of the tarsier genome pinpointing the peculiarity of these primates’ molecular evolution, e.g., a likely deficiency in IgE production (Wu et al. 2012) or fully functional color vision in extant tarsiers despite a nocturnal lifestyle (Melin et al. 2013).
References
Adkins RM, Honeycutt RL, Disotell TR (1996) Evolution of eutherian cytochrome c oxidase subunit II: heterogeneous rates of protein evolution and altered interaction with cytochrome c. Mol Biol Evol 13:1393–1404
Andrews TD, Easteal S (2000) Evolutionary rate acceleration of cytochrome c oxidase subunit I in simian primates. J Mol Evol 50:562–568
Andrews TD, Jermiin LS, Easteal S (1998) Accelerated evolution of cytochrome b in simian primates: adaptive evolution in concert with other mitochondrial proteins? J Mol Evol 47:249–257
Arnason E, Rand DM (1992) Heteroplasmy of short tandem repeats in mitochondrial DNA of Atlantic cod, Gadusmorhua. Genetics 132:211–220
Arnason U, Adegoke JA, Bodin K, Born EW, Esa YB et al (2002) Mammalian mitogenomic relationships and the root of the eutherian tree. Proc Natl Acad Sci USA 99:8151–8156
Bridle JR, Garn A-K, Monk KA, Butlin RK (2001) Speciation in Chitaura grasshoppers (Acrididae: Oxyinae) on the island of Sulawesi: colour patterns, morphology and contact zones. Biol J Linn Soc 72:373–390
Buroker NE, Brown JR, Gilbert TA, O’Hara PJ, Beckenbach AT et al (1990) Length heteroplasmy of sturgeon mitochondrial DNA: an illegitimate elongation model. Genetics 124:157–163
Casane D, Dennebouy N, de Rochambeau H, Mounolou JC, Monnerot M (1997) Nonneutral evolution of tandem repeats in the mitochondrial DNA control region of lagomorphs. Mol Biol Evol 14:779–789
Chatterjee HJ, Ho SYW, Barnes I, Groves C (2009) Estimating the phylogeny and divergence times of primates using a super matrix approach. BMC Evol Biol 9:259
da Fonseca RR, Johnson WE, O’Brien SJ, Ramos MJ, Antunes A (2008) The adaptive evolution of the mammalian mitochondrial genome. BMC Genomics 9:119
Damuth J (1981) Population density and body size in mammals. Nature 290:699–700
Damuth J (1991) Of size and abundance. Nature 351:268–269
Drummond AJ, Ashton B, Buxton S, Cheung M, Cooper A, et al (2011) Geneious v5.4. http://www.geneious.com. Accessed 30 Dec 2013
Drummond AJ, Suchard MA, Xie D, Rambaut A (2012) Bayesian phylogenetics with BEAUti and the BEAST 1.7. Mol Biol Evol 29:1969–1973
Efstratiadis A, Posakony JW, Maniatis T, Lawn RM, O’Connell C et al (1980) The structure and evolution of the human β-globin gene family. Cell 21:653–668
Eizirik E, Murphy WJ, O’Brien SJ (2001) Molecular dating and biogeography of the early placental mammal radiation. J Hered 92:212–219
Evans BJ, Brown RM, McGuire JA, Supriatna J, Andayani N et al (2003a) Phylogenetics of fanged frogs: testing biogeographical hypotheses at the interface of the Asian and Australian faunal zones. Syst Biol 52:794–819
Evans BJ, Supriatna J, Andayani N, Melnick DJ (2003b) Diversification of Sulawesi macaque monkeys: decoupled evolution of mitochondrial and autosomal DNA. Evolution 57:1931–1946
Evans BJ, Supriatna J, Andayani N, Setiadi MI, Cannatella DC et al (2003c) Monkeys and toads define areas of endemism on Sulawesi. Evolution 57:1436–1443
Faber JE, Stepien CA (1998) Tandemly repeated sequences in the mitochondrial DNA control region and phylogeography of the Pike-Perches Stizostedion. Mol Phylogenet Evol 10:310–322
Fabre PH, Rodrigues A, Douzery EJP (2009) Patterns of macroevolution among Primates inferred from a supermatrix of mitochondrial and nuclear DNA. Mol Phylogenet Evol 53:808–825
Fumagalli L, Taberlet P, Favre L, Hausser J (1996) Origin and evolution of homologous repeated sequences in the mitochondrial DNA control region of shrews. Mol Biol Evol 13:31–46
Ghivizzani SC, Mackay SL, Madsen CS, Laipis PJ, Hauswirth WW (1993) Transcribed heteroplasmic repeated sequences in the porcine mitochondrial DNA D-loop region. J Mol Evol 37:36–47
Goodman M, Porter CA, Czelusniak J, Page SL, Schneider H et al (1998) Toward a phylogenetic classification of primates based on DNA evidence complemented by fossil evidence. Mol Phylogenet Evol 9:585–598
Grant JR, Stothard P (2008) The CGView Server: a comparative genomics tool for circular genomes. Nucleic Acids Res 36:W181–W184
Groves C, Shekelle M (2010) The genera and species of Tarsiidae. Int J Primatol 31:1071–1082
Hall R (2001) Cenozoic reconstructions of SE Asia and the SW Pacific: changing patterns of land and sea. In: Metcalfe I, Smith JMB, Morwood M, Davidson I (eds) Faunal and floral migrations and evolution in SE Asia-Australasia. A.A. Balkema, Lisse, pp 35–56
Hartig G, Churakov G, Warren WC, Brosius J, Makałowski W, Schmitz J (2013) Retrophylogenomics place tarsiers on the evolutionary branch of anthropoids. Sci Rep 3:1756
Hasegawa M, Kishino H, Hayasaka K, Horai S (1990) Mitochondrial DNA evolution in primates: transition rate has been extremely low in the lemur. J Mol Evol 31:113–121
Hayasaka K, Gojobori T, Horai S (1988) Molecular phylogeny and evolution of primate mitochondrial DNA. Mol Biol Evol 5:626–644
Hoelzel AR, Lopez JV, Dover GA, O’Brien SJ (1994) Rapid evolution of a heteroplasmic repetitive sequence in the mitochondrial DNA control region of carnivores. J Mol Evol 39:191–199
Inoue JG, Miya M, Tsukamoto K, Nishida M (2001) Complete mitochondrial DNA sequence of Conger myriaster (Teleostei: Anguilliformes): novel gene order for vertebrate mitochondrial genomes and the phylogenetic implications for anguilliform families. J Mol Evol 52:311–320
Jameson NM, Hou Z-C, Sterner KN, Weckle A, Goodman M et al (2011) Genomic data reject the hypothesis of a prosimian primate clade. J Hum Evol 61:295–305
Levinson G, Gutman GA (1987) Slipped-strand mispairing: a major mechanism for DNA sequence evolution. Mol Biol Evol 4:203–221
Lohman DJ, de Bruyn M, Page T, von Rintelen K, Hall R et al (2011) Biogeography of the Indo-Australian Archipelago. Annu Rev Ecol Evol Syst 42:205–226
Matsui A, Rakotondraparany F, Munechika I, Hasegawa M, Horai S (2009) Molecular phylogeny and evolution of prosimians based on complete sequences of mitochondrial DNAs. Gene 441:53–66
Melin AD, Matsushita Y, Moritz GL, Dominy NJ, Kawamura S (2013) Inferred L/M cone opsin polymorphism of ancestral tarsiers sheds dim light on the origin of anthropoid primates. Proc R Soc Lond B Biol Sci 280:20130189
Mercer JM, Roth VL (2003) The effects of Cenozoic global change on squirrel phylogeny. Science 299:1568–1572
Meredith RW, Janečka JE, Gatesy J, Ryder OA, Fisher CA et al (2011) Impacts of the Cretaceous Terrestrial Revolution and KPg extinction on mammal diversification. Science 334:521–524
Merker S (2006) Habitat-specific ranging patterns of Dian’s tarsiers (Tarsius dianae) as revealed by radiotracking. Am J Primatol 68:111–125
Merker S, Groves CP (2006) Tarsius lariang: a new primate species from Western Central Sulawesi. Int J Primatol 27:465–485
Merker S, Driller C, Perwitasari-Farajallah D, Pamungkas J, Zischler H (2009) Elucidating geological and biological processes underlying the diversification of Sulawesi tarsiers. Proc Natl Acad Sci USA 106:8459–8464
Merker S, Driller C, Dahruddin H, Wirdateti, Sinaga W et al (2010) Tarsius wallacei: a new tarsier species from central Sulawesi occupies a discontinuous range. Int J Primatol 31:1107–1122
Mignotte F, Gueride M, Champagne AM, Mounolou JC (1990) Direct repeats in the non-coding region of rabbit mitochondrial DNA. Involvement in the generation of intra- and inter-individual heterogeneity. Eur J Biochem 194:561–571
Murphy WJ, Eizirik E, Johnson WE, Zhang YP, Ryder OA et al (2001) Molecular phylogenetics and the origins of placental mammals. Nature 409:614–618
Ohta T (1973) Slightly deleterious mutant substitutions in evolution. Nature 246:96–98
Page SL, Goodman M (2001) Catarrhine phylogeny: noncoding DNA evidence for a diphyletic origin of the mangabeys and for a human-chimpanzee clade. Mol Phylogenet Evol 18:14–25
Pereira F, Soares P, Carneiro J, Pereira L, Richards MB et al (2008) Evidence for variable selective pressures at a large secondary structure of the human mitochondrial DNA control region. Mol Biol Evol 25:2759–2770
Perelman P, Johnson WE, Roos C, Seuánez HN, Horvath JE et al (2011) A molecular phylogeny of living primates. PLoS Genet 7:e1001342
Pfenninger M, Bugert M (2001) Dynamic microsatellites in transcribed regions of gastropod mitochondrial 16S rDNA. Genome 44:163–166
Popadin K, Polishchuk LV, Mamirova L, Knorre D, Gunbin K (2007) Accumulation of slightly deleterious mutations in mitochondrial protein-coding genes of large versus small mammals. Proc Natl Acad Sci USA 104:13390–13395
Popadin KY, Nikolaev SI, Junier T, Baranova M, Antonarakis SE (2013) Purifying selection in mammalian mitochondrial protein-coding genes is highly effective and congruent with evolution of nuclear genes. Mol Biol Evol 30:347–355
Poux C, Douzery EJP (2004) Primate phylogeny, evolutionary rate variations, and divergence times: a contribution from the nuclear gene IRBP. Am J Phys Anthropol 124:1–16
Rand DM (1993) Endotherms, ectotherms, and mitochondrial genome-size variation. J Mol Evol 37:281–295
Rand DM (2001) The units of selection on mitochondrial DNA. Annu Rev Ecol Syst 32:415–448
Rand DM, Harrison RG (1989) Molecular population genetics of mtDNA size variation in crickets. Genetics 121:551–569
Rayko E, Goursot R, Cherif-Zahar B, Melis R, Bernardi G (1988) Regions flanking ori sequences affect the replication efficiency of the mitochondrial genome of ori+ petite mutants from yeast. Gene 63:213–226
Rozen S, Skaletsky HJ (2000) Primer3 on the WWW for general users and for biologist programmers. In: Krawetz S, Misener S (eds) Bioinformatics methods and protocols: methods in molecular biology. Humana Press, Totowa, pp 365–386
Saccone C, Pesole G, Sbisá E (1991) The main regulatory region of mammalian mitochondrial DNA: structure–function model and evolutionary pattern. J Mol Evol 33:83–91
Sbisà E, Tanzariello F, Reyes A, Pesole G, Saccone C (1997) Mammalian mitochondrial D-loop region structural analysis: identification of new conserved sequences and their functional and evolutionary implications. Gene 205:125–140
Schattner P, Brooks AN, Lowe TM (2005) The tRNAscan-SE, snoscan and snoGPS web servers for the detection of tRNAs and snoRNAs. Nucleic Acids Res 33:W686–W689
Schmitz J, Ohme M, Zischler H (2001) SINE insertions in cladistic analyses and the phylogenetic affiliations of Tarsius bancanus to other primates. Genetics 157:777–784
Schmitz J, Ohme M, Zischler H (2002) The complete mitochondrial sequence of Tarsius bancanus: evidence for an extensive nucleotide compositional plasticity of primate mitochondrial DNA. Mol Biol Evol 19:544–553
Shekelle M, Leksono SM (2004) Strategi konservasi di Pulau Sulawesi dengan menggunakan Tarsius sebagai flagship species [Conservation strategy in Sulawesi Island using Tarsius as flagship species]. Biota 9:1–10
Shekelle M, Groves C, Merker S, Supriatna J (2008a) Tarsius tumpara: a new tarsier species from Siau Island, North Sulawesi. Primate Conserv 23:55–64
Shekelle M, Morales JC, Niemitz C, Ichwan LL, Melnick D (2008b) Distribution of tarsier haplotypes for some parts of northern and central Sulawesi. In: Shekelle M, Maryanto I, Groves C, Schulze H, Fitch-Snyder H (eds) Primates of the oriental night. LIPI Press, Jakarta, pp 51–69
Shekelle M, Meier R, Wahyu I, Ting N (2010) Molecular phylogenetics and chronometrics of Tarsiidae based on 12S mtDNA haplotypes: evidence for Miocene origins of crown tarsiers and numerous species within the Sulawesian clade. Int J Primatol 31:1083–1106
Springer MS, Meredith RW, Gatesy J, Emerling CA, Park J et al (2012) Macroevolutionary dynamics and historical biogeography of primate diversification inferred from a species supermatrix. PLoS One 7:e49521
Tamura K, Peterson D, Peterson N, Stecher G, Nei M et al (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Wai T, Teoli D, Shoubridge EA (2008) The mitochondrial DNA genetic bottleneck results from replication of a subpopulation of genomes. Nat Genet 40:1484–1488
Werle E, Schneider C, Renner M, Völker M, Fiehn W (1994) Convenient single-step, one tube purification of PCR products for direct sequencing. Nucleic Acids Res 22:4354–4355
White EP, Morgan Ernest SK, Kerkhoff AJ, Enquist BJ (2007) Relationships between body size and abundance in ecology. Trends Ecol Evol 22:323–330
Wilkinson GS, Chapman AM (1991) Length and sequence variation in evening bat D-loop mtDNA. Genetics 128:607–617
Wilkinson GS, Mayer F, Kerth G, Petri B (1997) Evolution of repeated sequence arrays in the D-loop region of bat mitochondrial DNA. Genetics 146:1035–1048
Woolley S, Johnson J, Smith MJ, Crandall KA, McClellan DA (2003) TreeSAAP: selection on amino acid properties using phylogenetic trees. Bioinformatics 19:671–672
Wu PC, Chen JB, Kawamura S, Roos C, Merker S et al (2012) The IgE gene in primates exhibits extraordinary evolutionary diversity. Immunogenetics 64:279–287
Yang Z (2007) PAML 4: a program package for phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Zietkiewicz E, Richer C, Labuda D (1999) Phylogenetic affinities of tarsier in the context of primate Alu repeats. Mol Phylogenet Evol 11:77–83
Acknowledgments
We thank the Indonesian authorities LIPI, RISTEK, PHKA, and BKSDA for granting research, capture, and export permits and the Primate Research Center at Bogor Agricultural University for providing administrative support. Nicolas Galtier and an anonymous reviewer provided valuable comments on an earlier draft of the manuscript. Sample collection, molecular analyses, and manuscript submission were supported through research grants from the Deutsche Forschungsgemeinschaft (DFG, Me2730/1-2, Me2730/1-3, and Me2730/2-1); the molecular work was partly supported by the research-funding program “LOEWE—Landes-Offensive zur Entwicklung Wissenschaftlich-ökonomischer Exzellenz” of Hesse’s Ministry of Higher Education, Research, and the Arts.
Conflict of interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Merker, S., Thomas, S., Völker, E. et al. Control Region Length Dynamics Potentially Drives Amino Acid Evolution in Tarsier Mitochondrial Genomes. J Mol Evol 79, 40–51 (2014). https://doi.org/10.1007/s00239-014-9631-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-014-9631-2