Abstract
The salivary androgen-binding proteins (ABPs) are members of the secretoglobin gene family present in mammals. Each ABP is a heterodimer assembled as an ABPA subunit encoded by an Abpa gene and linked by disulfide bridges to an ABPBG subunit encoded by an Abpbg gene. The ABP dimers are secreted into the saliva of mice and then transferred to the pelage after grooming and subsequently to the environment allowing an animal to mark territory with a biochemical signal. The putative role of the mouse salivary ABPs is that of pheromones mediating mate selection resulting in assortative mating in the Mus musculus species complex. We focused on comparing patterns of molecular evolution between the Abpa genes expressed in the submaxillary glands of species of New World and Old World muroids. We found that in both sets of rodents the Abpa genes expressed in the submaxillary glands appear to be evolving under a similar evolutionary regime, with relatively high nonsynonymous substitution rates, suggesting that ABP might play a similar biological role in both systems. Thus, ABP could be involved with mate recognition and species isolation in New World as well as Old World muroids.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Speciation can be driven by the isolation of two populations through geographic, temporal, ecological, or behavioral barriers (see Coyne and Orr 2004 for an extended discussion). In many mammals olfaction is a dominant sensory modality and chemical cues can be used to convey information about individuality. In the common house mouse (Mus musculus) subspecies complex, the salivary androgen-binding proteins (ABPs) are hypothesized to be a component of such a cue, as they are thought to mediate mate recognition (Laukaitis et al. 1997; Talley et al. 2001). ABPs are secretoglobins (Klug et al. 2000; Laukaitis et al. 2005; Laukaitis and Karn 2005; Mukherjee and Chilton 2000) present in the saliva following expression within the submaxillary, sublingual, and parotid glands (Dlouhy et al. 1986; Laukaitis et al. 2005). The putative biological function of mouse salivary ABPs is that of a pheromone, mediating mate selection resulting in assortative mating and incipient reinforcement at the edges of the house mouse hybrid zone in Europe (Bímová et al. 2005, 2011).
ABP is secreted into the saliva and transferred to the pelage and environment after grooming, allowing an animal to mark territory with a biochemical signal. From a genomic standpoint, the mouse ABP system was thought to consist of three single copy genes, Abpa, Abpb, and Abpg, encoding for the three separate subunits, ABPA, ABPB, and ABPG, respectively (Dlouhy et al. 1987; Karn and Laukaitis 2003). Genomic comparisons, however, revealed a more complex pattern (Emes et al. 2004; Laukaitis et al. 2008; Karn and Laukaitis 2009). Most mammals have a single Abpa gene encoding an ABPA protein and a single Abpbg gene encoding an ABPBG protein in the Abp locus, however, rat and house mouse possess multiple paralogs in the corresponding location (Laukaitis et al. 2008; Karn and Laukaitis 2009). In the case of house mouse there are 64 Abp paralogs (30 Abpa and 34 Abpbg genes) over a 3 megabase (Mb) region, whereas the rat genome has 6 paralogs (3 Abpa and 3 Abpbg paralogs). Analyses of intron sequences suggest that these expansions occurred independently in these two species (Laukaitis et al. 2008). In house mouse, the Abpa27 paralog has been the most studied gene in the Abp locus. The translated gene ABPA27 forms a dimer with either ABPBG26 or ABPBG27 via disulfide bridges and is expressed in the submaxillary glands of both males and females. Evidence suggests the ABPA27 subunit plays a significant role in conspecific recognition and mate selection (Bímová et al. 2005; Hwang et al. 1997; Karn et al. 2002; Laukaitis et al. 1997; Talley et al. 2001). Interestingly, recent studies also suggest that additional Abpbg paralogs secreted into the saliva of house mouse, specifically Abpbg26 and Abpbg27 show patterns of molecular variation suggesting adaptive evolution (Laukaitis et al. 2012).
Protein coding genes associated with reproductive and chemosensory roles have a tendency to evolve rapidly and display signatures of positive selection (Duret and Mouchiroud 2000; Kosiol et al. 2008; Park et al. 2011; Swanson and Vacquier 2002; Torgerson et al. 2002). In line with these expectations, comparisons among different Mus sp. alleles revealed high rates of nonsynonymous to synonymous substitutions, and comparative analyses among lineages within the Mus species complex have shown that distinct Abpa27 alleles are fixed in the different subspecies of M. musculus, and molecular evolution analyses of mouse Abpa27 sequences detected strong signals of Darwinian selection (Emes et al. 2004; Hwang et al. 1997; Karn and Nachman 1999; Karn et al. 2002). Therefore the combination of controlled mate choice experiments and the observed genetic signatures consistent with positive selection suggests that ABPs probably play a role in olfactory communication, assortative mating, and incipient reinforcement of reproductive isolation (Bímová et al. 2005, 2011; Karn and Dlouhy 1991; Karn et al. 2002; Laukaitis et al. 2005). The expression and evolution of ABP have been extensively studied in Mus (Dlouhy et al. 1987; Emes et al. 2004; Karn et al. 2010; Laukaitis et al. 2008; Laukaitis and Karn 2005; Karn and Laukaitis 2009). However, few data have been presented for other rodent genera: Karn and Dlouhy (1991) extracted and identified ABP in the saliva for New World rodents, and Laukaitis et al. (2008) identified two putative paralogs of Abpa in Apodemus sylvaticus.
In this study, our objective was to explore whether the patterns of variation in the Abpa genes that encode for the ABPA subunit expressed in the submaxillary glands are similar in New and Old Word rodents. In particular, we focused on comparing patterns of molecular evolution between the Abpa27 gene in house mouse, with their functional counterparts in other Old and New World rodent species. To do so, we analyzed partial sequence data from mRNA transcripts isolated from submaxillary glands of selected New and Old World muroids and used maximum likelihood methods to characterize patterns of molecular evolution. Our results indicate that the Abpa genes expressed in the submaxillary glands are evolving rapidly in both rodent groups, and that a similar set of codon positions appears to be under positive selection in the two systems. Given that proteins involved in species isolation tend to evolve rapidly, our findings would suggest that ABPs might play similar roles related to maintaining species boundaries in both sets of species.
Materials and Methods
Data Collection
Representatives of New World muroids were collected from West Texas and Ecuador and Old World muroid representative rodents were collected from northern Ukraine. Voucher specimens are stored in the Natural Science Research Laboratory collections at the Texas Tech Museum. The submaxillary gland was removed from the animal and stored at −70 °C. Isolation of mRNA, cDNA preparation, PCR amplification of expressed Abpa, and sequencing of amplicons followed Wickliffe et al. (2002). In addition, we obtained additional known expressed Abpa sequences from Spermophilus tridecemlineatus, Mus sp., and Rattus norvegicus from Karn et al. (2002) and Laukaitis et al. (2008) (Supplementary File 1). Mus spicilegus was not included from Karn et al. (2002) given the exact sequence identity to M. macedonicus. In the case of heterozygous individuals, haplotypes were resolved using PHASE version 2.1 (Stephens and Donnelly 2003). Previously unpublished Abpa sequences were deposited in GenBank under the accession numbers JX275970–JX275986.
Sequence Alignment and Analysis
Partial Abpa cDNA sequences were aligned by codons with MUSCLE (Edgar 2004) as implemented in MEGA 5 (Tamura et al. 2011). We explored alignment sensitivity by comparing the MUSCLE alignment to the results of ClustalW (Thompson et al. 1994), MAFFT (Katoh et al. 2005), and PRANK (Löytynoja and Goldman 2005). Alternative alignment strategies yielded the same alignment, which was used for downstream analyses. The intraspecific number of haplotypes, nucleotide diversity, and the uncorrected average pairwise number of differences for nucleotide and amino acid changes were determined in MEGA 5. Intrageneric numbers of nonsynonymous and synonymous changes were counted within Apodemus, Mus, Peromyscus, and Reithrodontomys. Once redundant alleles were removed from the alignment, we reconstructed phylogenetic relationships using maximum likelihood as implemented in Treefinder version March 2011 (Jobb et al. 2004), and we evaluated support for the nodes with 1,000 bootstrap pseudoreplicates. We used the “propose model” tool of Treefinder to select the best-fit models of nucleotide substitution, with an independent model at each codon position in nucleotide analyses. Model selection was based on the Akaike information criterion with correction for small sample size.
We then estimated patterns of molecular evolution using the maximum likelihood approach described by Goldman and Yang (1994) as implemented in CODEML in PAML (Yang 2007). In order to estimate the putative role of negative and positive selection, we compared the rate of nonsynonymous substitution per nonsynonymous site (d N) to the rate of synonymous substitution per synonymous site (d S). The d N/d S ratio (also labeled as ω) can be used to measure the selective regime of a given codon, as similar rates of nonsynonymous and synonymous substitution (ω ≈ 1) are indicative of neutral evolution, an excess of synonymous mutations (ω < 1) is indicative of purifying, or negative selection, and an excess of nonsynonymous mutations (ω > 1) is indicative of positive Darwinian selection, or adaptive evolution. We compared models that allow ω to vary among codons in the alignment (M0 vs. M3, M1a vs. M2a, M7 vs. M8, and M8a vs. M8). In all cases, we used likelihood ratio tests (LRTs) to compare nested sets of model (Yang 1998). Bayesian Empirical Bayes (BEB) was used to calculate the posterior probabilities for sites under positive selection in models M2a and M8 (Yang et al. 2005). These analyses were performed separately for New World and Old World muroids, because orthology between Abpa27 in Mus and our New World taxa cannot be guaranteed. To account for the potential problem generated by intraspecific polymorphism, these analyses were repeated using only one randomly selected representative from each species. Phylogenetic trees were reconstructed from each subset under the same parameters as the full data set.
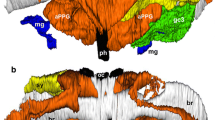
Residues were considered to be under selection if a position was predicted to be under positive selection with posterior probability >0.9 in one model and >0.5 in one other model. We predicted the Mus ABPA27 amino acid structure using Phyre2 (Kelly and Sternberg 2009). Residues under selection in New World rodents and Old World rodents were mapped onto the structure. Swiss-PDBviewer (Guex and Peitsch 1997) was used to manipulate the protein structure and the 3D image was rendered with POV-Ray (http://www.povray.org). Relative solvent accessibilities of residues were calculated based on criteria given in Emes et al. (2004). Relative accessibility values were divided into three categories, buried (<9 % relative accessibility), intermediate (9–35 % relative accessibility), and exposed (>35 % relative accessibility) (Emes et al. 2004; Rost and Sander 1994).
PolyPhen-2 (Polymorphism Phenotyping ver. 2.0; http://genetics.bwh.harvard.edu/pph2/index.shtml), an in silico tool for predicting structural and functional effects of amino acid substitutions on proteins, was used to explore the possible impacts of amino acid variants that appeared to be under strong, positive selection within and among New World and Old World rodent lineages (Adzhubei et al. 2010). Default options were used for all analyses.
Results
Description of Data
We obtained sequences corresponding to 213 bp of the 279 nucleotides of coding sequence of Abpa transcripts from submaxillary glands from 34 rodent specimens. Twenty-seven samples were collected from New World species, 9 Peromyscus leucopus, 6 Peromyscus maniculatus, 3 Reithrodontomys fulvescens, 1 Akodon aerosus, 6 Sigmodon hispidus, 2 Oligoryzomys microtis; and six samples were collected from Old World species, 1 Microtus oeconomus, 1 Apodemus agrarius, 1 Apodemus flavicollis, and 3 Apodemus sylvaticus (Supplementary Table 1). The Apodemus sequences were previously isolated and described in Wickliffe et al. (2002). Even though M. oeconomus was collected in northern Ukraine, the genus is more closely related to New World muroid rodents. Therefore, the M. oeconomus Abpa cDNA sequence was compared to the remaining New World muroid rodents in our study. The Abpa sequence of S. tridecemlineatus reported by Laukaitis et al. (2008), which corresponds to a single-copy gene in the spetri2 assembly of the squirrel genome, was included as an outgroup for phylogenetic analyses. All alignment strategies yielded similar results, with no gaps or premature stop codons found among cDNA-derived sequences (Supplementary File 1). Intraspecific pairwise comparisons of the number of nucleotide substitutions ranged from 0 to 14 and from 0 to 12 when comparing amino acid sequences (Table 1). Despite the relatively small sample sizes, we found that A. sylvaticus, P. leucopus, P. maniculatus, and R. fulvescens were polymorphic for the Abpa gene in our sample, whereas there was no sequence variation among the six specimens of Sigmodon hispidus surveyed (Table 1).
We then estimated phylogenetic relationships among the different alleles using maximum likelihood, excluding redundant haplotypes. In the resulting tree the New World and Old World muroid sequences fell in reciprocally monophyletic clades, as expected given current estimates of organismal phylogeny (Fig. 1; Jansa and Weksler 2004; Steppan et al. 2004). In general, the sequences of given genera were monophyletic, with the exception of the rat Abpa paralogs (Fig. 1). Phylogenies based on intronic sequence suggest that the presence of multiple paralogs of Abpa in rat and mouse derived from lineage-specific expansions (Laukaitis et al. 2008). Our results differ from those of Laukaitis et al. (2008) as the rat paralogs did not form a monophyletic clade. However, an approximately unbiased topology test (Shimodaira 2002) could not discriminate between the maximum-likelihood tree and a tree where the rat paralogs were constrained to monophyly. This issue warrants further attention once intronic sequences from the Abp genes of a wider selection of Old World muroid rodents become available.
Phylogenetic reconstructions from the 210 bp fragment of Abpa. a Maximum likelihood phylogeny for all newly isolated Abpa cDNA sequences, and previously published Old World orthologs and paralogs, excluding redundant haplotypes. b Maximum likelihood phylogeny for the Old World Abpa sequences included in the molecular evolution analyses and c maximum likelihood phylogeny for the New World Abpa sequences included in the molecular evolution analyses
Patterns of Molecular Evolution
As a starting point, we compared nonsynonymous and synonymous substitution rates in a pairwise manner. We detected an excess of nonsynonymous substitutions relative to synonymous substitutions in intrageneric and intraspecific comparisons within the Old World and New World muroid groups. This is especially noticeable in comparisons within R. fulvescens (Supplementary Table 2). In line with these results, our estimates of d N/d S (=ω) for the whole fragment sequenced were high for both data sets, 0.9 for New World muroids and 1.18 for Old World rodents (Supplementary Table 3). In general, estimates of d N/d S averaged over all sites in excess of 0.5 have been considered as suggestive of positive Darwinian selection (Swanson et al. 2004).
We then compared different models of molecular evolution that explore variation in ω among codons in a tree-based approach. We first explored whether there was evidence of variation in ω among codons. We compared models M0, which assigns the same value of ω to all codons with M3, a model that groups codons in three separate classes with independent estimates of ω for each class. There was significant variation in ω, as the LRT-rejected M0 in favor of M3 in both cases (Table 2). We then looked for evidence of positive Darwinian selection among our samples by comparing models that allow a class of sites to have ω > 1, M2a and M8, with the corresponding models that restrict all ω estimates to be ≤1, M1a, M7, and M8a. In all comparisons, M1a versus M2a, M7 versus M8, and M8a versus M8, the models that allow a class of sites to have ω > 1 were favored by the corresponding LRTs (Table 2). Under the criteria given, 5 and 12 sites were found to be evolving under positive selection in New World and Old World muroids, respectively (Table 2; Supplementary Table 3). Similar results were obtained when we reran the analyses using only one representative from each species (Table 2). Most of the sites were found to be in exposed regions of the ABPA27 structure in both the New World and Old World groups (Supplementary Table 4; Fig. 2). We then focused on amino acid variants at sites 68 and 69 [90 and 91 in the accessioned amino acid sequence for M. musculus (http://www.ncbi.nlm.nih.gov/protein/NP_033726.1)] as these sites exhibited marked positive selection in the New World and somewhat in the Old World muroids. In silico simulations with PolyPhen-2 suggest that changes in either of these sites do not appear to alter the structure or putative function of the Abpa.
Discussion
In house mouse, the ABP heterodimers present in the saliva appear to play a behavioral role in reproductive isolation (Laukaitis et al. 1997; Talley et al. 2001; Laukaitis et al. 2012). Most of these studies have focused on the Abpa27 gene, which is expressed in the submaxillary glands in both male and female Mus musculus (Laukaitis et al. 2005). In the M. musculus subspecies complex, several lines of evidence suggest salivary ABP plays an important role in assortative mating. First, alternative alleles are fixed in each M. musculus subspecies at the Abpa27 locus (Hwang et al. 1997; Karn and Dlouhy 1991). Second, in laboratory experiments, female mice preferred to associate and mate with males of their own Abpa27 genotype significantly more often than with males carrying a different allele (Laukaitis et al. 1997; Talley et al. 2001). Lastly, comparisons of evolutionary rates among Abpa27 alleles in different species of mice revealed a large excess of nonsynonymous substitutions over synonymous substitutions, consistent with positive Darwinian selection (Hwang et al. 1997; Karn and Nachman 1999; Karn et al. 2002).
Because the rat and mouse Abpa repertoires derived from largely independent sets of duplications, resolving orthology for the expressed genes and predicted proteins can be difficult. However, the evidence at hand indicates that all of the Abpa genes in mammals were derived from the single copy ancestral Abpa gene present in the common ancestor of placental mammals. The most recent assessments of variation in the gene complement of the Abp gene family (Emes et al. 2004; Laukaitis et al. 2008; Karn and Laukaitis 2009) indicate that the duplications that gave rise to the presence of multiple Abpa genes in mouse genome are specific to the mouse lineage (see Fig. 1), therefore, we infer all mouse Abpa paralogs are co-orthologs with the ancestral single copy Abpa gene. We have no evidence of the presence of multiple Abpa paralogs in New World rodents; thus, we assumed that Abpa gene from New World rodents is an ortholog of the ancestral single-copy gene. Because of this shared ancestry and tissue-specific patterns of expression, we assumed that the Abpa27 genes of house mouse were functionally equivalent to the Abpa genes expressed in the submaxillary glands of other rodents. To ensure this, we extracted and sequenced cDNA from the salivary glands in New World rodent group using primers designed to amplify Abpa27 in Mus (Hwang et al. 1997). If the proposed role of the APBA subunit of New World rodents is to act as a mate choice pheromone, as the ABPA27 subunit does in Mus, we would expect to observe similar patterns of molecular evolution in New and Old World rodents. In our case, we addressed this question by comparing patterns of molecular evolution between Old World and New World muroids.
Our results suggest that both the New and Old World muroid Abpa sequences share the signature of positive Darwinian selection. In both sets of sequences we found high rates of nonsynonymous substitutions within and among genera (Supplementary Table 2), consistent with positive selection. In particular, all analyses indicate that a similar subset of positions appears to be under positive selection within these two separate rodent lineages. These results suggest that changes in a few key residues might play a significant role in the evolution of this protein. Also, a prediction of solvent accessibility is consistent with previous results where the majority of sites under positive selection are mostly in exposed regions of the protein (Fig. 2; Supplementary Table 4). Because there is strong evidence that Abpa27 plays an important role in speciation among Old World muroids, we speculate that the ABPA subunits might be playing a similar role among the New World rodents studied.
Interestingly, the patterns of intraspecific variation we identified are not entirely consistent with those reported for the M. musculus subspecies complex. The signal of positive selection detected among New World muroids is not as strong as the one detected among Old World rodents. There were five sites under selection inferred among New World rodents compared to the 12 sites detected in the Old World muroids. In addition, despite our limited sample size, we found significant levels of intraspecific variation in species of the genera Apodemus, Peromyscus and Reithrodontomys (Supplementary Table 2). Thus, the data would suggest that the putative role of the Abpa gene in New World muroids might be slightly different relative to the M. musculus complex where different alleles segregate with different subspecies (Bímová et al. 2005; Bímová et al. 2011). The fact that alternative variants of the Abpa27 paralog are fixed in the two subspecies M. m. musculus and M. m. domesticus and that the ABPA27 subunit is apparently involved with reinforcing reproductive barriers in the Central Europe hybrid zone between these two subspecies could account for the higher rate of molecular evolution in this system relative to New World rodents (Bímová et al. 2005; Bímová et al. 2011).
In rodents, chemical cues and pheromones play a central role in individual recognition as evidenced by the approximately 1,000 functional olfactory receptors and 212 vomeronasal receptors found in the mouse genome (Shi and Zhang 2009; Zhang and Firestein 2002). Currently, the primary hypotheses proposed to explain patterns of ABP evolution are related to assortative mating, and the coevolution of the ABP pheromone and vomeronasal (VNO) receptors of the V1R receptor family (Karn et al. 2010). As suggested by Karn et al. (2010), it may be that these two systems, ABP and V1R receptors, are coevolving to promote and maintain species boundaries. It would be interesting to identify the V1Rs involved in this putative interaction, and evaluate whether positive selection is also acting on their evolution.
References
Adzhubei IA, Schmidt S, Peshkin L, Ramensky VE, Gerasimova A, Bork P, Kondrashov AS, Sunyaev SR (2010) A method and server for predicting damaging missense mutations. Nat Methods 7:248–249
Bímová B, Karn RC, Piálek J (2005) The role of salivary androgen-binding protein in reproductive isolation between two subspecies of house mouse: Mus musculus musculus and Mus musculus domesticus. Biol J Linn Soc 84:349–361
Bímová BV, Macholán M, Baird SJE, Munclinger P, Dufková P, Laukaitis CM, Karn RC, Luzynski K, Tucker PK, Piálek J (2011) Reinforcement selection acting on the European house mouse hybrid zone. Mol Ecol 20:2403–2424
Coyne JA, Orr HA (2004) Speciation. Sinauer Associates, Sunderland
Dlouhy SR, Nichols WC, Karn RC (1986) Production of an antibody to mouse salivary androgen binding protein (ABP) and its use in identifying a prostate protein produced by a gene distinct from Abp. Biochem Genet 24:743–763
Dlouhy SR, Taylor BA, Karn RC (1987) The genes for mouse salivary androgen-binding protein (ABP) subunits alpha and gamma are located on chromosome 7. Genetics 115:535–543
Duret L, Mouchiroud D (2000) Determinants of substitution rates in mammalian genes: expression pattern affects selection intensity but not mutation rate. Mol Biol Evol 17:68–74
Edgar RC (2004) MUSCLE: multiple sequence alignment with high accuracy and high throughput. Nucleic Acids Res 32:1792–1797
Emes RD, Riley MC, Laukaitis CM, Goodstadt L, Karn RC, Ponting CP (2004) Comparative evolutionary genomics of androgen-binding protein genes. Genome Res 14:1516–1529
Goldman N, Yang Z (1994) A codon-based model of nucleotide substitution for protein-coding DNA sequences. Mol Biol Evol 11:725–736
Guex N, Peitsch MC (1997) SWISS-MODEL and the Swiss-Pdb Viewer: an environment for comparative protein modeling. Electrophoresis 18:2714–2723
Hwang JM, Hofstetter JR, Bonhomme F, Karn R (1997) The microevolution of mouse salivary androgen-binding protein (ABP) paralleled subspeciation of Mus musculus. J Hered 88:93–97
Jansa SA, Weksler M (2004) Phylogeny of muroid rodents: relationships within and among major lineages as determined by IRBP gene sequences. Mol Phylogenet Evol 31:256–276
Jobb G, Von Haeseler A, Strimmer K (2004) TREEFINDER: a powerful graphical analysis environment for molecular phylogenetics. BMC Evol Biol 4:18
Karn RC, Dlouhy SR (1991) Salivary androgen-binding protein variation in Mus and other rodents. J Hered 82:453–458
Karn RC, Laukaitis CM (2003) Characterization of two forms of mouse salivary androgen-binding protein (ABP): implications for evolutionary relationships and ligand-binding function. Biochemistry 42:7162–7170
Karn RC, Laukaitis CM (2009) The mechanism of expansion and the volatility it created in three pheromone gene clusters in the mouse (Mus musculus) genome. Genome Biol Evol 1:494–503
Karn RC, Nachman MW (1999) Reduced nucleotide variability at an androgen-binding protein locus (Abpa) in house mice: evidence for positive natural selection. Mol Biol Evol 16:1192–1197
Karn RC, Orth A, Bonhomme F, Boursot P (2002) The complex history of a gene proposed to participate in a sexual isolation mechanism in house mice. Mol Biol Evol 19:462–471
Karn RC, Young JM, Laukaitis CM (2010) A candidate subspecies discrimination system involving a vomeronasal receptor gene with different alleles fixed in M. m. domesticus and M. m. musculus. PLoS One 5:e12638
Katoh K, Kuma K, Toh H, Miyata T (2005) MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res 33:511–518
Kelly LA, Sternberg MJE (2009) Protein structure prediction on the web: a case study using the Phyre server. Nat Protoc 4:363–371
Klug J, Beier HM, Bernard A, Chilton BS, Fleming TP, Lehrer RI, Miele L, Pattabiraman N, Singh G (2000) Uteroglobin/Clara cell 10-kDa family of proteins: nomenclature committee report. Ann N Y Acad Sci 923:348–354
Kosiol C, Vinař T, Da Fonseca RR, Hubisz MJ, Bustamante CD, Nielsen R, Siepel A (2008) Patterns of positive selection in six mammalian genomes. PLoS Genet 4:e1000144
Laukaitis CM, Karn RC (2005) Evolution of the secretoglobins: a genomic and proteomic view. Biol J Linn Soc 84:493–501
Laukaitis CM, Critser ES, Karn RC (1997) Salivary androgen-binding protein (ABP) mediates sexual isolation in Mus musculus. Evolution 51:2000–2005
Laukaitis CM, Dlouhy SR, Emes RD, Ponting CP, Karn RC (2005) Diverse spatial, temporal, and sexual expression of recently duplicated androgen-binding protein genes in Mus musculus. BMC Evol Biol 5:40
Laukaitis CM, Heger A, Blakley TD, Munclinger P, Ponting CP, Karn RC (2008) Rapid bursts of androgen-binding protein (Abp) gene duplication occurred independently in diverse mammals. BMC Evol Biol 8:46
Laukaitis CM, Mauss C, Karn RC (2012) Congenic strain analysis reveals genes that are rapidly evolving components of a prezygotic isolation mechanism mediating incipient reinforcement. PLoS One 7:e35898
Löytynoja A, Goldman N (2005) An algorithm for progressive multiple alignment of sequences with insertions. Proc Natl Acad Sci USA 102:10557–10562
Mukherjee AB, Chilton BS (2000) The uteroglobin/Clara cell protein family. Ann N Y Acad Sci 923:1–358
Park SH, Podlaha O, Grus WE, Zhang J (2011) The microevolution of V1r vomeronasal receptor genes in mice. Genome Biol Evol 3:401–412
Rost B, Sander C (1994) Conservation and prediction of solvent accessibility in protein families. Proteins Struct Funct Bioinform 20:216–226
Shi P, Zhang J (2009) Extraordinary diversity of chemosensory receptor gene repertoires among vertebrates. Results Probl Cell Differ 47:1–23
Shimodaira H (2002) An approximately unbiased test of phylogenetic tree selection. Syst Biol 51:492–508
Stephens M, Donnelly P (2003) A comparison of bayesian methods for haplotype reconstruction from population genotype data. Am J Hum Genet 73:1162–1169
Steppan SJ, Adkins RM, Anderson J (2004) Phylogeny and divergence-date estimates of rapid radiations in muroid rodents based on multiple nuclear genes. Syst Biol 53:533–553
Swanson WJ, Vacquier VD (2002) The rapid evolution of reproductive proteins. Nat Rev Genet 3:137–144
Swanson WJ, Wong A, Wolfner MF, Aquadro CF (2004) Evolutionary expressed sequence tag analysis of Drosophila female reproductive tracts identifies genes subjected to positive selection. Genetics 168:1457–1465
Talley HM, Laukaitis CM, Karn RC (2001) Female preference for male saliva: implications for sexual isolation of Mus musculus subspecies. Evolution 55:631–634
Tamura K, Peterson D, Peterson N, Stecher G, Nei M, Kumar S (2011) MEGA5: molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol Biol Evol 28:2731–2739
Thompson JD, Higgins DG, Gibson TJ (1994) CLUSTAL W: improving the sensitivity of progressive multiple sequence alignment through sequence weighting, position-specific gap penalties and weight matrix choice. Nucleic Acids Res 22:4673–4680
Torgerson DG, Kulathinal RJ, Singh RS (2002) Mammalian sperm proteins are rapidly evolving: evidence of positive selection in functionally diverse genes. Mol Biol Evol 19:1973–1980
Wickliffe JK, Lee VH, Smith E, Tandler B, Phillips CJ (2002) Gene expression, cell localization, and evolution of rodent submandibular gland androgen-binding protein. Eur J Morphol 40:257–260
Yang Z (1998) Likelihood ratio tests for detecting positive selection and application to primate lysozyme evolution. Mol Biol Evol 15:568–573
Yang Z (2007) PAML 4: phylogenetic analysis by maximum likelihood. Mol Biol Evol 24:1586–1591
Yang Z, Wong WSW, Nielsen R (2005) Bayes empirical Bayes inference of amino acid sites under positive selection. Mol Biol Evol 22:1107–1118
Zhang X, Firestein S (2002) The olfactory receptor gene superfamily of the mouse. Nat Neurosci 5:124–133
Acknowledgments
We thank R. Baker, R. Chesser, B. Rodgers, M. Bondarkov, and S. Gaschak for access to collecting sites in northern Ukraine as part of another research project. We also want to thank two anonymous reviewers whose comments and suggestions greatly improved the quality of this manuscript. All field and laboratory studies were conducted with appropriate permits and approved institutional protocols and in accord with the NIH Guidelines for the Care and Use of Laboratory Animals. Research was partially funded by Texas Tech University funding to CJP and directed funding for the DOE (C. J. Phillips, R. Chesser, R. Baker, co-PIs). FGH acknowledges Grant support from the National Science Foundation (EPS-0903787).
Author information
Authors and Affiliations
Corresponding author
Electronic Supplementary Material
Below is the link to the electronic supplementary material.
Rights and permissions
About this article
Cite this article
Vandewege, M.W., Phillips, C.J., Wickliffe, J.K. et al. Evolution of the ABPA Subunit of Androgen-Binding Protein Expressed in the Submaxillary Glands in New and Old World Rodent Taxa. J Mol Evol 76, 324–331 (2013). https://doi.org/10.1007/s00239-013-9561-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00239-013-9561-4