Abstract
The rise of antibiotic-resistant pathogens focuses our attention on the source of antibiotic resistance genes, on the existence of these genes in environments exposed to little or no antibiotics, and on the relationship between resistance genes found in the clinic and those encountered in non-clinical settings. Here, we address the evolutionary history of a class of resistance genes, the SHV β-lactamases. We focus on bla SHV genes isolated both from clinical and non-clinical sources and show that clinically important resistance determinants arise repeatedly from within a diverse pool of bla SHV genes present in the environment. While our results argue against the notion of a single common origin for all clinically derived bla SHV genes, we detect a characteristic selective signature shaping this protein in clinical environments. This clinical signature reveals the joint action of purifying and positive selection on specific residues, including those known to confer extended-spectrum activity. Surprisingly, antibiotic resistance genes isolated from non-clinical—and presumably antibiotic-free—settings also experience the joint action of purifying and positive selection. The picture that emerges undercuts the notion of a separate reservoir of antibiotic resistance genes confined only to clinical settings. Instead, we argue for the presence of a single extensive and variable pool of antibiotic resistance genes present in the environment.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Clinically relevant antibiotic resistance, first noted soon after the introduction of penicillin for the treatment of wound-related infections in WWII, has increased dramatically over the past 70 years. In that time, our knowledge of the diversity and abundance of resistance mechanisms has expanded considerably, and their presence has led to the current situation: increasing morbidity and mortality due to bacterial infections, increased cost and duration of hospital stays, and a rise in the number of infections that respond to few, if any, existing antibiotics (Bush et al. 2011).
The causes underlying this increase in resistant bacterial pathogens have been extensively scrutinized. They range from the overuse of antibiotics in clinical and agricultural settings to the extensive traffic in resistance determinants (both within and across species boundaries) mediated by a variety of mobile genetic elements (Blaser 2011; Bush et al. 2011; Stokes and Gillings 2011). In response to the rapid spread of these resistance determinants, surveillance networks monitoring the dynamics and spread of resistance are now in place (Grundmann et al. 2011). While these efforts have revealed the speed with which a single resistant clonal lineage or resistance mechanism can spread around the world, the origins of resistance mechanisms remain comparatively unaddressed.
We are particularly interested in understanding the source of the diversity of resistance determinants seen in clinical isolates of pathogenic bacteria. Numerous studies have suggested that these genes arise in a clinical setting in response to human-mediated antibiotic usage (Costelloe et al. 2010; Medeiros 1997; Wright 2010). More recent studies reveal that the environment can also serve as a reservoir for resistance determinants (Allen et al. 2010; Davies and Davies 2010). Here, we explore the relationship between environmentally and clinically derived resistance determinants by examining their molecular evolutionary history.
Our analysis focuses on a specific class of antibiotic resistance genes, the bla SHV genes, which encode β-lactamases. Based on conserved and distinctive amino acid motifs, SHV is considered as a Class A group 2b enzyme (Bush and Jacoby 2010). This enzyme, encoded by a gene that is often located on a plasmid, is of particular interest because of its wide phylogenetic distribution within the Enterobacteriaceae and its presence in a range of clinically relevant pathogens, including Acinetobacter baumannii, Salmonella enterica, Shigella dysenteriae, Yersinia pestis, and Pseudomonas aeruginosa. Functionally, SHV is one of the three major extended-spectrum β-lactamase (ESBL) producers (Drawz and Bonomo 2010). Our attention was thus directed to this class of resistance genes because of their widespread distribution, clinical importance, and the availability of a significant number of sequences in the National Center for Biotechnology Information (NCBI) database.
An understanding of the evolutionary history of bla SHV depends on a dataset that fully captures its sequence diversity. While studies exist that identify this gene in the environment, none provide sequence data for those isolates (Chikwendu et al. 2011; Costa et al. 2006; Girlich et al. 2011; Literak et al. 2010). We address this deficiency here by reporting a set of 28 bla SHV sequences (15 of which are non-redundant) identified by screening 109 strains isolated from fecal samples of Australian placental and marsupial mammals, collected between 1993 and 1997 in environments far from human habitation (Gordon and FitzGibbon 1999). We also report a contemporaneous set of 40 clinically derived sequences (24 of which are non-redundant) of isolates collected from hospitals in Sydney, Australia (Gordon et al. 2005). We complete the dataset by including a sample of 71 sequences that reflect the diversity of non-redundant, full-length bla SHV clinical sequences present in the NCBI database (Online Resource 1). We note that no bla SHV environmental sequences were present in the NCBI database.
Materials and Methods
Bacterial Strains
The bacterial strains in this study were obtained from the Australian Enteric Collection of Dr. David Gordon (Australian National University) (Gordon and FitzGibbon 1999), and were isolated between 1993 and 1997 either from fecal samples of mammals in the Australian outback (regions with little to no human-association) or from hospital patients in Sydney, Australia and screened for the bla SHV gene. Sequences for hospital-derived bla SHV genes (n = 40) were kindly provided by Dr. David Gordon.
Genotypic Characterization
DNA was isolated from bacterial strains with the DNeasy® Tissue Kit (Qiagen, Valencia, CA, USA). Strains were PCR-screened using a standard protocol with primers specific for the bla SHV gene (Online Resource 2). PCR products were purified using the QIAquick® PCR Purification Kit (Qiagen, Valencia, CA, USA) and sequenced with Big Dye® Sequencing Mix V3.1 (Applied Biosystems, Foster City, CA, USA) under standard conditions and run on an ABI 3100 Genetic Analyzer.
Database Sequences
bla SHV sequences were retrieved from NCBI using a non-redundant protein BLAST search queried with Misc025. All sequences with an identity ≥97 % and an E-value ≤10−180 were retrieved; partial sequences were excluded. The origin of every sequence (clinical or environmental) was ascertained using NCBI annotations. Sequences of unknown origin were excluded. Only one identical sequence per origin was gathered, yielding a total of 71 sequences. Duplicate sequences were only retained when they were derived from different environments (See Online Resource 1 for strains included in data analysis). Sequences were aligned using ClustalW2.
Phylogenetic Reconstruction
A Bayesian tree for the bla SHV nucleotide sequences was constructed using MrBayes (Ronquist and Huelsenbeck 2003), (GTR model, across-site rate variation = Invgamma, outgroup accession number AY743416.1). 3 × 106 generations were run with a sampling of trees every 1,000 generations, resulting in convergence and a 0.027 average standard deviation of split frequencies. The “Sump burnin” and “Sumt burnin” options in MrBayes result in the discarding of the first 25 % of the generated samples. Posterior probabilities are reported for any clade falling below 80 %.
Selection Analysis
Nonsynonymous (d N) and synonymous (d S) substitution rates were estimated using the Selecton Server (Stern et al. 2007). Clinical and environmental sequences were treated as separate samples. Branch length optimization was selected and all analyses were run using both model 8 (positive selection) and model 8a (null model). Selected sites were mapped onto the crystal structure of the SHV-1 β-lactamase (PDB 1SHV) (Kuzin et al. 1999). Likelihood values are reported in Online Resource 3.
Results and Discussion
The sequences we examine in this study shed light on the extraordinary diversity of SHV resistance determinants. More than 500 bla SHV sequences, encoding over 150 unique protein sequences, have been deposited in NCBI (2012). Of the Australian environmentally derived bla SHV sequences reported here (n = 28), nine nucleotide and two protein sequences were novel to NCBI. In the Australian clinically derived sequences (n = 24), 12 nucleotide and two protein sequences were also new to NCBI. The Australian environmental and clinical sets share only one previously undescribed SHV protein sequence.
We subjected these bla SHV sequences (n = 110) to two distinct analyses. In the first of these, sequences were aligned and used to generate Bayesian rooted phylogenetic trees based on their nucleotide sequences (Ronquist and Huelsenbeck 2003). The prevailing view concerning the evolution of resistance genes argues that human-mediated selection is the primary driver (Costelloe et al. 2010; Medeiros 1997; Wright 2010), leading to the phylogenetic prediction depicted in Fig. 1a, where all clinically derived SHV sequences form a distinct subclade on the bla SHV phylogeny. Our results contradict that prediction, and instead yield the consensus tree shown in Fig. 1b: a tree with many unresolved nodes, and multiple mixed subclades composed of both Australian environmental and NCBI/Australian clinical sequences. Environmental sequences are distributed throughout the tree; while some subclades are composed exclusively of clinical isolates, this likely reflects the oversampling of clinical isolates and their overrepresentation in the database. Despite the uncertainty surrounding the tree topology, the environmental and clinical bla SHV sequences do not give rise to two distinct subclusters.
Bayesian polar trees of bla SHV clinical and environmental sequences. a Hypothetical tree displaying a clear separation of clinically and environmentally derived sequences. b Actual consensus tree of clinically and environmentally derived bla SHV sequences. There is no clear, definable structure to the tree (largely unresolved) as well as no segregation between those sequences derived from clinical isolates and those derived from environmental isolates. Taxon labels: Black Australian environmental sequences; Light gray Australian clinical sequences; Dark gray NCBI clinical sequences. Posterior probabilities for nodes that were less than 80 % are noted with bullet (•)
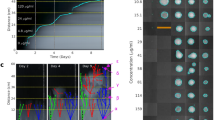
We further explore the relationship between clinical and environmental sequences by examining the footprint of selection on both the datasets. The sequences were subjected to a Bayesian codon-based selection analysis, using estimates of synonymous (d S) and nonsynonymous (d N) substitution rates to detect both the presence and character (positive or purifying) of selection. The results of this analysis are shown in Fig. 2a–c. Panel (a) shows the d N/d S ratio across the entire protein for both the NCBI/Australian clinical and environmental SHV datasets. Panels (b) (NCBI/Australian clinical sequences) and (c) (Australian environmental sequences) show selection results mapped directly onto the resolved crystal structure of the SHV protein (PDB 1SHV) (Kuzin et al. 1999). Space-filled sites highlight amino acid positions at which positive selection is detected. Sites under strong purifying selection are shown in gray. The extent of purifying and positive selection detected suggests that bla SHV genes are not evolving neutrally. We are led to conclude that the encoded proteins play a significant role in the fitness of their bearers, both in settings characterized by high antibiotic use and in environments essentially devoid of human antibiotic use.
Selection analysis results for SHV β-lactamase. a d N/d S ratios at each amino acid position for environmental and clinical datasets. A ratio >1 (dashed line) indicates positive selection. b Clinical selection results mapped onto SHV-1 structure (P < 0.001). c Environmental selection results mapped onto SHV-1 structure (P = 0.010). Backbone colors ranging from light to dark indicate sites of purifying selection (strength of selection increasing with color saturation). Space-filled sites show where positive selection has been detected. Site 31 is the only site where positive selection has been verified by confidence intervals. Positions are numbered based on our alignments; their correspondence to the Ambler numbering system is specified in Online Resource 4
In the NCBI/Australian clinical dataset, positive selection is detected at positions 3, 31, 142, 234, and 235. Two of these sites (positions 234 and 235) occur around the active site and have been associated with ESBL activity (Bradford 2001). Substitutions at site 234 have also been implicated in the binding of larger β-lactam antibiotics, such as cephems with bulky side chains (Lee et al. 1991; Nukaga et al. 2003). In addition, positive selection is detected at site 142 where a study reports a direct link to carbapenemase activity, though reasons for this are not clear (Mendonca et al. 2008; Poirel et al. 2003).
Surprisingly, the Australian environmental dataset revealed a higher proportion of residues under positive selection when compared to their NCBI/Australian clinical counterparts (ten and five residues, respectively) (Online Resource 4). For the environmental dataset, positive selection is detected at positions 3, 14, 31, 65, 125, 137, 140, 152, 191, and 272. Only one site of positive selection (position 65) is detected adjacent to the active-site serine; mutations at this site have been shown to confer resistance to inhibitors such as clavulanic acid (Randegger and Hächler 2001). This observation suggests that positive selection near the active site, generally associated with clinical levels of antibiotic exposure, is not limited to the clinic. Other sites where positive selection is detected are distributed throughout the protein and do not cluster in any particular structural subdomain.
In contrast with the similarity of the sites undergoing purifying selection seen in both samples, there are only two sites (3 and 31) undergoing positive selection shared by the clinical and environmental datasets. Position 3 is not involved in the enzymatic activity of the protein as it is cleaved in the formation of the mature protein (Rice et al. 2000). Position 31 exhibits the highest d N/d S ratio for both clinical and environmental datasets, but its functional significance remains unclear; although it has been suggested that mutations at this site lead to a conformational change in the protein (Nüesch-Inderbinen et al. 1995). Despite some overlap, the marked differences between environmental and clinical SHV samples in the sites undergoing positive selection suggest that our environmental samples are not simply SHV alleles shaped in the clinic that have recently made their way into the broader environment.
Our selection analysis reveals that, in both the environmental and clinical samples, multiple sites distributed around the protein are undergoing selective scrutiny. Coupled with the clear phylogenetic signal we describe, the picture that emerges undercuts the notion of a separate reservoir of antibiotic resistance genes confined only to clinical settings. Instead, we argue for the presence of a single extensive and variable pool of antibiotic resistance genes present in the environment. The bla SHV genes seen in clinical settings thus represent a recurrent sampling of this pool of resistance determinants. This scenario argues for the importance of extensive and systematic sampling of antibiotic resistance determinants in non-clinical settings. Clinically relevant pathogens appear to have a vast and as yet largely uncharacterized cornucopia of resistance determinants at their disposal. A more comprehensive exploration of the environmental reservoir is likely to shed light on the emergence and evolution of genes conferring resistance to antibiotics.
References
Allen HK, Donato J, Wang HH, Cloud-Hansen KA, Davies J, Handelsman J (2010) Call of the wild: antibiotic resistance genes in natural environments. Nat Rev Microbiol 8(4):251–259. doi:10.1038/nrmicro2312
Blaser M (2011) Antibiotic overuse: stop the killing of beneficial bacteria. Nature 476:393–394
Bradford PA (2001) Extended-spectrum beta-lactamases in the 21st century: characterization, epidemiology, and detection of this important resistance threat. Clin Microbiol Rev 14(4):933–951. doi:10.1128/CMR.14.4.933-951.2001
Bush K, Jacoby GA (2010) Updated functional classification of beta-lactamases. Antimicrob Agents Chemother 54(3):969–976. doi:10.1128/AAC.01009-09
Bush K, Courvalin P, Dantas G, Davies J, Eisenstein B, Huovinen P, Jacoby GA, Kishony R, Kreiswirth BN, Kutter E, Lerner SA, Levy S, Lewis K, Lomovskaya O, Miller JH, Mobashery S, Piddock LJ, Projan S, Thomas CM, Tomasz A, Tulkens PM, Walsh TR, Watson JD, Witkowski J, Witte W, Wright G, Yeh P, Zgurskaya HI (2011) Tackling antibiotic resistance. Nat Rev Microbiol 9(12):894–896. doi:10.1038/nrmicro2693
Chikwendu CI, Ibe SN, Okpokwasili GC (2011) Detection of bla SHV and bla TEM beta-lactamase genes in multi-resistant pseudomonas isolates from environmental sources. Afri J Microbiol Res 5(15):2067–2074
Costa D, Poeta P, Sa′enz Y, Vinue L, Rojo-Bezares B, Jouini A, Zarazaga M, Rodrigues J, Torres C (2006) Detection of Escherichia coli harbouring extended-spectrum β-lactamases of the CTX-M, TEM and SHV classes in faecal samples of wild animals in Portugal. J Antimicrob Chemother 58(6):1311–1312. doi:10.1093/jac/dkl414
Costelloe C, Metcalfe C, Lovering A, Mant D, Hay AD (2010) Effect of antibiotic prescribing in primary care on antimicrobial resistance in individual patients: systematic review and meta-analysis. British Med J 340(c2096):1120. doi:10.1136/bmj.c2220;10.1136/bmj.c2553;10.1136/bmj.c2096
Davies J, Davies D (2010) Origins and evolution of antibiotic resistance. Microbiol Mol Biol Rev 74(3):417–433. doi:10.1128/MMBR.00016-10
Drawz SM, Bonomo RA (2010) Three decades of beta-lactamase inhibitors. Clin Microbiol Rev 23(1):160–201. doi:10.1128/CMR.00037-09
Girlich D, Poirel L, Nordmann P (2011) Diversity of clavulanic acid-inhibited extended-spectrum beta-lactamases in Aeromonas spp. from the Seine River, Paris, France. Antimicrob Agents Chemother 55(3):1256–1261. doi:10.1128/AAC.00921-10
Gordon DM, FitzGibbon F (1999) The distribution of enteric bacteria from Australian mammals: host and geographical effects. Microbiology 145:2663–2671
Gordon DM, Stern SE, Collignon PJ (2005) Influence of the age and sex of human hosts on the distribution of Escherichia coli ECOR groups and virulence traits. Microbiology 151(Pt 1):15–23. doi:10.1099/mic.0.27425-0
Grundmann H, Klugman KP, Walsh T, Ramon-Pardo P, Sigauque B, Khan W, Laxminarayan R, Heddini A, Stelling J (2011) A framework for global surveillance of antibiotic resistance. Drug Resist Updat 14(2):79–87. doi:10.1016/j.drup.2011.02.007
Kuzin AP, Nukaga M, Nukaga Y, Hujer AM, Bonomo RA, Knox JR (1999) Structure of the SHV-1 beta-lactamase. Biochemistry 38(18):5720–5727. doi:10.1021/bi990136d
Lee K, Hopkins JD, Syvanen M, O’Brien TF (1991) Gly-238-Ser substitution changes the substrate specificity of the SHV class A β-lactamases. Proteins 11(1):45–51
Literak I, Dolejska M, Radimersky T, Klimes J, Friedman M, Aarestrup FM, Hasman H, Cizek A (2010) Antimicrobial-resistant faecal Escherichia coli in wild mammals in central Europe: multiresistant Escherichia coli producing extended-spectrum beta-lactamases in wild boars. J Appl Microbiol 108(5):1702–1711. doi:10.1111/j.1365-2672.2009.04572.x
Medeiros AA (1997) Evolution and dissemination of beta-lactamases accelerated by generations of beta-lactam antibiotics. Clin Infect Dis 24(Suppl 1):S19–S45
Mendonca N, Manageiro V, Robin F, Salgado MJ, Ferreira E, Canica M, Bonnet R (2008) The Lys234Arg substitution in the enzyme SHV-72 is a determinant for resistance to clavulanic acid inhibition. Antimicrob Agents Chemother 52(5):1806–1811. doi:10.1128/AAC.01381-07
Nüesch-Inderbinen MT, Hächler H, Kayser FH (1995) New system based on site-directed mutagenesis for highly accurate comparison of resistance levels conferred by SHV beta-lactamases. Antimicrob Agents Chemother 39(8):1726–1730. doi:10.1128/aac.39.8.1726
Nukaga M, Mayama K, Hujer AM, Bonomo RA, Knox JR (2003) Ultrahigh resolution structure of a Class A β-Lactamase: on the mechanism and specificity of the extended-spectrum SHV-2 enzyme. J Mol Biol 328(1):289–301. doi:10.1016/s0022-2836(03)00210-9
Poirel L, Heritier C, Podglajen I, Sougakoff W, Gutmann L, Nordmann P (2003) Emergence in Klebsiella pneumoniae of a chromosome-encoded SHV beta-lactamase that compromises the efficacy of imipenem. Antimicrob Agents Chemother 47(2):755–758. doi:10.1128/aac.47.2.755-758.2003
Randegger CC, Hächler H (2001) Amino acid substitutions causing inhibitor resistance in TEM β-lactamases compromise the extended-spectrum phenotype in SHV extended-spectrum β-lactamases. J Antimicrob Chemother 47:547–554
Rice LB, Carias LL, Hujer AM, Bonafede M, Hutton R, Hoyen C, Bonomo RA (2000) High-level expression of chromosomally encoded SHV-1 beta -lactamase and an outer membrane protein change confer resistance to ceftazidime and piperacillin- tazobactam in a clinical isolate of Klebsiella pneumoniae. Antimicrob Agents Chemother 44(2):362–367. doi:10.1128/aac.44.2.362-367.2000
Ronquist F, Huelsenbeck JP (2003) MrBayes 3: Bayesian phylogenetic inference under mixed models. Bioinformatics 19(12):1572–1574. doi:10.1093/bioinformatics/btg180
Stern A, Doron-Faigenboim A, Erez E, Martz E, Bacharach E, Pupko T (2007) Selecton 2007: advanced models for detecting positive and purifying selection using a Bayesian inference approach. Nucleic Acids Res 35(Web Server issue):W506–W511. doi:10.1093/nar/gkm382
Stokes HW, Gillings MR (2011) Gene flow, mobile genetic elements and the recruitment of antibiotic resistance genes into gram-negative pathogens. Fems Microbiol Rev 35(5):790–819. doi:10.1111/j.1574-6976.2011.00273.x
Wright GD (2010) Q&A: antibiotic resistance: where does it come from and what can we do about it? BMC Biology 8:123–128
Acknowledgments
This work was supported by National Institutes of Health 5R01GM068657-04 and National Institutes of Health American Reinvestment and Recovery Act of 2009 Supplement 3R01GM068657-04S1. We thank Dr. David Gordon for sharing his unpublished sequences and for his support during the collection phase of this project. We also thank Dr. Junhyong Kim for statistical advice.
Conflict of Interest
The authors declare that they have no conflict of interest.
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Online Resource 1
bla SHV Sequences Included in Data Analysis
Online Resource 2
Primers Targeting the bla SHV Gene
Online Resource 3
Selection Analysis Likelihood Values
Online Resource 4
Sites of Positive Selection
Rights and permissions
About this article
Cite this article
Dorit, R.L., Roy, C.M., Robinson, S.M. et al. The Evolutionary Histories of Clinical and Environmental SHV β-Lactamases are Intertwined. J Mol Evol 76, 388–393 (2013). https://doi.org/10.1007/s00239-013-9574-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-013-9574-z